Abstract
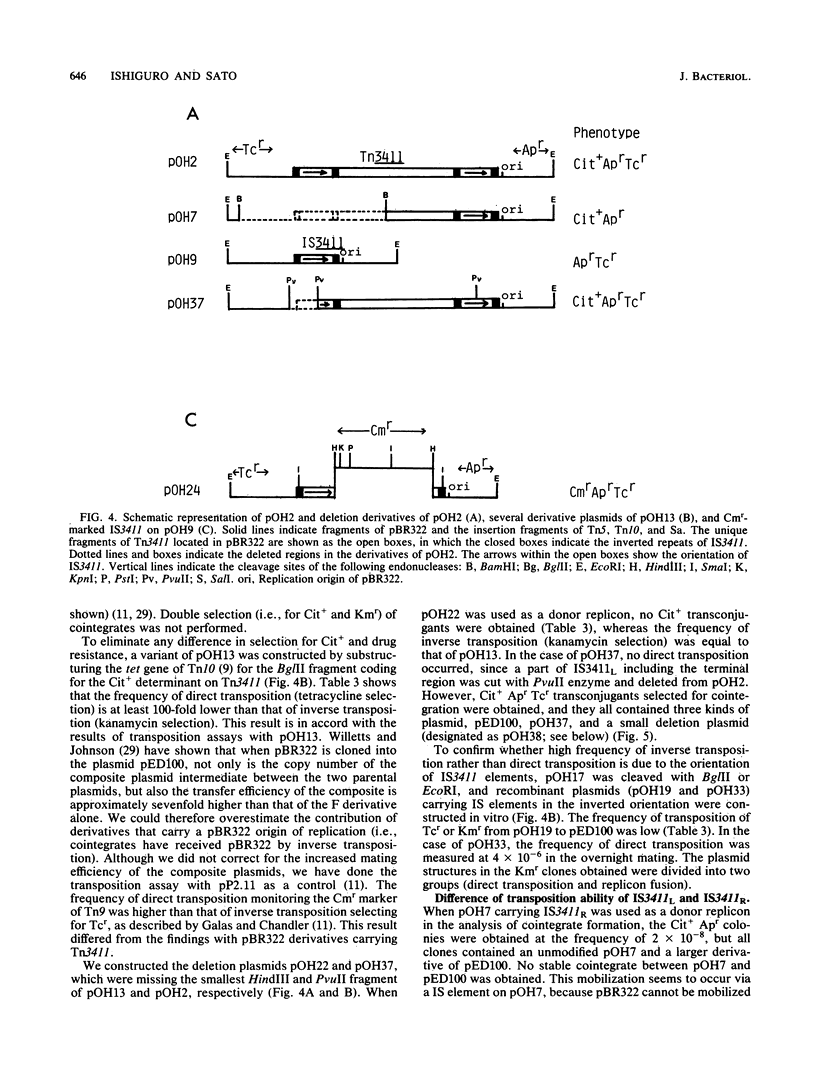
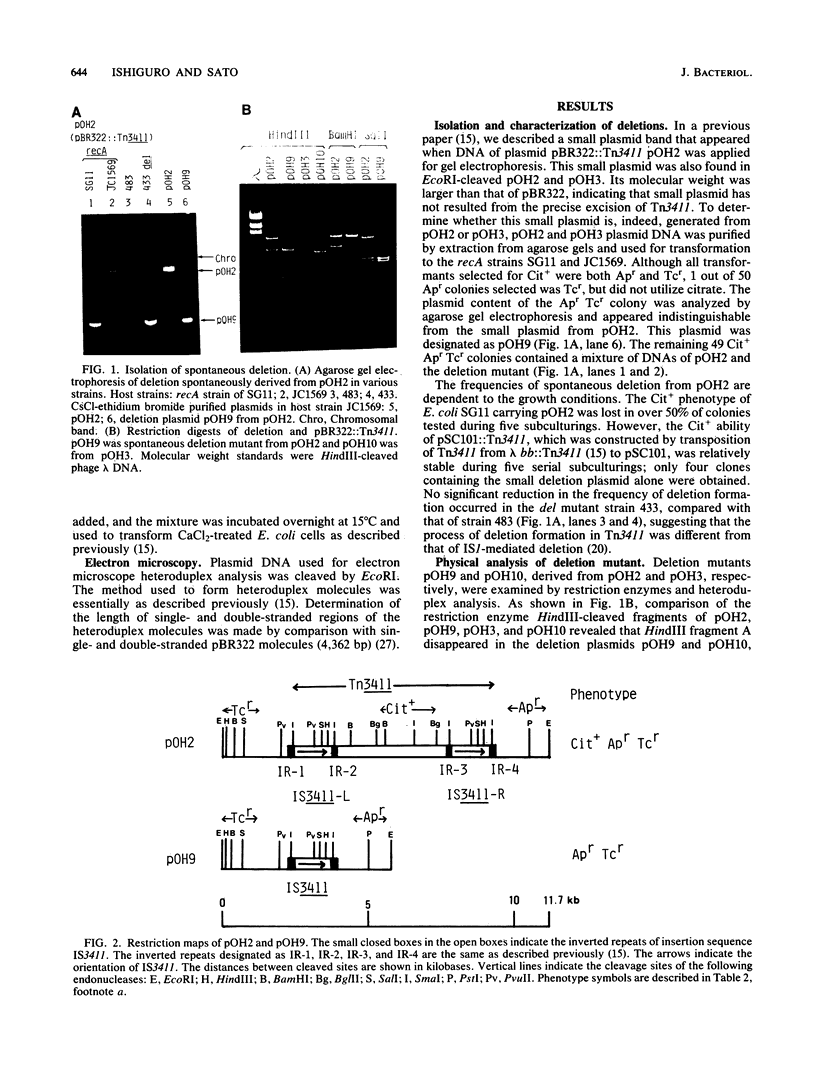
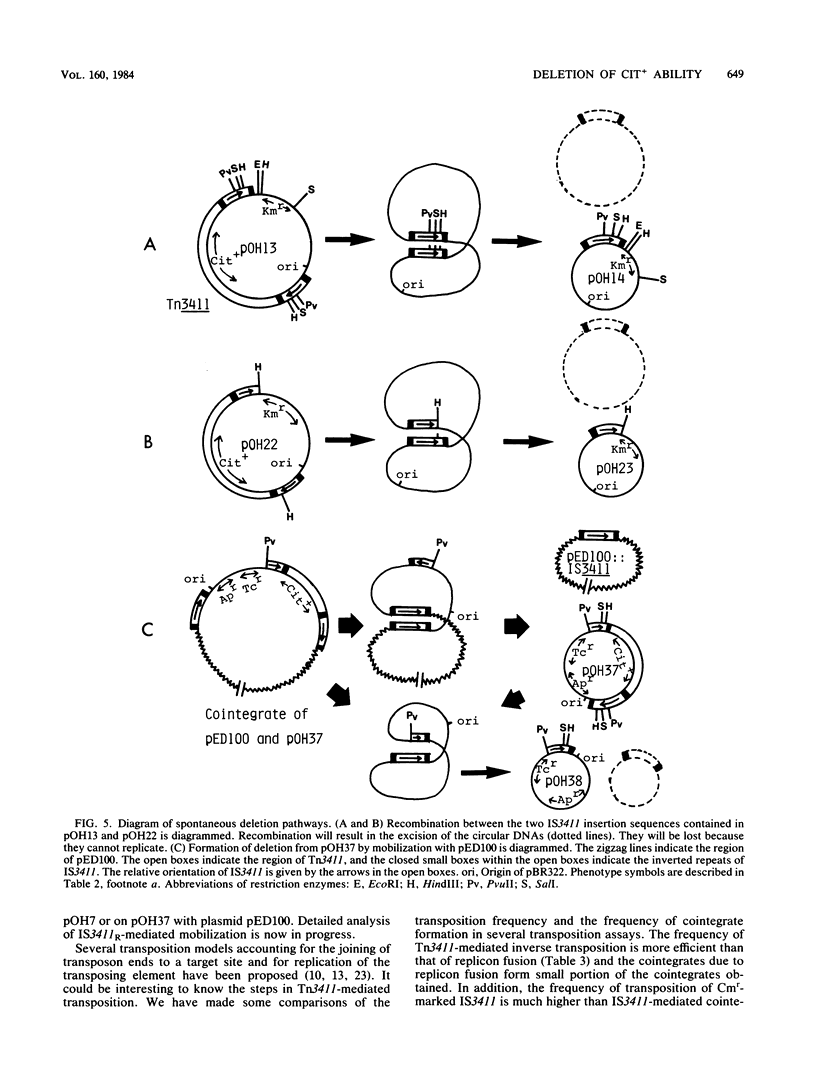
The citrate utilization (Cit+) transposon Tn3411 was shown to be flanked by directly repeated sequences (IS3411L and IS3411R) by restriction enzyme analysis and electron microscope observation. Cit- deletion mutants were frequently found to be generated in pBR322::Tn3411 by intramolecular recombination between the two copies of IS3411. The flanking IS3411 elements of Tn3411 were shown to be functional insertion sequences by Tn3411-mediated direct and inverse transposition. Tn3411-mediated inverse transposition from pBR322::Tn3411 to the F-plasmid derivative pED100 occurred more efficiently than that of direct transposition of the Cit+ determinant. This was thought to be due to the differential transposability of IS3411L and IS3411R in the transposition process. The frequency of transposition of IS3411 marked with a chloramphenicol resistance determinant was much higher than IS3411-mediated cointegrate formation, suggesting that replicon fusions are not essential intermediates in the transposition process of Tn3411 or IS3411. Spontaneous deletions occurred with high frequency in recA hosts. The spontaneous deletion promoted by homologous recombination between two IS3411 elements in Tn3411 was examined with deletion mutants.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albertini A. M., Hofer M., Calos M. P., Miller J. H. On the formation of spontaneous deletions: the importance of short sequence homologies in the generation of large deletions. Cell. 1982 Jun;29(2):319–328. doi: 10.1016/0092-8674(82)90148-9. [DOI] [PubMed] [Google Scholar]
- Bachmann B. J., Low K. B., Taylor A. L. Recalibrated linkage map of Escherichia coli K-12. Bacteriol Rev. 1976 Mar;40(1):116–167. doi: 10.1128/br.40.1.116-167.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg D. E., Johnsrud L., McDivitt L., Ramabhadran R., Hirschel B. J. Inverted repeats of Tn5 are transposable elements. Proc Natl Acad Sci U S A. 1982 Apr;79(8):2632–2635. doi: 10.1073/pnas.79.8.2632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Transposable elements. Cell. 1980 Jul;20(3):579–595. doi: 10.1016/0092-8674(80)90305-0. [DOI] [PubMed] [Google Scholar]
- Chandler M., Roulet E., Silver L., Boy de la Tour E., Caro L. Tn10 mediated integration of the plasmid R100.1 into the bacterial chromosome: inverse transposition. Mol Gen Genet. 1979 May 23;173(1):23–30. doi: 10.1007/BF00267687. [DOI] [PubMed] [Google Scholar]
- Cohen S. N. Transposable genetic elements and plasmid evolution. Nature. 1976 Oct 28;263(5580):731–738. doi: 10.1038/263731a0. [DOI] [PubMed] [Google Scholar]
- Collins J., Volckaert G., Nevers P. Precise and nearly-precise excision of the symmetrical inverted repeats of Tn5; common features of recA-independent deletion events in Escherichia coli. Gene. 1982 Jul-Aug;19(1):139–146. doi: 10.1016/0378-1119(82)90198-6. [DOI] [PubMed] [Google Scholar]
- Cornelis G., Ghosal D., Saedler H. Tn951: a new transposon carrying a lactose operon. Mol Gen Genet. 1978 Apr 6;160(2):215–224. doi: 10.1007/BF00267484. [DOI] [PubMed] [Google Scholar]
- Foster T. J., Davis M. A., Roberts D. E., Takeshita K., Kleckner N. Genetic organization of transposon Tn10. Cell. 1981 Jan;23(1):201–213. doi: 10.1016/0092-8674(81)90285-3. [DOI] [PubMed] [Google Scholar]
- Galas D. J., Chandler M. On the molecular mechanisms of transposition. Proc Natl Acad Sci U S A. 1981 Aug;78(8):4858–4862. doi: 10.1073/pnas.78.8.4858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galas D. J., Chandler M. Structure and stability of Tn9-mediated cointegrates. Evidence for two pathways of transposition. J Mol Biol. 1982 Jan 15;154(2):245–272. doi: 10.1016/0022-2836(82)90063-8. [DOI] [PubMed] [Google Scholar]
- Garaev M. M., Bobkov A. F., Bobkova A. F., Kalinin V. N., Smirnov V. D., Khudakov YuE, Tikchonenko T. I. The site-specific deletion in plasmid pBR322. Gene. 1982 Apr;18(1):21–28. doi: 10.1016/0378-1119(82)90052-x. [DOI] [PubMed] [Google Scholar]
- Harshey R. M., Bukhari A. I. A mechanism of DNA transposition. Proc Natl Acad Sci U S A. 1981 Feb;78(2):1090–1094. doi: 10.1073/pnas.78.2.1090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Ishiguro N., Sato G., Sasakawa C., Danbara H., Yoshikawa M. Identification of citrate utilization transposon Tn3411 from a naturally occurring citrate utilization plasmid. J Bacteriol. 1982 Mar;149(3):961–968. doi: 10.1128/jb.149.3.961-968.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones I. M., Primrose S. B., Ehrlich S. D. Recombination between short direct repeats in a recA host. Mol Gen Genet. 1982;188(3):486–489. doi: 10.1007/BF00330053. [DOI] [PubMed] [Google Scholar]
- Kleckner N. Transposable elements in prokaryotes. Annu Rev Genet. 1981;15:341–404. doi: 10.1146/annurev.ge.15.120181.002013. [DOI] [PubMed] [Google Scholar]
- LENNOX E. S. Transduction of linked genetic characters of the host by bacteriophage P1. Virology. 1955 Jul;1(2):190–206. doi: 10.1016/0042-6822(55)90016-7. [DOI] [PubMed] [Google Scholar]
- Rothstein S. J., Reznikoff W. S. The functional differences in the inverted repeats of Tn5 are caused by a single base pair nonhomology. Cell. 1981 Jan;23(1):191–199. doi: 10.1016/0092-8674(81)90284-1. [DOI] [PubMed] [Google Scholar]
- Sasakawa C., Berg D. E. IS50-mediated inverse transposition. Discrimination between the two ends of an IS element. J Mol Biol. 1982 Aug 5;159(2):257–271. doi: 10.1016/0022-2836(82)90495-8. [DOI] [PubMed] [Google Scholar]
- Shapiro J. A. Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1933–1937. doi: 10.1073/pnas.76.4.1933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- So M., Heffron F., McCarthy B. J. The E. coli gene encoding heat stable toxin is a bacterial transposon flanked by inverted repeats of IS1. Nature. 1979 Feb 8;277(5696):453–456. doi: 10.1038/277453a0. [DOI] [PubMed] [Google Scholar]
- Stanisich V. A., Bennett P. M., Richmond M. H. Characterization of a translocation unit encoding resistance to mercuric ions that occurs on a nonconjugative plasmid in Pseudomonas aeruginosa. J Bacteriol. 1977 Mar;129(3):1227–1233. doi: 10.1128/jb.129.3.1227-1233.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starlinger P. IS elements and transposons. Plasmid. 1980 May;3(3):241–259. doi: 10.1016/0147-619x(80)90039-6. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Tait R. C., Lundquist R. C., Kado C. I. Genetic map of the crown gall suppressive IncW plasmid pSa. Mol Gen Genet. 1982;186(1):10–15. doi: 10.1007/BF00422905. [DOI] [PubMed] [Google Scholar]
- Willetts N., Johnson D. pED100, a conjugative F plasmid derivative without insertion sequences. Mol Gen Genet. 1981;182(3):520–522. doi: 10.1007/BF00293948. [DOI] [PubMed] [Google Scholar]




