Abstract
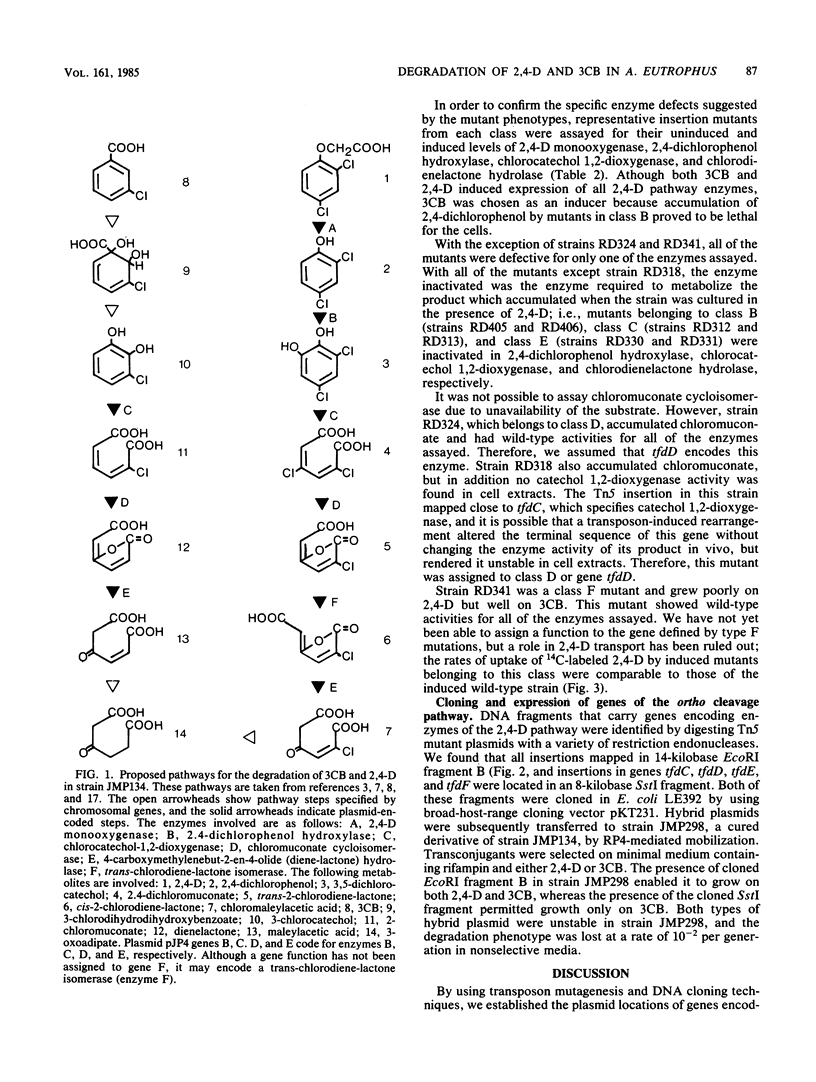
Plasmid pJP4 permits its host bacterium, strain JMP134, to degrade and utilize as sole sources of carbon and energy 3-chlorobenzoate and 2,4-dichlorophenoxyacetic acid (R. H. Don and J. M. Pemberton, J. Bacteriol. 145:681-686, 1981). Mutagenesis of pJP4 by transposons Tn5 and Tn1771 enabled localization of five genes for enzymes involved in these catabolic pathways. Four of the genes, tfdB, tfdC, tfdD, and tfdE, encoded 2,4-dichlorophenol hydroxylase, dichlorocatechol 1,2-dioxygenase, chloromuconate cycloisomerase, and chlorodienelactone hydrolase, respectively. No function has been assigned to the fifth gene, tfdF, although it may encode a trans-chlorodiene-lactone isomerase. Inactivation of genes tfdC, tfdD, and tfdE, which encode the transformation of dichlorocatechol to chloromaleylacetic acid, prevented host strain JMP134 from degrading both 3-chlorobenzoate and 2,4-dichlorophenoxyacetic acid, which indicates that the pathways for these two substrates utilize common enzymes for the dissimilation of chlorocatechols. Studies with cloned catabolic genes from pJP4 indicated that whereas all essential steps in the degradation of 2,4-dichlorophenoxyacetic acid are plasmid encoded, the conversion of 3-chlorobenzoate to chlorocatechol is specified by chromosomal genes.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berg D. E., Weiss A., Crossland L. Polarity of Tn5 insertion mutations in Escherichia coli. J Bacteriol. 1980 May;142(2):439–446. doi: 10.1128/jb.142.2.439-446.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Don R. H., Pemberton J. M. Properties of six pesticide degradation plasmids isolated from Alcaligenes paradoxus and Alcaligenes eutrophus. J Bacteriol. 1981 Feb;145(2):681–686. doi: 10.1128/jb.145.2.681-686.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorn E., Hellwig M., Reineke W., Knackmuss H. J. Isolation and characterization of a 3-chlorobenzoate degrading pseudomonad. Arch Microbiol. 1974;99(1):61–70. doi: 10.1007/BF00696222. [DOI] [PubMed] [Google Scholar]
- Dorn E., Knackmuss H. J. Chemical structure and biodegradability of halogenated aromatic compounds. Substituent effects on 1,2-dioxygenation of catechol. Biochem J. 1978 Jul 15;174(1):85–94. doi: 10.1042/bj1740085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorn E., Knackmuss H. J. Chemical structure and biodegradability of halogenated aromatic compounds. Two catechol 1,2-dioxygenases from a 3-chlorobenzoate-grown pseudomonad. Biochem J. 1978 Jul 15;174(1):73–84. doi: 10.1042/bj1740073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans W. C., Smith B. S., Fernley H. N., Davies J. I. Bacterial metabolism of 2,4-dichlorophenoxyacetate. Biochem J. 1971 May;122(4):543–551. doi: 10.1042/bj1220543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher P. R., Appleton J., Pemberton J. M. Isolation and characterization of the pesticide-degrading plasmid pJP1 from Alcaligenes paradoxus. J Bacteriol. 1978 Sep;135(3):798–804. doi: 10.1128/jb.135.3.798-804.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilardi G. L. Characterization of nonfermentative nonfastidious gram negative bacteria encountered in medical bacteriology. J Appl Bacteriol. 1971 Sep;34(3):623–644. doi: 10.1111/j.1365-2672.1971.tb02326.x. [DOI] [PubMed] [Google Scholar]
- Hansen J. B., Olsen R. H. Isolation of large bacterial plasmids and characterization of the P2 incompatibility group plasmids pMG1 and pMG5. J Bacteriol. 1978 Jul;135(1):227–238. doi: 10.1128/jb.135.1.227-238.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Murray N. E., Brammar W. J., Murray K. Lambdoid phages that simplify the recovery of in vitro recombinants. Mol Gen Genet. 1977 Jan 7;150(1):53–61. doi: 10.1007/BF02425325. [DOI] [PubMed] [Google Scholar]
- Schmidt E., Knackmuss H. J. Chemical structure and biodegradability of halogenated aromatic compounds. Conversion of chlorinated muconic acids into maleoylacetic acid. Biochem J. 1980 Oct 15;192(1):339–347. doi: 10.1042/bj1920339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wigmore G. J., Ribbons D. W. p-Cymene pathway in Pseudomonas putida: selective enrichment of defective mutants by using halogenated substrate analogs. J Bacteriol. 1980 Aug;143(2):816–824. doi: 10.1128/jb.143.2.816-824.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]


