Abstract
We developed novel plasmids and T-DNA binary vectors that incorporate a modified and more useful form of the superpromoter. The superpromoter consists of a trimer of the octopine synthase transcriptional activating element affixed to the mannopine synthase2′ (mas2′) transcriptional activating element plus minimal promoter. We tested a superpromoter-β-glucuronidaseA fusion gene in stably transformed tobacco (Nicotiana tabacum) and maize (Zea mays) plants and in transiently transformed maize Black Mexican Sweet protoplasts. In both tobacco and maize, superpromoter activity was much greater in roots than in leaves. In tobacco, superpromoter activity was greater in mature leaves than in young leaves, whereas in maize activity differed little among the tested aerial portions of the plant. When compared with other commonly used promoters (cauliflower mosaic virus 35S, mas2′, and maize ubiquitin), superpromoter activity was approximately equivalent to those of the other promoters in both maize Black Mexican Sweet suspension cells and in stably transformed maize plants. The addition of a maize ubiquitin intron downstream of the superpromoter did not enhance activity in stably transformed maize.
The availability of convenient vectors harboring a strong promoter that is active in all or most cells of different plant species would be useful for a variety of applications in plant molecular biology. We previously described a novel synthetic promoter consisting of a trimer of the octopine synthase (ocs) transcriptional activating element (ocs activator) linked to the mannopine synthase2′ (mas2′) activator-promoter region (Ni et al., 1995). Initial studies in tobacco (Nicotiana tabacum) indicated that this promoter, called the superpromoter, could direct expression of GUS activity to a level 2- to 20-fold higher than the commonly used enhanced double cauliflower mosaic virus (CaMV) 35S promoter (Ni et al., 1995). The activity of the superpromoter was highest in roots, but also was high in leaves and stems.
The superpromoter was originally created by ligating three ocs activator fragments (positions −333 to −116 relative to the transcription start site [Leisner and Gelvin, 1988]) from the ocs gene to the mas2′ activator-promoter region (−318 to +65 relative to the transcription start site [Ellis et al., 1984]), all from the Agrobacterium tumefaciens Ti-plasmid pTiA6. The construction of the original superpromoter resulted in the repeated presence within the promoter of several commonly used restriction endonuclease sites (BamHI, EcoRI, HindIII), as well as the presence of the restriction endonuclease sites PstI and XhoI. This feature precluded easily linking genes to the promoter. In addition, it complicated further analysis of T-DNA insertions in the plant genome. We therefore modified the superpromoter, eliminating most of these internal restriction endonuclease sites. This modified superpromoter (MSP) formed the basis for the construction of several novel plant expression and T-DNA binary vectors. Here, we describe these vectors and the use of the superpromoter to promote reporter-gusA gene expression in tobacco and maize (Zea mays).
RESULTS AND DISCUSSION
Construction of Superpromoter Cassettes in pUC119
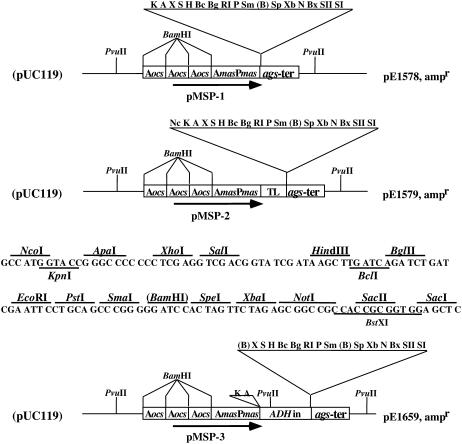
To provide researchers with convenient tools for various applications in plant gene expression, we constructed expression cassettes containing the MSP. The plasmid pUC119 (Vieira and Messing, 1987; GenBank accession no. U07650) formed the basis for these initial cassettes (the pMSP series), whose maps are shown in Figure 1.
Figure 1.
Maps of the pUC119-based pMSP series of plasmids. pExxxx numbers at the right of each plasmid map indicate Gelvin laboratory stock numbers. ampr, Plasmid confers resistance to ampicillin upon host bacteria; Aocs, ocs transcriptional activating element; AmasPmas, mas2′-activating and promoter elements; ags-ter, poly(A) addition signal from the agropine synthase gene; ADHin, intron from the maize alcohol dehydrogenase I gene. Restriction endonuclease sites: A, ApaI; B, BamHI; Bc, BclI; Bg, BglII; Bx, BstXI; H, HindIII; N, NotI; Nc, NcoI; P, PstI; RI, EcoRI; SI, SacI; SII, SacII; S, SalI; Sm, SmaI; Sp, SpeI; Xb, XbaI; X, XhoI. Restriction endonuclease sites within parentheses are not unique to the plasmid. The sequence below pMSP-2 shows the TL reading frame following the NcoI site that is at the 3′ end of the TEV TL leader sequence.
Plasmid pMSP-1 contains the MSP followed by a multiple cloning site (MCS) and a poly(A) addition signal. Translational (TL) enhancers from plant viruses (e.g. the ω-element from Tobacco mosaic virus [Gallie and Walbot, 1992] and the TL enhancer element from Tobacco etch virus [TEV; Restrepo et al., 1990]) may enhance the level of gene expression 2- to 3-fold (Holtorf et al., 1995). We constructed a second cassette, pMSP-2, with the TL enhancer element from TEV linked downstream of the superpromoter. The presence of introns within the 5′-untranslated region of genes often enhances expression in maize (Mascarenhas et al., 1990) and other cereals (Cornejo et al., 1993). We therefore created a third cassette, pMSP-3, containing a maize adh1 intron following the superpromoter for improved expression in monocot plant species.
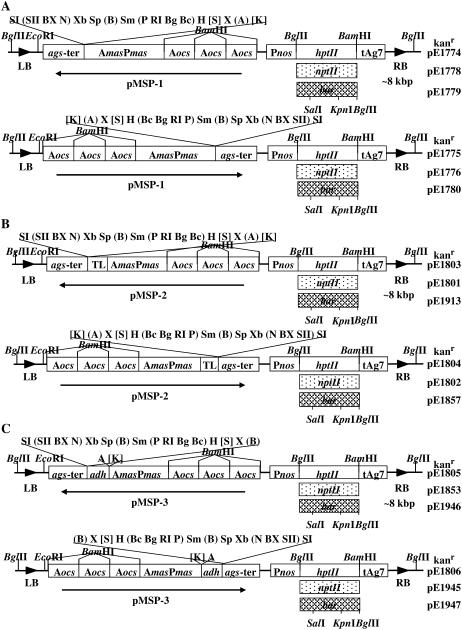
We subsequently constructed plant transformation vectors based upon pMSP-1, pMSP-2, and pMSP-3 in the T-DNA binary vectors pGPTV-KAN, pGPTV-HPT, and pGPTV-BAR (Becker et al., 1992). Figure 2 shows maps of these vectors.
Figure 2.
Maps of the T-DNA binary vectors based upon the pMSP series of plasmids. pExxxx numbers at the right of each plasmid map indicate Gelvin laboratory stock numbers. kanr, Plasmid confers resistance to kanamycin upon host bacteria; Pnos, nos promoter; tAg7, poly(A) addition signal for T-DNA gene 7; hptII, gene conferring resistance to hygromycin; nptII, gene conferring resistance to kanamycin; bar, gene conferring resistance to phosphinothricin/Basta/Bialophos. Other symbols and restriction endonuclease sites are as in the legend to Figure 1. The bar gene contains KpnI and SalI sites; therefore, these sites are not unique to T-DNA binary vectors containing the bar gene (indicated by [K] and [Sl]). A, T-DNA binary vectors based on the pMSP-1 series of plasmids. B, T-DNA binary vectors based upon the pMSP-2 series of plasmids. C, T-DNA binary vectors based on the pMSP-3 series of plasmids.
In summary, we modified the original superpromoter (Ni et al., 1995) by eliminating most of the commonly used restriction endonuclease sites from within the promoter sequence. We constructed 21 new vectors based upon pUC119 and the T-DNA binary vector pGPTV, containing the MSP. These vectors have the following features: (1) MSP; (2) various plant regulatory sequences for improved gene expression (either a maize adh1 intron for expression in monocot plant species or a TL enhancer from TEV for expression in dicot plant species); (3) MCS with eight to 16 unique restriction endonuclease sites (depending upon the vector) downstream of the promoter for cloning genes and more facile analysis of T-DNA insertions in the plant genome; (4) agropine or nopaline synthase (nos) poly(A) addition signals; and (5) markers for binary vectors for selection in plants using kanamycin, hygromycin, or phosphinothricin/Basta/Bialophos.
Activity of the Superpromoter in Transgenic Tobacco and Maize
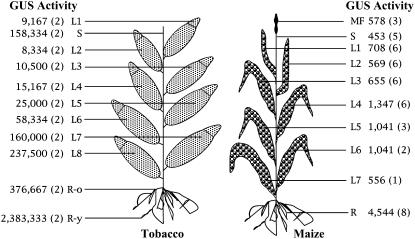
We previously showed that in transgenic tobacco, a superpromoter-gusA construction was approximately 5-fold more active in roots than in leaves (Ni et al., 1995). We carried out a more detailed investigation of the activity of the superpromoter in various organs of stably transformed tobacco and maize plants containing a superpromoter-gusA expression cassette. These tests were conducted before construction of the series of superpromoter vectors described above; the constructs therefore did not have the exact same configuration as the current vectors. In addition, our original constructions contained (when applicable) the maize ubiquitin intron, rather than the maize adh1 intron incorporated into the current set of vectors. However, tests in transgenic tobacco indicated that the current superpromoter configuration behaves in a manner almost identical to that of the older version of the superpromoter (data not shown). Figure 3 shows that in tobacco, GUS activity was greater in older (lower) leaves than in younger (upper) leaves. Interestingly, this is opposite to the expression pattern directed by the CaMV 35S promoter, which directs expression of transgenes most strongly in younger, meristematic tissues (Williamson et al., 1989). Conversely, GUS activity in young root tips was much higher than that in older portions of the roots. The pattern of expression in transgenic maize, however, was different. There was relatively little difference in GUS activity among younger or older leaves. As with the situation in transgenic tobacco, GUS activity in transgenic maize roots was considerably greater than that in leaves.
Figure 3.
GUS activity (pmol/min/mg protein) directed by the superpromoter in various tobacco and maize tissues and organs. L1, L2, etc., Leaf number; MF, maize flower; R, root; R-o, old root section; R-y, young root section; S, stem section. Numbers in parentheses indicate the number of samples analyzed for each tissue or organ.
Comparison of the Activities of the Superpromoter and Other Promoters in Maize
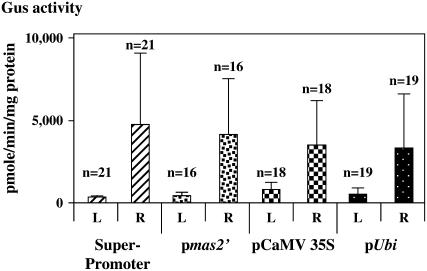
We previously showed that the superpromoter was as strong as or stronger than several commonly used promoters in transgenic tobacco (Ni et al., 1995). We therefore asked how the activity of the superpromoter compared with that of other promoters in monocots. Figure 4 shows a comparison of the relative strengths of the superpromoter, the mas2′ promoter, the CaMV 35S promoter (−800 from pBI121; Jefferson et al., 1987), and the maize ubiquitin promoter in transgenic maize. Within a factor of 2, all of these transcriptional regulatory sequences promoted approximately the same level of GUS activity in leaves, as well as in roots. These data indicate that the superpromoter functions as well as other commonly used promoters in these maize tissues.
Figure 4.
Comparison of the relative strengths of various promoters in the leaves and roots of transgenic maize plants. Ubi, Maize ubiquitin promoter; n, number of individual transgenic plants analyzed. The large sd most likely reflects the position effect of T-DNA integration upon transgene activity in independent transgenic events.
Response of the Superpromoter and Other Promoters to the Presence of Introns
The presence of introns enhances the activity of promoters for gene expression, especially in monocots (Mascarenhas et al., 1990; Cornejo et al., 1993). We therefore tested whether the presence of introns would enhance the activity of the superpromoter and several commonly used promoters in maize. Figure 5 shows the constructions that we tested. Each set of constructions contains either the superpromoter, the mas2′ promoter, the CaMV 35S promoter, or the maize ubiquitin promoter. Set I contains no intron in the gusA gene, set II contains the potato (Solanum tuberosum) ST-LS1 intron (Vancanneyt et al., 1990) in the gusA coding sequence, and set III contains a maize ubiquitin intron preceding the gusA coding sequence. As a control, we also generated a promoterless gusA construction.
Figure 5.
Constructions used for maize BMS protoplast transfection experiments. nos, Nopaline synthase poly(A) addition signal sequence; pUbi, maize ubiquitin promoter.
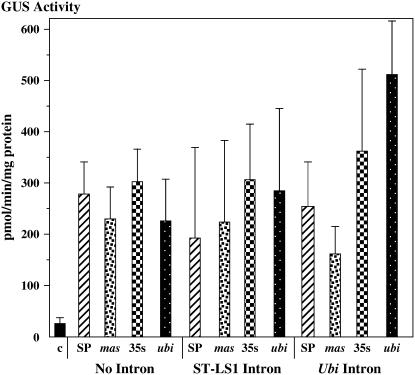
We first tested these constructions in transiently transformed maize Black Mexican Sweet (BMS) protoplasts. Within a factor of 2.5, all of these constructions (except the control, promoterless gusA gene, which yielded only a background level of activity) elicited approximately the same amount of GUS activity. Figure 6 shows the average data from four independent experiments. Thus, in transient expression assays, all of these promoter-intron combinations functioned equally well in maize suspension cells.
Figure 6.
Comparison of the relative strengths of various promoters (lacking or containing introns in the gusA gene) in transiently transfected maize BMS protoplasts. Numbers represent the average of four independent experiments with sd. C, Control construction lacking a promoter; mas, mas2′ promoter; 35S, CaMV 35S promoter; sp, superpromoter.
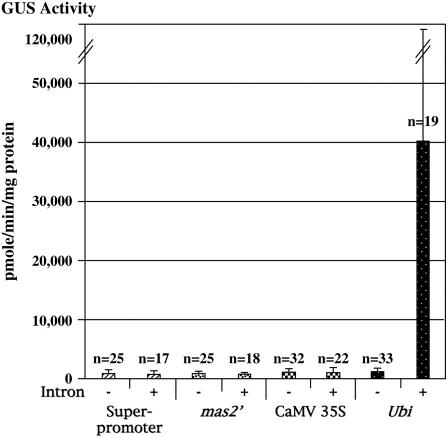
Promoters may have different activities, depending on whether they direct expression from nonintegrated transgenes in transiently transformed cells or from integrated transgenes that may be under transcriptional constraints of assembled chromatin (e.g. see Frisch et al., 1995). Strong expression of a linked CHLORAMPHENICOL ACETYLTRANSFERASE (CAT) reporter gene directed by a maize ubiquitin promoter plus ubiquitin leader intron in electroporated maize BMS cells has previously been described by Christensen et al. (1992). Enhancement of gene expression, resulting from increased stability of cytoplasmic mRNA, has also been reported as a result of incorporating the maize Adh1 intron into reporter constructions (Callis et al., 1987; Luehrsen and Walbot, 1994). We therefore tested, in stably transformed maize plants, the response of each of four promoters to the presence of the maize ubiquitin intron. Interestingly, the activity of only the ubiquitin promoter increased in maize leaves in the presence of the ubiquitin intron (Fig. 7). This response was up to 55-fold, indicating that, among the eight promoter-intron combinations tested, the strongest transcriptional regulatory sequence in maize is clearly the maize ubiquitin promoter followed by the maize ubiquitin intron.
Figure 7.
Effect of the maize ubiquitin intron upon promoter activity in stably transformed maize leaves. n, Number of individual transgenic plants analyzed; −, constructions lacking the maize ubiquitin intron; +, constructions containing the maize ubiquitin intron. The large sd in the data from maize plants containing the maize ubiquitin promoter plus intron construction most likely reflects the position effect of T-DNA integration upon transgene activity in independent transgenic events.
Our results contrast with those of Callis et al. (1987), who demonstrated that introduction of the Adh1 first intron into constructions containing either the CaMV 35S or nos promoters increased activity of a linked CAT gene 8- and 170-fold, respectively. Addition of a maize Bronze1 (Bz1) intron to the CaMV 35S promoter also stimulated CAT activity. This stimulation was associated with increased steady-state levels of CAT mRNA. However, these authors utilized Adh1 and Bz1 introns, rather than the ubiquitin intron used in our study, and they performed their analyses in transiently transfected maize BMS protoplasts. It is possible that the maize ubiquitin promoter plus intron combination functions optimally in stably transformed maize tissues, a system not tested by Callis et al. (1987).
Use of the Superpromoter in Various Plant Species
We reconfigured the original superpromoter (Ni et al., 1995) such that it could easily be utilized for gene expression in a number of plant species. Over approximately the past 10 years, our laboratory has distributed the various superpromoter constructions to dozens of laboratories. In our laboratory, the superpromoter has routinely and effectively been used to drive both transient and stable transgene expression in tobacco plants and Bright Yellow-2 cell suspensions (Ni et al., 1995; He et al., 1996; Narasimhulu et al., 1996; Kononov et al., 1997; Mysore et al., 1998; Veena et al., 2003; Lee and Gelvin, 2004), Arabidopsis (Arabidopsis thaliana) plants and cell suspensions (Nam et al., 1997, 1998, 1999; Mysore et al., 2000; Yi et al., 2002; Zhu et al., 2003; Gaspar et al., 2004; Hwang and Gelvin, 2004; Hwang et al., 2006; Crane and Gelvin, 2007; Kim et al., 2007), maize plants and BMS cell suspensions (Narasimhulu et al., 1996), and Brassica napus plants (S. Johnson and S.B. Gelvin, unpublished data). Others have shown that transcriptional regulatory sequences very similar to those of the superpromoter appear to be less susceptible to silencing than are other commonly used promoters, including the CaMV 35S and cassava vein mosaic virus promoters (De Bolle et al., 2003; Butaye et al., 2004).
The vectors described in this article can be obtained by contacting Dr. Stanton B. Gelvin (gelvin@bilbo.bio.purdue.edu) following completion of a Materials Transfer Agreement.
MATERIALS AND METHODS
Superpromoter and Vector Constructions
The original superpromoter contains a trimer of the ocs activator sequence (Aocs), cloned as a HindIII fragment upstream of the mas2′ activator plus promoter in pE1120. We modified the original superpromoter as follows.
We removed the superpromoter from pE1120 by partial digestion with HindIII plus complete digestion with XbaI. We cloned this fragment into the HindIII and XbaI sites of pBluescript KS(−) to generate the plasmid pE1037. Digestion of pE1037 with HindIII removed the three ocs activator sequences from the superpromoter. The single HindIII site within this resulting plasmid (pE1048) was converted into a BamHI site by filling in the overhanging nucleotides using Klenow fragment of DNA polymerase followed by annealing a BamHI linker, generating pE1049. We likewise converted the HindIII sites flanking Aocs into BamHI sites. We cloned this new Aocs fragment into pE1049. Screening of the resulting colonies yielded insertions of a single Aocs fragment (in either orientation), insertions of a dimer of the Aocs fragment (both in correct or both in inverted orientation), and a trimer of the Aocs fragment (all three in inverted orientation; pE1054).
We continued to modify pE1054, first by removing a PstI site within the superpromoter and subsequently transferring the newly MSP region into the SalI and XbaI sites of pUC119, generating pUΔP (pE1466). We next removed a XhoI site from the superpromoter region of pE1466 by filling in the XhoI site using Klenow fragment of DNA polymerase, generating pUΔPΔX (pE1467). We digested pE1467 with HindIII and SalI, filled in these sites using Klenow fragment, and ligated the blunted ends to remove sites (HindIII, SphI, PstI, BspMI, and SalI) between HindIII and SalI, generating pUΔPΔX-1.
We digested pUΔPΔX-1 with EcoRI, filled in the overhanging ends using Klenow fragment, and ligated into this site a 401-bp HincII-EcoRV fragment containing the agrocinopine synthase (ags) terminator (ags-ter) from pTiA6, generating pUΔPΔX-2 (pE1694). We digested pUΔPΔX-2 with XbaI and SmaI, filled in the overhanging ends using Klenow fragment, and self ligated the molecules, removing the XbaI, BamHI, and SmaI sites and generating the plasmid pUΔPΔX-3 (pE1695).
We digested pBluescript KS(−) with EcoRV and inserted a linker containing BglII and BclI sites. This plasmid, called pBB (pE1505), contains a new MCS region. We digested pUΔPΔX-3 with KpnI and SacI and inserted the MCS region from pBB using KpnI and SacI, generating pMSP-1 (pE1578).
Constructions in Escherichia coli were generated in strain DH10B. T-DNA binary vectors were introduced into Agrobacterium tumefaciens EHA101 (Hood et al., 1986) or EHA105 (Hood et al., 1993). E. coli strains were grown in Luria-Bertani medium containing the appropriate antibiotics (ampicillin, 100 μg/mL; kanamycin, 25 μg/mL). A. tumefaciens was grown in yeast (Saccharomyces cerevisiae) extract peptone-rich or Agrobacterium minimal medium (Lichtenstein and Draper, 1986) containing the appropriate antibiotics (rifampicin, 10 μg/mL; kanamycin, 100 μg/mL for plates, 25 μg/mL for liquid growth).
Maize BMS Suspension Culture and Protoplast Isolation
Maize (Zea mays) BMS cells were grown at room temperature with shaking (150 rpm) in medium containing Murashige and Skoog salts (Gibco), 20 g/L Suc, 2 mg/L 2,4-dichlorophenoxy acetic acid, 200 mg/L inositol, 130 mg/L Asn, 1.3 mg/L niacin, 0.25 mg/L thiamine, 0.25 mg/L pyridoxine, and 0.25 mg/L calcium pantothenate. Cells were isolated by centrifugation (1,000 rpm for 2 min) and suspended in isolation solution medium (Murashige and Skoog medium containing 0.4 m mannitol, 50 mm CaCl2, 10 mm sodium acetate, 15 mm 2-mercaptoethanol, 0.5 mg/L thiamine, and 2 mg/L 2,4-dichlorophenoxyacetic acid) containing 2% cellulase (Dyadic International), 0.1% pectolyase (Sigma), and 0.1% bovine serum albumin. BMS cells were incubated for 2 h with shaking at 80 rpm. Protoplasts were harvested, purified by centrifugation at 1,000 rpm for 10 min, and suspended in 50 mL electroporation solution (Murashige and Skoog plus 0.4 m mannitol, 10 mm HEPES [pH 9.6], 130 mm KCl, and 4 mm CaCl2). Electroporation was carried out using a Bio-Rad GenePulser at 200 V, capacitance 1,200, and 100-ms discharge. Following electroporation, cells were transferred to petri dishes containing EPS with 0.5 m mannitol and incubated in the dark at room temperature until assayed for GUS activity 18 h later.
Transformation Procedures
Transgenic tobacco (Nicotiana tabacum) plants were generated and maintained as previously described (Ni et al., 1995). Transgenic maize plants (genotype Hi-II) were generated as previously described (Frame et al., 2000).
GUS Activity Assays
Leaf explants were generated using a paper punch and assayed for GUS activity fluorimetrically as previously described (Jefferson et al., 1987).
DNA sequences of the superpromoter regions of pMSP-1, pMSP-2, and pMSP-3 can be found as GenBank accession numbers EU181145 (pMSP-1), EU181146 (pMSP-2), and EU181147 (pMSP-3).
This work was supported by the Biotechnology Research and Development Corporation, the Corporation for Plant Biotechnology Research, and the National Science Foundation (Plant Genome grant no. 0110023).
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Stanton B. Gelvin (gelvin@bilbo.bio.purdue.edu).
Open Access articles can be viewed online without a subscription.
References
- Becker D, Kemper E, Schell J, Masterson R (1992) New plant binary vectors with selectable markers located proximal to the left T-DNA border. Plant Mol Biol 20 1195–1197 [DOI] [PubMed] [Google Scholar]
- Butaye KMJ, Goderis IJWM, Wouters PFJ, Pues JMTG, Delaure SL, Broekaert WF, Depicker A, Cammue BPA, DeBolle FC (2004) Stable high-level transgene expression in Arabidopsis thaliana using gene silencing mutants and matrix attachment regions. Plant J 39 440–449 [DOI] [PubMed] [Google Scholar]
- Callis J, Fromm M, Walbot V (1987) Introns increase gene expression in cultured maize cells. Genes Dev 1 1183–1200 [DOI] [PubMed] [Google Scholar]
- Christensen AH, Sharrock RA, Quail PH (1992) Maize polyubiquitin genes: structure, thermal perturbation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol Biol 18 675–689 [DOI] [PubMed] [Google Scholar]
- Cornejo MJ, Luth D, Blankenship KM, Anderson OD, Blechl AE (1993) Activity of maize ubiquitin promoter in transgenic rice. Plant Mol Biol 23 567–581 [DOI] [PubMed] [Google Scholar]
- Crane YM, Gelvin SB (2007) RNAi-mediated gene silencing reveals involvement of Arabidopsis chromatin-related genes in Agrobacterium-mediated root transformation. Proc Natl Acad Sci USA 104 15156–15161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Bolle MFC, Butaye KMJ, Coucke WJW, Goderis IJWM, Wouters PFJ, van Boxel N, Broekaert WF, Commue BPA (2003) Analysis of the influence of promoter elements and a matrix attachment region on the inter-individual variation of transgene expression in populations of Arabidopsis thaliana. Plant Sci 164 169–179 [Google Scholar]
- Ellis JG, Ryder MH, Tate ME (1984) Agrobacterium tumefaciens TR-DNA encodes a pathway for agropine biosynthesis. Mol Gen Genet 195 466–473 [Google Scholar]
- Frame BR, Zhang H, Cocciolone SM, Sidorenko LV, Dietrich CR, Pegg SE, Zhen S, Schnable PS, Wang K (2000) Production of transgenic maize from bombarded type II callus: effect of gold particle size and callus morphology on transformation efficiency. In Vitro Cell Dev Biol Plant 36 21–29 [Google Scholar]
- Frisch DA, van der Geest AHM, Dias K, Hall TC (1995) Chromosomal integration is required for spatial regulation of expression from the β-phaseolin promoter. Plant J 7 503–512 [Google Scholar]
- Gallie DR, Walbot V (1992) Identification of the motifs within the tobacco mosaic virus 5′-leader responsible for enhancing translation. Nucleic Acids Res 20 4631–4638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaspar YM, Nam J, Schultz CJ, Lee LY, Gilson PR, Gelvin SB, Bacic A (2004) Characterization of the Arabidopsis lysine-rich arabinogalactan-protein AtAGP17 mutant (rat1) that results in a decreased efficiency of Agrobacterium transformation. Plant Physiol 135 2162–2171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He S, Abad A, Gelvin S, Mackenzie S (1996) A CMS-associated mitochondrial protein causes pollen disruption in transgenic tobacco. Proc Natl Acad Sci USA 93 11763–11768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holtorf S, Apel K, Bohlmann H (1995) Comparison of different constitutive and inducible promoters for the overexpression of transgenes in Arabidopsis thaliana. Plant Mol Biol 29 637–646 [DOI] [PubMed] [Google Scholar]
- Hood EE, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2 208–218 [Google Scholar]
- Hood EE, Helmer GL, Fraley RT, Chilton MD (1986) The hypervirulence of Agrobacterium tumefaciens A281 is encoded in a region of pTiBo542 outside of T-DNA. J Bacteriol 168 1291–1301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang HH, Gelvin SB (2004) Plant proteins that interact with VirB2, the Agrobacterium tumefaciens pilin protein, mediate plant transformation. Plant Cell 16 3148–3167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang HH, Mysore S, Gelvin SB (2006) Transgenic Arabidopsis plants expressing Agrobacterium tumefaciens VirD2 protein are less susceptible to Agrobacterium transformation. Mol Plant Pathol 7 473–484 [DOI] [PubMed] [Google Scholar]
- Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6 3901–3907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim SI, Veena, Gelvin SB (2007) Genome-wide analysis of Agrobacterium T-DNA integration sites in the Arabidopsis genome generated under non-selective conditions. Plant J 51 779–791 [DOI] [PubMed] [Google Scholar]
- Kononov ME, Bassuner B, Gelvin SB (1997) Integration of T-DNA binary vector “backbone” sequences into the tobacco genome: evidence for multiple complex patterns of integration. Plant J 11 945–957 [DOI] [PubMed] [Google Scholar]
- Lee LY, Gelvin SB (2004) Osa protein constitutes a strong oncogenic suppression system that can block vir-dependent transfer of IncQ plasmids between Agrobacterium cells, and the transfer of T-DNA and IncQ plasmids to plant cells. J Bacteriol 186 7254–7261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leisner SM, Gelvin SB (1988) Structure of the octopine synthase upstream activator sequence. Proc Natl Acad Sci USA 85 2553–2557 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lichtenstein C, Draper J (1986) Genetic engineering of plants. In DM Glover, ed, DNA Cloning. A Practical Approach. IRL Press, Washington, DC, pp 67–119
- Luehrsen KR, Walbot V (1994) Addition of A- and U-rich sequence increases the splicing efficiency of a deleted form of a maize intron. Plant Mol Biol 24 449–463 [DOI] [PubMed] [Google Scholar]
- Mascarenhas D, Mettler IJ, Pierce DA, Lowe HW (1990) Intron-mediated enhancement of heterologous gene expression in maize. Plant Mol Biol 15 913–920 [DOI] [PubMed] [Google Scholar]
- Mysore KS, Bassuner B, Deng Xb, Darbinian NS, Motchoulski A, Ream W, Gelvin SB (1998) Role of the Agrobacterium tumefaciens VirD2 protein in T-DNA transfer and integration. Mol Plant-Microbe Interact 11 668–683 [DOI] [PubMed] [Google Scholar]
- Mysore KS, Nam J, Gelvin SB (2000) An Arabidopsis histone H2A mutant is deficient in Agrobacterium T-DNA integration. Proc Natl Acad Sci USA 97 948–953 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nam J, Matthysse AG, Gelvin SB (1997) Differences in susceptibility of Arabidopsis ecotypes to crown gall disease may result from a deficiency in T-DNA integration. Plant Cell 9 317–333 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nam J, Mysore KS, Gelvin SB (1998) Agrobacterium tumefaciens transformation of the radiation hypersensitive Arabidopsis thaliana mutants uvh1 and rad5. Mol Plant Microbe Interact 11 1136–1141 [DOI] [PubMed] [Google Scholar]
- Nam J, Mysore KS, Zheng C, Knue MK, Matthysse AG, Gelvin SB (1999) Identification of T-DNA tagged Arabidopsis mutants that are resistant to transformation by Agrobacterium. Mol Gen Genet 261 429–438 [DOI] [PubMed] [Google Scholar]
- Narasimhulu SB, Deng XB, Sarria R, Gelvin SB (1996) Early transcription of Agrobacterium T-DNA genes in tobacco and maize. Plant Cell 8 873–886 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni M, Cui D, Einstein J, Narasimhulu S, Vergara CE, Gelvin SB (1995) Strength and tissue specificity of chimeric promoters derived from the octopine and mannopine synthase genes. Plant J 7 661–676 [Google Scholar]
- Restrepo MA, Freed DD, Carrington JC (1990) Nuclear transport of plant potyviral proteins. Plant Cell 2 987–998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vancanneyt G, Schmidt R, O'Connor-Sanchez A, Willmitzer L, Rocha-Sosa M (1990) Construction of an intron-containing marker gene: splicing of the intron in transgenic plants and its use in monitoring early events in Agrobacterium-mediated plant transformation. Mol Gen Genet 220 245–250 [DOI] [PubMed] [Google Scholar]
- Veena, Jiang H, Doerge RW, Gelvin SB (2003) Transfer of T-DNA and Vir proteins to plant cells by Agrobacterium tumefaciens induces expression of host genes involved in mediating transformation and suppresses host defense gene expression. Plant J 35 219–236 [DOI] [PubMed] [Google Scholar]
- Vieira J, Messing J (1987) Production of single-stranded plasmid DNA. Methods Enzymol 153 3–11 [DOI] [PubMed] [Google Scholar]
- Williamson JD, Hirsch-Wyncott ME, Larkins BA, Gelvin SB (1989) Differential accumulation of a transcript driven by the CaMV 35S promoter in transgenic tobacco. Plant Physiol 90 1570–1576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi H, Mysore KS, Gelvin S (2002) Expression of the Arabidopsis histone H2A-1 gene correlates with susceptibility to Agrobacterium transformation. Plant J 32 285–298 [DOI] [PubMed] [Google Scholar]
- Zhu Y, Nam J, Humara JM, Mysore KS, Lee LY, Cao H, Valentine L, Li J, Kaiser AD, Kopecky AL, et al (2003) Identification of Arabidopsis rat mutants. Plant Physiol 132 494–505 [DOI] [PMC free article] [PubMed] [Google Scholar]