Abstract
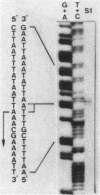
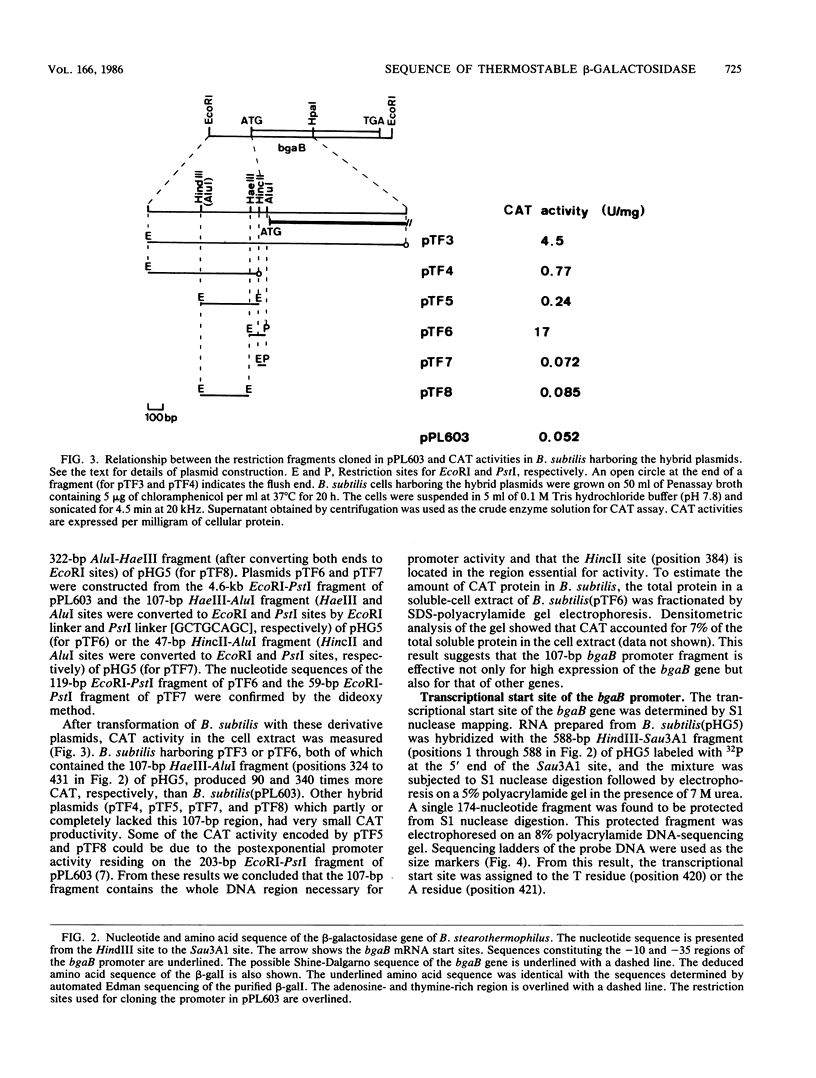
The nucleotide sequence of the bgaB gene, which encodes the thermostable beta-galactosidase I of Bacillus stearothermophilus, and its flanking region was determined. A 2,016-base-pair open reading frame observed was concluded to be for beta-galactosidase I (Mr 78,051) from observations that the amino acid composition of the enzyme and the sequence of 14 amino acids from the amino-terminus of the enzyme coincided with those deduced from this open frame. A 107-base-pair HaeIII-AluI fragment just upstream of the estimated Shine-Dalgarno sequence of the bgaB gene had promoter activity toward cat-86 (chloramphenicol acetyltransferase gene) and produced the enzyme at a level equivalent to 7% of the total cellular protein of B. subtilis. From the base sequence of this DNA region and the transcriptional start site determined by S1 nuclease mapping, the -35 and -10 sequences are estimated to be TTGACA and TAATTT, respectively, which are similar to the consensus sequence of B. subtilis sigma 43 RNA polymerase.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- Breunig K. D., Dahlems U., Das S., Hollenberg C. P. Analysis of a eukaryotic beta-galactosidase gene: the N-terminal end of the yeast Kluyveromyces lactis protein shows homology to the Escherichia coli lacZ gene product. Nucleic Acids Res. 1984 Mar 12;12(5):2327–2341. doi: 10.1093/nar/12.5.2327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brosius J., Cate R. L., Perlmutter A. P. Precise location of two promoters for the beta-lactamase gene of pBR322. S1 mapping of ribonucleic acid isolated from Escherichia coli or synthesized in vitro. J Biol Chem. 1982 Aug 10;257(15):9205–9210. [PubMed] [Google Scholar]
- Buvinger W. E., Riley M. Nucleotide sequence of Klebsiella pneumoniae lac genes. J Bacteriol. 1985 Sep;163(3):850–857. doi: 10.1128/jb.163.3.850-857.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Béguin P., Cornet P., Aubert J. P. Sequence of a cellulase gene of the thermophilic bacterium Clostridium thermocellum. J Bacteriol. 1985 Apr;162(1):102–105. doi: 10.1128/jb.162.1.102-105.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang S., Cohen S. N. High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet. 1979 Jan 5;168(1):111–115. doi: 10.1007/BF00267940. [DOI] [PubMed] [Google Scholar]
- Duvall E. J., Williams D. M., Mongkolsuk S., Lovett P. S. Regulatory regions that control expression of two chloramphenicol-inducible cat genes cloned in Bacillus subtilis. J Bacteriol. 1984 Jun;158(3):784–790. doi: 10.1128/jb.158.3.784-790.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fowler A. V., Smith P. J. The active site regions of lacZ and ebg beta-galactosidases are homologous. J Biol Chem. 1983 Sep 10;258(17):10204–10207. [PubMed] [Google Scholar]
- Gilman M. Z., Chamberlin M. J. Developmental and genetic regulation of Bacillus subtilis genes transcribed by sigma 28-RNA polymerase. Cell. 1983 Nov;35(1):285–293. doi: 10.1016/0092-8674(83)90231-3. [DOI] [PubMed] [Google Scholar]
- Herrchen M., Legler G. Identification of an essential carboxylate group at the active site of lacZ beta-galactosidase from Escherichia coli. Eur J Biochem. 1984 Feb 1;138(3):527–531. doi: 10.1111/j.1432-1033.1984.tb07947.x. [DOI] [PubMed] [Google Scholar]
- Hirata H., Negoro S., Okada H. High Production of Thermostable beta-Galactosidase of Bacillus stearothermophilus in Bacillus subtilis. Appl Environ Microbiol. 1985 Jun;49(6):1547–1549. doi: 10.1128/aem.49.6.1547-1549.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirata H., Negoro S., Okada H. Molecular basis of isozyme formation of beta-galactosidases in Bacillus stearothermophilus: isolation of two beta-galactosidase genes, bgaA and bgaB. J Bacteriol. 1984 Oct;160(1):9–14. doi: 10.1128/jb.160.1.9-14.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunkapiller M. W., Hewick R. M., Dreyer W. J., Hood L. E. High-sensitivity sequencing with a gas-phase sequenator. Methods Enzymol. 1983;91:399–413. doi: 10.1016/s0076-6879(83)91038-8. [DOI] [PubMed] [Google Scholar]
- Ikai A. Thermostability and aliphatic index of globular proteins. J Biochem. 1980 Dec;88(6):1895–1898. [PubMed] [Google Scholar]
- Kalnins A., Otto K., Rüther U., Müller-Hill B. Sequence of the lacZ gene of Escherichia coli. EMBO J. 1983;2(4):593–597. doi: 10.1002/j.1460-2075.1983.tb01468.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- McLaughlin J. R., Murray C. L., Rabinowitz J. C. Unique features in the ribosome binding site sequence of the gram-positive Staphylococcus aureus beta-lactamase gene. J Biol Chem. 1981 Nov 10;256(21):11283–11291. [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Moran C. P., Jr, Lang N., LeGrice S. F., Lee G., Stephens M., Sonenshein A. L., Pero J., Losick R. Nucleotide sequences that signal the initiation of transcription and translation in Bacillus subtilis. Mol Gen Genet. 1982;186(3):339–346. doi: 10.1007/BF00729452. [DOI] [PubMed] [Google Scholar]
- Murray C. L., Rabinowitz J. C. Nucleotide sequences of transcription and translation initiation regions in Bacillus phage phi 29 early genes. J Biol Chem. 1982 Jan 25;257(2):1053–1062. [PubMed] [Google Scholar]
- Negoro S., Taniguchi T., Kanaoka M., Kimura H., Okada H. Plasmid-determined enzymatic degradation of nylon oligomers. J Bacteriol. 1983 Jul;155(1):22–31. doi: 10.1128/jb.155.1.22-31.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piggot P. J., Hoch J. A. Revised genetic linkage map of Bacillus subtilis. Microbiol Rev. 1985 Jun;49(2):158–179. doi: 10.1128/mr.49.2.158-179.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw W. V. Chloramphenicol acetyltransferase from chloramphenicol-resistant bacteria. Methods Enzymol. 1975;43:737–755. doi: 10.1016/0076-6879(75)43141-x. [DOI] [PubMed] [Google Scholar]
- Sprague K. U., Steitz J. A., Grenley R. M., Stocking C. E. 3' terminal sequences of 16S rRNA do not explain translational specificity differences between E. coli and B. stearothermophilus ribosomes. Nature. 1977 Jun 2;267(5610):462–465. doi: 10.1038/267462a0. [DOI] [PubMed] [Google Scholar]
- Stokes H. W., Betts P. W., Hall B. G. Sequence of the ebgA gene of Escherichia coli: comparison with the lacZ gene. Mol Biol Evol. 1985 Nov;2(6):469–477. doi: 10.1093/oxfordjournals.molbev.a040372. [DOI] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Tsunasawa S., Kondo J., Sakiyama F. Isocratic separation of PTH-amino acids at picomole level by reverse-phase HPLC in the presence of sodium dodecylsulfate. J Biochem. 1985 Feb;97(2):701–704. doi: 10.1093/oxfordjournals.jbchem.a135108. [DOI] [PubMed] [Google Scholar]
- Williams D. M., Duvall E. J., Lovett P. S. Cloning restriction fragments that promote expression of a gene in Bacillus subtilis. J Bacteriol. 1981 Jun;146(3):1162–1165. doi: 10.1128/jb.146.3.1162-1165.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]