Abstract
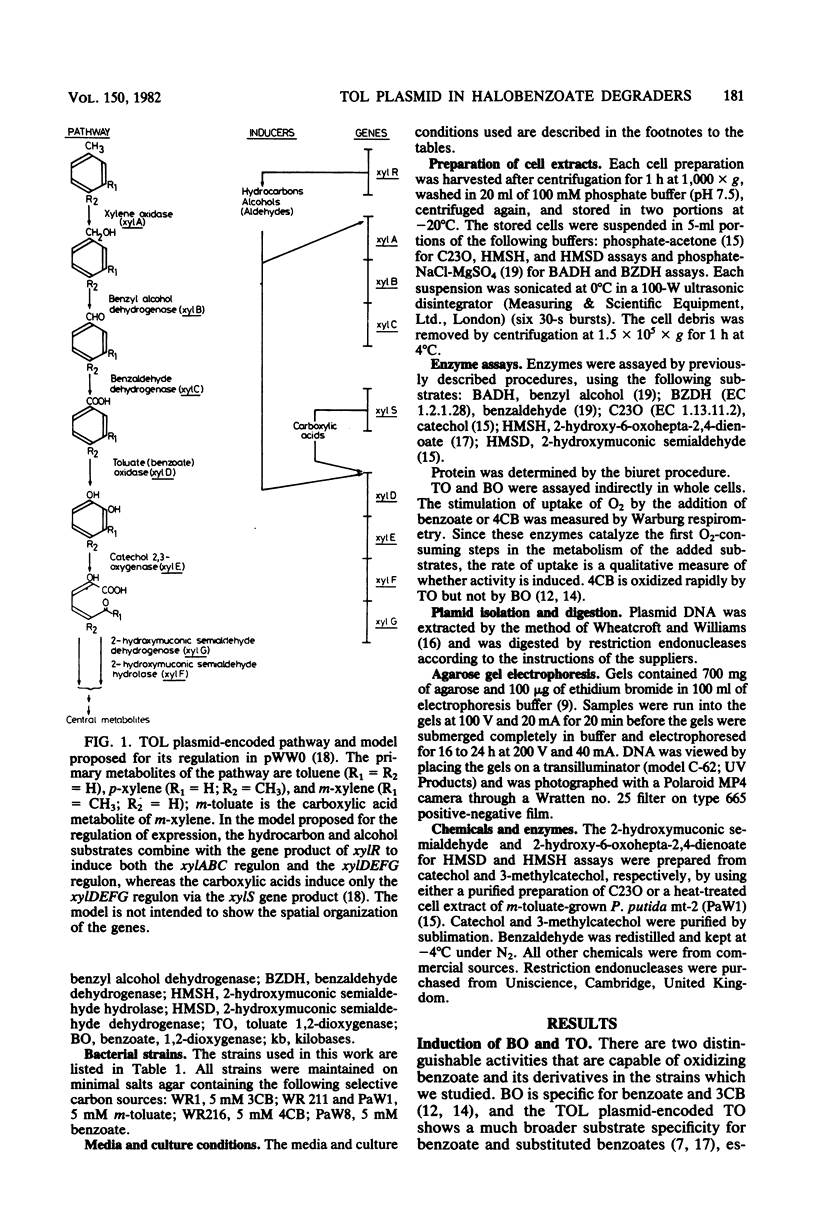
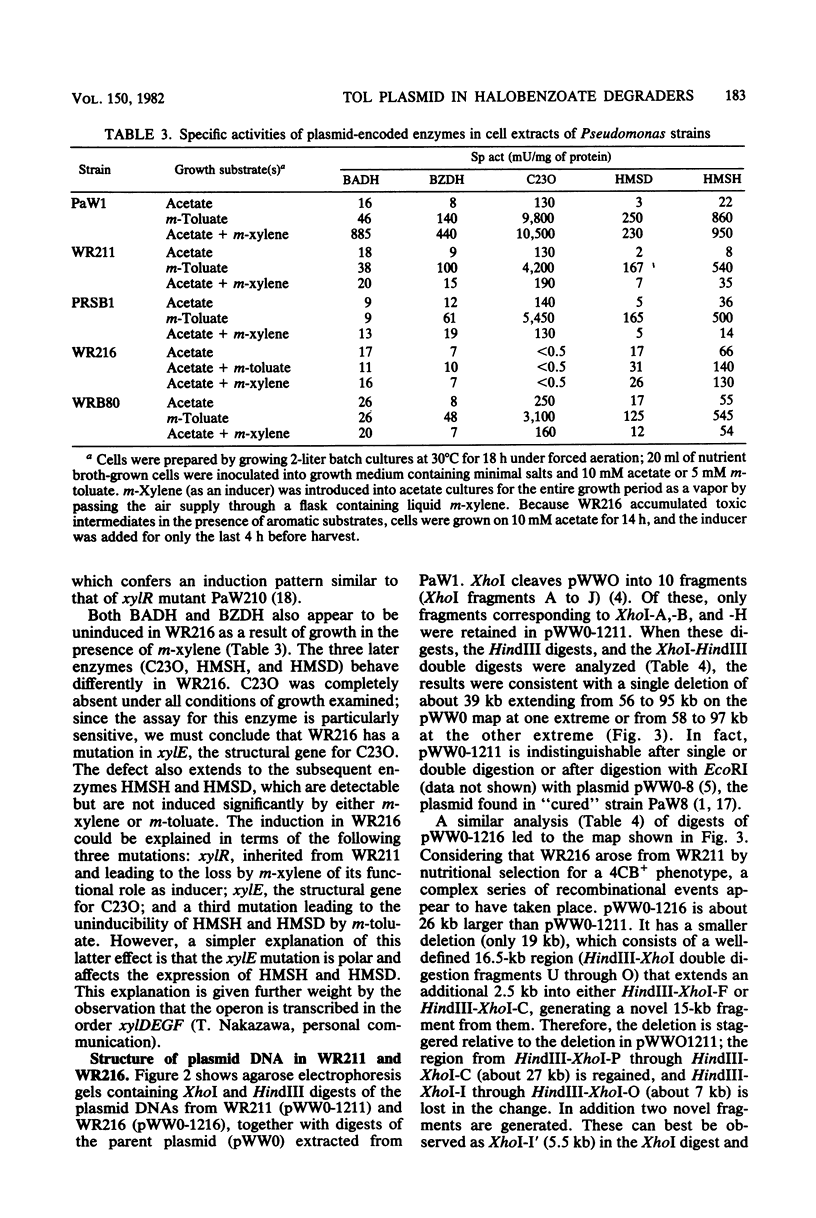
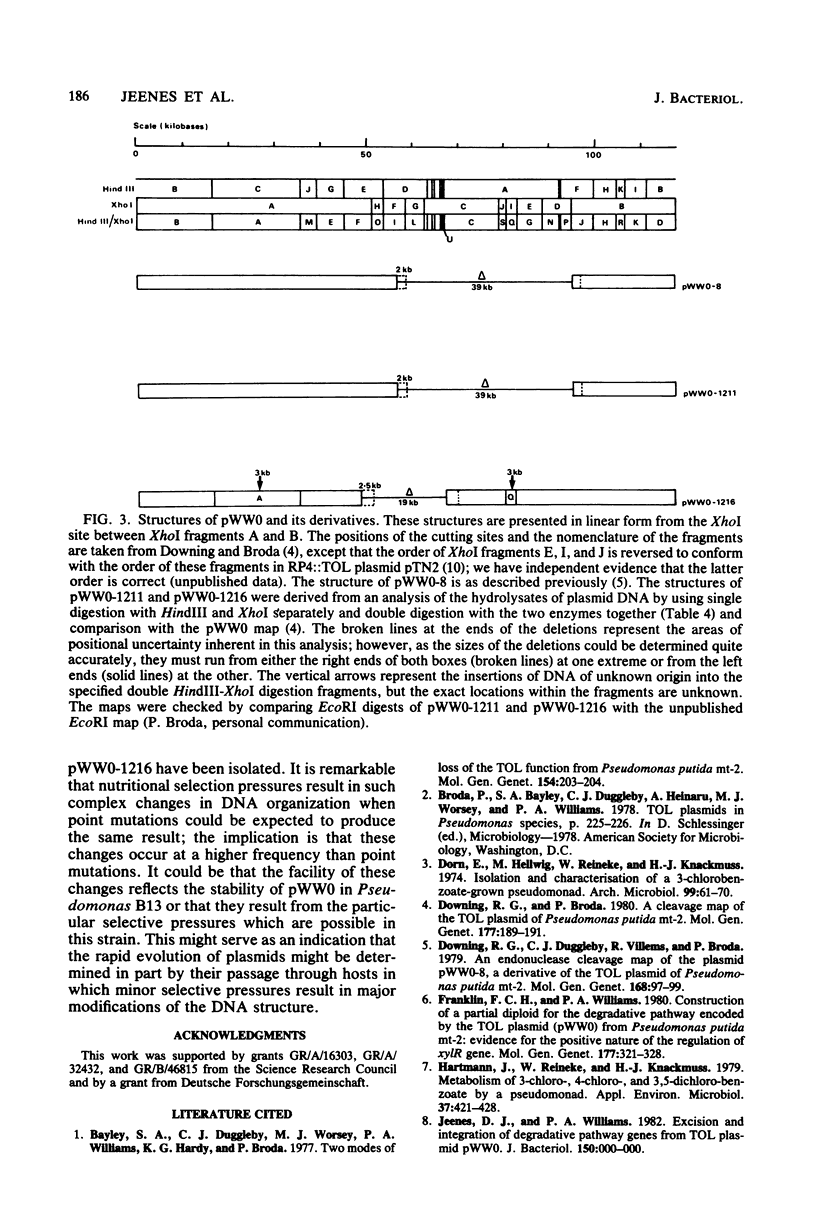
WR211 and WR216 are derivatives of halobenzoate-degrading Pseudomonas sp. strain B13 into which the 117-kilobase TOL degradative plasmid pWW0 has been transferred from Pseudomonas putida mt-2. WR211 has lost the ability to grow on the TOL-specific substrate m-xylene but retains the ability to grow on its metabolite, m-toluate. An analysis of the induction of enzymes was consistent with WR211 carrying a nonfunctional regulatory gene, xy1R, WR216 is a spontaneous derivative of WR211 which grows on one of the TOL substrates and yet expresses the nonspecific toluate oxidase, which enables it to grow on the novel substrate 4-chlorobenzoate. In addition to the xy1R lesion inherited from WR211, WR216 appears to carry a mutation in the structural gene for catechol 2,3-oxygenase, xy1E. The plasmids in both strains were analyzed by restriction endonuclease digestion. pWW0-1211 in WR211 has a large deletion (39 kilobases) compared with pWW0 and appears to be identical to a previously described plasmid (pWW0-8) which encodes none of the TOL degradative functions. pWW0-1216 in WR216 has undergone a major structural reorganization relative to its parent, pWW0-1211. This plasmid has a smaller deletion (19 kilobases), which is staggered relative to the deletion in pWW0-1211, and in addition it has two 3-kilobase insertions of unknown origin, one of which appears to cause the xylE mutation.
Full text
PDF



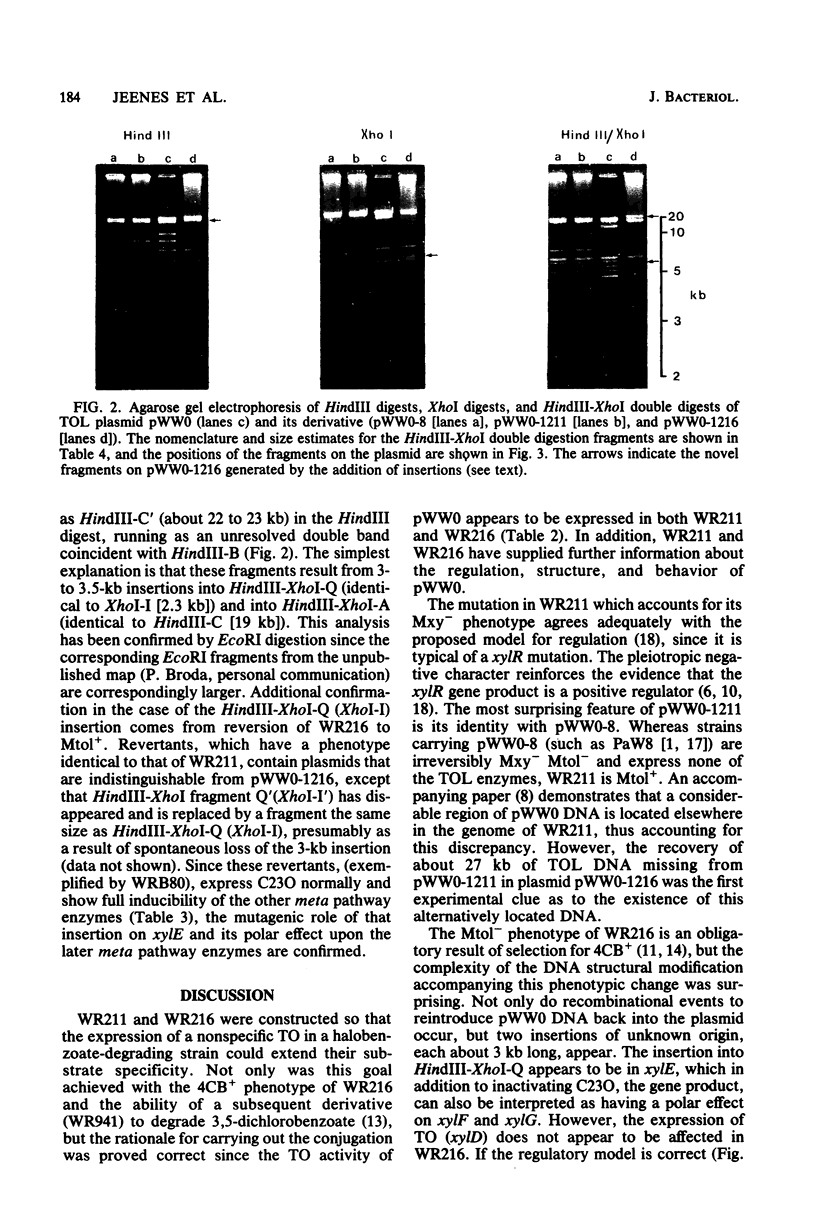



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bayley S. A., Duggleby C. J., Worsey M. J., Williams P. A., Hardy K. G., Broda P. Two modes of loss of the Tol function from Pseudomonas putida mt-2. Mol Gen Genet. 1977 Jul 20;154(2):203–204. doi: 10.1007/BF00330838. [DOI] [PubMed] [Google Scholar]
- Dorn E., Hellwig M., Reineke W., Knackmuss H. J. Isolation and characterization of a 3-chlorobenzoate degrading pseudomonad. Arch Microbiol. 1974;99(1):61–70. doi: 10.1007/BF00696222. [DOI] [PubMed] [Google Scholar]
- Downing R. G., Duggleby C. J., Villems R., Broda P. An endonuclease cleavage map of the plasmid pWWO-8, a derivative of the TOL plasmid of Pseudomonas putida mt-2. Mol Gen Genet. 1979 Jan 5;168(1):97–99. doi: 10.1007/BF00267938. [DOI] [PubMed] [Google Scholar]
- Downing R., Broda P. A cleavage map of the TOL plasmid of Pseudomonas putida mt-2. Mol Gen Genet. 1979;177(1):189–191. doi: 10.1007/BF00267270. [DOI] [PubMed] [Google Scholar]
- Franklin F. C., Williams P. A. Construction of a partial diploid for the degradative pathway encoded by the TOL plasmid (pWWO) from Pseudomonas putida mt-2: evidence for the positive nature of the regulation by the xyIR gene. Mol Gen Genet. 1980 Jan;177(2):321–328. doi: 10.1007/BF00267445. [DOI] [PubMed] [Google Scholar]
- Hartmann J., Reineke W., Knackmuss H. J. Metabolism of 3-chloro-, 4-chloro-, and 3,5-dichlorobenzoate by a pseudomonad. Appl Environ Microbiol. 1979 Mar;37(3):421–428. doi: 10.1128/aem.37.3.421-428.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers J. A., Sanchez D., Elwell L. P., Falkow S. Simple agarose gel electrophoretic method for the identification and characterization of plasmid deoxyribonucleic acid. J Bacteriol. 1976 Sep;127(3):1529–1537. doi: 10.1128/jb.127.3.1529-1537.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakazawa T., Inouye S., Nakazawa A. Physical and functional mapping of RP4-TOL plasmid recombinants: analysis of insertion and deletion mutants. J Bacteriol. 1980 Oct;144(1):222–231. doi: 10.1128/jb.144.1.222-231.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reineke W., Knackmuss H. J. Chemical structure and biodegradability of halogenate aromatic compounds. Substituent effects on 1,2-dioxygenation of benzoic acid. Biochim Biophys Acta. 1978 Sep 6;542(3):412–423. doi: 10.1016/0304-4165(78)90372-0. [DOI] [PubMed] [Google Scholar]
- Reineke W., Knackmuss H. J. Construction of haloaromatics utilising bacteria. Nature. 1979 Feb 1;277(5695):385–386. doi: 10.1038/277385a0. [DOI] [PubMed] [Google Scholar]
- Reineke W., Knackmuss H. J. Hybrid pathway for chlorobenzoate metabolism in Pseudomonas sp. B13 derivatives. J Bacteriol. 1980 May;142(2):467–473. doi: 10.1128/jb.142.2.467-473.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sala-Trepat J. M., Evans W. C. The meta cleavage of catechol by Azotobacter species. 4-Oxalocrotonate pathway. Eur J Biochem. 1971 Jun 11;20(3):400–413. doi: 10.1111/j.1432-1033.1971.tb01406.x. [DOI] [PubMed] [Google Scholar]
- Wheatcroft R., Williams P. A. Rapid methods for the study of both stable and unstable plasmids in Pseudomonas. J Gen Microbiol. 1981 Jun;124(2):433–437. doi: 10.1099/00221287-124-2-433. [DOI] [PubMed] [Google Scholar]
- Williams P. A., Murray K. Metabolism of benzoate and the methylbenzoates by Pseudomonas putida (arvilla) mt-2: evidence for the existence of a TOL plasmid. J Bacteriol. 1974 Oct;120(1):416–423. doi: 10.1128/jb.120.1.416-423.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worsey M. J., Franklin F. C., Williams P. A. Regulation of the degradative pathway enzymes coded for by the TOL plasmid (pWWO) from Pseudomonas putida mt-2. J Bacteriol. 1978 Jun;134(3):757–764. doi: 10.1128/jb.134.3.757-764.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worsey M. J., Williams P. A. Metabolism of toluene and xylenes by Pseudomonas (putida (arvilla) mt-2: evidence for a new function of the TOL plasmid. J Bacteriol. 1975 Oct;124(1):7–13. doi: 10.1128/jb.124.1.7-13.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]



