Abstract
Insertion of the transposable deoxyribonucleic acid sequence that specifies the TEM beta-lactamase (TnA) occurred in at least 19 sites on the 5.5 x 10(6)-dalton plasmid RSF1010. There was no significant difference in the frequency of transposition or in the distribution of TnA insertion sites for recombinant plasmids isolated from recombination-proficient (rec+) or recombination-deficient (rec-) bacterial host cells. The site and orientation of TnA insertions were determined by both heteroduplex analysis and enzymatic digestion with restriction endonucleases. Insertion in the gene encoding for sulfonamide resistance occurred without circular permutation in one or the other of two distinct orientations. Insertions in orientation P were strongly polar on distal gene expression, whereas insertions in orientation M were mutagenic but not polar. In addition, we have observed that TnA elements from different R plasmids show fine structural heterogeneity, and that TnA insertion at a site adjacent to the origin of replication causes an increase in plasmid copy number.
Full text
PDF





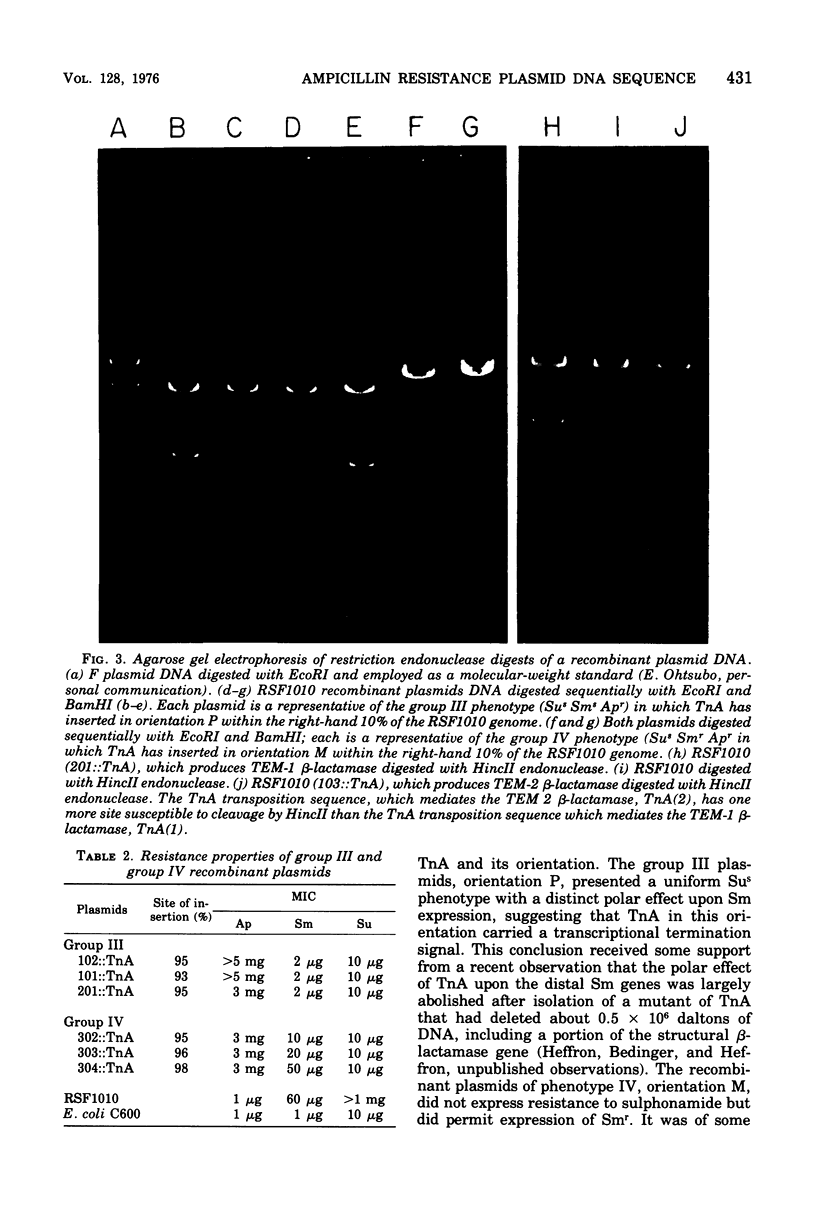



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bachmann B. J. Pedigrees of some mutant strains of Escherichia coli K-12. Bacteriol Rev. 1972 Dec;36(4):525–557. doi: 10.1128/br.36.4.525-557.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barth P. T., Grinter N. J. Comparison of the deoxyribonucleic acid molecular weights and homologies of plasmids conferring linked resistance to streptomycin and sulfonamides. J Bacteriol. 1974 Nov;120(2):618–630. doi: 10.1128/jb.120.2.618-630.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennett P. M., Richmond M. H. Translocation of a discrete piece of deoxyribonucleic acid carrying an amp gene between replicons in Eschericha coli. J Bacteriol. 1976 Apr;126(1):1–6. doi: 10.1128/jb.126.1.1-6.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg D. E., Davies J., Allet B., Rochaix J. D. Transposition of R factor genes to bacteriophage lambda. Proc Natl Acad Sci U S A. 1975 Sep;72(9):3628–3632. doi: 10.1073/pnas.72.9.3628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B. Nature of Col E 1 plasmid replication in Escherichia coli in the presence of the chloramphenicol. J Bacteriol. 1972 May;110(2):667–676. doi: 10.1128/jb.110.2.667-676.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elwell L. P., De Graaff J., Seibert D., Falkow S. Plasmid-linked ampicillin resistance in haempohilus influenza type b. Infect Immun. 1975 Aug;12(2):404–410. doi: 10.1128/iai.12.2.404-410.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottesman M. M., Rosner J. L. Acquisition of a determinant for chloramphenicol resistance by coliphage lambda. Proc Natl Acad Sci U S A. 1975 Dec;72(12):5041–5045. doi: 10.1073/pnas.72.12.5041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedges R. W., Jacob A. E., Datta N., Coetzee J. N. Properties of plasmids produced by recombination between R factors of groups J and FII. Mol Gen Genet. 1975 Oct 22;140(4):289–302. doi: 10.1007/BF00267320. [DOI] [PubMed] [Google Scholar]
- Hedges R. W., Jacob A. E. Transposition of ampicillin resistance from RP4 to other replicons. Mol Gen Genet. 1974;132(1):31–40. doi: 10.1007/BF00268228. [DOI] [PubMed] [Google Scholar]
- Heffron F., Rubens C., Falkow S. Translocation of a plasmid DNA sequence which mediates ampicillin resistance: molecular nature and specificity of insertion. Proc Natl Acad Sci U S A. 1975 Sep;72(9):3623–3627. doi: 10.1073/pnas.72.9.3623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heffron F., Sublett R., Hedges R. W., Jacob A., Falkow S. Origin of the TEM-beta-lactamase gene found on plasmids. J Bacteriol. 1975 Apr;122(1):250–256. doi: 10.1128/jb.122.1.250-256.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirsch H. J., Starlinger P., Brachet P. Two kinds of insertions in bacterial genes. Mol Gen Genet. 1972;119(3):191–206. doi: 10.1007/BF00333858. [DOI] [PubMed] [Google Scholar]
- Hu S., Otsubo E., Davidson N., Saedler H. Electron microscope heteroduplex studies of sequence relations among bacterial plasmids: identification and mapping of the insertion sequences IS1 and IS2 in F and R plasmids. J Bacteriol. 1975 May;122(2):764–775. doi: 10.1128/jb.122.2.764-775.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan E., Saedler H., Starlinger P. O0 and strong-polar mutations in the gal operon are insertions. Mol Gen Genet. 1968;102(4):353–363. doi: 10.1007/BF00433726. [DOI] [PubMed] [Google Scholar]
- Kleckner N., Chan R. K., Tye B. K., Botstein D. Mutagenesis by insertion of a drug-resistance element carrying an inverted repetition. J Mol Biol. 1975 Oct 5;97(4):561–575. doi: 10.1016/s0022-2836(75)80059-3. [DOI] [PubMed] [Google Scholar]
- Kopecko D. J., Cohen S. N. Site specific recA--independent recombination between bacterial plasmids: involvement of palindromes at the recombinational loci. Proc Natl Acad Sci U S A. 1975 Apr;72(4):1373–1377. doi: 10.1073/pnas.72.4.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai C. J., Nathans D. Deletion mutants of simian virus 40 generated by enzymatic excision of DNA segments from the viral genome. J Mol Biol. 1974 Oct 15;89(1):179–193. doi: 10.1016/0022-2836(74)90169-7. [DOI] [PubMed] [Google Scholar]
- Landy A., Ruedisueli E., Robinson L., Foeller C., Ross W. Digestion of deoxyribonucleic acids from bacteriophage T7, lambda, and phi 80h with site-specific nucleases from Hemophilus influenzae strain Rc and strain Rd. Biochemistry. 1974 May 7;13(10):2134–2142. doi: 10.1021/bi00707a022. [DOI] [PubMed] [Google Scholar]
- Lee H. J., Otsubo E., Deonier R. C., Davidson N. Electron microscope heteroduplex studies of sequence relations among plasmids of Escherichia coli. V. ilv+ Deletion mutants of F14. J Mol Biol. 1974 Nov 15;89(4):585–597. doi: 10.1016/0022-2836(74)90037-0. [DOI] [PubMed] [Google Scholar]
- Malamy M. H., Fiandt M., Szybalski W. Electron microscopy of polar insertions in the lac operon of Escherichia coli. Mol Gen Genet. 1972;119(3):207–222. doi: 10.1007/BF00333859. [DOI] [PubMed] [Google Scholar]
- Matthew M., Hedges R. W. Analytical isoelectric focusing of R factor-determined beta-lactamases: correlation with plasmid compatibility. J Bacteriol. 1976 Feb;125(2):713–718. doi: 10.1128/jb.125.2.713-718.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meynell E., Datta N. Mutant drug resistant factors of high transmissibility. Nature. 1967 May 27;214(5091):885–887. doi: 10.1038/214885a0. [DOI] [PubMed] [Google Scholar]
- Ptashne K., Cohen S. N. Occurrence of insertion sequence (IS) regions on plasmid deoxyribonucleic acid as direct and inverted nucleotide sequence duplications. J Bacteriol. 1975 May;122(2):776–781. doi: 10.1128/jb.122.2.776-781.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reif H. J., Saedler H. IS1 is involved in deletion formation in the gal region of E. coli K12. Mol Gen Genet. 1975;137(1):17–28. doi: 10.1007/BF00332538. [DOI] [PubMed] [Google Scholar]
- Ruby C., Novick R. P. Plasmid interactions in Staphylococcus aureus: nonadditivity of compatible plasmid DNA pools. Proc Natl Acad Sci U S A. 1975 Dec;72(12):5031–5035. doi: 10.1073/pnas.72.12.5031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saedler H., Reif H. J., Hu S., Davidson N. IS2, a genetic element for turn-off and turn-on of gene activity in E. coli. Mol Gen Genet. 1974;132(4):265–289. doi: 10.1007/BF00268569. [DOI] [PubMed] [Google Scholar]
- Shapiro J. A. Mutations caused by the insertion of genetic material into the galactose operon of Escherichia coli. J Mol Biol. 1969 Feb 28;40(1):93–105. doi: 10.1016/0022-2836(69)90298-8. [DOI] [PubMed] [Google Scholar]
- Sharp P. A., Hsu M. T., Otsubo E., Davidson N. Electron microscope heteroduplex studies of sequence relations among plasmids of Escherichia coli. I. Structure of F-prime factors. J Mol Biol. 1972 Nov 14;71(2):471–497. doi: 10.1016/0022-2836(72)90363-4. [DOI] [PubMed] [Google Scholar]
- So M., Gill R., Falkow S. The generation of a ColE1-Apr cloning vehicle which allows detection of inserted DNA. Mol Gen Genet. 1975 Dec 30;142(3):239–249. doi: 10.1007/BF00425649. [DOI] [PubMed] [Google Scholar]
- Wilson G. A., Young F. E. Isolation of a sequence-specific endonuclease (BamI) from Bacillus amyloliquefaciens H. J Mol Biol. 1975 Sep 5;97(1):123–125. doi: 10.1016/s0022-2836(75)80028-3. [DOI] [PubMed] [Google Scholar]




