Abstract
Rates of transcriptions of the human immunodeficiency virus are greatly increased by the viral trans activator Tat. In vitro, Tat binds to the 5' bulge of the trans-activation response (TAR) RNA stem-loop, which is present in all viral transcripts. In human cells, the central loop in TAR and its cellular RNA-binding proteins are also critical for the function of Tat. Previously, we demonstrated that in rodent cells (CHO cells), but not in those which contain the human chromosome 12 (CHO12 cells), Tat-TAR interactions are compromised. In this study, we examined the roles of the bulge and loop in TAR in Tat trans activation in these cells. Whereas low levels of trans activation depended solely on interactions between Tat and the bulge in CHO cells, high levels of trans activation depended also on interactions between Tat and the loop in CHO12 cells. Since the TAR loop binding proteins in these two cell lines were identical and different from their human counterpart, the human chromosome 12 does not encode TAR loop binding proteins. In vivo binding competition studies with TAR decoys confirmed that the binding of Tat to TAR is more efficient in CHO12 cells. Thus, the protein(s) encoded on human chromosome 12 helps to tether Tat to TAR via its loop, which results in high levels of trans activation.
Full text
PDF

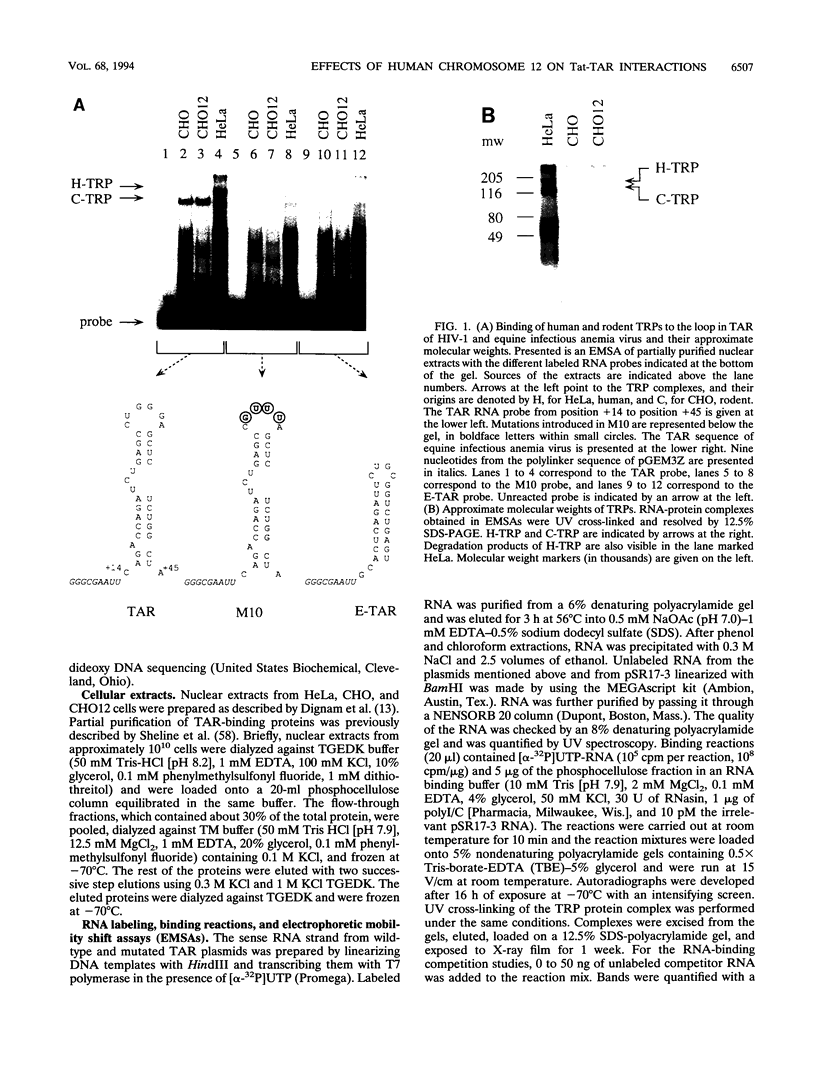


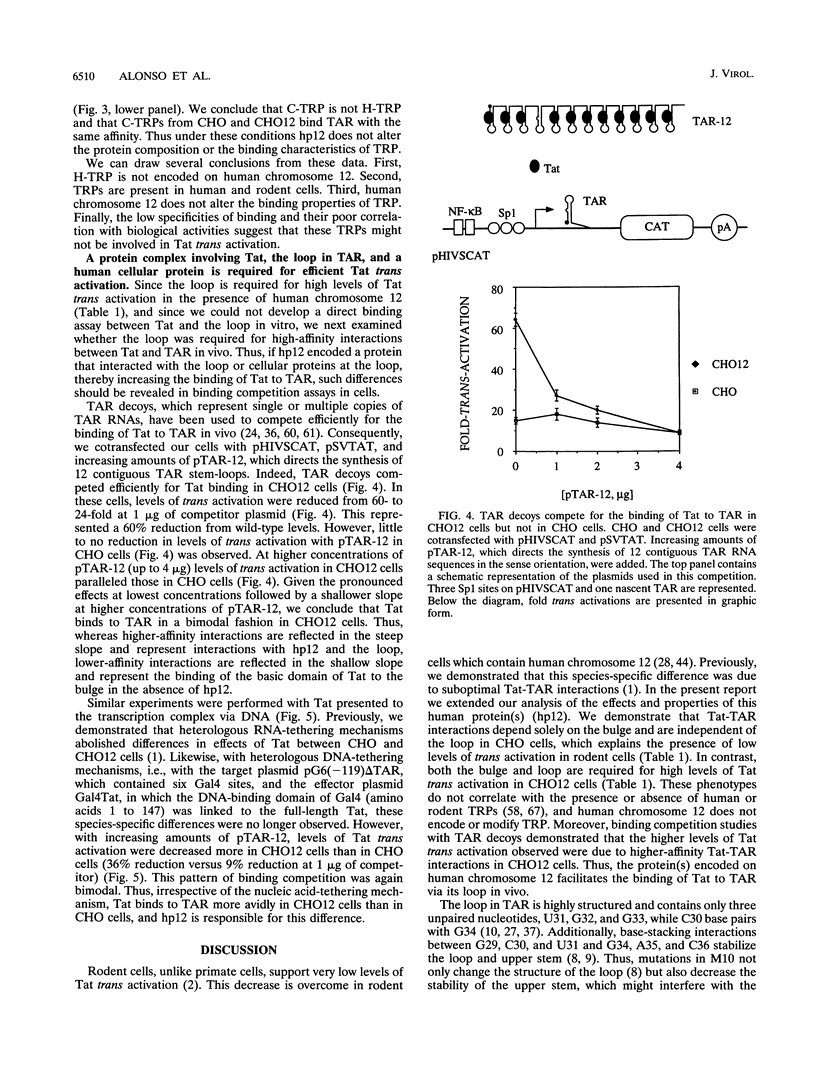



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alonso A., Derse D., Peterlin B. M. Human chromosome 12 is required for optimal interactions between Tat and TAR of human immunodeficiency virus type 1 in rodent cells. J Virol. 1992 Jul;66(7):4617–4621. doi: 10.1128/jvi.66.7.4617-4621.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barry P. A., Pratt-Lowe E., Unger R. E., Luciw P. A. Cellular factors regulate transactivation of human immunodeficiency virus type 1. J Virol. 1991 Mar;65(3):1392–1399. doi: 10.1128/jvi.65.3.1392-1399.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berkhout B., Jeang K. T. Detailed mutational analysis of TAR RNA: critical spacing between the bulge and loop recognition domains. Nucleic Acids Res. 1991 Nov 25;19(22):6169–6176. doi: 10.1093/nar/19.22.6169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berkhout B., Jeang K. T. trans activation of human immunodeficiency virus type 1 is sequence specific for both the single-stranded bulge and loop of the trans-acting-responsive hairpin: a quantitative analysis. J Virol. 1989 Dec;63(12):5501–5504. doi: 10.1128/jvi.63.12.5501-5504.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calnan B. J., Biancalana S., Hudson D., Frankel A. D. Analysis of arginine-rich peptides from the HIV Tat protein reveals unusual features of RNA-protein recognition. Genes Dev. 1991 Feb;5(2):201–210. doi: 10.1101/gad.5.2.201. [DOI] [PubMed] [Google Scholar]
- Calnan B. J., Tidor B., Biancalana S., Hudson D., Frankel A. D. Arginine-mediated RNA recognition: the arginine fork. Science. 1991 May 24;252(5009):1167–1171. doi: 10.1126/science.252.5009.1167. [DOI] [PubMed] [Google Scholar]
- Churcher M. J., Lamont C., Hamy F., Dingwall C., Green S. M., Lowe A. D., Butler J. G., Gait M. J., Karn J. High affinity binding of TAR RNA by the human immunodeficiency virus type-1 tat protein requires base-pairs in the RNA stem and amino acid residues flanking the basic region. J Mol Biol. 1993 Mar 5;230(1):90–110. doi: 10.1006/jmbi.1993.1128. [DOI] [PubMed] [Google Scholar]
- Colvin R. A., Garcia-Blanco M. A. Unusual structure of the human immunodeficiency virus type 1 trans-activation response element. J Virol. 1992 Feb;66(2):930–935. doi: 10.1128/jvi.66.2.930-935.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Critchley A. D., Haneef I., Cousens D. J., Stockley P. G. Modeling and solution structure probing of the HIV-1 TAR stem-loop. J Mol Graph. 1993 Jun;11(2):92-7, 124. doi: 10.1016/0263-7855(93)87002-m. [DOI] [PubMed] [Google Scholar]
- Delling U., Reid L. S., Barnett R. W., Ma M. Y., Climie S., Sumner-Smith M., Sonenberg N. Conserved nucleotides in the TAR RNA stem of human immunodeficiency virus type 1 are critical for Tat binding and trans activation: model for TAR RNA tertiary structure. J Virol. 1992 May;66(5):3018–3025. doi: 10.1128/jvi.66.5.3018-3025.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Derse D., Carvalho M., Carroll R., Peterlin B. M. A minimal lentivirus Tat. J Virol. 1991 Dec;65(12):7012–7015. doi: 10.1128/jvi.65.12.7012-7015.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dignam J. D., Lebovitz R. M., Roeder R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 1983 Mar 11;11(5):1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall C., Ernberg I., Gait M. J., Green S. M., Heaphy S., Karn J., Lowe A. D., Singh M., Skinner M. A. HIV-1 tat protein stimulates transcription by binding to a U-rich bulge in the stem of the TAR RNA structure. EMBO J. 1990 Dec;9(12):4145–4153. doi: 10.1002/j.1460-2075.1990.tb07637.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Endo S., Kubota S., Siomi H., Adachi A., Oroszlan S., Maki M., Hatanaka M. A region of basic amino-acid cluster in HIV-1 Tat protein is essential for trans-acting activity and nucleolar localization. Virus Genes. 1989 Nov;3(2):99–110. doi: 10.1007/BF00125123. [DOI] [PubMed] [Google Scholar]
- Feinberg M. B., Baltimore D., Frankel A. D. The role of Tat in the human immunodeficiency virus life cycle indicates a primary effect on transcriptional elongation. Proc Natl Acad Sci U S A. 1991 May 1;88(9):4045–4049. doi: 10.1073/pnas.88.9.4045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng S., Holland E. C. HIV-1 tat trans-activation requires the loop sequence within tar. Nature. 1988 Jul 14;334(6178):165–167. doi: 10.1038/334165a0. [DOI] [PubMed] [Google Scholar]
- Fiddes J. C., Goodman H. M. The cDNA for the beta-subunit of human chorionic gonadotropin suggests evolution of a gene by readthrough into the 3'-untranslated region. Nature. 1980 Aug 14;286(5774):684–687. doi: 10.1038/286684a0. [DOI] [PubMed] [Google Scholar]
- Garcia J. A., Harrich D., Pearson L., Mitsuyasu R., Gaynor R. B. Functional domains required for tat-induced transcriptional activation of the HIV-1 long terminal repeat. EMBO J. 1988 Oct;7(10):3143–3147. doi: 10.1002/j.1460-2075.1988.tb03181.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia J. A., Harrich D., Soultanakis E., Wu F., Mitsuyasu R., Gaynor R. B. Human immunodeficiency virus type 1 LTR TATA and TAR region sequences required for transcriptional regulation. EMBO J. 1989 Mar;8(3):765–778. doi: 10.1002/j.1460-2075.1989.tb03437.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gatignol A., Buckler-White A., Berkhout B., Jeang K. T. Characterization of a human TAR RNA-binding protein that activates the HIV-1 LTR. Science. 1991 Mar 29;251(5001):1597–1600. doi: 10.1126/science.2011739. [DOI] [PubMed] [Google Scholar]
- Gatignol A., Kumar A., Rabson A., Jeang K. T. Identification of cellular proteins that bind to the human immunodeficiency virus type 1 trans-activation-responsive TAR element RNA. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7828–7832. doi: 10.1073/pnas.86.20.7828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaynor R., Soultanakis E., Kuwabara M., Garcia J., Sigman D. S. Specific binding of a HeLa cell nuclear protein to RNA sequences in the human immunodeficiency virus transactivating region. Proc Natl Acad Sci U S A. 1989 Jul;86(13):4858–4862. doi: 10.1073/pnas.86.13.4858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham G. J., Maio J. J. RNA transcripts of the human immunodeficiency virus transactivation response element can inhibit action of the viral transactivator. Proc Natl Acad Sci U S A. 1990 Aug;87(15):5817–5821. doi: 10.1073/pnas.87.15.5817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gunning P., Leavitt J., Muscat G., Ng S. Y., Kedes L. A human beta-actin expression vector system directs high-level accumulation of antisense transcripts. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4831–4835. doi: 10.1073/pnas.84.14.4831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harper J. W., Logsdon N. J. Refolded HIV-1 tat protein protects both bulge and loop nucleotides in TAR RNA from ribonucleolytic cleavage. Biochemistry. 1991 Aug 13;30(32):8060–8066. doi: 10.1021/bi00246a026. [DOI] [PubMed] [Google Scholar]
- Hart C. E., Galphin J. C., Westhafer M. A., Schochetman G. TAR loop-dependent human immunodeficiency virus trans activation requires factors encoded on human chromosome 12. J Virol. 1993 Aug;67(8):5020–5024. doi: 10.1128/jvi.67.8.5020-5024.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hart C. E., Ou C. Y., Galphin J. C., Moore J., Bacheler L. T., Wasmuth J. J., Petteway S. R., Jr, Schochetman G. Human chromosome 12 is required for elevated HIV-1 expression in human-hamster hybrid cells. Science. 1989 Oct 27;246(4929):488–491. doi: 10.1126/science.2683071. [DOI] [PubMed] [Google Scholar]
- Hauber J., Cullen B. R. Mutational analysis of the trans-activation-responsive region of the human immunodeficiency virus type I long terminal repeat. J Virol. 1988 Mar;62(3):673–679. doi: 10.1128/jvi.62.3.673-679.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hauber J., Perkins A., Heimer E. P., Cullen B. R. Trans-activation of human immunodeficiency virus gene expression is mediated by nuclear events. Proc Natl Acad Sci U S A. 1987 Sep;84(18):6364–6368. doi: 10.1073/pnas.84.18.6364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kao S. Y., Calman A. F., Luciw P. A., Peterlin B. M. Anti-termination of transcription within the long terminal repeat of HIV-1 by tat gene product. Nature. 1987 Dec 3;330(6147):489–493. doi: 10.1038/330489a0. [DOI] [PubMed] [Google Scholar]
- Kato H., Sumimoto H., Pognonec P., Chen C. H., Rosen C. A., Roeder R. G. HIV-1 Tat acts as a processivity factor in vitro in conjunction with cellular elongation factors. Genes Dev. 1992 Apr;6(4):655–666. doi: 10.1101/gad.6.4.655. [DOI] [PubMed] [Google Scholar]
- Kuppuswamy M., Subramanian T., Srinivasan A., Chinnadurai G. Multiple functional domains of Tat, the trans-activator of HIV-1, defined by mutational analysis. Nucleic Acids Res. 1989 May 11;17(9):3551–3561. doi: 10.1093/nar/17.9.3551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laspia M. F., Rice A. P., Mathews M. B. HIV-1 Tat protein increases transcriptional initiation and stabilizes elongation. Cell. 1989 Oct 20;59(2):283–292. doi: 10.1016/0092-8674(89)90290-0. [DOI] [PubMed] [Google Scholar]
- Laspia M. F., Rice A. P., Mathews M. B. Synergy between HIV-1 Tat and adenovirus E1A is principally due to stabilization of transcriptional elongation. Genes Dev. 1990 Dec;4(12B):2397–2408. doi: 10.1101/gad.4.12b.2397. [DOI] [PubMed] [Google Scholar]
- Lisziewicz J., Rappaport J., Dhar R. Tat-regulated production of multimerized TAR RNA inhibits HIV-1 gene expression. New Biol. 1991 Jan;3(1):82–89. [PubMed] [Google Scholar]
- Loret E. P., Georgel P., Johnson W. C., Jr, Ho P. S. Circular dichroism and molecular modeling yield a structure for the complex of human immunodeficiency virus type 1 trans-activation response RNA and the binding region of Tat, the trans-acting transcriptional activator. Proc Natl Acad Sci U S A. 1992 Oct 15;89(20):9734–9738. doi: 10.1073/pnas.89.20.9734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo Y., Madore S. J., Parslow T. G., Cullen B. R., Peterlin B. M. Functional analysis of interactions between Tat and the trans-activation response element of human immunodeficiency virus type 1 in cells. J Virol. 1993 Sep;67(9):5617–5622. doi: 10.1128/jvi.67.9.5617-5622.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo Y., Peterlin B. M. Juxtaposition between activation and basic domains of human immunodeficiency virus type 1 Tat is required for optimal interactions between Tat and TAR. J Virol. 1993 Jun;67(6):3441–3445. doi: 10.1128/jvi.67.6.3441-3445.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marciniak R. A., Garcia-Blanco M. A., Sharp P. A. Identification and characterization of a HeLa nuclear protein that specifically binds to the trans-activation-response (TAR) element of human immunodeficiency virus. Proc Natl Acad Sci U S A. 1990 May;87(9):3624–3628. doi: 10.1073/pnas.87.9.3624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marciniak R. A., Sharp P. A. HIV-1 Tat protein promotes formation of more-processive elongation complexes. EMBO J. 1991 Dec;10(13):4189–4196. doi: 10.1002/j.1460-2075.1991.tb04997.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muesing M. A., Smith D. H., Cabradilla C. D., Benton C. V., Lasky L. A., Capon D. J. Nucleic acid structure and expression of the human AIDS/lymphadenopathy retrovirus. Nature. 1985 Feb 7;313(6002):450–458. doi: 10.1038/313450a0. [DOI] [PubMed] [Google Scholar]
- Newstein M., Stanbridge E. J., Casey G., Shank P. R. Human chromosome 12 encodes a species-specific factor which increases human immunodeficiency virus type 1 tat-mediated trans activation in rodent cells. J Virol. 1990 Sep;64(9):4565–4567. doi: 10.1128/jvi.64.9.4565-4567.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nodwell J. R., Greenblatt J. The nut site of bacteriophage lambda is made of RNA and is bound by transcription antitermination factors on the surface of RNA polymerase. Genes Dev. 1991 Nov;5(11):2141–2151. doi: 10.1101/gad.5.11.2141. [DOI] [PubMed] [Google Scholar]
- Pavlakis G. N., Felber B. K. Regulation of expression of human immunodeficiency virus. New Biol. 1990 Jan;2(1):20–31. [PubMed] [Google Scholar]
- Puglisi J. D., Chen L., Frankel A. D., Williamson J. R. Role of RNA structure in arginine recognition of TAR RNA. Proc Natl Acad Sci U S A. 1993 Apr 15;90(8):3680–3684. doi: 10.1073/pnas.90.8.3680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puglisi J. D., Tan R., Calnan B. J., Frankel A. D., Williamson J. R. Conformation of the TAR RNA-arginine complex by NMR spectroscopy. Science. 1992 Jul 3;257(5066):76–80. doi: 10.1126/science.1621097. [DOI] [PubMed] [Google Scholar]
- Rappaport J., Lee S. J., Khalili K., Wong-Staal F. The acidic amino-terminal region of the HIV-1 Tat protein constitutes an essential activating domain. New Biol. 1989 Oct;1(1):101–110. [PubMed] [Google Scholar]
- Rosen C. A., Sodroski J. G., Haseltine W. A. The location of cis-acting regulatory sequences in the human T cell lymphotropic virus type III (HTLV-III/LAV) long terminal repeat. Cell. 1985 Jul;41(3):813–823. doi: 10.1016/s0092-8674(85)80062-3. [DOI] [PubMed] [Google Scholar]
- Rounseville M. P., Kumar A. Binding of a host cell nuclear protein to the stem region of human immunodeficiency virus type 1 trans-activation-responsive RNA. J Virol. 1992 Mar;66(3):1688–1694. doi: 10.1128/jvi.66.3.1688-1694.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roy S., Delling U., Chen C. H., Rosen C. A., Sonenberg N. A bulge structure in HIV-1 TAR RNA is required for Tat binding and Tat-mediated trans-activation. Genes Dev. 1990 Aug;4(8):1365–1373. doi: 10.1101/gad.4.8.1365. [DOI] [PubMed] [Google Scholar]
- Roy S., Parkin N. T., Rosen C., Itovitch J., Sonenberg N. Structural requirements for trans activation of human immunodeficiency virus type 1 long terminal repeat-directed gene expression by tat: importance of base pairing, loop sequence, and bulges in the tat-responsive sequence. J Virol. 1990 Mar;64(3):1402–1406. doi: 10.1128/jvi.64.3.1402-1406.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruben S., Perkins A., Purcell R., Joung K., Sia R., Burghoff R., Haseltine W. A., Rosen C. A. Structural and functional characterization of human immunodeficiency virus tat protein. J Virol. 1989 Jan;63(1):1–8. doi: 10.1128/jvi.63.1.1-8.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadaie M. R., Benter T., Wong-Staal F. Site-directed mutagenesis of two trans-regulatory genes (tat-III,trs) of HIV-1. Science. 1988 Feb 19;239(4842):910–913. doi: 10.1126/science.3277284. [DOI] [PubMed] [Google Scholar]
- Selby M. J., Bain E. S., Luciw P. A., Peterlin B. M. Structure, sequence, and position of the stem-loop in tar determine transcriptional elongation by tat through the HIV-1 long terminal repeat. Genes Dev. 1989 Apr;3(4):547–558. doi: 10.1101/gad.3.4.547. [DOI] [PubMed] [Google Scholar]
- Selby M. J., Peterlin B. M. Trans-activation by HIV-1 Tat via a heterologous RNA binding protein. Cell. 1990 Aug 24;62(4):769–776. doi: 10.1016/0092-8674(90)90121-t. [DOI] [PubMed] [Google Scholar]
- Sheline C. T., Milocco L. H., Jones K. A. Two distinct nuclear transcription factors recognize loop and bulge residues of the HIV-1 TAR RNA hairpin. Genes Dev. 1991 Dec;5(12B):2508–2520. doi: 10.1101/gad.5.12b.2508. [DOI] [PubMed] [Google Scholar]
- Southgate C. D., Green M. R. The HIV-1 Tat protein activates transcription from an upstream DNA-binding site: implications for Tat function. Genes Dev. 1991 Dec;5(12B):2496–2507. doi: 10.1101/gad.5.12b.2496. [DOI] [PubMed] [Google Scholar]
- Sullenger B. A., Gallardo H. F., Ungers G. E., Gilboa E. Analysis of trans-acting response decoy RNA-mediated inhibition of human immunodeficiency virus type 1 transactivation. J Virol. 1991 Dec;65(12):6811–6816. doi: 10.1128/jvi.65.12.6811-6816.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sullenger B. A., Gallardo H. F., Ungers G. E., Gilboa E. Overexpression of TAR sequences renders cells resistant to human immunodeficiency virus replication. Cell. 1990 Nov 2;63(3):601–608. doi: 10.1016/0092-8674(90)90455-n. [DOI] [PubMed] [Google Scholar]
- Sumner-Smith M., Roy S., Barnett R., Reid L. S., Kuperman R., Delling U., Sonenberg N. Critical chemical features in trans-acting-responsive RNA are required for interaction with human immunodeficiency virus type 1 Tat protein. J Virol. 1991 Oct;65(10):5196–5202. doi: 10.1128/jvi.65.10.5196-5202.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiley L. S., Brown P. H., Cullen B. R. Does the human immunodeficiency virus Tat trans-activator contain a discrete activation domain? Virology. 1990 Oct;178(2):560–567. doi: 10.1016/0042-6822(90)90354-t. [DOI] [PubMed] [Google Scholar]
- Weeks K. M., Ampe C., Schultz S. C., Steitz T. A., Crothers D. M. Fragments of the HIV-1 Tat protein specifically bind TAR RNA. Science. 1990 Sep 14;249(4974):1281–1285. doi: 10.1126/science.2205002. [DOI] [PubMed] [Google Scholar]
- Weeks K. M., Crothers D. M. RNA recognition by Tat-derived peptides: interaction in the major groove? Cell. 1991 Aug 9;66(3):577–588. doi: 10.1016/0092-8674(81)90020-9. [DOI] [PubMed] [Google Scholar]
- Wright C. M., Felber B. K., Paskalis H., Pavlakis G. N. Expression and characterization of the trans-activator of HTLV-III/LAV virus. Science. 1986 Nov 21;234(4779):988–992. doi: 10.1126/science.3490693. [DOI] [PubMed] [Google Scholar]
- Wu F., Garcia J., Sigman D., Gaynor R. tat regulates binding of the human immunodeficiency virus trans-activating region RNA loop-binding protein TRP-185. Genes Dev. 1991 Nov;5(11):2128–2140. doi: 10.1101/gad.5.11.2128. [DOI] [PubMed] [Google Scholar]





