Abstract
A secondary structure in the 5' noncoding region of avian retrovirus RNA, called the U5-leader stem, was shown previously to have a role in initiation of reverse transcription (D. Cobrinik, L. Soskey, and J. Leis, J. Virol. 62:3622-3630, 1988). We now show that an additional RNA secondary structure near the U5 terminus, called the U5-IR stem, is also important for reverse transcription. Mutations that disrupt the U5-IR stem cause a replication defect associated with both a decrease in synthesis of viral DNA in infected cells and a decrease in initiation of reverse transcription in melittin-permeabilized virions. Structure-compensating base substitutions in the U5-IR restore reverse transcription efficiency. In viral DNA, U5-IR sequences are included in the U5 terminal region that functions as a viral integration donor site. When base substitutions are introduced into these sequences, a reduced efficiency of integration in vitro and in vivo is observed. These observations indicate that U5-IR sequences have a structural role in reverse transcription of viral RNA and a sequence-specific role in the integration of viral DNA.
Full text
PDF




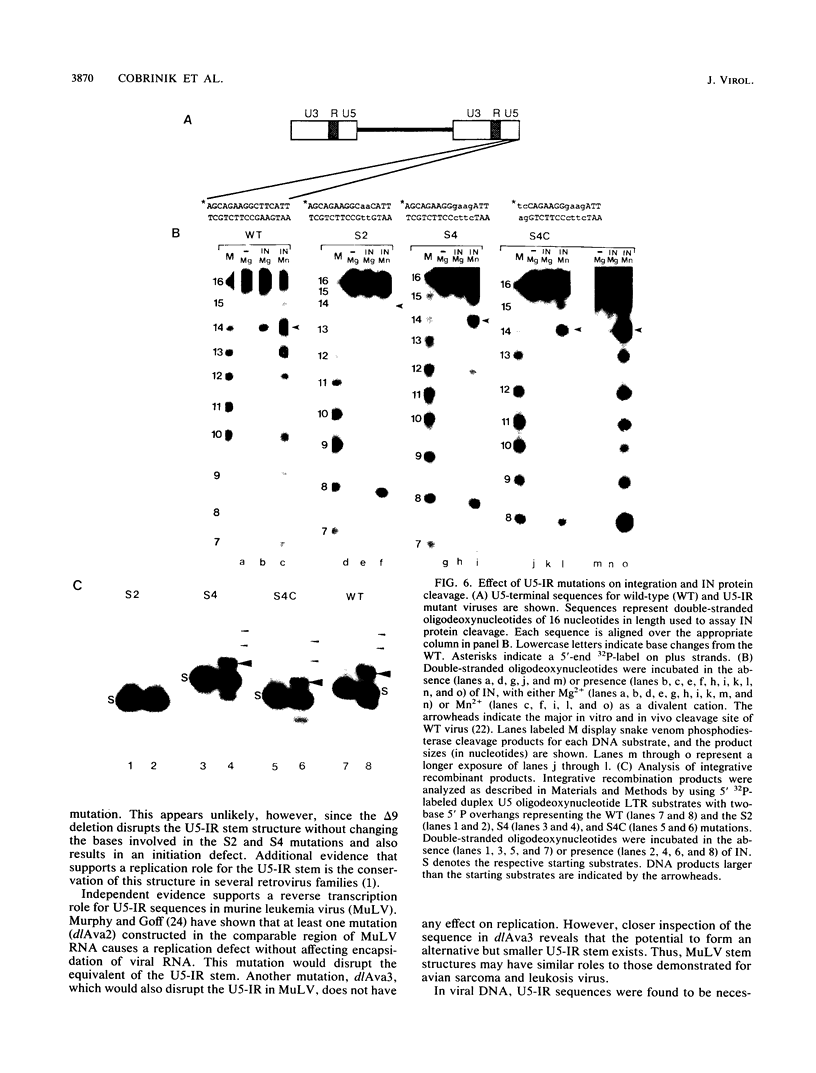


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andino R., Rieckhof G. E., Baltimore D. A functional ribonucleoprotein complex forms around the 5' end of poliovirus RNA. Cell. 1990 Oct 19;63(2):369–380. doi: 10.1016/0092-8674(90)90170-j. [DOI] [PubMed] [Google Scholar]
- Bilofsky H. S., Burks C., Fickett J. W., Goad W. B., Lewitter F. I., Rindone W. P., Swindell C. D., Tung C. S. The GenBank genetic sequence databank. Nucleic Acids Res. 1986 Jan 10;14(1):1–4. doi: 10.1093/nar/14.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boone L. R., Skalka A. M. Viral DNA synthesized in vitro by avian retrovirus particles permeabilized with melittin. I. Kinetics of synthesis and size of minus- and plus-strand transcripts. J Virol. 1981 Jan;37(1):109–116. doi: 10.1128/jvi.37.1.109-116.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown P. O., Bowerman B., Varmus H. E., Bishop J. M. Correct integration of retroviral DNA in vitro. Cell. 1987 May 8;49(3):347–356. doi: 10.1016/0092-8674(87)90287-x. [DOI] [PubMed] [Google Scholar]
- Cobrinik D., Katz R., Terry R., Skalka A. M., Leis J. Avian sarcoma and leukosis virus pol-endonuclease recognition of the tandem long terminal repeat junction: minimum site required for cleavage is also required for viral growth. J Virol. 1987 Jun;61(6):1999–2008. doi: 10.1128/jvi.61.6.1999-2008.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cobrinik D., Soskey L., Leis J. A retroviral RNA secondary structure required for efficient initiation of reverse transcription. J Virol. 1988 Oct;62(10):3622–3630. doi: 10.1128/jvi.62.10.3622-3630.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colicelli J., Goff S. P. Mutants and pseudorevertants of Moloney murine leukemia virus with alterations at the integration site. Cell. 1985 Sep;42(2):573–580. doi: 10.1016/0092-8674(85)90114-x. [DOI] [PubMed] [Google Scholar]
- Colicelli J., Goff S. P. Sequence and spacing requirements of a retrovirus integration site. J Mol Biol. 1988 Jan 5;199(1):47–59. doi: 10.1016/0022-2836(88)90378-6. [DOI] [PubMed] [Google Scholar]
- Cordell B., Swanstrom R., Goodman H. M., Bishop J. M. tRNATrp as primer for RNA-directed DNA polymerase: structural determinants of function. J Biol Chem. 1979 Mar 25;254(6):1866–1874. [PubMed] [Google Scholar]
- Craigie R., Fujiwara T., Bushman F. The IN protein of Moloney murine leukemia virus processes the viral DNA ends and accomplishes their integration in vitro. Cell. 1990 Aug 24;62(4):829–837. doi: 10.1016/0092-8674(90)90126-y. [DOI] [PubMed] [Google Scholar]
- Dahlberg J. E., Sawyer R. C., Taylor J. M., Faras A. J., Levinson W. E., Goodman H. M., Bishop J. M. Transcription of DNA from the 70S RNA of Rous sarcoma virus. I. Identification of a specific 4S RNA which serves as primer. J Virol. 1974 May;13(5):1126–1133. doi: 10.1128/jvi.13.5.1126-1133.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darlix J. L., Zuker M., Spahr P. F. Structure-function relationship of Rous sarcoma virus leader RNA. Nucleic Acids Res. 1982 Sep 11;10(17):5183–5196. doi: 10.1093/nar/10.17.5183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Craigie R. Integration of mini-retroviral DNA: a cell-free reaction for biochemical analysis of retroviral integration. Proc Natl Acad Sci U S A. 1989 May;86(9):3065–3069. doi: 10.1073/pnas.86.9.3065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagino-Yamagishi K., Donehower L. A., Varmus H. E. Retroviral DNA integrated during infection by an integration-deficient mutant of murine leukemia virus is oligomeric. J Virol. 1987 Jun;61(6):1964–1971. doi: 10.1128/jvi.61.6.1964-1971.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haseltine W. A., Kleid D. G., Panet A., Rothenberg E., Baltimore D. Ordered transcription of RNA tumor virus genomes. J Mol Biol. 1976 Sep 5;106(1):109–131. doi: 10.1016/0022-2836(76)90303-x. [DOI] [PubMed] [Google Scholar]
- Haseltine W. A., Maxam A. M., Gilbert W. Rous sarcoma virus genome is terminally redundant: the 5' sequence. Proc Natl Acad Sci U S A. 1977 Mar;74(3):989–993. doi: 10.1073/pnas.74.3.989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967 Jun 14;26(2):365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- Hughes S. H., Greenhouse J. J., Petropoulos C. J., Sutrave P. Adaptor plasmids simplify the insertion of foreign DNA into helper-independent retroviral vectors. J Virol. 1987 Oct;61(10):3004–3012. doi: 10.1128/jvi.61.10.3004-3012.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katz R. A., Merkel G., Kulkosky J., Leis J., Skalka A. M. The avian retroviral IN protein is both necessary and sufficient for integrative recombination in vitro. Cell. 1990 Oct 5;63(1):87–95. doi: 10.1016/0092-8674(90)90290-u. [DOI] [PubMed] [Google Scholar]
- Katzman M., Katz R. A., Skalka A. M., Leis J. The avian retroviral integration protein cleaves the terminal sequences of linear viral DNA at the in vivo sites of integration. J Virol. 1989 Dec;63(12):5319–5327. doi: 10.1128/jvi.63.12.5319-5327.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullis K. B., Faloona F. A. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 1987;155:335–350. doi: 10.1016/0076-6879(87)55023-6. [DOI] [PubMed] [Google Scholar]
- Murphy J. E., Goff S. P. Construction and analysis of deletion mutations in the U5 region of Moloney murine leukemia virus: effects on RNA packaging and reverse transcription. J Virol. 1989 Jan;63(1):319–327. doi: 10.1128/jvi.63.1.319-327.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panganiban A. T., Temin H. M. The terminal nucleotides of retrovirus DNA are required for integration but not virus production. Nature. 1983 Nov 10;306(5939):155–160. doi: 10.1038/306155a0. [DOI] [PubMed] [Google Scholar]
- Roth M. J., Schwartzberg P. L., Goff S. P. Structure of the termini of DNA intermediates in the integration of retroviral DNA: dependence on IN function and terminal DNA sequence. Cell. 1989 Jul 14;58(1):47–54. doi: 10.1016/0092-8674(89)90401-7. [DOI] [PubMed] [Google Scholar]
- Schroeder W., Bade E. G., Delius H. Participation of Escherichia coli DNA in the replication of temperate bacteriophage Mu1. Virology. 1974 Aug;60(2):534–542. doi: 10.1016/0042-6822(74)90347-x. [DOI] [PubMed] [Google Scholar]
- Schwartz D. E., Tizard R., Gilbert W. Nucleotide sequence of Rous sarcoma virus. Cell. 1983 Mar;32(3):853–869. doi: 10.1016/0092-8674(83)90071-5. [DOI] [PubMed] [Google Scholar]
- Schwartzberg P., Colicelli J., Goff S. P. Construction and analysis of deletion mutations in the pol gene of Moloney murine leukemia virus: a new viral function required for productive infection. Cell. 1984 Jul;37(3):1043–1052. doi: 10.1016/0092-8674(84)90439-2. [DOI] [PubMed] [Google Scholar]
- Temin H. M., Baltimore D. RNA-directed DNA synthesis and RNA tumor viruses. Adv Virus Res. 1972;17:129–186. doi: 10.1016/s0065-3527(08)60749-6. [DOI] [PubMed] [Google Scholar]