Abstract
PutA is a novel flavoprotein in Escherichia coli that switches from a transcriptional repressor to a membrane-bound proline catabolic enzyme. Previous crystallographic studies of the PutA proline dehydrogenase (PRODH) domain under oxidizing conditions revealed that the FAD N(5) and ribityl 2′- OH form hydrogen bonds with Arg431 and Arg556, respectively. Here we identify molecular interactions in the PutA PRODH active site that underlie redox-dependent functional switching of PutA. We report that reduction of the PRODH domain induces major structural changes in the FAD cofactor, including a 22° bend of the isoalloxazine ring along the N(5)-N(10) axis, crankshaft rotation of the upper part of the ribityl chain, and formation of a new hydrogen bond network involving the ribityl 2′- OH, FAD N(1) atom, and Gly435. The roles of the FAD 2′-OH group and the FAD N(5)-Arg431 hydrogen bond pair in regulating redox-dependent PutA-membrane associations were tested using FAD analogues and site-directed mutagenesis. Kinetic membrane-binding measurements and cell-based reporter gene assays of modified PutA proteins show that disrupting the FAD N(5)-Arg431 interaction impairs reductive activation of PutA-membrane binding. We also show that the FAD-2′-OH acts as a redox-sensitive toggle switch that controls PutA-membrane binding. These results illustrate a new versatility of the ribityl chain in flavoprotein mechanisms.
Proline utilization A (PutA) from Escherichia coli and other Gram negative bacteria is a large multifunctional protein that uniquely combines enzymatic and transcriptional regulatory activities within a single 1320 amino acid polypeptide (1-4). As an enzyme, PutA peripherally associates with the inner cytoplasmic membrane to catalyze the four-electron oxidation of proline to glutamate via the coordinated actions of separate flavin-dependent proline dehydrogenase (PRODH) and NAD+-dependent Δ1-pyrroline-5-carboxylate dehydrogenase (P5CDH) domains involving residues 263-612 and 650-1130, respectively (5, 6). An N-terminal ribbon-helix-helix motif (residues 2-43) endows PutA with DNA-binding activity enabling PutA to also function as a cytosolic autogenous transcriptional repressor of the proline utilization (put) genes putA and putP (encodes a high affinity proline transporter) (7-9). To fulfill its mutually exclusive roles as a transcriptional repressor and membrane-bound proline catabolic enzyme, PutA undergoes proline-dependent functional switching. Proline reduction of the FAD cofactor induces PutA-membrane binding which consequently disrupts the PutA-DNA complex equilibrium and activates put gene expression (10-14).
To reveal the mechanism by which FAD reduction drives tight PutA-membrane associations and therefore regulates PutA function, we have begun to explore molecular interactions between the FAD cofactor and active site residues in the PRODH domain of PutA. Previously, the structure of a truncated form of oxidized PutA containing residues 86-669 (PutA86-669) was solved by X-ray crystallography (2.0-2.1 Å) in complex with different active site inhibitors such as L-lactate (Ki = 1.4 mM) and L-tetrahydro-2-furoic acid (L-THFA, Ki = 0.2 mM) (15). Important ion pair interactions are evident between the substrate carboxylate group and the guanidinium groups of Arg555 and Arg556. Also, Arg556 is within hydrogen bonding distance of the 2′-OH ribityl group of the FAD cofactor. Another notable interaction is between Arg431 and the N(5) atom of the isoalloxazine ring. Because electron density changes dramatically across the N(1)-N(5) enediamine system upon FAD reduction, we hypothesized that Arg431 may have a role in enabling PutA to sense redox changes in the FAD cofactor.
In this study, redox-dependent structural changes in the FAD cofactor were identified by comparing crystal structures of oxidized and dithionite reduced PutA86-669. Substantial movement of the 2′-OH group of the ribityl moiety was observed in the dithionite treated crystals relative to oxidized PutA86- 669 resulting in disruption of the Arg556-N(2)H-O(2′)-FAD interaction and the formation of a new hydrogen bond network between the ribityl 2′-OH group, the N(1) atom of the isoalloxazine ring, the ribityl 4′-OH, and Gly435. We subsequently analyzed the impact of these interactions on PutA function by measuring the kinetic membrane binding properties of PutA reconstituted with FAD analogues and matching PutA site directed mutants by surface plasmon resonance (SPR). Results are presented that provide an example of a unique mechanism by which the FAD cofactor controls membrane binding of a large multifunctional enzyme.
Materials and Methods
Materials
2′-deoxy- and 5-deaza-riboflavin were generous gifts from Dr. Marian T. Stankovich (University of Minnesota). 2′-deoxy-FAD and 5-deaza-FAD were synthesized from the respective riboflavin analogs and purified as previously described (16-18). All chemicals and buffers were from Sigma unless stated otherwise. The oligonucleotides ′5- CGTCGCCGTCTGATGATTATGCTGGTGAAAGGCGCGTAC-3′ and ′5-GGCGTATCTGGTGCGTATGCTGCTGGAAAACGGTGC-3′ (IDT, Inc.) were used for generating PutA mutants R431M and R556M, respectively, by QuikChange (Stratagene) site directed mutagenesis. The mutants were confirmed by DNA sequencing in the DNA Core Facility at the University of Nebraska-Lincoln. Lipid vesicles were prepared from E. coli lipid polar extracts (Avanti Polar Lipids) as previously described (19).
Structure determination of reduced PutA86-669
Crystals of oxidized PutA86-669 were grown in sitting drops in the presence of PEG3350 and citrate buffer at pH 5.6-6.0 as described previously (15, 20). The crystals were stabilized by transfer into 20-50 μL of cryo-buffer, which consisted of 24% PEG 3350, 15% PEG 200, and 0.1 M citrate buffer at pH 5.7. The crystalline enzyme was reduced by adding a few grains of solid sodium dithionite to the liquid surrounding the stabilized crystals. The reaction of sodium dithionite with PutA86-669 bleached the deep yellow color of the crystals indicating that FAD was reduced. The reduced crystals were picked up with Hampton mounting loops and plunged into liquid nitrogen to trap the reduced enzyme conformation.
The structure of reduced PutA86-669 was determined to 2.3 Å resolution using a diffraction data set collected from one crystal at beamline 19ID of the Advanced Photon Source. Integration and scaling of the data were performed with HKL (21). Reduced PutA86-669 crystals have space group I222 with unit cell dimensions of a = 73.1 Å, b = 141.4 Å, c = 145.7 Å, and one protein molecule per asymmetric unit. For reference, the oxidized crystals also have space group I222 and unit cell dimension of a = 72.9 Å, b = 141.1 Å, and c = 145.4 Å (15). Thus, reduction by dithionite did not change the crystal lattice appreciably. See Table 1 for a summary of data processing statistics.
Table 1.
X-ray Diffraction Data Collection and Refinement Statisticsa
| Diffraction resolution (Å) | 50 - 2.3 (2.38 - 2.30) |
| No. of observations | 189645 |
| No. of unique reflections | 33078 |
| Completeness (%) | 98 (94) |
| Rsym | 0.071 (0.465) |
| Average I/σ | 0.071 (0.465) |
| No. of non-hydrogen atoms | 3633 |
| No. of protein residues | 450 |
| No. of water molecules | 117 |
| Rcryst/Rfree | 0.209 (0.256) / 0.257 (0.280) |
| RMSD bond lengths (Å) / angles (deg)b | 0.015 / 1.5 |
| Ramachandran plot (% of residues)c | |
| most favored regions | 94.1 |
| additional allowed regions | 5.9 |
| Average B-factors (Å2) | |
| Protein | 40 |
| FAD | 35 |
| SO2 | 50 |
| Water | 45 |
| PDB accession code | 2FZM |
The starting model for refinement was a previously solved structure of oxidized PutA86-669 (PDB code 1TJ0) with solvent, FAD and bound L-lactate removed (15). Refinement calculations were performed with CNS (22) and REFMAC5 (23). Model building was done with O (24). The test set of reflections (5%) used for cross validation matched the one used for refinement of 1TJ0.
The refined structure consists of 450 protein residues, one FAD cofactor, 117 water molecules, and one SO2 ligand bound in the substrate-binding pocket. As with previously solved structures of oxidized PutA86-669, the β8α8 FAD-binding domain is highly ordered but electron density for residues outside of this domain is very weak indicating significant disorder. In the present structure of reduced PutA86-669, residues 148-161, 185-243, and 611-669 are disordered. See Table 1 for a summary of refinement statistics.
PutA protein purification and characterization
Site-directed mutants R431M and R556M were generated in full-length PutA. Full-length recombinant wild-type PutA and PutA mutants R431M and R556 were purified as N-terminal 6xHis tag proteins as described (13). The dimeric forms of the proteins were collected by size-exclusion chromatography (Superdex-200 column) and were used for all of the SPR experiments. Full-length PutA apoprotein was prepared and reconstituted with either FAD, 2′-deoxy-FAD or 5-deaza-FAD as described (18). PutA86-669 was used for the X-ray crystallography studies and PutA86-630 was used for potentiometric measurements of 2′-deoxy-FAD. PutA86-669 and PutA86-630 were purified as previously described (25). PutA86-630 apoprotein was prepared and reconstituted with 2′-deoxy-FAD as described (18). Protein concentrations were determined using bicinchoninic acid reagents (Pierce) with bovine serum albumin as the standard and spectrophotometrically using a molar extinction coefficient of 12,700 M−1 cm−1 at 451 nm (26). Kinetic parameters for PRODH activity were determined as described earlier (26). DNA-binding activity of each PutA protein was analyzed by gel mobility shift assays as previously described (26). Proline and sodium dithionite titrations of PutA proteins were performed under anaerobic conditions in a nitrogen atmosphere in a glove box (Belle Technology) as previously described (26). Potentiometric titrations of 2′-deoxy-FAD PutA86-630 were performed as described previously under a nitrogen atmosphere (27-29). Potential values are reported relative to the normal hydrogen electrode.
Cell-based reporter gene assays
The put control DNA region (putC, 419 bp) was positioned upstream of the lacZ gene in the vector pRS415, a generous gift from Dr. Robert W. Simons (University of California, Los Angeles) using EcoRI and BamHI. Unique Eco52I and SalI sites were then introduced at the 5′ and 3′ ends of the putC:lacZ fusion by PCR, respectively, and used to subclone the putC:lacZ reporter gene into the low copy vector pACYC184 to generate the putC:lacZ reporter construct. The genes encoding full-length wild-type PutA and PutA R431M were cloned downstream of the lac promoter in a compatible pUC18 vector using NcoI and HindIII. Previously, a unique NcoI site was introduced into pUC18 immediately upstream of the multiple cloning site by QuikChange site-directed mutagenesis to optimize expression of the putA genes. Each of the above constructs was confirmed by DNA sequencing.
To test the proline-dependent activation of the putC:lacZ reporter gene, E. coli strain JT31 putA−lacZ− was transformed with the putC:lacZ construct and with either wild-type PutA (pUC18-PutA) or PutA R431M (pUC18-R431M). For control assays, cells were transformed with a pUC18 empty vector which lacks the putA gene and has the lacZ gene interrupted by a putC insertion in the multiple cloning site as previously described (7). E. coli cells containing the putC:lacZ construct and the different pUC18 test vectors were grown in minimal medium at 37 °C with ampicillin (50 μg/ml), chloroamphenicol (30 μg/ml), kanamycin (40 μg/ml) and with and without proline or L-THFA to OD at 600 nm ~ 1.0. Pelleted cells were then resuspended and broken using the bacterial protein extraction reagent from Pierce (20 mM Tris-HCL, pH 7.5). β-Galactosidase activity assays were then performed as previously described by measuring the increase in absorbance at 420 nm (7).
PutA-membrane association kinetics
The kinetics of PutA-membrane binding were performed on a Biacore 2000 instrument at 25 °C as previously described in the Genomics Core Facility in the Center for Biotechnology at the University of Nebraska (13, 19). The running buffer used in all SPR experiments was 10 mM N-(2-hydroxyethyl)piperazine-N′-2-ethanesulfonic acid containing 150 mM NaCl at pH 7.4 (HEPES-N buffer). E. coli polar lipid vesicles were loaded onto a L1 sensor chip at 5 μl/min during which the vesicles merge into a bilayer membrane on the surface (30). After coating with lipid vesicles, the surface was washed with 10 mM NaOH (60 μl/min) resulting in a total response (RU) for lipid loading of about 4500 RU. Nonspecific protein-lipid interactions were blocked by injection of bovine serum albumin (0.1 mg/ml) onto the sensor surface prior to injection of the PutA proteins.
The effects of proline and sodium dithionite on PutA-lipid binding were tested by adding the reagents to the PutA samples 15 min prior to injection. The proline concentrations used were 5 mM for reconsituted PutA, 10 mM for PutA R431M and 2′-deoxy-FAD-PutA, 100 mM for 5-deaza-FAD-PutA, and 400 mM for PutA R556M. Dithionite was used at 5 mM for all of the PutA proteins except 5-deaza-FAD-PutA which was supplemented with 50 mM sodium dithionite. Sodium dithionite was added to the PutA samples under a nitrogen atmosphere in a glovebox. SPR analysis of PutA-lipid binding in the presence of dithionite was then performed under anaerobic conditions as described (13). For kinetic analysis, PutA samples were injected at a flow rate of 60 μl/min but different flow rates from 20 – 80 μl/min were also used to confirm there were no mass transfer problems. The association and dissociation phases were monitored for 120 s and 300 s, respectively. The sensorgrams of PutA-lipid associations were analyzed by Biaevaluation 4.1 software as previously described (19). Changes in refractive index due to buffer changes were subtracted prior to kinetic analysis. Global fitting to a 1:1 Langmuir PutA-membrane binding model was used to calculate the association (ka) and dissociation (kd) rate constants, and the dissociation constant (KD) was calculated from KD = kd/ka. Complete kinetic parameters are reported in the supporting information in Table S1.
Results
X-ray crystal structure of reduced PutA86-669
To gain insights into structural changes in the PRODH domain that are caused by reduction of the FAD cofactor, PutA86-669 crystals were reduced with sodium dithionite. As expected, the structure of reduced PutA86-669 exhibits the α8β8 fold that was observed for oxidized PutA86-669 (Figure 1). The FAD cofactor is bound on the carboxyl terminal ends of the strands of the barrel. The re face of the FAD isoalloxazine packs tightly against strands 4-6 of the barrel, while the si face helps form the substrate binding pocket. No significant differences in amino acid side chain conformations were found between the oxidized and dithionite-reduced forms of PutA86-669.
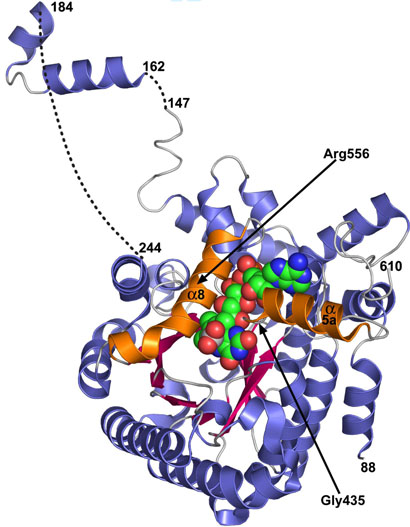
FIGURE 1.
Ribbon drawing of dithionite-reduced PutA86-669. β-strands of the β8α8 barrel are colored pink and α-helices are colored blue. The two helices α8 and α5a are highlighted in orange. The FAD cofactor (green) and bound SO22- (gray/red) are drawn in CPK mode. The dashed curves represent disordered residues. Selected residue numbers are indicated.
Electron density maps of dithionite-reduced PutA86-669 clearly indicated a ligand larger than water bound in the active site (Figure 2A). The ligand was modeled as SO2 (presumably hyposulfite, SO2 2−) and was likely generated by oxidation of dithionite (S2O42−). SO2 binds at the location occupied by the carboxyl group of competitive inhibitors such as L-lactate and L-THFA. The SO2 ligand forms hydrogen bonds with the side chains of Arg555, Arg556, and Lys329. Analogous interactions are formed by inhibitor carboxyl groups in structures of oxidized PutA86-669 bound to L-lactate and L-THFA (15).
FIGURE 2.
Three views of the active site of dithionite-reduced PutA86-669. A) Highlighting interactions with SO22-, B) conformation of FAD 2′-OH, and C) nonplanar isoalloxazine. In each panel, the red dashed lines denote electrostatic interactions (hydrogen bonds, ion pairs) and the blue cage represents a simulated annealing σA-weighted mFo-DFc electron density map contoured at 3.5σ.
Although no cshanges were observed in the overall PutA86-669 protein fold, significant conformational changes occurred in the FAD cofactor upon reduction. Figure 3 compares the structures of the FAD cofactor bound to PutA86-669 before and after treatment with sodium dithionite. In the oxidized form, 2′-OH group of the ribityl moiety points toward the guanidinium group of Arg556 and a hydrogen bond is formed between the 2′-OH and Arg556 (see green dotted line in Figure 3). When FAD is reduced by dithionite, a crankshaft motion occurs in the upper part of the ribityl chain, which causes the 2′-OH group to rotate by approximately 90°. This motion brings the 2′-OH group within hydrogen bonding distance of the N(1) atom of the isoalloxazine ring (2.7 Å), the 4′-OH of the FAD ribityl (2.7 Å), and the N atom of Gly435 (3.0 Å), suggesting the formation of two new favorable intra-FAD interactions and one new favorable protein-FAD interaction (Figures 2B and 3). As a consequence, the Arg556-N(2)H-O(2′)-FAD hydrogen bond, which is observed in the oxidized enzyme, is broken in the reduced enzyme.
FIGURE 3.
Stereo view of conformational differences between dithionite-reduced PutA86-669 and oxidized ligand-bound PutA86-669. Reduced PutA86-669 with bound SO22- is shown in white. Oxidized PutA86-669 with bound substrate analogue L-THFA is colored green. Black dotted lines indicate hydrogen bonds observed in both structures. Green dotted lines indicate the hydrogen bond between Arg556 and 2′-OH observed only in the oxidized enzyme. Orange dotted lines indicate hydrogen bonds unique to the reduced enzyme.
In addition to movement of the 2′-OH group, the FAD isoalloxazine bends dramatically (22°) along the N(5)-N(10) axis compared to the oxidized planar conformation (Figure 2C). The bend is generated by 1-Å movement of the N(5)-N(10) axis toward the si face (proline binding face) while the dimethyl benzene and pyrimidine rings remain relatively stationary. This motion decreases the nonpolar interactions between the protein and the flavin by disrupting the intimate packing of Val433, which is part of β-strand 5, against the re side of the FAD (Figure 3). In the oxidized state, side chain atoms of Val433 are within 3.6 Å of FAD atoms N5, N10 and C1′. In the reduced state, however, these distances are increased to 4.0 - 4.9 Å, which is outside van der Waals contact.
We are aware of only one other flavoenzyme that exhibits comparable structural changes in the FAD cofactor upon reduction, E. coli thioredoxin reductase (31). In E. coli thioredoxin reductase, reduction of the FAD cofactor results in formation of a new intramolecular FAD-N(1)H-O(2′) hydrogen bond and a 34° bend of the isoalloxazine.
FAD N(5)-Arg431 hydrogen bond
To explore the mechanism by which the FAD redox state regulates PutA-membrane binding, we first examined the role of the N(5)-Arg431 interaction which is found in both the oxidized and dithionite-reduced crystal structures of PutA86-669 (Figure 3). PutA was reconstituted with 5-deaza-FAD which replaces the N(5) atom with carbon in the isoalloxazine ring. Prior to characterizing 5-deaza-FAD PutA, the properties of apoPutA and PutA reconstituted with native FAD were determined. As expected, apoPutA has minimal PRODH activity (< 3%) relative to the holoprotein. The DNA-binding activity of apoPutA is similar to wild-type PutA (KD ~ 50 nM for the PutA-DNA complex) as previously reported (11, 26). SPR analysis of apoPutA shows it binds the lipid bilayer surface with KD =1.7 ± 0.8 nM (Table 2 and Figure S1). Upon reconstitution of PutA with normal FAD, enzyme activity is restored and DNA-binding activity remains fully functional. In addition, reconstituted PutA does not bind to the membrane surface in the oxidized state (Table 2). In the presence of proline or dithionite, however, reconstituted FAD-PutA binds tightly to the membrane similarly to wild-type PutA with upper limits of KD < 0.01 nM for the PutA-lipid complex (Table 2, Figure 4 and Figure S2) (13). Thus, regulation of PutA-membrane binding is fully restored by reconstituting with FAD.
Table 2.
Equilibrium Constants of Full-length PutA Binding to E. coli Polar Lipid Vesicles Determined on Biacore 2000a
| PutA Protein | no reductant |
KD (nM) proline |
dithionite |
|---|---|---|---|
| Apoprotein | 1.7 ± 0.8 | ndb | No binding |
| Reconstituted PutA | No binding | <0.01 | <0.01 |
| 5-deaza-FAD-PutA | No binding | No binding | No binding |
| PutA R431M | No binding | No binding | No binding |
| 2′-deoxy-FAD-PutA | 0.5 ± 0.1 | 1.1 ± 0.3 | No binding |
| PutA R556M | 1.6 ± 0.3 | 1.0 ± 0.2 | <0.01 |
KD values were determined in HEPES-N buffer (pH 7.4) at 25 °C.
Not determined. Complete kinetic parameters are provided in Table S1 as supporting information. The concentrations of proline and dithionite used for the analysis of each protein are described in the material and methods.
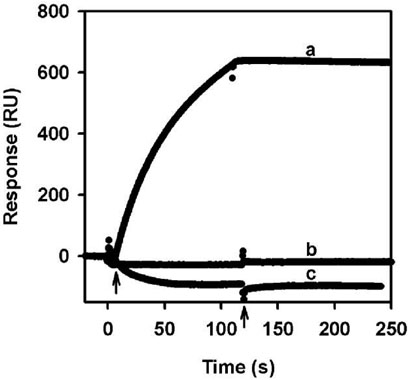
FIGURE 4.
SPR sensorgrams of reduced PutA proteins injected onto a L1 chip coated with E. coli polar lipid vesicles. Sensorgrams (a) 100 nM reconstituted FAD PutA (5 mM dithionite), (b) 100 nM R431M (5 mM dithionite), and (c) 100 nM 5-deaza-FAD PutA (50 mM dithionite), respectively. PutA proteins were injected at 60 μL/min in the presence of sodium dithionite in HEPES-N buffer (pH 7.4) with arrows indicating the beginning and end of the protein sample injection. No lipid interactions are observed with R431M and 5-deaza-FAD PutA.
Next, PutA was reconstituted with 5-deaza-FAD and as anticipated was devoid of PRODH activity. The spectrum of 5-deaza-FAD PutA is characterized by two main absorbance peaks at 401 and 350 nm as previously shown (18). Reconstituting PutA with 5-deaza-FAD did not perturb the DNA-binding properties of PutA. SPR analysis of 5-deaza-FAD PutA revealed that PutA reconstituted with 5-deaza-FAD does not interact with lipids in the oxidized state or under conditions that have been shown to reduce the 5-deaza-FAD analogue (Table 2 and Figure 4) (18). The N(5) position was then probed by replacing Arg431 with Met. The PutA mutant R431M has a higher Km (174 ± 32 mM proline) and lower kcat (1.2 ± 0.06 s−1) for PRODH activity than wild-type PutA (Km = 100 mM proline and kcat = 7 s−1) (27). The DNA-binding activity of PutA R431M is similar to that of wild-type PutA. As observed with wild-type PutA, the FAD cofactor in PutA R431M is reduced by dithionite and proline (Figure S3). Reduction of PutA R431M, however, fails to induce lipid binding (Table 2). Figure 4 shows a SPR sensorgram of PutA R431M injected onto the lipid bilayer surface in the presence of 5 mM dithionite. Clearly, the N(5)-Arg431 interaction is critical for FAD reduction to trigger PutA-membrane binding.
Cell-based reporter gene assays were then performed with PutA R431M to test whether the lack of membrane binding impacts the functional switching of PutA. Figure 5 shows that both wild-type PutA and the R431M mutant repress expression of the lacZ reporter gene by nearly 70% relative to control cells (pUC18 empty vector). In cells containing wild-type PutA, repression of the lacZ gene is relieved by adding proline with 50% activation of lacZ expression estimated at 0.28 ± 0.04 mM proline. Addition of L-THFA to the medium does not increase β-galactosidase activity consistent with proline reduction of the FAD cofactor being essential for tight PutA-membrane binding and PutA functional switching (Figure 5) (13). In contrast to wild-type PutA, proline has no affect on lacZ expression in cells containing the PutA R431M mutant. Therefore, the failure of FAD reduction to induce PutA R431M-membrane binding correlates with the loss of proline-dependent transcriptional activation of the putC:lacZ reporter gene in vivo.
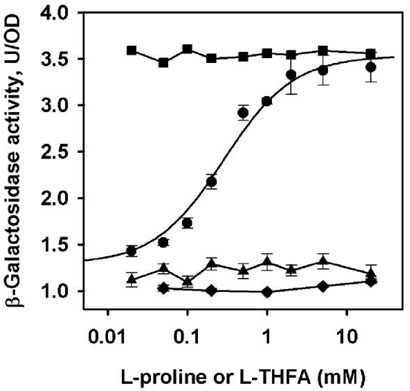
FIGURE 5.
Proline-dependent transcriptional activation of the putC:lacZ reporter gene. β-galactosidase activity from E. coli strain JT31 putA− lacZ− containing the putC:lacZ reporter construct and either pUC18 empty vector (■), pUC18-PutA (●), or pUC18-PutA R431M (♦) grown in medium supplemented with increasing amounts of proline. Also shown is β-galactosidase activity from cells containing putC:lacZ and pUC18-PutA grown in medium supplemented with increasing amounts of LTHFA (▲). Bars represent the ± standard errors of the mean from at least three independent experiments. Solid curve is the fit of the increase in β-galactosidase activity as a function of proline concentration in cells containing pUC18-PutA to the equation Pobs = Pmax * [pro]/(Keq + [pro]).
Role of the FAD ribityl 2′-OH group
The FAD 2′-OH group was examined by reconstituting PutA with 2′-deoxy-FAD and replacing Arg556 with Met in the PutA mutant R556M. PutA reconstituted with 2′-deoxy-FAD exhibits kinetic parameters (Km = 103 ± 11 mM proline and kcat = 4 ± 0.2 s-−) that are similar to wild-type PutA and two main flavin absorbance peaks at 451 nm and 380 nm (Figure S4). Reconstituting PutA with 2′-deoxy-FAD also did not change its DNA-binding properties. 2′-deoxy-FAD PutA, however, is only partially reduced (54%) by 10 mM proline under anaerobic conditions (Figure S4). Electrochemical characterization of 2′-deoxy-FAD bound to a truncated PutA protein (PutA86-630) similar to that used for the crystallization studies confirmed a lower redox potential of the 2′-deoxy-FAD cofactor. A reduction potential (Em) value of −0.111 V (pH 7.0) was determined for 2′-deoxy-FAD PutA86-630, a value which is 35 mV more negative than that previously reported for the normal FAD cofactor in PutA (Figure S5) (26, 27). The Em for 2′-deoxy-FAD in solution is 13 mV more negative than normal FAD in solution, thus, the difference in Em between PutA-bound FAD and 2′-deoxy-FAD is likely caused by altered protein-FAD interactions (32). The more negative Em value of 2′-deoxy-FAD PutA relative to wild-type PutA indicates that interactions formed by the 2′-OH group such as hydrogen bonds with FAD N(1), FAD 4′-OH, and Gly435, help stabilize the anionic hydroquinone state of PutA. It also suggests that a water molecule is not able to replace the 2′-OH hydrogen bond with the FAD N(1) position in PutA.
SPR measurements of 2′-deoxy-FAD PutA revealed that its membrane binding behavior resembles that of apoPutA. Oxidized 2′-deoxy-FAD PutA binds to the lipid surface with KD=0.5 ± 0.1 nM for the protein-lipid complex (Table 2 and Figure 6A). In the presence of proline, similar binding is observed (Table 2). Because proline only partially reduces 2′-deoxy-FAD, the membrane binding behavior of 2′- deoxy-FAD PutA was also examined in the presence of dithionite under anaerobic conditions. Surprisingly, no significant lipid binding was observed in the sensorgrams (Table 2). We also observed that sodium dithionite disrupts apoPutA membrane binding (Table 2). Thus, the loss of 2′-deoxy-FAD PutA-lipid interactions in the presence of dithionite is not redox related but possibly due to diminished electrostatic protein-membrane interactions.
The interaction of the 2′-OH group with R556 was then examined using the matching PutA mutant R556M. The PRODH catalytic turnover number of PutA R556M was estimated to be ~ 1.4 s−1 while a lower limit of 1 M proline was estimated for the Km value. The large increase in Km confirms previous X-ray crystallographic data of PutA86-669 that suggested the primary function of Arg556 is to form ion pair interactions with the carboxylate group of proline (5, 15). Consistent with the elevated Km value, high proline concentrations are required to reduce PutA R556M which has a similar flavin absorbance spectrum to wild-type PutA (Figure S6). PutA R556M binding to the put intergenic DNA is similar to wild-type PutA with an overall KD of 50 ± 8 nM for the PutA R556M-DNA complex (data not shown). As observed with 2′-deoxy-FAD-PutA, PutA R556M binds similarly to the lipid surface under oxidizing conditions (Table 2 and Figure 6B) and in the presence of proline (Table 2). However, sodium dithionite induces tight membrane binding comparable to that of wild-type PutA (Table 2 and Figure 6C) (13).
FIGURE 6.
SPR kinetic analyses of oxidized 2′-deoxy-FAD-PutA, oxidized PutA R556M and reduced PutA R556M binding to E. coli polar lipids at 25 °C in HEPES-N buffer (pH 7.4). Sensorgrams of increasing concentrations of A) oxidized 2′-deoxy-FAD-PutA (12.5, 25, 50, 100, and 200 nM) and B) oxidized PutA R556M (6.25, 12.5, 25, 50, and 100 nM). C) From bottom to top, sensorgrams of increasing concentrations of PutA R556M (6.25, 12, 25, 50, and 100 nM) in the presence of 5 mM sodium dithionite. For all sensorgrams, association phase (1) corresponds to injection of the PutA protein at 60 μl/min for 120 s and the dissociation phase (2) corresponds to the flow of HEPES-N buffer at 60 μl/min for 300 s. The data were fit by global analysis to 1:1 Langmuir binding isotherm. Signals from the control surface have been subtracted.
discussion
A redox mechanism for the regulation of PutA intracellular location and function was first proposed by Wood (10). In our study, two significant redox-linked structural changes of the FAD cofactor were identified in the dithionite reduced crystal structure of PutA86-669. First, the reduced FAD cofactor adopts a highly nonplanar conformation with a bending angle about the N(5)-N(10) axis of 22°. Second, the 2′-OH ribityl group forms new hydrogen bonds to the N(1) position of the isoalloxazine ring, the ribityl 4′-OH and Gly435, resulting in disruption of the R556-N(2)H-O(2′)-FAD hydrogen bond that is observed in all oxidized PutA86-669 crystal structures solved to date. Also, in the structure of reduced FAD there is decreased contact surface area between Val433 and the re face of the FAD cofactor and consequently less support from β-strand 5. We hypothesize that these structural changes occur during proline reduction of FAD in full-length PutA and have a fundamental role in the functional switching mechanism of PutA. Because the three-dimensional structure of full-length PutA from E. coli is not yet known, insights into how reduction of the FAD cofactor may affect the architectural arrangement of the different PutA domains are limited at this time.
The nonplanar structure of the reduced FAD cofactor in PutA is commonly called the “butterfly” conformation. The magnitude of the bend in the pyrazine ring of dithionite treated PutA86-669 is among the largest observed in reduced flavins with the greatest bending angle for reduced flavin being reported at 34° in E. coli thioredoxin reductase (31). The energetically favored bending angle in reduced neutral flavin has been estimated at 15° and 27°, thus, the bend observed in the reduced FAD cofactor of PutA appears to be within an energetically optimized range (33-35). Previous electrochemical studies of PutA, however, indicate reduced FAD is in the anionic form (27). Theoretical studies based on models have suggested that the anionic form of reduced flavin is less energetically favored to adopt a nonplanar conformation than neutral reduced flavin (35). Therefore, the intramolecular FAD 2′-OH-N(1) hydrogen bond in the reduced state of PutA may help stabilize the nonplanar conformation.
The second dominant feature in the active site of reduced PutA86-669 involves the formation of a new hydrogen bond network anchored by the FAD 2′-OH ribityl. The FAD 2′-OH ribityl group is likely a key component in the functional switching mechanism of PutA as indicated by our SPR study of 2′-deoxy-FAD PutA and PutA R556M. Since both proteins lack the 2′-OH-Arg556 hydrogen bond and bind lipids in the oxidized state, the 2′-OH-Arg556 hydrogen bond functions as a constraint that prevents oxidized PutA from binding to the membrane. But this interaction apparently is not part of the redoxactivated molecular switch that drives tight PutA-membrane associations. R556M, which retains the 2′-OH ribityl hydrogen bond network of the reduced enzyme (Figure 3), forms a tight protein-lipid complex (KD < 0.01 nM) in the presence of dithionite (Figure 6C). In contrast, 2′-deoxy-FAD PutA does not exhibit membrane-binding in the presence of dithionite. Therefore, the hydrogen bonding formed between FAD-2′-OH-N(1), 4′-OH, and Gly435 in the reduced FAD cofactor must be critical for inducing the tight membrane-binding observed with reduced PutA.
Our structural and biochemical data suggest a hypothesis for how the 2′-OH ribityl group could serve a dual role in PutA functional switching. As shown in Figure 3, the FAD 2′-OH ribityl group toggles between two secondary structure elements on opposite sides of the barrel, the α8 and α5a helices. In the oxidized enzyme, the 2′-OH ribityl hydrogen bonds to Arg556 of the α8 helix while in the reduced enzyme the 2′-OH ribityl forms a hydrogen bond with the backbone N atom of Gly435 located on a loop between β-strand 5 and the α5a helix. The result is the transfer of a cofactor-protein hydrogen bond from the α8-helix to the β-α5a loop in the reduced state. The α8 and α5a helices are atypical in the β8α8 barrel structure implying a unique function in PutA. Thus, the 2′-OH ribityl hydrogen bond network involving Gly435 may help unleash tight membrane-binding in the reduced state. X-ray crystal structures of various flavoenzymes show that ribityl hydroxyl groups of FAD participate in a variety of structural and catalytically important hydrogen bonds (15, 31, 36-41). Here, we report a new example of the 2′-OH ribityl group acting as a molecular switch that responds to electronic changes in the FAD cofactor to control PutA-membrane binding activity. The unique function of the FAD 2′-OH group uncovered in this study further exemplifies the remarkable ability of the ribityl moiety to play versatile roles in flavoprotein mechanisms.
Unlike the 2′-OH-Arg556 hydrogen bond, the FAD N(5)-Arg431 interaction does not constrain oxidized PutA-membrane binding activity, however, it has a profound impact on the reductive activation of PutA-membrane associations. The FAD N(5)-Arg431 hydrogen bond is clearly pivotal for FAD reduction to trigger membrane binding as reduced 5-deaza-FAD PutA and PutA R431M do not bind lipids. Furthermore, in vivo transcription assays with cells containing wild-type PutA show activation of lacZ expression by proline while in cells containing PutA R431M, no proline-dependent increase in β-galactosidase activity is observed. Together, the SPR and reporter gene assays demonstrate that the FAD N(5)-Arg431 hydrogen bond has a critical role in the functional switching mechanism of PutA which hinges on PutA-membrane associations. We presently are not able to speculate how signals are transmitted beyond Arg431 to membrane-binding residues since the PutA-membrane binding domain is not known. Molecular dissection of PutA, however, suggests a membrane-binding domain is located in the C-terminal region of PutA (unpublished data). Whether reductive activation of PutA-membrane binding involves changes in the FAD-N(5)-Arg431 interaction is also uncertain. Reduction of the PutA86-669 crystal elicited changes exclusively in the FAD cofactor, with a similar distance (3.1 – 3.2 Å) between the FAD N(5) and Arg431 NH(1) atom in the oxidized and reduced enzymes. Another caveat is that we have only assessed structural differences between ligand-bound oxidized PutA and reduced PutA. SPR has shown ligand-bound oxidized PutA exhibits lipid binding that is > 300-fold weaker than reduced PutA (13). Figure 5 indicates that the membrane-binding induced by nonreducing ligands (e.g., L-THFA) is not adequate for relieving PutA repression of the lacZ reporter gene. Consequently, the ligand-bound oxidized PutA86-669 structure may represent an intermediate conformation en route from the “off” to the “fully on” reduced membrane-binding state. Even so, this study has identified key FAD interactions involved in the reductive activation of PutA-membrane binding activity.
Supplementary Material
ACKNOWLEDGMENTS
We thank Stephan L. Ginell for assistance with data collection at APS beamline 19-ID. Use of the Argonne National Laboratory Structural Biology Center beamlines at the Advanced Photon Source was supported by the U. S. Department of Energy, Office of Biological and Environmental Research, under Contract No. W-31-109-ENG-38.
ABBREVIATIONS
- FAD
flavin adenine dinucleotide
- PutA
proline utilization A
- PRODH
proline dehydrogenase
- SPR
surface plasmon resonance
- HEPES
N-(2-hydroxyethyl)piperazine-N′-2-ethanesulfonic acid
- L-THFA
L-tetrahydro-2-furoic acid
- Em
reduction potential
Footnotes
SUPPORTING INFORMATION AVAILABLE
Table S1 and Figures S1-S6. This material is available free of charge via the Internet at http://pubs.acs.org.
This research was supported by NIH Grants GM061068 (DFB) and GM065546 (JJT) and the University of Nebraska Agricultural Research Division, supported in part by funds provided through the Hatch Act. This publication was also made possible by NIH Grant Number P20 RR-017675-02 from the National Center for Research Resources.
REFERENCES
- 1.Brown E, Wood JM. Redesigned purification yields a fully functional PutA protein dimer from Escherichia coli. J. Biol. Chem. 1992;267:13086–13092. [PubMed] [Google Scholar]
- 2.Menzel R, Roth J. Enzymatic properties of the purified putA protein from Salmonella typhimurium. J. Biol. Chem. 1981;256:9762–9766. [PubMed] [Google Scholar]
- 3.Menzel R, Roth J. Regulation of genes for proline utilization in Salmonella typhimurium: autogenous repression by the putA gene product. J. Mol. Biol. 1981;148:21–44. doi: 10.1016/0022-2836(81)90233-3. [DOI] [PubMed] [Google Scholar]
- 4.Vílchez S, Manzanera M, Ramos J. Control of expression of divergent Pseudomonas putida put promoters for proline catabolism. Appl. Environ. Microbiol. 2000;66:5221–5225. doi: 10.1128/aem.66.12.5221-5225.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lee YH, Nadaraia S, Gu D, Becker DF, Tanner JJ. Structure of the proline dehydrogenase domain of the multifunctional PutA flavoprotein. Nat. Struct. Biol. 2003;10:109–114. doi: 10.1038/nsb885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ling M, Allen SW, Wood JM. Sequence analysis identifies the proline dehydrogenase and pyrroline-5-carboxylate dehydrogenase domains of the multifunctional Escherichia coli PutA protein. J. Mol. Biol. 1994;245:950–956. doi: 10.1006/jmbi.1994.1696. [DOI] [PubMed] [Google Scholar]
- 7.Gu D, Zhou Y, Kallhoff V, Baban B, Tanner JJ, Becker DF. Identification and characterization of the DNA-binding domain of the multifunctional PutA flavoenzyme. J. Biol. Chem. 2004;279:31171–31176. doi: 10.1074/jbc.M403701200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen C-C, Wilson TH. Solubilization and functional reconstitution of the proline transport system of Escherichia coli. J. Biol. Chem. 1986;261:2599–2604. [PubMed] [Google Scholar]
- 9.Larson JD, Jenkins JL, Schuermann JP, Zhou Y, Becker DF, Tanner JJ. Crystal structures of the DNA-binding domain of Escherichia coli proline utilization A flavoprotein and analysis of the role of Lys9 in DNA recognition. Protein Sci. 2006;15:2630–41. doi: 10.1110/ps.062425706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wood J. Membrane association of proline dehydrogenase in Escherichia coli is redox dependent. Proc. Natl. Acad. Sci. USA. 1987;84:373–377. doi: 10.1073/pnas.84.2.373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Brown ED, Wood JM. Conformational change and membrane association of the PutA protein are coincident with reduction of its FAD cofactor by proline. J. Biol. Chem. 1993;268:8972–8979. [PubMed] [Google Scholar]
- 12.Surber MW, Maloy S. Regulation of flavin dehydrogenase compartmentalization: requirements for PutA-membrane association in Salmonella typhimurium. Biochim. Biophys. Acta. 1999;1421:5–18. doi: 10.1016/s0005-2736(99)00104-2. [DOI] [PubMed] [Google Scholar]
- 13.Zhang W, Zhou Y, Becker DF. Regulation of PutA-membrane associations by flavin adenine dinucleotide reduction. Biochemistry. 2004;43:13165–13174. doi: 10.1021/bi048596g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Muro-Pastor AM, Ostrovsky P, Maloy S. Regulation of gene expression by repressor localization: Biochemical evidence that membrane and DNA binding by the PutA protein are mutually exclusive. J. Bacteriol. 1997;179:2788–2791. doi: 10.1128/jb.179.8.2788-2791.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhang M, White TA, Schuermann JP, Baban BA, Becker DF, Tanner JJ. Structures of the Escherichia coli PutA proline dehydrogenase domain in complex with competitive inhibitors. Biochemistry. 2004;43:12539–12548. doi: 10.1021/bi048737e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Nakagawa S, Hagihara T, Fujio T, Aisaka K. Metaphosphate-dependent phosphorylation of riboflavin to FMN by Corynebacterium ammoniagenes. Appl. Microbiol. Biotechnol. 1995;43:325–329. [Google Scholar]
- 17.Efimov I, Kuusk V, Zhang X, McIntire WS. Proposed steady-state kinetic mechanism for Corynebacterium ammoniagenes FAD synthetase produced by Escherichia coli. Biochemistry. 1998;37:9716–23. doi: 10.1021/bi972817j. [DOI] [PubMed] [Google Scholar]
- 18.Zhu W, Becker DF. Flavin redox state triggers conformational changes in the PutA protein from Escherichia coli. Biochemistry. 2003;42:5469–5477. doi: 10.1021/bi0272196. [DOI] [PubMed] [Google Scholar]
- 19.Zhang W, Krishnan N, Becker DF. Kinetic and thermodynamic analysis of Bradyrhizobium japonicum PutA-membrane associations. Arch. Biochem. Biophys. 2006;445:174–83. doi: 10.1016/j.abb.2005.10.022. [DOI] [PubMed] [Google Scholar]
- 20.Zhang M, Tanner JJ. Detection of L-lactate in polyethylene glycol solutions confirms the identity of the active-site ligand in a proline dehydrogenase structure. Acta Crystallogr. D Biol. Crystallogr. 2004;60:985–6. doi: 10.1107/S0907444904003786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 22.Brünger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang JS, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. 1998;D54:905–921. doi: 10.1107/s0907444998003254. [DOI] [PubMed] [Google Scholar]
- 23.Winn MD, Isupov MN, Murshudov GN. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 2001;57:122–33. doi: 10.1107/s0907444900014736. [DOI] [PubMed] [Google Scholar]
- 24.Jones TA, Zou J-Y, Cowan SW, Kjeldgaard M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. 1991;A47:110–119. doi: 10.1107/s0108767390010224. [DOI] [PubMed] [Google Scholar]
- 25.Zhu W, Becker DF. Exploring the proline-dependent conformational change in the multifunctional PutA flavoprotein by tryptophan fluorescence spectroscopy. Biochemistry. 2005;44:12297–12306. doi: 10.1021/bi051026b. [DOI] [PubMed] [Google Scholar]
- 26.Becker DF, Thomas EA. Redox properties of the PutA protein from Escherichia coli and the influence of the flavin redox state on PutA-DNA interactions. Biochemistry. 2001;40:4714–4722. doi: 10.1021/bi0019491. [DOI] [PubMed] [Google Scholar]
- 27.Vinod MP, Bellur P, Becker DF. Electrochemical and functional characterization of the proline dehydrogenase domain of the PutA flavoprotein from Escherichia coli. Biochemistry. 2002;41:6525–6532. doi: 10.1021/bi025706f. [DOI] [PubMed] [Google Scholar]
- 28.Baban BA, Vinod MP, Tanner JJ, Becker DF. Probing a hydrogen bond pair and the FAD redox properties in the proline dehydrogenase domain of Escherichia coli PutA. Biochem. Biophys. Acta. 2004;1701:49–59. doi: 10.1016/j.bbapap.2004.06.001. [DOI] [PubMed] [Google Scholar]
- 29.Stankovich MT. An anaerobic spectroelectrochemical cell for studying the spectral and redox properties of flavoproteins. Anal. Biochem. 1980;109:295–308. doi: 10.1016/0003-2697(80)90652-1. [DOI] [PubMed] [Google Scholar]
- 30.Erb EM, Chen X, Allen S, Roberts CJ, Tendler SJB, Davies MC, Forsen S. Characterization of the surfaces generated by liposome binding to the modified dextran matrix of a surface plasmon resonance sensor chip. Anal. Biochem. 2000;280:29–35. doi: 10.1006/abio.1999.4469. [DOI] [PubMed] [Google Scholar]
- 31.Lennon BW, Williams CH, Jr., Ludwig ML. Crystal structure of reduced thioredoxin reductase from Escherichia coli: structural flexibility in the isoalloxazine ring of the flavin adenine dinucleotide cofactor. Protein Sci. 1999;8:2366–79. doi: 10.1110/ps.8.11.2366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Murthy YV, Massey V. Chemical modification of the N-10 ribityl side chain of flavins. Effects on properties of flavoprotein disulfide oxidoreductases. J. Biol. Chem. 1995;270:28586–94. doi: 10.1074/jbc.270.48.28586. [DOI] [PubMed] [Google Scholar]
- 33.Dixon DA, Lindner DL, Branchaud B, Lipscomb WN. Conformations and electronic structures of oxidized and reduced isoalloxazine. Biochemistry. 1979;18:5770–5. doi: 10.1021/bi00593a004. [DOI] [PubMed] [Google Scholar]
- 34.Moonen CT, Vervoort J, Muller F. Carbon-13 nuclear magnetic resonance study on the dynamics of the conformation of reduced flavin. Biochemistry. 1984;23:4868–72. doi: 10.1021/bi00316a008. [DOI] [PubMed] [Google Scholar]
- 35.Zheng YJ, Ornstein RL. A theoretical study of the structures of flavin in different oxidation and protonation states. J. Am. Chem. Soc. 1996;118:9402–9408. [Google Scholar]
- 36.Karplus PA, Schulz GE. Substrate binding and catalysis by glutathione reductase as derived from refined enzyme: substrate crystal structures at 2 Å resolution. J. Mol. Biol. 1989;210:163–80. doi: 10.1016/0022-2836(89)90298-2. [DOI] [PubMed] [Google Scholar]
- 37.Mattevi A, Schierbeek AJ, Hol WG. Refined crystal structure of lipoamide dehydrogenase from Azotobacter vinelandii at 2.2 Å resolution. A comparison with the structure of glutathione reductase. J. Mol. Biol. 1991;220:975–94. doi: 10.1016/0022-2836(91)90367-f. [DOI] [PubMed] [Google Scholar]
- 38.Kim JJ, Wu J. Structure of the medium-chain acyl-CoA dehydrogenase from pig liver mitochondria at 3-Å resolution. Proc. Natl. Acad. Sci. U S A. 1988;85:6677–81. doi: 10.1073/pnas.85.18.6677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kim JJ, Wang M, Paschke R. Crystal structures of medium-chain acyl-CoA dehydrogenase from pig liver mitochondria with and without substrate. Proc. Natl. Acad. Sci. U S A. 1993;90:7523–7. doi: 10.1073/pnas.90.16.7523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Roberts DL, Frerman FE, Kim JJ. Three-dimensional structure of human electron transfer flavoprotein to 2.1-Å resolution. Proc. Natl. Acad. Sci. U S A. 1996;93:14355–60. doi: 10.1073/pnas.93.25.14355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Engst S, Vock P, Wang M, Kim JJ, Ghisla S. Mechanism of activation of acyl-CoA substrates by medium chain acyl-CoA dehydrogenase: interaction of the thioester carbonyl with the flavin adenine dinucleotide ribityl side chain. Biochemistry. 1999;38:257–67. doi: 10.1021/bi9815041. [DOI] [PubMed] [Google Scholar]
- 42.Engh RA, Huber R. Accurate bond and angle parameters for X-ray protein structure refinement. Acta Crystallogr. 1991;A47:392–400. [Google Scholar]
- 43.Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 1993;26:283–291. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.