Abstract
Prion infection is characterized by the conversion of host cellular prion protein (PrPC) into disease-related conformers (PrPSc) and can be arrested in vivo by passive immunization with anti-PrP monoclonal antibodies. Here, we show that the ability of an antibody to cure prion-infected cells correlates with its binding affinity for PrPC rather than PrPSc. We have visualized this interaction at the molecular level by determining the crystal structure of human PrP bound to the Fab fragment of monoclonal antibody ICSM 18, which has the highest affinity for PrPC and the highest therapeutic potency in vitro and in vivo. In this crystal structure, human PrP is observed in its native PrPC conformation. Interactions between neighboring PrP molecules in the crystal structure are mediated by close homotypic contacts between residues at position 129 that lead to the formation of a 4-strand intermolecular β-sheet. The importance of this residue in mediating protein–protein contact could explain the genetic susceptibility and prion strain selection determined by polymorphic residue 129 in human prion disease, one of the strongest common susceptibility polymorphisms known in any human disease.
Keywords: Creutzfeldt–Jakob disease, PrP–Fab complex, monoclonal antibody, prion therapeutics
Prion diseases are associated with the accumulation in the brain of a misfolded protease-resistant glycoprotein known as PrPSc. The native cellular form of this protein (PrPC) is ubiquitously expressed at high levels in the central nervous system, lymphoreticular tissue, and at neuromuscular junctions, and is tethered at the cell surface by a glycosylphosphatidylinositol (GPI) anchor (1). The disease-related PrPSc is derived from PrPC by posttranslational processes involving conformational change and aggregation. According to the “protein-only” hypothesis (2), PrPSc is propagated by serving as a template for the autocatalytic recruitment of PrPC and is the principal, possibly the sole, constituent of the transmissible agent or prion (3). Susceptibility to human prion diseases depends on the expression of PrPC (4–6), and it has been shown that targeting PrPC is an effective therapeutic strategy. In animal models of established neuroinvasive prion disease, abrogation of neuronal PrPC synthesis, using transgenic manipulation (7, 8) or RNA interference (9), can prevent clinical disease onset and reverse early pathological changes. Hence, it is reasonable to focus on the PrPC form owing to its role as the well-characterized “substrate” for prion propagation. Moreover, the fact that PrPC is converted to a radically different conformation during prion propagation means that any molecule that binds specifically to PrPC must stabilize the native fold, inhibiting conformation change and, consequently, prion propagation (10, 11) without the requirement for interference in PrP synthesis. We would therefore expect that antibodies specifically binding to PrPC should inhibit prion propagation, with efficacies correlating to their PrPC binding affinity.
Here, we show that, for a selection of anti-PrP antibodies, there is such a correlation between affinity for PrPC and therapeutic potency in a cell model of prion infection. We also present the crystal structure of PrP in complex with the most effective therapeutic antibody, which gives us detailed information on the intermolecular interactions both between antibody and PrP and between PrP molecules.
Results
Antibodies with the Highest Affinity for PrPC Are the Most Therapeutically Active.
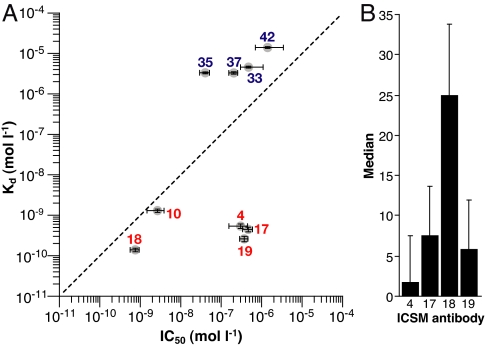
The ICSM monoclonal antibodies we used were raised to either α-PrP or β-PrP, 2 different conformations of recombinant PrP [see supporting information (SI) Table S1]. The α-PrP species is formed by oxidative refolding of recombinant PrP and has the same, predominantly α-helical conformation as PrPC, whereas the β-PrP species is a reduced and acidified form of the recombinant protein with features that resemble PrPSc, including a high content of β-sheet and propensity to aggregate (12, 13). The α and β conformations elicit different immune responses in mice (14), and, whereas antibodies raised against both recombinant species generally recognize both PrPC and denatured PrPSc, only those raised against β-PrP can detect native PrPSc (15). To test the hypothesis that antibodies specifically binding to PrPC inhibit prion propagation with efficacies that quantitatively correlate with their PrPC binding affinity, we examined the relationship between dissociation constants for α-PrP (as measured by ELISA) and IC50s for treatment of prion-infected ScN2a cells for a series of ICSM antibodies (Fig. 1A; see also Fig. S1 and Table S1). As expected, the monoclonal antibodies raised against α-PrP (ICSMs 4, 10, 17, 18, and 19) interact strongly with recombinant α-PrP, with Kd values in the 0.1–10 nM range, whereas those raised against β-PrP (ICSMs 33, 35, 37, and 42) have affinities some 4 orders of magnitude weaker, with Kd values typically in the 10 μM range. These antibodies raised against β-PrP are also ineffective in halting prion propagation, implying that targeting this PrPSc-like conformation is not an effective way of curing the infection.
Fig. 1.
The relationship between antibody affinity and ability to inhibit prion propagation for the ICSM antibodies. (A) IC50s (M) for 3-day treatment of chronically infected N2a cells with antibody are plotted against Kd (M) for recognition of recombinant PrP by antibody as measured by ELISA (Fig. S1 and Table S1). Antibodies raised to α-PrP are shown in red and to β-PrP in blue. Equivalence between IC50 and Kd is shown by the dashed diagonal line. (B) Median count of FACS measurements (n = 3) for the antibodies indicated, detected with FITC-conjugated anti-mouse secondary antibody on NS0 mouse cells.
Superimposed on the graph in Fig. 1A is the trend line that represents a quantitative equivalence between the affinity for α-PrP and the IC50 for inhibiting prion propagation. The data for most of the antibodies lie reasonably close to this line, supporting the contention that the ability to inhibit PrPSc propagation is correlated with binding affinity for the PrPC-type conformation. The antibodies ICSM 4, 17, and 19, however, clearly form a subgroup that does not follow this trend, i.e., despite their high affinity for PrP, they are poor inhibitors of PrPSc propagation. In one case, the explanation is straightforward; the ICSM 4 epitope on PrP spans the sites of N-glycosylation on mature PrPC (15), consequently ICSM 4 binds tightly only to the unglycosylated protein (16). Also, data presented by Leclerc et al. (17) on mature PrPC show that an epitope near to the recognition sites for ICSM 17 and 19 becomes inaccessible when the protein is at the cell surface.
To test for accessibility in situ, we used flow cytometry to probe the affinities of this subset of antibodies for mature, cell-surface PrPC rather than recombinant PrP using ICSM 18 as a positive control (Fig. 1B). ICSMs 4, 17, and 19 gave weak signals in the FACS analysis compared with ICSM 18, indicating that their interaction is impaired and that the ability to recognize cell-surface PrPC is important for inhibiting PrPSc formation in the cell system. The lower affinities of ICSMs 17 and 19 for mature PrPC must result from the occlusion of their binding epitopes in mature membrane-associated PrPC, as has been seen previously (17) and attributed to interaction with other ligands or to dimerization.
ICSM 18 was our most potent antibody, both as an inhibitor of prion propagation (IC50 0.7 nM) and in its affinity for PrPC (Kd 0.1 nM). Significantly, this antibody is highly effective in treating prion-infected mice (18). In mice treated with ICSM 18, peripheral PrPSc levels and prion infectivity were markedly reduced, even when the antibody was administered as PrPSc reached maximal levels in the spleen. Furthermore, animals in which the treatment was continued remained healthy for >300 days after untreated animals succumbed to disease (18).
Crystal Structure of PrP in Complex with ICSM 18-Fab.
We visualized the interaction between PrP and ICSM 18 by determining the structure of truncated human PrP119-231 complexed with a Fab fragment of the antibody to 2.9 Å resolution (Fig. 2 and Table 1; see also SI Text and Fig. S2 for the quality of the electron density of the structure). The overall fold of the human PrP globular domain in the complex reported here is similar to that of the C-terminal domain of human PrP in the NMR structure (19) but is different from that in the domain-swapped dimer (20), where helix 3 is swapped between the 2 molecules of the dimer, and an intramolecular disulfide bridge (Cys-179-Cys-214) is formed between helix 2 (residues 172–179) and helix 3 (200–223).
Fig. 2.
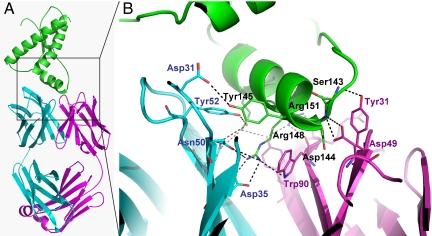
The complex between recombinant PrP119-231 and the ICSM 18-Fab as determined by X-ray crystallography. (A) PrP119-231 is shown in green with the heavy and light chains of the Fab in cyan and magenta, respectively. (B) Expanded view of the PrP/Fab interface. The participating PrP residues are labeled in black and those from the Fab heavy and light chains in blue and magenta, respectively. Potential hydrogen bonds are shown as dashed lines.
Table 1.
Summary of crystallographic data collection and refinement statistics
| FAB18 | FAB18-PrP | |
|---|---|---|
| Space group | P212121 | P6322 |
| Resolution, Å | 22–1.57 (1.63–1.57) | 42–2.9 (3.0–2.9) |
| Completeness, % | 95 (70.0) | 97.7 (83.9) |
| R merge | 7.8 (38.3) | 15.6 (79.2) |
| 〈I〉/σI last shell | 2.35 | 2.0 |
| Redundancy | 6.7 (3.1) | 12.6 (6.9) |
| Overall reflections | 1,080,709 | 482,099 |
| Unique reflections | 54,085 | 14,267 |
| Wilson B-factor, Å2 | 17.8 | 62 |
| Unit cell (Å, °) | ||
| a, Å | 36.7 | 126.1 |
| b, Å | 84.7 | 126.1 |
| c, Å | 127.8 | 134.1 |
| α = β, ° | 90 | 90 |
| γ, ° | 90 | 120 |
| Solvent, % | 47.0 | 46.0 |
| No. of FAB molecules | 1 | 1 |
| No. of PrP molecules | 1 | |
| Final R crystal, % | 18.3 | 21.0 |
| R-free, % | 21.7 | 26.9 |
| ESU, Å, based on R value | 0.094 | 1.4 |
| No. of all atoms | 4,083 | 4,185 |
| No. of water molecules | 728 | 55 |
| No. of calcium ions | 1 | |
| No. of sulfate ions | 1 | |
| Average B-factor, Å2 | 18.3 | 40.0 |
The ICSM 18 epitope is formed by residues spanning the whole of helix 1 (H1, residues 143–156), confirming previous epitope mapping experiments (14). The Fab–PrP interface buries ≈900 Å2 (15%) of the solvent accessible surface area of PrP, and there are several hydrogen bonds and salt bridges between PrP and both the light (L) and heavy (H) chains of the variable domain of the Fab molecule (Fig. 2B and Table 2). H1 has been proposed as a site for β-sheet transformation that may promote PrPSc formation (21, 22) and mutational analysis of PrP in cell-free conversion assays highlight this helix as the initiation site for the conversion of PrPC to the proteinase resistant form (23). The extensive contacts observed in this crystal structure would provide a significant stabilizing effect on the helix and would restrict its involvement in secondary-structure changes. Indeed, the average temperature factor for H1 (36 Å2) in the complex suggests that intermolecular contacts stabilize this region of PrP relative to the overall structure (40 Å2; see Fig. S3), but it is worthy of note that this region is well defined in the majority of PrPC structures derived from a range of species (19, 24, 25). Conversely, the most disordered part of the molecule is at the C-terminal end of H2, the N-terminal end of H3, and the short loop connecting them (residues 188–201, average B-factor 56 Å2, Figs. S2 and S3). This disordered segment of the human PrPC structure has also been suggested as a possible site for propagating α → β transitions that promote PrPSc formation (20, 25, 26). In our complex, part of this segment, extending from Asn-197 to Met-205, creates an interface with the H chain of Fab that buries 140 Å2 of the PrP surface and 200 Å2 of the Fab surface. The physical proximity of the Fab H chain to a region of PrP that is a possible site for β-sheet formation suggests that the complex may provide structural inhibition by burying the “active” residues at this interface. Structural changes to the Fab molecule upon PrP binding are shown in Fig. S4.
Table 2.
Summary of close contacts between PrP and ICSM 18-Fab in the crystal structure
| PrP | FAB18 light chain | Distance, Å | PrP | FAB18 heavy chain | Distance, Å |
|---|---|---|---|---|---|
| Ser143 N | Tyr31 OH | 3.4 | Tyr145 OH | Asp31 OD2 | 3.4 |
| Asp144 N | Asp49 OD1 | 3.2 | Arg148 NH2 | Asp35 OD1 | 2.7 |
| Asp144 OD2 | His33 NE2 | 3.3 | Arg148 NH1 | Asp35 OD2 | 3.2 |
| Arg151 NH2 | Tyr95 OH | 3.1 | Glu152 OE1 | Tyr52 OH | 2.7 |
| Arg151 NH1 | Arg91 O | 3.2 | Glu152 OE2 | Asn50 OD1 | 3.3 |
| Arg151 NH2 | Trp90 O | 2.9 | Lys204 NZ | Tyr101 OH | 3.6 |
Discussion
Many anti-PrP monoclonal antibodies are therapeutically active in cellular models of prion propagation (27–35) and in animals (18, 36, 37). Our results indicate a clear correlation between ability to inhibit PrPSc propagation and binding affinity for a PrPC-type conformation for therapeutic antibodies. This is consistent with observations that antibody efficacy is determined by cell-surface PrP recognition (32) and retention (31) but rather contrary to interpretations that efficacy is primarily determined by antibody recognition epitopes (28) or their ability to recognize PrPSc as well as PrPC (33). Our most potent antibody in both prion inhibition and PrPC affinity, ICSM 18, is also highly effective in treating prion-infected mice (18), and we have no evidence that the interaction has any distorting effect on the PrPC conformation as has been observed in some other cases (38, 39). The therapeutic value of targeting PrPC is further underlined by conditional knockout experiments showing that its abrogation has little detrimental effect on the adult animal (40).
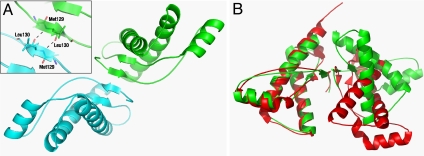
Prion propagation requires contact between PrP molecules, and in this respect, it is interesting to note that symmetry-related PrP molecules in the crystal interact by forming a 4-stranded antiparallel β-sheet (Fig. 3A) made up of the existing 2-stranded β-sheets of each PrP molecule, residues 129–131 and 161–163. An identical interaction was observed in the crystal structure of sheep PrP obtained in the absence of antibody and in a different crystallographic space group (25), which is shown overlaid with our human PrP structure in Fig. 3B. The coincidence of β-sheet structure between PrP molecules in different crystal structures suggests that it is not an artifact of crystal packing but might have greater biological significance for prion propagation, especially in view of the powerful genetic susceptibility and prion strain selection determined by polymorphic residue 129 in human prion disease (41). The common human methionine–valine polymorphism at residue 129 has a profound influence on prion pathogenesis; for instance, heterozygosity at PRNP codon 129 is highly protective against sporadic and acquired prion diseases in humans (42–44), delays age of onset of some inherited prion diseases (45), and appears to have been selected during the evolution of modern humans (46). Residue 129 also plays a key role in conformational selection of prion strains (41, 47). The bovine spongioform encephalopathy (BSE)-like strain causing vCJD appears incompatible with the valine isoform of human PrP (48); to date vCJD has occurred exclusively in individuals homozygous for methionine at PRNP codon 129 (49, 50). Moreover, the controlling influence of this residue is not limited to humans; polymorphism at the corresponding codon protects against chronic wasting disease, a prion disease of cervids (51, 52).
Fig. 3.
The interaction of PrP chains in the crystal. (A) Illustration of the intermolecular 4-stranded antiparallel β-sheet formed between neighboring PrP chains (in cyan and green) emphasizing residue 129 at the molecular interface (see Inset). (B) Superimposition of the ovine [red (25)] and human (green) PrP dimers from the respective crystal structures. Note the common occurrence of the 4-stranded intermolecular β-sheet.
Interestingly, the stability and physical properties of PrPC are not influenced by the residue 129 variation (53), so its effect must be realized in the physical properties and propagation of PrPSc. The only compelling explanation for the effects of this polymorphism is that the 129 region of the molecule is critical in determining the fold and the packing of protein chains in prion particles. Heterozygosity is protective owing to the requirement for sequence homogeneity in forming ordered, self-replicating particles (43) (analogous to protein crystallization), and prion strain susceptibility arises from the ease by which the methionine or the valine sequence adopts a given prion structure. The presence—in 2 completely different PrP crystal structures—of the same intermolecular 4-stranded sheet supports our proposal that such a structure is present in PrPSc and is propagated and stabilized by stacking to form the type of antiparallel zipper interactions that have been experimentally shown by Eisenberg and colleagues (54) to form cross-β amyloid spines. However, we point out that the formation of this β-sheet interface is not the only event necessary for PrPSc formation; it must also be coupled to gross rearrangement of the structure of component monomers (see Fig. S5). In this regard, it should be noted that antiparallel interactions are found in fibrils formed from short peptides (54) rather than those formed from proteins that, thus far, have been shown to be parallel (55). However, in some denaturing conditions large proteins have been seen to aggregate initially by antiparallel interactions that slowly rearrange to parallel on maturation (56). Nonetheless, if β-strand interactions in this region mediate PrP interactions during PrPSc propagation, then it is reasonable to propose that the exact geometry and packing of this segment of the chain varies between strains and that the nature of the 129 side chain has a strong selective effect on conformation. Such a mechanism would explain the potent effects of this human polymorphism at the level of fine structure within a prion particle.
Materials and Methods
Recombinant PrP and Antibodies.
Recombinant human PrP containing residues 91–231 (PrP91-231) or 119–231 (PrP119-231) was produced and purified as described (12). The ICSM antibodies were raised to recombinant PrP in either α- or β-conformation (12, 13), as listed in Table S1. The antibodies were characterized as previously described (14, 15) and were the gift of D-Gen Limited.
Determination of Antibody–PrP Affinities by ELISA.
Recombinant α-PrP91-231 [2.5 μg/mL in bicarbonate buffer (pH 9.6)] was immobilized in 96-well ELISA plates. After washes and blocking with RF10/Tween-20, the plates were incubated with serial dilutions (0.1 mM to 10 μM) of the ICSM antibodies, HRP-conjugated goat anti-mouse secondary antibody (Sigma) and substrate TMB (Sigma). The color reaction was stopped by addition of 3 M sulfuric acid before reading intensities at 450 nm. Intensities were averaged across replicates (n = 3) and fitted to a single site binding curve to derive Kds by using Grafit5 (Erithacus Software Ltd); see Fig. S1A.
Determination of IC50s for ICSM-Treated Cells.
Scrapie susceptible N2a cells (subclone “PK1”) (57) were cultured in OPTIMEM (Invitrogen) supplemented with 10% FCS (Perbio) and penicillin/streptomycin (Invitrogen). Cells chronically infected with RML prions were cultivated in 24-well plates with 1 mL per well media supplemented with ICSM antibodies at various concentrations for 3 days before assay for levels of proteinase K-resistant PrP. Briefly, cells were lysed in ice-cold lysis buffer, blotted onto nitrocellulose membrane in a BioRad ELIFA apparatus, treated in situ with proteinase K (10 μg/mL for 1 h at 37 °C) and denaturant (3 M guadinidium thiocyanate) before immunodetection using ICSM 18 (0.2 μg/mL), HRP-conjugated anti-mouse secondary antibody (Sigma), Super Signal West Pico ECL reagent (Pierce), and exposure on X-ray film (Kodak Biomax). PK-resistant signal intensities were quantified by using a Kodak ImageStation 440 CF, and data were averaged across replicates (n = 4) and normalized against 100% intensity in untreated control samples. Half-maximal inhibition constants (IC50) were derived by fitting of the averaged and normalized data by using Grafit5 (Erithacus Software Ltd); see Fig. S1B.
Flow Cytometry.
Suspensions of NS0 cells (2 × 106 cells per milliliter) were incubated with 1–10 μg/mL ICSM antibodies, washed in PBS, and resuspended in FITC-conjugated secondary antibody (Sigma) in PBS. Cells were further washed and resuspended in PBS for FACS measurements by using a FACSCalibur (Becton Dickinson).
Crystallization, Data Collection, Refinement, and Model Building.
The Fab fragment of ICSM 18 was prepared by limited papain digest of the mature antibody and purification by gel filtration (58). Before crystallization, ICSM 18-Fab was additionally purified on a S200 gel filtration column by using 150 mM NaCl and 50 mM Tris (pH 8.0) and concentrated to 100 mg/mL. Crystals of ICSM 18-Fab were obtained by using the sitting-drop vapor diffusion technique; droplets containing 20 mg/mL protein in 0.1 M magnesium chloride, 0.05 M Tris·HCl (pH 8.0), 10% wt/vol PEG3350 were equilibrated over wells containing 0.2 M magnesium chloride, 0.1 M Tris·HCl (pH 8.0), 20% wt/vol PEG3350. Crystals grew in 1 day to 0.2 × 0.05 × 0.05 mm. ICSM 18-Fab and PrP119-231 were mixed at a molar ratio of 1: 1 for preparation of the complex before crystallization. Crystals of the complex were obtained by using the hanging-drop vapor diffusion technique; droplets containing 20 mg/mL protein in 1.0 M ammonium sulfate, 0.05 M Tris (pH 8.0) were equilibrated over wells containing 2.0 M ammonium sulfate, 0.1 M Tris (pH 8.0). Crystals grew in 5 months to 0.1 × 0.05 × 0.05 mm.
Crystallographic Data Collection, Refinement, and Model Building.
Data were collected at 100 K with a MAR225 CCD detector by using beamline 10.1 at the Daresbury Laboratory Synchrotron Radiation Source, U.K. ICSM 18-Fab crystals were picked directly from the hanging drop for data collection, whereas for the ICSM 18-Fab–PrP complex, 3.4 M sodium malonate (pH 7.0) was used as a cryo protector. Data were processed with the program HKL2000 (59). Data collection statistics and cell parameters are summarized in Table 1. Structures were solved by the molecular replacement method by using the programs PHASER (60) for the complex and MOLREP (61) for ICSM 18 (see SI Text for ICSM 18 structure). An anti-PrP Fab structure (PDB ID: 2hh0) (62), was used as a starting model for ICSM 18-Fab. The variable and constant regions of the heavy and light chains were treated as independent rigid bodies before restrained refinement and model building cycles with addition of solvent. Subsequently, the refined structure of ICSM 18-Fab was used with the globular domain of sheep PrP (PDB ID: 1tpx) (21) as the starting point for solving the structure of the complex. All models were refined without NCS restrains by the maximum-likelihood method as implemented in REFMAC5 (63). Overall temperature factor refinement was used for the complex, and TLS refinement (64) was applied to both models, with each domain treated as a separate group, excluding the water molecules. This resulted in a drop in both R and R-free factors of nearly 5%. Model building was done with COOT (65). Solvent molecules were added to the model by using ARP (66), being accepted only when well-defined positive peaks were present in both 2Fo-Fc and Fo-Fc electron density maps and when they could form hydrogen bond(s) with either protein atoms or other water molecules. A summary of the structure refinement is given in Table 1. The final coordinates have been deposited in the Brookhaven Protein Data Bank.
Supplementary Material
Acknowledgments.
We thank our colleagues, particularly Drs. Pat Ridley and Tracy Turner for their encouragement, and Ray Young and Laszlo Hosszu for preparation of figures. The figures based on structural data were prepared by using Pymol (www.pymol.org). We are particularly indebted to Paul Quinn for constant help and support. This work was supported by the U.K. Medical Research Council. The structural work was supported by the Synchrotron Radiation Department at the Science and Technology Facilities Council, Daresbury Laboratory, and X-ray data were collected on beamline 10.1 at the Synchrotron Radiation Source, which was supported by Biotechnology and Biological Sciences Research Council Grant BB/E001971 (to S.S.H. and R.W.S.).
Footnotes
Conflict of interest statement: J.C. is a director and J.C., G.S.J. and A.R.C. are consultants and shareholders of D-Gen Limited, an academic spin-out company in the field of prion disease diagnosis, decontamination, and therapeutics, which markets several of the monoclonal antibodies used in this study.
This article is a PNAS Direct Submission.
Data deposition: The atomic coordinates have been deposited in the Brookhaven Protein Data Bank (accession nos. 2w9d and 2w9e).
This article contains supporting information online at www.pnas.org/cgi/content/full/0809170106/DCSupplemental.
References
- 1.Collinge J. Prion diseases of humans and animals: Their causes and molecular basis. Annu Rev Neurosci. 2001;24:519–550. doi: 10.1146/annurev.neuro.24.1.519. [DOI] [PubMed] [Google Scholar]
- 2.Griffith JS. Self replication and scrapie. Nature. 1967;215:1043–1044. doi: 10.1038/2151043a0. [DOI] [PubMed] [Google Scholar]
- 3.Prusiner SB. Novel proteinaceous infectious particles cause scrapie. Science. 1982;216:136–144. doi: 10.1126/science.6801762. [DOI] [PubMed] [Google Scholar]
- 4.Bueler H, et al. Mice devoid of PrP are resistant to scrapie. Cell. 1993;73:1339–1347. doi: 10.1016/0092-8674(93)90360-3. [DOI] [PubMed] [Google Scholar]
- 5.Sailer A, et al. No propagation of prions in mice devoid of PrP. Cell. 1994;77:967–968. doi: 10.1016/0092-8674(94)90436-7. [DOI] [PubMed] [Google Scholar]
- 6.Manson JC, et al. PrP gene dosage determines the timing but not the final intensity or distribution of lesions in scrapie pathology. Neurodegeneration. 1994;3:331–340. [PubMed] [Google Scholar]
- 7.Mallucci G, et al. Depleting neuronal PrP in prion infection prevents disease and reverses spongiosis. Science. 2003;302:871–874. doi: 10.1126/science.1090187. [DOI] [PubMed] [Google Scholar]
- 8.Mallucci G, et al. Targeting cellular prion protein reverses early cognitive deficits and neurophysiological dysfunction in prion-infected mice. Neuron. 2007;53:325–335. doi: 10.1016/j.neuron.2007.01.005. [DOI] [PubMed] [Google Scholar]
- 9.White MD, et al. Single treatment with RNAi against prion protein rescues early neuronal dysfunction and prolongs survival in mice with prion disease. Proc Natl Acad Sci USA. 2008;105:10238–10243. doi: 10.1073/pnas.0802759105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hosszu LLP, et al. Structural mobility of the human prion protein probed by backbone hydrogen exchange. Nat Struct Biol. 1999;6:740–743. doi: 10.1038/11507. [DOI] [PubMed] [Google Scholar]
- 11.Mallucci G, Collinge J. Rational targeting for prion therapeutics. Nat Rev Neurosci. 2005;6:23–34. doi: 10.1038/nrn1584. [DOI] [PubMed] [Google Scholar]
- 12.Jackson GS, et al. Multiple folding pathways for heterologously expressed human prion protein. Biochim Biophys Acta. 1999;1431:1–13. doi: 10.1016/s0167-4838(99)00038-2. [DOI] [PubMed] [Google Scholar]
- 13.Jackson GS, et al. Reversible conversion of monomeric human prion protein between native and fibrilogenic conformations. Science. 1999;283:1935–1937. doi: 10.1126/science.283.5409.1935. [DOI] [PubMed] [Google Scholar]
- 14.Khalili-Shirazi A, et al. Protein conformation significantly influences immune responses to prion protein. J Immunol. 2005;174:3256–3263. doi: 10.4049/jimmunol.174.6.3256. [DOI] [PubMed] [Google Scholar]
- 15.Khalili-Shirazi A, et al. Beta-PrP form of human prion protein stimulates production of monoclonal antibodies to epitope 91–110 that recognise native PrP(Sc) Biochim Biophys Acta. 2007;1774:1438–1450. doi: 10.1016/j.bbapap.2007.08.028. [DOI] [PubMed] [Google Scholar]
- 16.Khalili-Shirazi A, et al. PrP glycoforms are associated in a strain-specific ratio in native PrPSc. J Gen Virol. 2005;86:2635–2644. doi: 10.1099/vir.0.80375-0. [DOI] [PubMed] [Google Scholar]
- 17.Leclerc E, et al. Conformation of PrP(C) on the cell surface as probed by antibodies. J Mol Biol. 2003;326:475–483. doi: 10.1016/s0022-2836(02)01365-7. [DOI] [PubMed] [Google Scholar]
- 18.White AR, et al. Monoclonal antibodies inhibit prion replication and delay the development of prion disease. Nature. 2003;422:80–83. doi: 10.1038/nature01457. [DOI] [PubMed] [Google Scholar]
- 19.Zahn R, et al. NMR solution structure of the human prion protein. Proc Natl Acad Sci USA. 2000;97:145–150. doi: 10.1073/pnas.97.1.145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Knaus KJ, et al. Crystal structure of the human prion protein reveals a mechanism for oligomerization. Nat Struct Biol. 2001;8:770–774. doi: 10.1038/nsb0901-770. [DOI] [PubMed] [Google Scholar]
- 21.Eghiaian F, et al. Insight into the PrPC → PrPSc conversion from the structures of antibody-bound ovine prion scrapie-susceptibility variants. Proc Natl Acad Sci USA. 2004;101:10254–10259. doi: 10.1073/pnas.0400014101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Govaerts C, et al. Evidence for assembly of prions with left-handed β-helices into trimers. Proc Natl Acad Sci USA. 2004;101:8342–8347. doi: 10.1073/pnas.0402254101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Speare JO, et al. The role of helix 1 aspartates and salt bridges in the stability and conversion of prion protein. J Biol Chem. 2003;278:12522–12529. doi: 10.1074/jbc.M211599200. [DOI] [PubMed] [Google Scholar]
- 24.Riek R, et al. Prion protein NMR structure and familial human spongiform encephalopathies. Proc Natl Acad Sci USA. 1998;95:11667–11672. doi: 10.1073/pnas.95.20.11667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Haire LF, et al. The crystal structure of the globular domain of sheep prion protein. J Mol Biol. 2004;336:1175–1183. doi: 10.1016/j.jmb.2003.12.059. [DOI] [PubMed] [Google Scholar]
- 26.Dima RI, Thirumalai D. Probing the instabilities in the dynamics of helical fragments from mouse PrPC. Proc Natl Acad Sci USA. 2004;101:15335–15340. doi: 10.1073/pnas.0404235101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Enari M, Flechsig E, Weissmann C. Scrapie prion protein accumulation by scrapie-infected neuroblastoma cells abrogated by exposure to a prion protein antibody. Proc Natl Acad Sci USA. 2001;98:9295–9299. doi: 10.1073/pnas.151242598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Peretz D, et al. Antibodies inhibit prion propagation and clear cell cultures of prion infectivity. Nature. 2001;412:739–743. doi: 10.1038/35089090. [DOI] [PubMed] [Google Scholar]
- 29.Gilch S, et al. Polyclonal anti-PrP auto-antibodies induced with dimeric PrP interfere efficiently with PrPSc propagation in prion-infected cells. J Biol Chem. 2003;278:18524–18531. doi: 10.1074/jbc.M210723200. [DOI] [PubMed] [Google Scholar]
- 30.Perrier V, et al. Anti-PrP antibodies block PrP replication in prion-infected cell cultures by accelerating PrP degradation. J Neurochem. 2004;89:454–463. doi: 10.1111/j.1471-4159.2004.02356.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kim CL, et al. Cell-surface retention of PrPC by anti-PrP antibody prevents protease-resistant PrP formation. J Gen Virol. 2004;85:3473–3482. doi: 10.1099/vir.0.80113-0. [DOI] [PubMed] [Google Scholar]
- 32.Feraudet C, et al. Screening of 145 anti-PrP monoclonal antibodies for their capacity to inhibit PrPSc replication in infected cells. J Biol Chem. 2004;280:11247–11258. doi: 10.1074/jbc.M407006200. [DOI] [PubMed] [Google Scholar]
- 33.Beringue V, et al. PrPSc binding antibodies are potent inhibitors of prion replication in cell lines. J Biol Chem. 2004;279:39671–39676. doi: 10.1074/jbc.M402270200. [DOI] [PubMed] [Google Scholar]
- 34.Miyamoto K, et al. Inhibition of prion propagation in scrapie-infected mouse neuroblastoma cell lines using mouse monoclonal antibodies against prion protein. Biochem Biophys Res Commun. 2005;335:197–204. doi: 10.1016/j.bbrc.2005.07.063. [DOI] [PubMed] [Google Scholar]
- 35.Pankiewicz J, et al. Clearance and prevention of prion infection in cell culture by anti-PrP antibodies. Eur J Neurosci. 2006;23:2635–2647. doi: 10.1111/j.1460-9568.2006.04805.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sigurdsson EM, et al. Anti-prion antibodies for prophylaxis following prion exposure in mice. Neurosci Lett. 2003;336:185–187. doi: 10.1016/s0304-3940(02)01192-8. [DOI] [PubMed] [Google Scholar]
- 37.Song CH, et al. Effect of intraventricular infusion of anti-prion protein monoclonal antibodies on disease progression in prion-infected mice. J Gen Virol. 2008;89:1533–1544. doi: 10.1099/vir.0.83578-0. [DOI] [PubMed] [Google Scholar]
- 38.Yu X, Shibata T, Egelman EH. Identification of a defined epitope on the surface of the active RecA-DNA filament using a monoclonal antibody and three-dimensional reconstruction. J Mol Biol. 1998;283:985–992. doi: 10.1006/jmbi.1998.2141. [DOI] [PubMed] [Google Scholar]
- 39.Chavali GB, et al. The crystal structure of human angiogenin in complex with an antitumor neutralizing antibody. Structure (London) 2003;11:875–885. doi: 10.1016/s0969-2126(03)00131-x. [DOI] [PubMed] [Google Scholar]
- 40.Mallucci G, et al. Post-natal knockout of prion protein alters hippocampal CA1 properties, but does not result in neurodegeneration. EMBO J. 2002;21:202–210. doi: 10.1093/emboj/21.3.202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Collinge J, Clarke A. A general model of prion strains and their pathogenicity. Science. 2007;318:930–936. doi: 10.1126/science.1138718. [DOI] [PubMed] [Google Scholar]
- 42.Collinge J, Palmer MS, Dryden AJ. Genetic predisposition to iatrogenic Creutzfeldt–Jakob disease. Lancet. 1991;337:1441–1442. doi: 10.1016/0140-6736(91)93128-v. [DOI] [PubMed] [Google Scholar]
- 43.Palmer MS, et al. Homozygous prion protein genotype predisposes to sporadic Creutzfeldt–Jakob disease. Nature. 1991;352:340–342. doi: 10.1038/352340a0. [DOI] [PubMed] [Google Scholar]
- 44.Lee HS, et al. Increased susceptibility to Kuru of carriers of the PRNP 129 methionine/methionine genotype. J Infect Dis. 2001;183:192–196. doi: 10.1086/317935. [DOI] [PubMed] [Google Scholar]
- 45.Poulter M, et al. Inherited prion disease with 144 base pair gene insertion: I: Genealogical and molecular studies. Brain. 1992;115:675–685. doi: 10.1093/brain/115.3.675. [DOI] [PubMed] [Google Scholar]
- 46.Mead S, et al. Balancing selection at the prion protein gene consistent with prehistoric kuru-like epidemics. Science. 2003;300:640–643. doi: 10.1126/science.1083320. [DOI] [PubMed] [Google Scholar]
- 47.Collinge J, et al. Molecular analysis of prion strain variation and the aetiology of ‘new variant’ CJD. Nature. 1996;383:685–690. doi: 10.1038/383685a0. [DOI] [PubMed] [Google Scholar]
- 48.Wadsworth JD, et al. Human prion protein with valine 129 prevents expression of variant CJD phenotype. Science. 2004;306:1793–1796. doi: 10.1126/science.1103932. [DOI] [PubMed] [Google Scholar]
- 49.Collinge J, et al. Prion protein gene analysis in new variant cases of Creutzfeldt–Jakob disease. Lancet. 1996;348:56. doi: 10.1016/s0140-6736(05)64378-4. [DOI] [PubMed] [Google Scholar]
- 50.Wroe SJ, et al. Clinical presentation and pre-mortem diagnosis of variant Creutzfeldt–Jakob disease associated with blood transfusion: A case report. Lancet. 2006;368:2061–2067. doi: 10.1016/S0140-6736(06)69835-8. [DOI] [PubMed] [Google Scholar]
- 51.O'Rourke KI, et al. PrP genotypes of captive and free-ranging Rocky Mountain elk (Cervus elaphus nelsoni) with chronic wasting disease. J Gen Virol. 1999;80:2765–2769. doi: 10.1099/0022-1317-80-10-2765. [DOI] [PubMed] [Google Scholar]
- 52.Green KM, et al. The elk PRNP codon 132 polymorphism controls cervid and scrapie prion propagation. J Gen Virol. 2008;89:598–608. doi: 10.1099/vir.0.83168-0. [DOI] [PubMed] [Google Scholar]
- 53.Hosszu LL, et al. The residue 129 polymorphism in human prion protein does not confer susceptibility to CJD by altering the structure or global stability of PrPC. J Biol Chem. 2004;279:28515–28521. doi: 10.1074/jbc.M313762200. [DOI] [PubMed] [Google Scholar]
- 54.Sawaya MR, et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature. 2007;447:453–457. doi: 10.1038/nature05695. [DOI] [PubMed] [Google Scholar]
- 55.Wickner RB, et al. Protein inheritance (prions) based on parallel in-register beta-sheet amyloid structures. BioEssays. 2008;30:955–964. doi: 10.1002/bies.20821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Fabian H, et al. Early stages of misfolding and association of beta2-microglobulin: Insights from infrared spectroscopy and dynamic light scattering. Biochemistry. 2008;47:6895–6906. doi: 10.1021/bi800279y. [DOI] [PubMed] [Google Scholar]
- 57.Klohn P, et al. A quantitative, highly sensitive cell-based infectivity assay for mouse scrapie prions. Proc Natl Acad Sci USA. 2003;100:11666–11671. doi: 10.1073/pnas.1834432100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Harlow E, Lane D. Antibodies: A Laboratory Manual. Cold Spring Harbor, New York: Cold Spring Harbor Lab Press; 1988. [Google Scholar]
- 59.Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 60.McCoy AJ, et al. Phaser crystallographic software. J Appl Crystallogr. 2007;40:658–674. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Vagin AA, Teplyakov A. MOLREP: An automated program for molecular replacement. J Appl Crystallogr. 1997;30:1022–1025. [Google Scholar]
- 62.Luginbuhl B, et al. Directed evolution of an anti-prion protein scFv fragment to an affinity of 1 pM and its structural interpretation. J Mol Biol. 2006;363:75–97. doi: 10.1016/j.jmb.2006.07.027. [DOI] [PubMed] [Google Scholar]
- 63.Murshudov GN, Vagin AA, Dodson EJ. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D. 1997;53:240–255. doi: 10.1107/S0907444996012255. [DOI] [PubMed] [Google Scholar]
- 64.Winn MD, Isupov MN, Murshudov GN. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr D. 2001;57:122–133. doi: 10.1107/s0907444900014736. [DOI] [PubMed] [Google Scholar]
- 65.Emsley P, Cowtan K. Coot: Model building tools for molecular graphics. Acta Crystallogr D. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 66.Lamzin VS, Wilson KS. Automated refinement of protein models. Acta Crystallogr D. 1993;49:129–147. doi: 10.1107/S0907444992008886. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.