Abstract
A general chemoenzymatic method for the site-specific attachment of lipids to protein substrates is described. Sortase A is used to append short lipid-modified oligoglycine peptides to the C terminus of protein substrates bearing a five amino acid sortase A recognition sequence (LPETG). We demonstrate the attachment of a range of hydrophobic modifications in excellent yield (60–90%), including a simple step for removing the sortase enzyme post-reaction. Lipoproteins prepared using these procedures were subsequently shown to associate with mammalian cells in a lipid tail-dependent fashion, and localized to the plasma membrane and endosomes.
Introduction
Methods for the construction of protein-lipid conjugates are valuable tools for studying the role of post-translational lipid modification in controlling protein function and localization. Lipid attachment also provides a means for manipulating the properties of peptides and proteins that normally do not possess these types of modifications. Highlighting the utility of synthetic lipoproteins, Bertozzi and co-workers recently reported the synthesis of GFP containing GPI-anchor mimetics as a means for probing the role of individual sugar residues in this complex membrane anchor.1,2 Numerous reports have also shown that the addition of lipids and fatty acids can endow proteins and peptides with cell penetrating capabilities3–7, modulate immunogenicity8,9, and can provide anchors for incorporation into liposomes10,11.
With respect to full-size protein substrates, the attachment of lipids in site-specific fashion remains a significant bioconjugation challenge. Only a relatively limited set of chemical and chemoenzymatic strategies are available for this purpose. These include the use of enzymes such as myristoyl12,13 and prenyl transferases14–16, wherein the user is limited to the natural substrates of these enzymes or closely related analogoues. In the case of chemical labeling, a common strategy is cysteine derivatization through alkylation17,18 or disulfide formation19,20, an approach that requires site-directed mutagenesis to ensure that protein substrates contain only a single reactive cysteine residue. Among the available strategies for lipid attachment, expressed protein ligation (EPL)1,2,21–25 has proven to be the most versatile. For example, this method has been used to prepare GFP bearing a range of lipid modifications1,2,24 and to generate lipidated Ras25 and Rab22 proteins. However, EPL mandates the expression of protein substrates as fusions with large intein domains, a necessity that can reduce protein expression yields and may require careful selection of the intein domain to prevent premature intein cleavage and a corresponding drop in the yield of ligation products.26–29
With the goal of complementing existing strategies for lipoprotein synthesis, we have developed a general method for the site-specific installation of lipids and fatty acids using sortase-catalyzed transpeptidation (Scheme 1). This chemoenzymatic approach has seen increasing use as a method for protein engineering.30–38 Sortase A from Staphylococcus aureus recognizes a five amino acid sequence (LPXTG) near the C-terminus of its natural protein targets.39–41 The LPXTG motif has been successfully transposed onto unstructured regions of other proteins to generate new sortase substrates. The active site cysteine of sortase cleaves between the threonine and glycine residues of the recognition site to produce a thioester intermediate which is then attacked by the amino terminus of an oligoglycine nucleophile resulting in the formation of a new amide bond. The method is extremely versatile, stemming from the remarkable tolerance of the enzyme for substituents C-terminal to the oligoglycine unit. Synthetic nucleophiles, containing 1–5 glycine residues, have been decorated with a range of substituents including fluorophores33,34, photoaffinity probes34, peptide nucleic acids36, polymers35, solid supports32,35,42, or other polypeptides33,38 allowing the site-specific ligation of these moieties to peptide and protein substrates. Here we describe the synthesis of lipid-modified oligoglycine nucleophiles compatible with the sortase-mediated transpeptidation reaction, and demonstrate their high yielding ligation to a protein substrate. Lipoproteins prepared using this procedure were shown to strongly associate with mammalian cells in a lipid tail-dependent fashion, and were found to localize to the plasma membrane and endosomes.
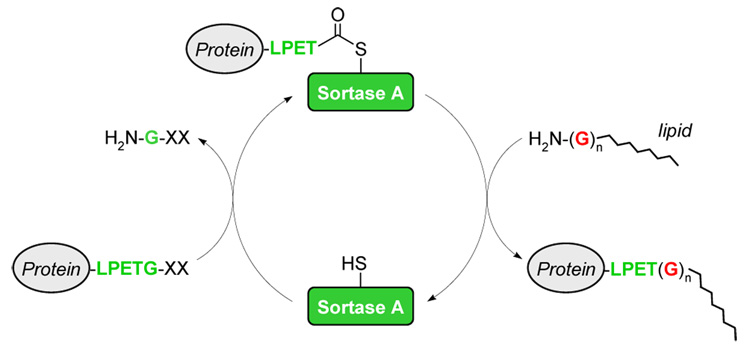
Scheme 1.
Site-specific lipid attachment through sortase-mediated transpeptidation.
Results and Discussion
Design and synthesis of lipid-modified triglycine nucleophiles
Nucleophiles compatible with sortase-mediated transpeptidation have the simple structural requirement of a stretch of glycine residues with a free amino terminus. While successful transpeptidation has been reported with nucleophiles containing anywhere from 1–5 glycines, maximum reaction rates are obtained when two or more glycines are present.32,38 It should be noted that primary alkyl amines have been shown to participate in the transpeptidation reaction31,35, though in general these appear to be less efficient than oligoglycine derivatives. With these considerations in mind, we synthesized a panel of lipid-modified triglycine nucleophiles using solid phase synthesis (Scheme 2).
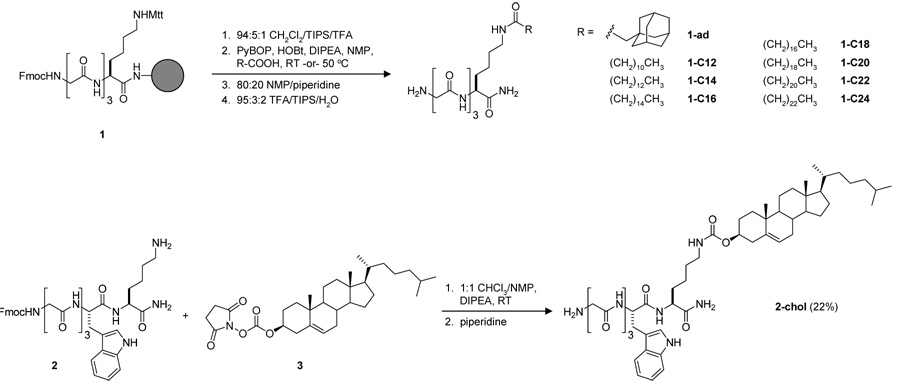
Scheme 2.
Synthesis of lipid-modified triglycine nucleophiles.
Resin-bound precursor 1 was prepared on Rink Amide resin using standard Fmoc chemistry. An orthogonally protected lysine residue was incorporated to allow selective liberation and subsequent acylation of the ε-amino group. Following removal of the 4-methyltrityl (Mtt) group, the lysine side chain was acylated with a range of hydrophobic carboxylic acids, including naturally occurring posttranslational modifications such as myristic (1-C14) and palmitic (1-C16) acid, as well as modifications not found in eukaryotic cells (1-ad). The couplings of C18, C20, C22, and C24 acids were performed at 50 °C to maintain the solubility of all the reaction components. Following Fmoc deprotection and resin cleavage, the identity of nucleophiles derived from 1 was confirmed by LC-ESI-MS (Figure S1). All peptides were obtained with good purity and did not require extensive purification prior to transpeptidation.
In addition to purely aliphatic substituents, we also prepared a cholesterol-modified triglycine nucleophile (2-chol, Scheme 2). Peptide 2 was prepared on Rink Amide resin and then cleaved from the solid support leaving the N-terminal Fmoc group intact. A tryptophan residue was included in this peptide as a UV active chromophore to aid in purification by HPLC. Peptide 2 was then reacted with cholesterol NHS-carbonate 3, followed by the addition of piperidine to remove the Fmoc group. The crude material was then purified by RP-HPLC to yield 2-chol in 22% yield.
Sortase-mediated ligation of triglycine nucleophiles to a protein substrate
With a series of lipidated nucleophiles in hand, we proceeded to explore their coupling to a protein substrate using sortase. A model protein consisting of eGFP equipped with a C-terminal LPETG sequence followed by a His6 affinity handle was expressed and purified from E. coli. Recombinant sortase A containing an N-terminal His6 tag was produced as described.40 At the outset of these studies, we decided to incorporate a procedure for removing the sortase enzyme after the reaction, similar to a purification scheme proposed by Parthasarathy et. al.35 The configuration of His6 tags on both sortase and the eGFP substrate provided a simple means for doing so, where Ni-NTA resin was added following tranpeptidation to retrieve the sortase enzyme as well as any unreacted eGFP-LPETG-His6. In this scenario, the tranpeptidation product should no longer be His6-tagged because residues C-terminal to the LPETG motif are lost in the course of transpeptidation, and thus this material will not be bound by Ni-NTA. As shown in Figure 1a, exposing eGFP-LPETG-His6 to 150 µM sortase and 2 mM triglycine (GGG) in Tris buffer (pH 7.5) containing 10 mM CaCl2 resulted in the appearance of a lower molecular species just below the band corresponding to sortase (lane 2). Excess sortase was used to drive the reaction to completion in a reasonable time frame. Near quantitative conversion of the input eGFP-LPETG-His6 protein (lane 1) was observed over the course of a 3 h incubation at 37 °C. A slurry of Ni-NTA resin containing 1 M NaCl and 40 mM imidazole was then added and the mixture was incubated for 2 h. The addition of imidazole and NaCl at the specified concentrations was critical to preventing nonspecific binding of the transpeptidation products. After filtration to remove Ni-NTA, the filtrate was analyzed by SDS-PAGE. A single polypeptide was observed with no evidence for the presence of residual sortase (lane 3). Characterization by ESI-MS confirmed that the protein observed in lane 3 was indeed the desired ligation product (Figure S2). Also, judging from the intensity of the band in lane 3 as compared to the corresponding eGFP bands in lanes 1 and 2, we were confident that very little material was lost over the course of the two step transpeptidation/sortase-depletion protocol.
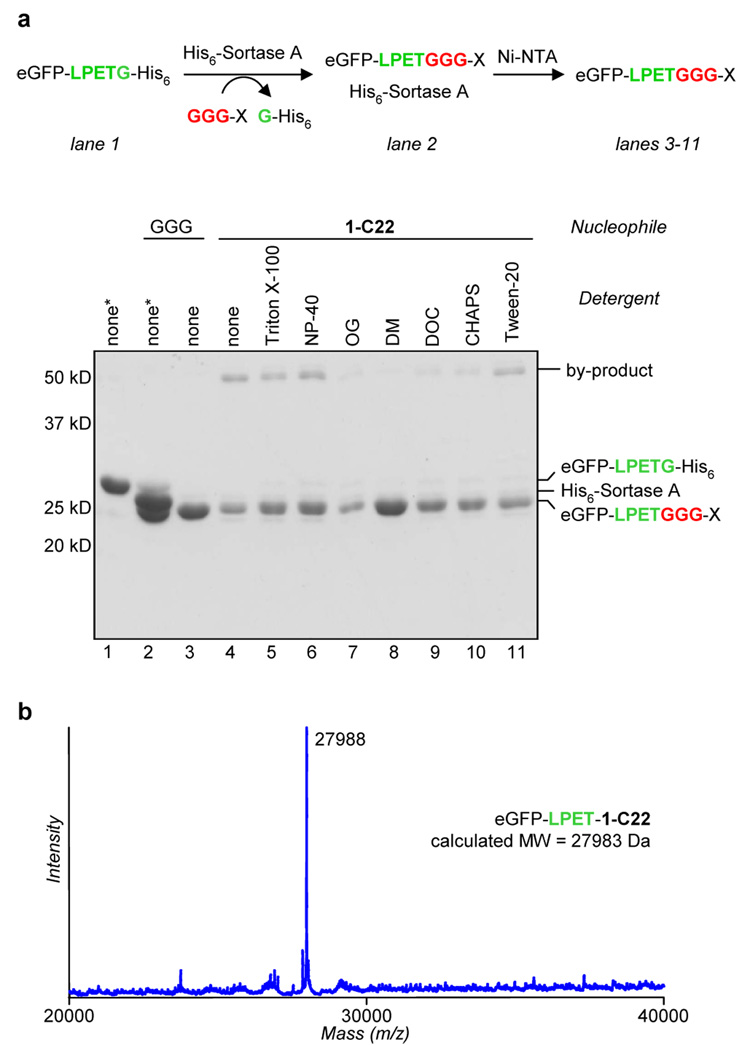
Figure 1.
(a) Optimization of a two step protocol for lipid ligation followed by removal of the sortase enzyme. Conditions: 77 µM eGFP-LPETG-His6, 150 µM sortase A, 2 mM nucleophile, 1% (w/v) detergent, 50 mM Tris pH 7.5, 150 mM NaCl, 10 mM CaCl2, 3 h at 37 °C. After transpeptidation, Ni-NTA resin was added as a slurry in 1 M NaCl and 40 mM imidazole and incubated for 2 h at RT. Abbreviations: OG = n-octyl glucoside, DM = n-dodecyl maltoside, DOC = deoxycholate. *Samples not treated with Ni-NTA resin. (b) ESI-MS spectrum of eGFP-LPET-1-C22 ligation product.
Initial attempts to ligate one of the lipid modified nucleophiles (1-C22) using the conditions described above yielded only small amounts of the putative transpeptidation product as well as an unidentified higher molecular weight species43 (Figure 1a, lane 4). Not surprisingly, the solubility of 1-C22 in aqueous buffer was extremely poor, and we suspected that this may limit the amount of nucleophile available in solution. To improve solubility we examined the effect of adding mild detergents to the reaction mixture. As shown in Figure 1a (lane 8), the addition of 1% (w/v) n-dodecyl maltoside afforded a significant increase in the level of transpeptidation product relative to the other detergents tested. This material was further characterized by ESI-MS and found to consist exclusively of the desired lipid-modified eGFP derivative (Figure 1b).
Using 1% (w/v) n-dodecyl maltoside to maintain nucleophile solubility, we were able to successfully ligate the entire panel of lipid-modified triglycine nucleophiles to our eGFP model substrate (Figure 2). In all cases, a single major protein that migrated at a lower apparent molecular weight than unreacted eGFP-LPETG-His6 (lane 1) was observed in the filtrate of reactions using lipidated nucleophiles (lanes 5–13). As expected, these samples also showed complete removal of the sortase enzyme. Coupling reactions using only triglycine (lanes 2–3) served as positive controls for transpeptidation. A reaction lacking an oligoglycine nucleophile resulted in slow hydrolysis of the acyl enzyme intermediate and the formation of a higher molecular weight by-product43 (lane 4). As an additional negative control, we confirmed that the formation of transpeptidation products was dependent on the presence of sortase (Figure S3). The identity of all eGFP conjugates was ultimately established by mass spectrometry (Table 1 and Figure S2).
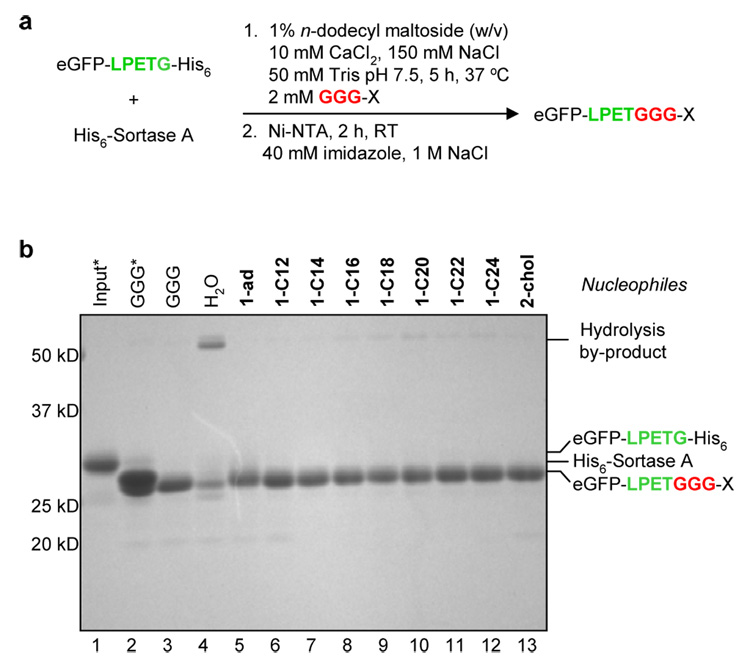
Figure 2.
Lipid attachment through sortase-catalyzed transpeptidation. (a) Optimized procedure for lipid coupling followed by removal of His6-tagged proteins with Ni-NTA resin. (b) SDS-PAGE analysis of lipid-modified eGFP following transpeptidation and depletion of His6-tagged proteins. [Input = eGFP-LPETG-His6 in the absence of sortase and nucleophile, *Samples incubated with Ni-NTA resin and 300 mM imidazole to block binding of His6-tagged proteins].
Table 1.
Mass spectral characterization and percent yield of eGFP-lipid conjugates.
| eGFP Conjugate | calculated (Da) | observed (Da) | % yield |
|---|---|---|---|
| eGFP-GGG | 27534 | 27531 | 93 |
| eGFP-1-ad | 27837 | 27835 | 87 |
| eGFP-1-C12 | 27843 | 27842 | 87 |
| eGFP-1-C14 | 27871 | 27870 | 72 |
| eGFP-1-C16 | 27899 | 27898 | 64 |
| eGFP-1-C18 | 27927 | 27926 | 57 |
| eGFP-1-C20 | 27955 | 27954 | 66 |
| eGFP-1-C22 | 27983 | 27982 | 83 |
| eGFP-1-C24 | 28011 | 28010 | 84 |
| eGFP-2-chol | 28260 | 28257 | 87 |
The eGFP chromophore provided a convenient method for quantifying the yield of the eGFP conjugates. By comparing the absorbance at 488 nm of the conjugates relative to the input eGFP-LPETG-His6 protein, we estimated that the lipid-modified protein conjugates were all obtained in excellent (~60–90%) yield (Table 1 and Figure S4). A reproducible drop in reaction yield for nucleophiles containing fatty acids of intermediate length (1-C16, 1-C18, 1-C20) was observed. Whether this reflects a drop in the efficiency of the actual transpeptidation reaction or limited solubility of the resulting lipoprotein conjugates, which could contribute to losses during the sortase depletion step, is presently unclear. It should be noted, however, that the purity of these eGFP conjugates, as determined by both SDS-PAGE (Figure 2) and ESI-MS (Figure S2), was not compromised.
Interaction of lipid-modified eGFP with mammalian cells
The presence of a hydrophobic tail on eGFP should render it capable of spontaneous association with biological membranes.1,2,24,44 As an initial investigation into the effect of the identity of the lipid tail on the ability of these lipoproteins to associate with cells, we incubated HeLa cells with lipid-modified eGFP (2.5 µg/mL) in serum-free medium for 1 h at 37 °C. After washing, the cells were analyzed by flow cytometry (Figure 3). In general, we observed a steady increase in cellular fluorescence with increasing length of the lipid tail. In the case of eGFP-1-C22, we recorded a 500-fold enhancement in mean cellular fluorescence relative to eGFP-GGG. Interestingly, eGFP-1-C24 consistently yielded lower association than eGFP-1-C22. For all samples cell viability was >94% as determined by exclusion of propidium iodide, indicative of a lack of cytotoxicity of the lipidated eGFP derivatives. Also, because the eGFP preparations contained low levels of free lipidated nucleophile (~1 µM in culture medium), we verified that this contaminant did not contribute to the observed increase in cellular fluorescence. eGFP-GGG in combination with 1 µM 1-C22, 1-C24, or 2-chol produced only modest increases in fluorescence, <5% of the magnitude observed for the corresponding covalently modified eGFP conjugates (Figure S5).
Figure 3.
Association of lipid-modified eGFP with HeLa cells. Cells were incubated with lipid-modified eGFP (2.5 µg/mL) for 1 h in serum-free medium and then analyzed by flow cytometry. Numbers in parentheses represent the fold enhancement in mean cellular fluorescence relative to eGFP-GGG. Histograms for eGFP-GGG (black) and eGFP-1-ad (gray) are overlapping.
A preliminary evaluation of the subcellular localization of eGFP-1-C22 and eGFP-2-chol was performed using spinning disk confocal microscopy. HeLa cells and U373 cells were incubated with 2.5 µg/mL of lipoprotein in serum-free medium, following by washing and live cell imaging. For both cell types we observed staining of the cell surface and well as the appearance of internal fluorescent structures (Figure 4). For all cells, filamentous membrane extensions, presumably filopodia, were evident in the fluorescent micrographs, indicating the presence of the eGFP lipoproteins on the plasma membrane. In U373 cells, distinct vesicular structures were observed at early (1 h) and later (5 h) time points, with a more pronounced presence of vesicles observed for eGFP-1-C22 than for eGFP-2-chol. In general, less internalization was observed for HeLa cells, though punctate internal staining could be observed with eGFP-1-C22 after a 5 h incubation in these cells. In the case of U373, we were also able to verify that eGFP-1-C22 had access to early/recycling endosomal compartments as determined by partial colocalization with transferrin (Figure 5).6,45 Colocalization between eGFP-2-chol and transferrin was not observed (Figure 5) for this cell line. Taken together, these results demonstrate that synthetic lipoproteins using sortase mediated transpeptidation can interact strongly with the plasma membrane and access endosomal compartments. The range of staining observed for our eGFP conjugates suggests that interactions with cells could be fine-tuned, and studies are underway to characterize the differences in the behavior of protein conjugates bearing different lipid anchors, specifically looking for differences in subcellular distribution and routes of endocytosis.
Figure 4.
Subcellular distribution of eGFP-1-C22 and eGFP-2-chol in U373 and HeLa cells. Cells were incubated with lipid-modified eGFP (2.5 µg/mL) at 37 °C in serum-free medium and then imaged live using spinning disc confocal microscopy. (a) U373 cells 1 h incubation, (b) U373 cells 5 h incubation, (c) HeLa cells 1.25 h incubation, (d) HeLa cells 5 h incubation.
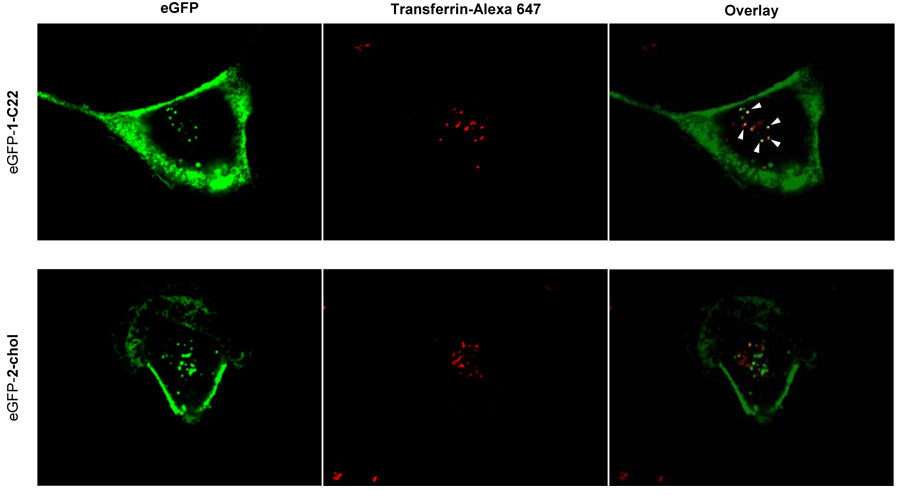
Figure 5.
eGFP-1-C22 is able to access early endosomal compartments as determined by partial colocalization with Transferrin-Alexa 647. Cells were incubated with eGFP-1-C22 (2.5 µg/mL) for 5 h in serum-free medium followed by the addition of Transferrin-Alexa 647 (100 µg/mL) during the final 15 minutes. Significant colocalization between eGFP-2-chol and Transferrin-Alexa 647 was not observed.
Conclusion
In summary, we have a developed a general strategy using sortase mediated transpeptidation as a means to install hydrophobic lipid modifications onto protein substrates in site-specific fashion. The ease of use of this method stands out as a significant asset. Protein substrates require only a five amino acid extension (LPETG) in order to serve as sortase substrates, a modest insertion that is not expected to impede the function of most proteins and should also have minimal impact on the expression yield of these polypeptides. Recombinant proteins containing the LPETG motif are also stable and unreactive until activated by the sortase enzyme. This affords a distinct advantage over the expressed protein ligation approach where premature cleavage of intein fusion proteins has been documented, leading to a reduction in the yield of final ligation products. The requisite lipid-modified nucleophiles compatible with sortase mediated transpeptidation also have a very straightforward structural requirement of a short stretch of glycine residues and are prepared by standard solid-phase synthesis. Therefore, both natural and non-natural hydrophobic modifications can be easily incorporated using this system.
While the requirement for the sortase enzyme itself could be perceived as a disadvantage of the transpeptidation strategy, it should noted that we routinely express batches with yields of >50 mg/L, and stock solutions of the enzyme can be stored for several months with no apparent degradation in enzymatic activity. Moreover, in this work we provided a convenient method for removing sortase following transpeptidation. The ready availability of sortase and ease of removal following transpeptidation are important features as the enzyme is relatively inefficient and excess sortase is typically employed to drive the labeling reactions to high levels of conversion in a reasonable period of time.
In this initial work, we used eGFP as our model protein substrate due to its natural fluorescence and verified that eGFP proteins lipidated through sortase mediated transpeptidation strongly associate with mammalian cells. We envision further applications of this chemical modification strategy to include anchorage of proteins to cellular membranes that do not normally occur on the cell surface to create artificial cell receptors. Additionally, because the transpeptidation strategy is able to attach extremely hydrophobic modifications, it could provide a new method for interfacing proteins with materials that possess poor water solubility, such as carbon nanotubes.
Supplementary Material
Full experimental details and characterization data for triglycine nucleophiles and protein conjugates. These materials are available free of charge via the Internet at http://pubs.acs.org.
Acknowledgment
This work was supported by grants from the National Institutes of Health (R01-AI057182, P01-CA100707, R01-AI033456, R21-EB008875) and a Clay Postdoctoral Fellowship (JMA). The authors gratefully acknowledge Eric Spooner for mass spectrometry expertise, and Maximilian W. Popp for help in preparing the eGFP-LPETG-His6 substrate.
References
- 1.Paulick MG, Forstner MB, Groves JT, Bertozzi CR. Proc. Natl. Acad. Sci.U.S.A. 2007;104:20332–20337. doi: 10.1073/pnas.0710139104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Paulick MG, Wise AR, Forstner MB, Groves JT, Bertozzi CR. J. Am. Chem. Soc. 2007;129:11543–11550. doi: 10.1021/ja073271j. [DOI] [PubMed] [Google Scholar]
- 3.Martin SE, Peterson BR. Bioconjug. Chem. 2003;14:67–74. doi: 10.1021/bc025601p. [DOI] [PubMed] [Google Scholar]
- 4.Boonyarattanakalin S, Athavankar S, Sun Q, Peterson BR. J. Am. Chem. Soc. 2006;128:4917–4917. doi: 10.1021/ja056126j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Covic L, Gresser AL, Talavera J, Swift S, Kuliopulos A. Proc. Natl. Acad. Sci. U.S.A. 2002;99:643–648. doi: 10.1073/pnas.022460899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sun Q, Cai S, Peterson BR. J. Am. Chem. Soc. 2008;130:10064–10065. doi: 10.1021/ja803380a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nelson AR, Borland L, Allbritton NL, Sims CE. Biochemistry. 2007;46:14771–14781. doi: 10.1021/bi701295k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Moyle PM, Toth I. Curr. Med. Chem. 2008;15:506–516. doi: 10.2174/092986708783503249. [DOI] [PubMed] [Google Scholar]
- 9.BenMohamed L, Wechsler SL, Nesburn AB. Lancet Infect. Dis. 2002;2:425–431. doi: 10.1016/s1473-3099(02)00318-3. [DOI] [PubMed] [Google Scholar]
- 10.Reulen SW, Brusselaars WW, Langereis S, Mulder WJ, Breurken M, Merkx M. Bioconjug. Chem. 2007;18:590–596. doi: 10.1021/bc0602782. [DOI] [PubMed] [Google Scholar]
- 11.Torchilin VP. Nat. Rev. Drug Discov. 2005;4:145–160. doi: 10.1038/nrd1632. [DOI] [PubMed] [Google Scholar]
- 12.Ha VL, Thomas GM, Stauffer S, Randazzo PA. Methods Enzymol. 2005;404:164–174. doi: 10.1016/S0076-6879(05)04016-4. [DOI] [PubMed] [Google Scholar]
- 13.Heal WP, Wickramasinghe SR, Bowyer PW, Holder AA, Smith DF, Leatherbarrow RJ, Tate EW. Chem. Commun. 2008:480–482. doi: 10.1039/b716115h. [DOI] [PubMed] [Google Scholar]
- 14.Kalinin A, Thoma NH, Iakovenko A, Heinemann I, Rostkova E, Constantinescu AT, Alexandrov K. Protein Expression and Purification. 2001;22:84–91. doi: 10.1006/prep.2001.1423. [DOI] [PubMed] [Google Scholar]
- 15.Gauchet C, Labadie GR, Poulter CD. J. Am. Chem. Soc. 2006;128:9274–9275. doi: 10.1021/ja061131o. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Duckworth BP, Zhang ZY, Hosokawa A, Distefano MD. ChemBioChem. 2007;8:98–105. doi: 10.1002/cbic.200600340. [DOI] [PubMed] [Google Scholar]
- 17.Breydo L, Sun Y, Makarava N, Lee CI, Novitskaia V, Bocharova O, Kao JP, Baskakov IV. Biochemistry. 2007;46:852–861. doi: 10.1021/bi061923v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bader B, Kuhn K, Owen DJ, Waldmann H, Wittinghofer A, Kuhlmann J. 2000;403:223–226. doi: 10.1038/35003249. [DOI] [PubMed] [Google Scholar]
- 19.Eberl H, Tittmann P, Glockshuber R. J. Biol. Chem. 2004;279:25058–25065. doi: 10.1074/jbc.M400952200. [DOI] [PubMed] [Google Scholar]
- 20.Hicks MR, Gill AC, Bath IK, Rullay AK, Sylvester ID, Crout DH, Pinheiro TJ. Febs. J. 2006;273:1285–1299. doi: 10.1111/j.1742-4658.2006.05152.x. [DOI] [PubMed] [Google Scholar]
- 21.Muir TW. Annu. Rev. Biochem. 2003;72:249–289. doi: 10.1146/annurev.biochem.72.121801.161900. [DOI] [PubMed] [Google Scholar]
- 22.Alexandrov K, Heinemann I, Durek T, Sidorovitch V, Goody RS, Waldmann H. J. Am. Chem. Soc. 2002;124:5648–5649. doi: 10.1021/ja017799e. [DOI] [PubMed] [Google Scholar]
- 23.Pellois JP, Muir TW. Angew. Chem. Int. Ed. Engl. 2005;44:5713–5717. doi: 10.1002/anie.200501244. [DOI] [PubMed] [Google Scholar]
- 24.Grogan MJ, Kaizuka Y, Conrad RM, Groves JT, Bertozzi CR. J. Am. Chem. Soc. 2005;127:14383–14387. doi: 10.1021/ja052407f. [DOI] [PubMed] [Google Scholar]
- 25.Gottlieb D, Grunwald C, Nowak C, Kuhlmann J, Waldmann H. Chem. Commun. 2006:260–262. doi: 10.1039/b511736d. [DOI] [PubMed] [Google Scholar]
- 26.Luka Z, Wagner C. Protein Expr. Purif. 2003;28:280–286. doi: 10.1016/s1046-5928(02)00710-6. [DOI] [PubMed] [Google Scholar]
- 27.Zhao W, Zhang Y, Cui C, Li Q, Wang J. Protein Sci. 2008;17:736–747. doi: 10.1110/ps.073383708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cui C, Zhao W, Chen J, Wang J, Li Q. Protein Expr. Purif. 2006;50:74–81. doi: 10.1016/j.pep.2006.05.019. [DOI] [PubMed] [Google Scholar]
- 29.Tan LP, Lue RY, Chen GY, Yao SQ. Bioorg. Med. Chem. Lett. 2004;14:6067–6070. doi: 10.1016/j.bmcl.2004.09.083. [DOI] [PubMed] [Google Scholar]
- 30.Pritz S, Kraetke O, Klose A, Klose J, Rothemund S, Fechner K, Bienert M, Beyermann M. Angew. Chem. Int. Ed. Engl. 2008;47:3642–3645. doi: 10.1002/anie.200705718. [DOI] [PubMed] [Google Scholar]
- 31.Samantaray S, Marathe U, Dasgupta S, Nandicoori VK, Roy RP. J. m. Chem. Soc. 2008;130:2132–2133. doi: 10.1021/ja077358g. [DOI] [PubMed] [Google Scholar]
- 32.Chan L, Cross HF, She JK, Cavalli G, Martins HFP, Neylon C. PLoS ONE. 2007;2:e1164. doi: 10.1371/journal.pone.0001164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tanaka T, Yamamoto T, Tsukiji S, Nagamune T. ChemBioChem. 2008;9:802–807. doi: 10.1002/cbic.200700614. [DOI] [PubMed] [Google Scholar]
- 34.Popp MW, Antos JM, Grotenbreg GM, Spooner E, Ploegh HL. Nat. Chem. Biol. 2007;3:707–708. doi: 10.1038/nchembio.2007.31. [DOI] [PubMed] [Google Scholar]
- 35.Parthasarathy R, Subramanian S, Boder ET. Bioconjug. Chem. 2007;18:469–476. doi: 10.1021/bc060339w. [DOI] [PubMed] [Google Scholar]
- 36.Pritz S, Wolf Y, Kraetke O, Klose J, Bienert M, Beyermann M. J. Org. Chem. 2007;72:3909–3912. doi: 10.1021/jo062331l. [DOI] [PubMed] [Google Scholar]
- 37.Mao H. Protein Expr. Purif. 2004;37:253–263. doi: 10.1016/j.pep.2004.06.013. [DOI] [PubMed] [Google Scholar]
- 38.Mao H, Hart SA, Schink A, Pollok BA. J. Am. Chem. Soc. 2004;126:2670–2671. doi: 10.1021/ja039915e. [DOI] [PubMed] [Google Scholar]
- 39.Kruger RG, Otvos B, Frankel BA, Bentley M, Dostal P, McCafferty DG. Biochemistry. 2004;43:1541–1551. doi: 10.1021/bi035920j. [DOI] [PubMed] [Google Scholar]
- 40.Ton-That H, Liu G, Mazmanian SK, Faull KF, Schneewind O. Proc. Natl. Acad. Sci. U.S.A. 1999;96:12424–12429. doi: 10.1073/pnas.96.22.12424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ton-That H, Mazmanian SK, Faull KF, Schneewind O. J. Biol. Chem. 2000;275:9876–9881. doi: 10.1074/jbc.275.13.9876. [DOI] [PubMed] [Google Scholar]
- 42.Clow F, Fraser JD, Proft T. Biotechnol. Lett. 2008;30:1603–1607. doi: 10.1007/s10529-008-9718-1. [DOI] [PubMed] [Google Scholar]
- 43.The high molecular weight by-product may correspond to the acyl enyme intermediate formed between sortase A and the eGFP substrate. This assignment implies that the affinity of the His6 tag on the N-terminus of sortase is impaired in the acyl enzyme intermediate because this species is not removed during depletion of His6-tagged proteins using N-NTA resin. Further characterization of this by-product was not performed.
- 44.Zacharias DA, Violin JD, Newton AC, Tsien RY. Science. 2002;296:913–916. doi: 10.1126/science.1068539. [DOI] [PubMed] [Google Scholar]
- 45.Sheff D, Pelletier L, O'Connell CB, Warren G, Mellman I. J. Cell. Biol. 2002;156:797–804. doi: 10.1083/jcb.20111048. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Full experimental details and characterization data for triglycine nucleophiles and protein conjugates. These materials are available free of charge via the Internet at http://pubs.acs.org.