Abstract
Our studies have shown that constitutive interactions between hyaluronan and CD44 on tumor cells induces various anti-apoptotic cell survival pathways through the formation of a multimeric signaling complex that contains activated receptor tyrosine kinases. Inhibition of the hyaluronan-CD44 interactions on tumor cells by hyaluronan-CD44 interaction antagonists suppresses these activities by disassembling the complex. Although the anti-tumor activity of hyaluronan-oligosaccharides, a hyaluronan-CD44 interaction antagonist, is effective in sensitizing tumor cells to chemotherapeutic agents and reducing tumor growth in xenografts, hyaluronan-oligosaccharide alone was not effective in reducing tumor progression in Apc Min/+ mice. We now show in vitro and in vivo that targeted inhibition of the expression of CD44v6 depletes the ability of the colon tumor cells to signal through hyaluronan-CD44v6 interactions. First, we cloned oligonucleotides coding CD44v6 shRNA into a conditionally silenced pSico vector. Second, using pSico-CD44v6 shRNA and a colon-specific Fabpl promoter-driven Cre recombinase expression vector packaged into transferrin-coated nanoparticles, we successfully delivered the CD44v6 shRNA within pre-neoplastic and neoplastic colon malignant cells. Third, using the Apc Min/+ mice model, we demonstrated that inhibition of the CD44v6 expression reduces the signaling through a hyaluronan/CD44v6-pErbB2-Cox-2 interaction pathway and reduced adenoma number and growth. Together, these data provide insight into the novel therapeutic strategies of short hairpin RNA/nanoparticle technology and its potential for silencing genes associated with colon tumor cells.
Extracellular matrix has a significant role in solid tumor growth (1). Hyaluronan (HA)3 is one of the constituents of extracellular matrix. HA is a high molecular weight glycosaminoglycan present in almost every tissue of vertebrates. It is concentrated in regions of high cell division and invasion. Like numerous extracellular matrix constituents, HA serves both structural and instructive roles in terms of cell signaling via HA receptors, mainly CD44, on the surface of most cells (2-4). However, when cells proliferate or migrate, e.g. in embryonic processes, tissue remodeling, inflammation, and diseases such as cancer and atherosclerosis, HA-induced signaling is activated (4-6). High levels of HA in tumors are a prognostic factor in several malignancies (7), and manipulations of HA production or interaction with cell surface receptors strongly influence tumor growth and metastasis (4, 6).
CD44 also has an important role in tumor progression (8). CD44 proteins exist in three states with respect to HA binding as follows: non-HA binding; nonbinding unless activated by physiological stimuli; or constitutively activated (9). Thus HA induces intracellular signaling when it binds to “constitutively activated” CD44 variants during cell dynamic processes, but it does not do so under conditions of adult tissue homeostasis (10, 11). The CD44 structure of normal cells is distinct from that of cancer cells, because under various physiological and pathological conditions, the local environmental pressure influences alternate splicing and post-translational modification to produce diversified CD44 molecules (8, 12). This diversification allows the production of specific targeting agents that will be useful for both diagnosis and therapy. Overexpression of the variant high molecular weight isoforms CD44v4-v7 and CD44v6-v9 in human lymphomas, colorectal adenocarcinomas, endometrial cancer, papillary thyroid carcinoma, lung and breast cancer, and metastasizing rat adenocarcinomas (13-20), as well as down-regulation of standard CD44 (CD44s), are postulated to result in increased tumorigenicity (21). Moreover, the systemic application of antibodies directed against the v6 epitope and the expression of antisense CD44v6 can retard tumor growth and block metastasis in vivo (13, 14), emphasizing the potential importance of CD44 variants as therapeutic targets in cancer, particularly colon cancer. Indeed overexpression of CD44 is an early event in the colorectal adenoma-carcinoma sequence (22-24). Thus, unlike the targeting of various cancer-promoting molecules, CD44 targeting provides a strong opportunity for discerning therapy in colon cancer cells.
Altered COX-2 gene expression occurs in human colon carcinomas, and Cox-2 antagonists inhibit colon cancer progression in animal models (25, 26). A recent study implicates COX-2 overexpression as a proximal mediator of CD44-dependent invasion in human non-small cell lung cancer and in human renal carcinoma cells (27, 28). Cox-2 and CD44v (23, 24, 29) are overexpressed in carcinogen-induced tumors, and our recent study demonstrates that HA-CD44 interactions constitutively regulate COX-2-induced cell survival in normal epithelial cells and colon carcinoma cells (30, 31).
Our previous studies demonstrated that antagonists of HA-CD44 interactions, i.e. HA oligomers and overexpression of the ectodomain of CD44 (soluble CD44) (32) that acts as a competitive decoy by binding to endogenous HA, inhibit cell survival pathway activities, including activation of several receptor tyrosine kinases, namely ERBB2, EGFR, IGF1Rβ, c-MET, and PDGFRβ, in several types of malignant colon, breast, and prostate carcinoma cells (33, 34). In our recent study we demonstrated that elevated HA in normal intestinal epithelial cells (HIEC6-HAS2) regulates several properties required for the transformed phenotype. Increased HA in these cells regulates expression and enzymatic activity of COX-2, activation of ErbB2 and AKT, and translocates β-catenin to the nucleus (30, 31). To explore the mechanism of constitutive HA-CD44 interaction and the consequent outcomes in cancer cells, we have shown that all four types of reagents, namely HA-oligosaccharides, anti-CD44 antibody, soluble CD44, or CD44 siRNA, block signaling responses in a variety of tumor cell types and also block activation of receptor tyrosine kinases (30, 31, 33, 34). Although the anti-tumor activity of HA-oligosaccharides is effective in sensitizing tumor cells to chemotherapeutic agents in cell culture models (34-36) and in reducing tumor growth in a subcutaneous in vivo xenograft model (37), HA-oligosaccharides alone are not effective in reducing tumor progression in an Apc Min/+ mice in vivo model.4 Thus, an alternative HA/CD44 antagonist is required to treat distant tumors. Because CD44 is present in the HA-induced signaling complex, we also used RNA interference to test the role of CD44 in these cancer cells (33, 34). All cancer cell lines (HCT 116 and HCA7 colon cancer cells, C4-2 and LNCaP prostate cancer cells, and MCF7/Adr breast cancer cells and TA3St mouse mammary carcinoma cells) transfected with a 21-nucleotide small interfering RNA (siRNA) targeted to CD44 greatly decreased activation of cell survival proteins and multiple receptor tyrosine kinases as well as CD44 and COX-2 expression when compared with the above cells transfected with scrambled control siRNA (30, 31, 34).
In mammalian cell culture, the transfection of 21-nucleotide double-stranded siRNA efficiently inhibits endogenous gene expression in a sequence-specific manner without induction of the interferon response (38). However, the phenotypic changes induced by siRNAs only persist 1 week because of lack of transfer of siRNA or dilution of siRNA concentration after each cell division, which limits their utility for use in inhibiting tumor progression. The technique of using shRNA in an expression vector is an alternative strategy to stably suppress gene expression, and such constructs with well defined initiation and termination sites have been used to produce various small RNA species that inhibit the expression of genes with diverse functions in mammalian cell lines (39). The conditional alteration of gene expression by the use of an shRNA expression vector holds potential promise for therapeutic approaches for silencing disease-causing genes provided that appropriate extracellular and intracellular nucleic acid delivery systems (vector systems) are available that offer efficient vehicles for stable complexation and protection of the nucleic acid. To reach the targeted tissue, vectors need to overcome a number of extracellular and intracellular barriers. Systemic targeting by viral vectors toward the desired tissue is difficult because the host immune responses activate viral clearance. Systemic administration of a large amount of adenovirus (e.g. into the liver) can be a serious health hazard that even caused the death of one patient (40).
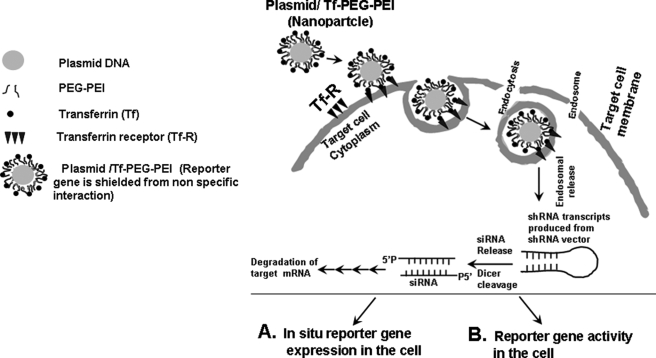
Nonviral vectors, such as positively charged PEI complexes, mediate unspecific interactions with non-target cells and blood components, which results in the rapid clearance from the circulation. These unfavorable effects can be minimized by “shielding” of the positive surface charge of the vectors with polyethylene glycol (PEG). PEGylation of PEI polyplexes can prevent the systemic degradation of the plasmid DNA and reduce the toxicity of polyplexes (41). To increase the transfection efficiency of the shielded particles (plasmid DNA/PEG-PEI), different targeting ligands, such as peptide, growth factors and proteins, or antibodies, have been incorporated into the vectors (42). One such targeting ligands is transferrin (Tf), an iron-transporting protein that is recognized by Tf receptors (Tf-R) present at high levels in the tumor cells (42, 43). In contrast, in nonproliferating cells, expression of Tf-R is low or undetectable. Association of Tf to polyplexes significantly enhances transfection efficiency by promoting the internalization of polyplexes (plasmid DNA/Tf-PEG-PEI (designated as nanoparticles throughout this study)) in dividing and nondividing cells (42). After cellular association of nanoparticles to the target cells, particles are internalized by receptor-mediated endocytosis (42). Fig. 1 illustrates that the uptake of nanoparticles carrying multiple functional domains (surface shielding particles Tf-PEG-PEI, tissue-specific promoter-driven Cre recombinase, and conditionally silenced plasmid) can overcome the intracellular barriers for successful delivery of the shRNA gene.
FIGURE 1.
Schematic illustration of cellular uptake of plasmid DNA/Tf-PEG-PEI (nanoparticles) polyplexes, their shielding from nonspecific interaction, and mechanism of action of shRNA. Internalization of PEG-shielded and Tf-R-targeted polyplexes into target cells occurs by receptor-mediated endocytosis after association of polyplex ligand Tf to Tf-R present on the target cell plasma membrane. Internalized particles are trafficked to endosomes followed by endosomal release of the particles and/or the nucleic acid into cytoplasm. Released siRNA will be induced to RNA-induced silencing complex and will be guided for cleavage of complementary target mRNA in the cytoplasm. siRNA (antisense) guide strand will direct the targeted RNAs to be cleaved by RNA endonuclease. Finally plasmid/Tf-PEG-PEI-nanoparticles delivery in the target cell shows reporter gene expression and activity.
We tested the effects of CD44v6 shRNA in vivo in a mouse model of human familial adenomatous polyposis (FAP) where adenomatous polyposis coli gene (Apc) is mutated. Most FAP patients carry truncation mutations in the N-terminal half (44). Several mouse models of FAP were constructed either by chemical mutagenesis of the Apc gene at codon 850 (Apc Min/+ mouse) or by homologous recombination in embryonic stem cells (knock-out strains) such as Apc 1638N, Apc 1638T, Apc 1309, Apc Δ716, and Apc Δ474 (45). Germ line mutagenesis of C57BL/6J (B6) males was done by N-ethyl N-nitrosourea, and the offspring of mutated B6 × AKR/J (AKR) females yielded mice that carry 100% of an autosomal dominant mutation at codon 850 in the Apc gene. These five strains have mutated Apc alleles that encode truncated Apc proteins of 850, 1638, 1309, 716, and 474 amino acids, respectively. These variable truncated products form dimers with wild-type Apc protein and show variable dominant negative activities. As a result, the number of polyps varies in different strains. For example, the number of polyps at 16 weeks of age are ∼100 in Apc Min/+, Apc Δ474, and Apc 1309 mice. A much greater number of polyps (300-400) are found in Apc Δ716 mice, whereas much fewer polyps (∼10) are found in Apc 638N mice (46). These mutations in the Apc allele and the phenotype variability observed in FAP patients allowed the establishment of genotype-phenotype correlations at the Apc locus resulting in multiple intestinal adenomas throughout the length of the small and large intestine (47). Interestingly, most polyps are found in the small intestine, although a small but significant number of polyps develop in the colon, a phenotype different from human FAP. Despite this caveat, the Apc Min/+ mouse offers the prospect to study intestinal tumors that are of the same genetic background as the host. Strong up-regulations of CD44, including both CD44s and CD44v6 encoded epitopes, and of Cox-2 were observed in aberrant crypt foci in Apc Min/+ mice (48, 49). Furthermore, CD44v proteins can form multimeric complexes in the plasma membrane, which dramatically enhances their HA binding capacity (11). Moreover, our results with Apc Min/+ mice confirm earlier findings that Tf-R is present at high levels in the tumor cells (42), which is crucial for active targeting of Tf-R by Tf-mediated CD44v6 shRNA delivery in tumor cells. For these reasons, the Apc Min/+ mouse model can be used for modulating adenoma growth by using HA-CD44v6 interaction antagonists as therapeutic agents in vivo.
In this study we tested whether HA regulates Cox-2 via its effects on a CD44v6 → ErbB2 → Cox-2 axis in colon cancer cells. More importantly we provide evidence that systemic application of (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles in the Apc Min/+ mouse model reduces intestinal tumor growth by perturbing CD44v6 containing isoform expression.
EXPERIMENTAL PROCEDURES
Materials
Apc 10.1, an intestinal cell line, was derived from Apc Min/+ mice and retains the host heterozygous Apc genotype (50). HCA7 clone 29 colon carcinoma cells were obtained from ECCA (United Kingdom). HT-29 cells were obtained from ATCC (Manassas, VA). Apc Min/+ mice (47) were obtained from the Colon Cancer Center, COBRE, at the University of South Carolina, Columbia, and The Jackson Laboratories (Bar Harbor, ME). The gene-switch reporter plasmid pSV(EGFP)/β-galactosidase (51) was obtained from Addgene Inc. (Cambridge, MA). The promoter of the liver form of the fatty acid-binding protein (Fabpl)-Cre plasmid (52) was a gift from Dr. J. I. Gordon (Washington University School of Medicine, St. Louis, MO). The COX-2-luc construct was a gift from Dr. R. DuBois (Vanderbilt-Ingram Cancer Center, Nashville, TN). The conditional silencing vector pSico and the conditional inactivating vector pSicoR (53) were gifts from Dr. T. Jacks (Massachusetts Institute of Technology, Cambridge, MA). X-Gal was purchased from Invitrogen. The pSV-β-galactosidase plasmid (designated as pSV-β-gal throughout the paper) was purchased from Promega. Anti-COX-2 antibody, anti-human Tf-R antibody (cross-reacts with mouse Tf-R), and anti-human CD44 antibody were purchased from Santa Cruz Biotechnology; anti-phospho-ErbB2/HER2 (Y1248) antibody was from Upstate Biotechnology, Inc. (Lake Placid, NY); and monoclonal anti-β-actin clone AC-15 antibody was purchased from Sigma. The restriction enzymes HpaI, XhoI, SacII, NotI, XhoI, and XbaI were purchased from New England Biolabs (Beverly, MA). Chemically competent DH5α cells were purchased from Invitrogen. Sephacryl S200 was purchased from Amersham Biosciences. ECL reagents were from Santa Cruz Biotechnology. The mouse HAS2 cDNA construct pCI-neo-HAS2 was obtained from Dr. A. Spicer (University of California, Davis, CA). pCI-neo and pSV-β-galactosidase plasmids were from Promega. Transferrin (molecular mass of 77 kDa) was purchased from Sigma; branched chain PEI (average molecular mass of 25 kDa) was from Sigma; and N-hydroxysuccinimide/PEG/maleimide (molecular mass of 3.40 kDa) was from Pierce. All other reagents were of analytical reagent grade.
RNA Silencing and Transfection with the Various Plasmids
The siRNA transfections were done at 100 pmol using Oligofectamine (Invitrogen) according to the manufacturer's instructions. Cells were transfected with the CD44v6 siRNA (scrambled (scr) siRNA as control) or plasmids (corresponding vector plasmid as control) in 6-well plates with cells at 70-90% confluence. The cells were then incubated at 37 °C in 5% CO2 for 24 h, replated in 150-mm dishes, and allowed to grow for 48 h in complete medium.
CD44v6 shRNA Cloning in pSico and pSicoR Vectors
We prepared a double-stranded cassette for CD44v6 shRNA following instructions from the Dr. Tyler Jacks laboratory (Massachusetts Institute of Technology). Using the CD44v6 siRNA sequence as described (54), sense and antisense oligonucleotides for the double-stranded cassette were designed. These oligonucleotides were synthesized and purified by Integrated DNA Technologies (Coralville, IA).
The pSico and pSicoR vectors (Fig. 2) were linearized by digesting with HpaI and XhoI restriction enzymes, and the purified linear vectors gave no colonies when transfected with competent DH5α. These linearized vectors were ligated to the double-stranded oligonucleotide cassette. Transformation of DH5α was done with the products of ligation following the manufacturer's instructions. The plasmids were prepared from cultures grown from ampicillin-resistant colonies using the Qiagen kit.
FIGURE 2.
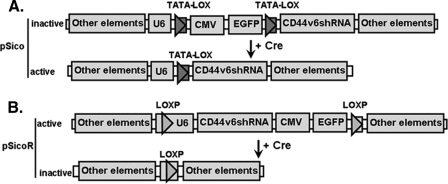
Description of pSico and pSicoR vectors (53). A, section of pSico vector. In pSico, the U6 promoter that drives the expression of shRNA is made conditional in two steps. First, bifunctional TATAlox boxes are placed in place of TATA box sites so that the resulting promoter retains transcriptional activity. Second, a cytomegalovirus (CMV)-enhanced EGFP stop/reporter cassette was inserted between two TATALox sites so that after Cre-mediated recombination the CMV-EGFP cassette will be excised to generate a functional U6 promoter. A T6 sequence is placed immediately upstream of U6 and serves as a stop signal of U6 promoter. Oligonucleotides coding shRNA are cloned into HpaI-XhoI-digested pSico vectors. Cre recombinase eliminates the CMV-EGFP cassette. Transcription of CD44v6 shRNA from the U6 promoter will ensue. B, section of pSicoR vector. In pSicoR the CMV-EGFP cassette is placed downstream of the U6 promoter and does not affect its activity. Two LoxP sites in the same orientation are present in this vector. Oligonucleotides coding CD44v6 shRNA are cloned into HpaI-XhoI-digested pSicoR vectors. The transcription from U6 promoter will produce CD44v6 shRNA constitutively in the absence of Cre recombinase. In the presence of Cre recombinase, the CD44v6 shRNA production unit and CMV-EGFP cassette are eliminated.
The purified plasmids were checked for the insert by digesting with SacII-NotI (for pSico-CD44v6 shRNA) and with XhoI-XbaI (for pSicoR-CD44v6 shRNA). Parallel control digestions were done with empty vectors. The digested products were separated in 2% agarose gels to identify the shifts in restriction digestion. The positive clones released a 710-bp fragment, which is 50 bp larger than the 660-bp fragment released by the empty pSico vector. Fragments of 400 and 350 bp were released by the pSicoR vector. The positive clones were grown, and plasmids were prepared as described previously (55).
Preparation of pSico-CD44v6 shRNA and pFabpl-Cre
All plasmid DNAs were amplified in DH5α and purified according to the protocols as described previously (55). The quantities and qualities of the purified plasmid DNAs were assessed by measuring absorbance at 260 and 280 nm (ratio was 1.80).
Preparation of Tf-coated PEG-PEI
The Tf-PEG-PEI was prepared as described previously (41). Briefly, transferrin was linked with N-hydroxysuccinimide/PEG/maleimide and then allowed to react with a mercaptopropionate-modified branched PEI to form Tf-PEG-PEI (molecular mass of ∼400 kDa). This conjugate was prepared and purified in the laboratory of Prof. E. Jabbari by Dr. Xuezhong He at the Chemical Engineering Department, University of South Carolina, Columbia.
Nanoparticle Formation
We prepared two types of nanoparticles: (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles and pSV-β-gal/nanoparticles as follows. The plasmid DNA was encapsulated in Tf-PEG-PEI. The final PEI nitrogen to DNA phosphate ratio of 6 was calculated from the concentrations of PEI and DNA (56). The pSico-CD44 shRNA (75 μg) and pFabpl-Cre (25 μg) were mixed in a total volume of 0.5 ml of HEPES-buffered glucose (5% glucose, 20 mm HEPES, pH 7.1). The Tf-PEG-PEI was dissolved in the same buffer (0.5 ml) and mixed with the plasmid mixture. The solutions (1 ml) were mixed by gentle pipetting and allowed to stand for 30 min at room temperature. The nanoparticles were purified by size exclusion chromatography in Sephacryl S 200 using a Bio-Rad column in a bed volume of 2.5 ml. Fractionation of the plasmid-nanoparticles was done as described previously (56). The elution of the nanoparticles was monitored at 254 nm. Fractions with major amounts of DNA were pooled and then quantified at 260 nm. The PEI content was analyzed by the trinitrobenzenesulfonic acid assay (57). The stability of the nanoparticles was determined by incubating in serum at 37 °C for 4 h, and it was found that the plasmids remained undegraded. Similarly, reporter plasmid pSV-β-gal-encapsulated nanoparticles (100 μg) were prepared and purified.
Nanoparticle Size Determination
The size distribution of nanoparticles was measured by dynamic light scattering with a NICOMP submicron particle sizer (NICOMP Particle Sizing Systems, Santa Barbara, CA). Five hundred μl of the diluted suspension was added to a culture tube and placed in the instrument cell holder. The scattered light intensity was inverted to size distribution by inverse Laplace transform using the CW370 software (NICOMP particle sizing systems). The particle size measurements were done in the Chemical Engineering Department, University of South Carolina, Columbia (courtesy of Prof. E. Jabbari). For the application, an in vivo nanoparticle size within a few hundred nanometers is a prerequisite (58, 59). As shown in Table 1, the mean size of our nanoparticles prepared for this study matches the published values (58, 59).
TABLE 1.
Size distribution of various plasmid-containing nanoparticles
The size distributions of plasmid/nanoparticles were measured by dynamic light scattering with a NICOMP submicron particle sizer as described under “Experimental Procedures.”
| Sample | Mean size | ±S.D. |
|---|---|---|
| nm | nm | |
| pSV-β-gal/nanoparticles | 35 | 20 |
| (pSico-CD44v6 shRNA + pFabpl-Cre)/nanoparticles | 210 | 130 |
Transfection of Cells with Plasmids
The validity of the pFabpl-Cre plasmid for cell transfection was done in two stages. First, the cells were transfected with the pSV-(EGFP)/β-galactosidase gene-switch reporter plasmid (Fig. 3) and divided into two groups. One group was used to detect green fluorescence, and the other group was divided and further transfected with pFabpl-Cre (positive control) or with pProbasin-Cre (negative control). In the appropriate cells, Cre recombinase was expressed, which removed the EGFP cassette within the loxP sites of the plasmid and allowed the SV promoter to initiate synthesis of the β-galactosidase. The cells were fixed in 0.2% glutaraldehyde. The activity of the enzyme in the intact cells was detected by the X-gal staining. Cells (1 × 104) were plated in a 24-well plate, and each cell line was plated in duplicate. The cells in each well were transfected with 1.0 μg of the pSV-(EGFP)/β-galactosidase gene-switch reporter plasmid using Lipofectamine (Invitrogen) following the manufacturer's protocol. Thirty six hours after transfection, the cells in the second well were further transfected with 1.0 μg of the pFabpl-Cre plasmid. Forty hours later, the cells in the duplicate well were fixed in 0.2% glutaraldehyde and stained with X-gal overnight at 37 °C. The cells in the first well were fixed in 4% paraformaldehyde and photographed for EGFP. The cells in the second well were photographed for X-gal. Second, we transfected cells with the pSV-β-gal/nanoparticles and pSV-β-gal via Lipofectamine (Invitrogen) following the manufacturer's instruction to compare the transfectibilty of Tf-coated nanoparticles. We measured the β-galactosidase enzyme activity in the cell extracts and stained the cells with X-gal.
FIGURE 3.
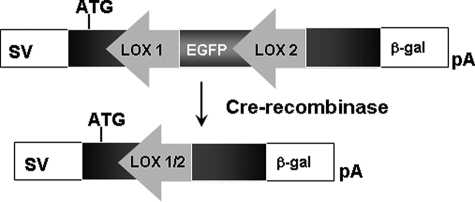
Validity of tissue-specific promoter using a gene-switch reporter construct. The specificity of promoter-driven Cre recombinase was determined by using a gene-switch reporter construct. The construct is an SV40 promoter-driven gene-switch plasmid containing floxed EGFP inserted into the open reading frame of the β-galactosidase (β-gal) gene resulting in appropriate translation of the EGFP gene showing green fluorescence but not the β-galactosidase gene. In the presence of appropriate Cre recombinase, the EGFP gene is excised, and the correct reading frame of the β-galactosidase gene is restored, which is now expressed, and blue color in Fig. 5 indicates that it can react with X-gal to produce blue dye.
β-Galactosidase Assay
Tissues were homogenized in the lysis buffer containing 10 mm CHAPS in citric acid (40 mm), disodium hydrogen phosphate (Na2HPO4) (40 mm), pH 7.3, 100 μg/ml phenylmethylsulfonyl fluoride. Similarly, the cells transfected with pSV-β-gal were extracted to compare the transfection efficiency. The homogenates were centrifuged at 12,000 × g for 15 min at 4 °C. The clear supernatants were used as the source of the enzyme. The protein concentrations of the extracts were determined by Folin phenol reagent (60).
The β-galactosidase activity was measured by determining the absorbance at 420 nm of the yellow product, o-nitrophenol, from the hydrolysis of a chromogenic substrate, o-nitrophenyl β-d-galactopyranoside. 150 μl of each extract was incubated with 150 μl of the assay buffer (citric acid (40 mm), Na2HPO4 (40 mm), pH 7.3, 300 mm NaCl, 4 mm MgCl2, 10 mm β-mercaptoethanol, and o-nitrophenyl β-d-galactopyranoside at 1.33 mg/ml). The mixture was vortexed and incubated at 37 °C for 30 min. The reaction was stopped by adding 700 μl of Tris (1 m), and the absorbance was measured at 420 nm in a spectrophotometer. One unit of β-galactosidase was the amount of enzyme that hydrolyzes 1 μmol of o-nitrophenyl β-d-galactopyranoside/min/mg of protein at 37 °C (molar extinction coefficient of o-nitrophenol is 4500 m-1 cm -1).
Preparation of Cell Lysates
Cells were washed in cold PBS two times (10 ml for 2 min), harvested with Versene, and then washed in cold PBS again. The cells were pelleted by centrifugation at 5000 × g at 4 °C. The pellet was treated with the lysis buffer containing 1% Nonidet P-40, 0.5 mm EGTA, 5 mm sodium orthovanadate, 10% (v/v) glycerol, 100 μg/ml phenylmethylsulfonyl fluoride, 1 μg/ml leupeptin, 1 μg/ml pepstatin A, 1 μg/ml aprotinin, and 50 mm HEPES, pH 7.5. The lysate was clarified by centrifugation at 12,000 × g for 10 min at 4 °C and then stored at -80 °C as described previously (34).
Reporter Gene Assay
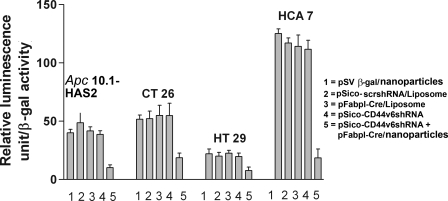
Apc 10.1-HAS2 clone cells and CT26, HT29, and HCA7 colon cancer cells were seeded in 24-well plates at 2 × 105 cells per well in complete medium. Twenty four h later, the cells were transfected by Lipofectamine with Cox-2 luciferase (1.0 μg per well). To normalize for transfection efficiency, all samples were co-transfected with pSV-β-gal (1.0 μg per well) and incubated for 12 h. Samples were transfected with pSico-scrshRNA, or pFabpl-Cre, or pSico-CD44v6 shRNA or treated with either pSV-β-gal/nanoparticle or (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles. After 72 h of incubation, cells were extracted with reporter lysis buffer (Promega). The luciferase activity in the extracts was then measured with luciferase substrate (Promega) in a Sirius luminometer. The β-galactosidase activity in the corresponding extracts was measured using o-nitrophenyl β-d-galactopyranoside as substrate as described above. The results were expressed as the ratio of relative luminescent units to β-galactosidase activity.
SDS-PAGE
Western blotting of cell lysates was done as described earlier (34). Intensities of the bands were quantified by densitometry.
Immunohistochemistry
Immunohistochemical detection of Cox-2 and pErbB2 in sections from paraffin-embedded intestine tissue with and without embedded tumors from various treatment groups was done using a standard avidin-biotin-peroxidase complex method as described by the manufacturer (Vector Laboratories). Goat polyclonal antibody against COX-2 and rabbit polyclonal antibody against pErbB2 (Santa Cruz Biotechnology) were used. As controls, duplicate sections were treated with isotype-matched immunoglobulin. Hyaluronan was detected in the sections using a biotinylated hyaluronan-binding protein and a standard avidin-biotin-peroxidase complex method as a probe as described previously (61). As controls for hyaluronan detection, duplicate sections were treated with testicular hyaluronidase before staining for hyaluronan.
Animal Experiments
Animals—Apc Min/+ mice were obtained from an in-house colony established in the Center for Colon Cancer Research, University of South Carolina, Columbia. They were genotyped by using allele-specific PCR analysis of tail DNA (62). Mice were randomly selected on sex and body weight bases, divided into groups, housed four per cage, and provided with standard rodent chow (Harland Teklad Rodent Diet 8604, Harland Teklad, Madison, WI) and water ad libitum. Three groups of Apc Min/+ mice (10 mice/group) were used. The Institutional Care and Use Committee of the University of South Carolina approved the animal protocol.
Study Design—The 12-week-old Apc Min/+ mice were divided into three groups as follows: group 1, plasmid (pSV-β-gal); group 2, pSV-β-gal/nanoparticles (100 μg); group 3, (pSico-CD44 shRNA (75 μg) plus pFabpl-Cre (25 μg))/nanoparticles. Thus, the animals in group 1 received 100 μg of naked plasmid/mouse, group 2 received 100 μg of plasmid nanoparticles/mouse, and group 3 received a total of 100 μg mixture of plasmids nanoparticles/mouse. Each group received four intraperitoneal injections for 10 days every other day.
Autopsy—Mice were euthanized with isoflurane in a vented chemical hood followed by cervical dislocation. Livers, spleens, kidneys, and lungs were harvested and frozen immediately in dry ice. The intestines of animals of various treatment groups were opened longitudinally using a small scissors. Small intestines from duodenum to cecum were removed from each animal and soaked in cold phosphate-buffered saline (PBS). The lumen of each intestine was cleaned by flushing with cold PBS to remove debris. Each intestine was divided into three segments as follows: near the stomach (duodenum), the middle (jejunum), and the far third (ileum) for assessing the adenoma load. Each intestinal segment was examined under a dissection microscope to count the number of adenomas, which were graded as >1 mm or <1 mm. After counting the tumors, the flattened intestines from various treatment groups, pSV-β-gal (group 1), pSV-β-gal/nanoparticles (group 2), and pSico-CD44v6 shRNA+pFabpl-Cre/nanoparticles (group 3), were divided into three equal parts as follows. One part from the groups 1 and 2 animals was rolled into “Swiss rolls” and fixed in OCT on dry ice and used for the detection of β-galactosidase by lacZ staining in the tissue. A second part of the flattened intestine from groups 1 to 3 was used for analysis of Cox-2 and pErbB2 in the paraffin sections. From the third part of groups 1-3, mucosa and enterocytes were removed from segments of intestine by lightly scraping the mucosal surface with the edge of a microscope slide, and tumors were excised and pooled. The tumor and adjacent normal tissues were subsequently used for measuring β-galactosidase activity. Lysates from these tumors and adjacent normal tissues were processed for Western blot analysis. RNA was extracted from these tumor and adjacent normal tissues and analyzed by RT-PCR.
X-Gal Staining of β-Galactosidase in Transfected Cell Lines and in Frozen Sections of Intestines from Apc Min/+ Mice
Transfected cell lines and the frozen sections were stained according to a protocol from the Welcome Trust Sanger Institute. The cell lines having β-galactosidase were fixed for 15 min in a fixative buffer (100 mm phosphate buffer, pH 7.3) supplemented with 5 mm EGTA, 2 mm MgCl2, and 0.2% glutaraldehyde. The cells were then washed twice with wash buffer (100 mm phosphate buffer, pH 7.3, supplemented with 2 mm MgCl2) for 5 min in each wash. Finally, the cells were incubated in X-gal staining buffer ((100 mm phosphate buffer, pH 7.3) supplemented with 2 mm MgCl2, 5 mm potassium ferrocyanide, 5 mm potassium ferricyanide, and 1 mg/ml X-gal) in a volume sufficient to cover the cells. The plates were covered and incubated overnight at 37 °C inside a humidified chamber. The sections were counterstained with eosin. The blue stains in cells were visualized under an inverted microscope (Leica DM1RB) and digitally photographed.
Statistics
Statistical analysis was performed between two groups by unpaired two tailed student's t test or unpaired t test with Welch corrections and among multiple groups by one-way ANOVA followed by Neuman-Keuls multiple comparison test. Statistical significance was determined and expressed as p < 0.05.
RESULTS
Transfection of Apc 10.1 Cells with HAS2-neo Up-regulates Synthesis of CD44v6 Variants—Fig. 4A (lanes 1 and 2) shows Western blot analyses of lysates of cultures of nonmalignant Apc 10.1 cells transfected with either vector-neo (control) or HAS2-neo prepared as described under “Experimental Procedures.” CD44v6 protein expression is markedly increased in the HAS2-neo lysate, whereas the standard form CD44s is not.
FIGURE 4.
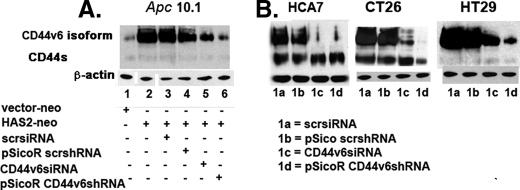
Inhibition of CD44 variants by CD44v6 siRNA and CD44v6 shRNA. A, Apc 10.1 cells were transfected with vector-neo or HAS2-neo and co-transfected with control scrsiRNA, or control pSicoR-scrshRNA, or CD44v6 siRNA, or pSicoR-CD44v6 shRNA plasmids. Transient transfection was carried out at 100 pmol of siRNA using Oligofectamine or 0.5 μg of shRNA using Lipofectamine (Invitrogen) according to the manufacturer's instructions. Cells were transfected with the siRNA or shRNA in 6-well plates with cells at 70-90% confluence. The cells were then incubated at 37 °C in 5% CO2 for 24 h, replated in 150-mm dishes, and allowed to grow for 48 h in complete medium as described under “Experimental Procedures.” B, similarly, HCA7, CT26, and HT29 colon cancer cells were transfected with the plasmids 1a-1d as indicated. The transfectants were grown for 72 h after transfections. Western blots of cell lysates were done with human CD44 and β-actin (loading control) antibodies.
CD44v6 siRNA and CD44v6 shRNA Inhibit CD44v6 Expression—CD44v6 siRNA duplexes and CD44v6 shRNA in U6 promoter-based pSicoR vectors (53) were prepared as described under “Experimental Procedures.” Cultures of Apc 10.1 cells were transfected with HAS2-neo (Fig. 4A), and cultures of HCA7, CT26, and HT29 colon cancer cells (Fig. 4B) were transfected with the vectors and CD44 siRNA (100 pmol, 72 h) as described under “Experimental Procedures.” Western blots in Fig. 4 show that the control scrambled siRNA and vectors containing scrambled shRNA (scrsiRNA and pSicoR-scrshRNA; Fig. 4, A, lanes 3 and 4, and B, lanes 1a and 1b) do not inhibit protein expression of CD44v6 variants or CD44s significantly. It is important to mention that this CD44v6 shRNA silences the CD44 variants 6 through 9 (v6-v9) mRNA (detected by RT-PCR) in the colon cancer cells (data not shown). Both CD44v6 siRNA and pSicoR-CD44v6 shRNA transfections significantly decreased protein expression of CD44v6 and CD44v6-containing variants (70-90%) with little or no effect on protein expression of CD44s (Fig. 4, A, lanes 5 and 6, and B, lanes 1c and 1d). These results indicate that the pSicoR-CD44v6 shRNA plasmid is as effective, if not more so, as the synthetic CD44v6 siRNA in silencing mRNA of CD44v6 and CD44v6-containing variants.
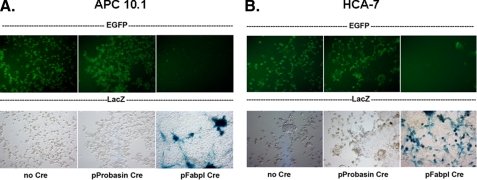
Expression of Cre Recombinase Depends on a Cell-specific Promoter—A Cre/Lox-induced translational gene-switch plasmid, pSV-EGFP-β-gal, as described previously (51), was used to test the specificity of Cre recombinase plasmid. As seen in Fig. 3, the pSV-EGFP-β-gal plasmid contains a reporter gene construct in which the SV40 promoter drives the expression of the EGFP gene flanked by Lox sites in the absence of Cre recombinase, whereas the SV40 promoter drives the expression of the β-galactosidase gene (lacZ gene) in the presence of Cre recombinase by restoring the reading frame of β-galactosidase. Apc 10.1 and HCA7 cells were transfected with the pSV-EGFP-β-gal plasmid, either alone (Fig. 5, no Cre) or with a Cre plasmid containing a prostate-specific promoter (Fig. 5, pProbasin-Cre), or with pFabpl-Cre (52), which produces Cre recombinase driven by the colon-specific Fabpl promoter exclusively in colon epithelial cells and cancer cells derived from them (Fig. 5, pFabpl-Cre). The upper panels of Fig. 5 show that green fluorescence protein EGFP is expressed in response to the pSV-EGFP-β-gal plasmid alone, and the bottom panels of Fig. 5 show that Cre recombinase is not actively produced in the presence of pProbasin-Cre. However, green fluorescence protein is not expressed when co-transfected with pFabpl-Cre, which actively produces Cre recombinase in these cells. The bottom panels of Fig. 5 show that β-galactosidase is present only in the cultures co-transfected with pFabpl-Cre. These results show that co-transfection with pFabpl-Cre excised the Lox-EGFP-Lox cassette with concomitant expression of the lacZ gene. The sporadic expression of lacZ was due to the nature of transfection by Lipofectamine. Similarly, lacZ gene expression was observed in both the cell lines when pSV-EGFP-β-gal was co-transfected with pCMV-Cre, a plasmid that drives the expression of Cre recombinase in a nonspecific manner (51) (data not shown). These results demonstrate that cell-specific Cre expression can be used to activate our gene of interest (CD44v6 shRNA) cloned into the conditional silencing vector pSico (gift from Dr. T. Jacks, Massachusetts Institute of Technology) (53).
FIGURE 5.
Tissue-specific gene expression with a gene-switch reporter construct. Transient transfection was carried out at 0.5 μg of plasmid using Lipofectamine (Invitrogen) according to the manufacturer's instructions. Cells were transfected with the plasmid in 6-well plates with cells at 70-90% confluence as described under “Experimental Procedures.” The cells were then incubated at 37 °C in 5% CO2 for 24 h, replated in 150-mm dishes, and allowed to grow for 48 h in complete medium. Apc 10.1 and HCA7 cells transfected with the pSV-EGFP-β-gal show EGFP expression in most, if not all, cells (upper left panels of A and B). Co-transfection with the nonspecific promoter-driven Cre cDNA plasmid (pProbasin-Cre) shows the same result (upper middle panels of A and B). Co-transfection with the tissue-specific promoter-driven Cre plasmid (pFabpl-Cre) eliminates the EGFP expression (upper right panels of A and B) with concomitant expression of β-galactosidase as demonstrated by X-gal staining (lower right panels of A and B). Cultures without the tissue-specific promoter did not express β-galactosidase (lower left and middle panels of A and B).
Expression of the Tf-R in Cancer Cells—The Western blot analyses in Fig. 6 show that Tf-R is highly expressed in the cancer cell lines CT26, HT29, and HCA7 (Fig. 6, lanes 3-5), ∼10-fold more than in the noncancerous Apc 10.1 cells (Fig. 6, lane 1). However, Tf-R is expressed at the same high level when the Apc 10.1 cells are transfected with HAS2-neo cDNA and cultured for 72 h (Fig. 6, lane 2). The cancer cells and the HAS2-transfected cells also synthesize much higher levels of hyaluronan than the nontransfected Apc 10.1 cells (50). Furthermore, HAS2-transfected Apc 10.1 cells have malignant properties in contrast to the nontransfected parent cells (50). These results show that Tf-R expression correlates with high level synthesis of hyaluronan and with malignancy in these cells and therefore is a good candidate for cell-specific targeting.
FIGURE 6.
Elevated expression of HAS2 stimulates expression of Tf-R protein in the colon pre-neoplastic (Apc 10. 1) cells and is present at high levels in the cancer cells. Apc 10.1 cells were transfected with the vector-neo (lane 1) or HAS2-neo cDNA (lane 2). These transfectants as well as HCA7, CT26, and HT29 colon cancer cells were grown for 72 h, and Western blots of cell lysates were analyzed for transferrin receptor and β-actin as a loading control. The data presented in this figure are representative of three independent experiments.
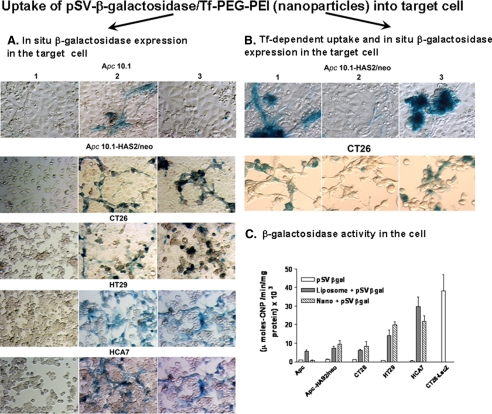
Tissue-specific Delivery of pSV-β-galactosidase Plasmid/Nanoparticles in Vitro—Nanoparticles with encapsulated pSV-β-gal were prepared with transferrin on their surface (pSV-β-gal/nanoparticles) as described under “Experimental Procedures.” The Apc 10.1 cells, cancer cells, and Apc 10.1-HAS2 cells were transfected with the nanoparticles (8 μg of DNA/ml), or the pSV-β-gal alone, or the plasmid with liposome at the same DNA concentrations. pSV-β-gal alone did not show any galactosidase activity (Fig. 7A, column 1 micrographs). pSV-β-gal with both liposome (Fig. 7A, column 2 micrographs) and nanoparticle encapsulation (Fig. 7A, column 3 micrographs) showed widespread β-galactosidase activity (Fig. 7, blue dye deposits) in cancer cells and Apc 10.1-HAS2 cells that express high Tf-R (Fig. 6). On the other hand, Apc 10.1 cells, a low Tf-R strain (Fig. 6), showed no or very insignificant β-galactosidase activity (Fig. 7A, column 3 micrograph for Apc 10.1 cells with very low blue dye deposits). These results implicate the presence of high levels of Tf-R for preferential uptake of pSV-β-gal/nanoparticles and the expression of β-galactosidase in cancer cells and Apc 10.1-HAS2 cells. We further confirmed the selectivity of transfection of Tf-coated nanoparticles for Tf-R by blocking the Tf-R on the Apc 10.1-HAS2 cells and CT26 cells with the Tf-R antibody prior to transfection with pSV-β-gal/nanoparticles (Fig. 7B, column 3 with pSV-β-gal/nanoparticles) versus column 2 (Tf-R antibody plus pSV-β-gal/nanoparticles); column 1 represents liposome transfection of pSV-β-gal in the micrographs. A similar result was obtained when the cells were transfected with pSV-β-gal/nanoparticles in the presence of a saturating concentration of Tf for its receptor (data not shown). In addition, the results also indicate that at 8 μg of DNA/ml, the pSV-β-gal/nanoparticles have equivalent or higher transfection efficiency than liposome transfection with pSV-β-gal directly. Nanoparticles with lower DNA concentrations were less effective than liposome transfection (data not shown).
FIGURE 7.
Delivery of pSV-β-galactosidase-nanoparticles in cell culture. A, in situ β-galactosidase expression in the target cell. The Apc 10.1 cells, Apc 10.1-HAS2 clone, and the CT26, HT29, and HCA7 cells were treated for 48 h with the pSV-β-gal alone (panel 1), or with the pSV-β-gal with liposome (panel 2), or with the pSV-β-gal/nanoparticles (35 nm average diameter, 8 μg of pSV-β-gal/ml) (panel 3). The average size of the pSV-β-gal/nanoparticles is ∼35 nm ± 20 nm (Table 1). The transfected cells were fixed in 0.2% glutaraldehyde in PBS and washed twice in PBS as described under “Experimental Procedures.” The cells were treated with a β-galactosidase staining solution as described under “Experimental Procedures ” and digitally photographed. B, transferrin-dependent uptake and in situ β-galactosidase expression in the target cell. The Apc 10.1-HAS2 cells were transfected with the pSV-β-gal with liposome (panel 1), or treated with Tf-R antibody and followed by transfection with the pSV-β-gal/nanoparticles (8 μg pSV-β-gal/ml) (panel 2), or treated with the pSV-β-gal/nanoparticles (8 μg pSV-β-gal/ml) alone (panel 3). The transfected cells were fixed in 0.2% glutaraldehyde in PBS and washed twice in PBS as described under “Experimental Procedures.” The cells were treated with a β-galactosidase staining solution as described under “Experimental Procedures ” and digitally photographed. C, cell-free extracts of parallel cultures were prepared in 10 mm CHAPS buffer and assayed for β-galactosidase activity using o-nitrophenyl β-d-galactopyranoside as substrate as described under “Experimental Procedures.” The results are expressed as micromoles of o-nitrophenol formed per min/mg protein and represent ± S.D. of triplicate assays from the untransfected, or liposome-transfected, or nanoparticles-treated cultures for each cell type.
Cell-free extracts of identical cultures were prepared, and β-galactosidase activity was measured as described under “Experimental Procedures.” The graph in Fig. 7C shows that the cultures treated with pSV-β-gal plasmids alone had barely detectable levels of β-galactosidase activity. In contrast, the pSV-β-gal delivered with liposome or in encapsulated nanoparticles showed significant and nearly equivalent increases in activity. From the in vitro transfection experiment, we found that the half-life of β-galactosidase expression is ∼30 h. The results in Fig. 7C demonstrate that the transfection efficiency of the pSV-β-gal/nanoparticle in CT26, Apc 10.1-HAS2, HT29, and HCA7 cells were 8.5-, 10-, 28-, and 36-fold, respectively, whereas that of the pSV-β-gal/liposome in these cells were 6.0-, 7.0-, 20-, and 45-fold, respectively, compared with the pSV-β-gal plasmid alone. The transfection efficiency with the HCA7 cells as measured by the enzyme activity was nearly as high as the control lacZ stable transfectant of CT26 cells (Fig. 7C).
The high transfection efficiency of the nanoparticles as shown in Fig. 7 suggests that they have low or no toxicity and that they are delivered effectively into the cells through uptake by the high concentration of Tf-R on their surfaces (Fig. 6). After uptake, the nanoparticles must be released from endosomes into the cytoplasm where the pSV-β-gal cDNA is protected from cytosol nuclease by the PEG coat in the polyplex. Random migration into the nucleus during cell division would then allow transcription with the production of the β-galactosidase (Fig. 7).
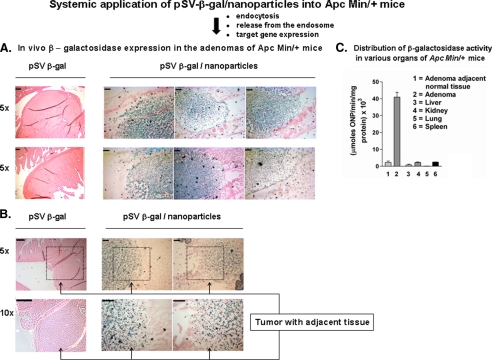
Tissue-specific Delivery of the pSV-β-gal/Nanoparticles in Vivo—Adenomas and adjacent normal tissue were isolated from Apc Min/+ mice and lysates prepared as described under “Experimental Procedures.” Fig. 8 shows Western blot analyses of the lysates for Tf-R and β-actin as a loading control. Similar to the differences in Tf-R in the Apc 10.1 cells compared with the cancer cells, the normal tissue showed much less Tf-R than in the adenoma. This result provides evidence that transferrin on the nanoparticles can preferentially bind to the adenoma cells. We tested this by comparing nanoparticle delivery of pSV-β-gal plasmid with the plasmid alone in Apc Min/+ mice bearing intestinal adenomas. Ten 7-8-week-old mice in each group were given intraperitoneal doses of either the nanoparticles (pSV-β-gal/nanoparticles) or the plasmid alone (100 μg/100 μl of plasmid DNA) every other day for 10 days as described under “Experimental Procedures.” Fig. 9 shows sections of six representative adenomas that stain for β-galactosidase (blue dye deposits), whereas the adjacent tissues do not show significant amounts of stain. All 10 animals treated with the nanoparticles had stained adenomas, whereas no adenomas in the animals treated with the pSV-β-gal alone showed staining (Fig. 9A, pink color with black outline). The graph in Fig. 9 shows β-galactosidase activity in extracts of the adenomas, in adjacent normal mucosal tissue, and in other organs isolated from mice treated with the nanoparticles. The in vivo transfection into adenomas by pSVβ-gal/nanoparticle was much greater (>90%, Fig. 9, A and C) than the small to negligible levels in the adjacent normal tissue (Fig. 9B) and the other organs (not shown). This result indicates that nanoparticles carrying plasmids accumulate into the cells of adenomas by enhanced permeation and retention effects. These results validate the Tf-decorated nanoparticle delivery system into adenomas via the Tf-R in vivo.
FIGURE 8.
Elevated expression of Tf-R in Apc Min/+ mice adenomas compared with adjacent normal colon mucosa. The tissue extracts from adenomas and adjacent normal tissue were processed for Western blot and immunoblotted with Tf-R antibody and β-actin (as loading control). The data presented in this figure are representative of three independent experiments.
FIGURE 9.
Delivery of pSV-β-gal nanoparticles in Apc Min/+ mice. A, in vivo β-galactosidase expression in the adenomas of Apc Min/+ mice. Ten Apc Min/+ mice received pSV-β-gal plasmids (100 μg/100 μl, intraperitoneally) alone and 10 received pSV-β-galactosidase (pSV-β-gal)/nanoparticles (100 μg/100 μl, intraperitoneally) every other day. Mice showed no sign of toxicity throughout the experiment. At day 10, mice were euthanized with isoflurane in a vented chemical hood followed by cervical dislocation. Livers, spleens, kidneys, and lungs were harvested and frozen immediately in dry ice as described under “Experimental Procedures.” After counting the tumors, the flattened intestines from various treatment groups such as pSV-β-gal (group 1), pSV-β-gal/nanoparticles (group 2) were rolled into Swiss rolls and fixed in OCT on dry ice. Frozen sections carrying the adenomas were stained with X-gal (blue dye deposits around the tumor tissue at ×5 magnification) and eosin, whereas the tumor bearing animals treated with pSV-β-gal showing no lacZ stain were outlined with a black line. To visualize whether low lacZ expression occurs in the surrounding tissue, the sections were examined at low (×5) and high (×10) magnifications. Scale bars, 200 μm (B). The selected portions in black square area at ×5 magnification are shown at ×10 magnification. The normal mucosal epithelium (at ×10 magnification) shows little or no lacZ expression (B). Scale bars, 200 μm. C, β-galactosidase activity in adenomas, adjacent normal tissue and other organs of Apc Min/+ mice. From the second part of the groups 1-3, mucosa and enterocytes were removed from segments of intestine by lightly scraping the mucosal surface with the edge of a microscope slide, and tumors were excised and pooled. The tumor and adjacent normal tissues were subsequently used for measuring β-galactosidase activity as described under “Experimental Procedures.” The enzyme assays in the tissue extracts were done in three sets of individual experiments. The graph shows a representative result (average of n = 3 ± S.D.).
Silencing the CD44v6 Gene in Vitro Efficiently Regulates the Cell Survival Protein and COX-2 Reporter Gene Expressions—We have shown that endogenous HA-CD44 interactions activate an ErbB2 signaling pathway that stimulates cell survival/cell proliferation through COX-2 induction in hyaluronan-overexpressing HIEC6 intestinal epithelial cells and HCA7 colon cancer cells (30, 31). This suggests that hyaluronan interaction with CD44v6 and CD44v6-containing variants, which are significantly elevated in colon cancer (18, 63), may account for constitutively increased pathologic induction of COX-2 in colon cancer, a pathway that is mediated by ErbB2/PI3K/β-catenin signaling (30, 31). If true, silencing the expression of CD44v6 and CD44v6-containing variants by using the Tf-nanoparticle delivery of CD44v6 shRNA within the tumor cell may inhibit the cell survival pathway and adenoma growth in Apc Min/+ mice. The delivery system consists of three components as follows: 1) Tf-PEG-PEI; 2) the pSico-CD44v6 shRNA (Fig. 2A); and 3) pFabpl-Cre (52). In principle, the Tf on the surfaces of the nanoparticles will deliver the plasmids primarily inside the tumor cells, which express high levels of Tf-R (see Fig. 8). The plasmids entering the cells become active only in intestine/colon tumor cells and work in trans, i.e. Cre recombinase produced from pFabpl-Cre acts on pSico-CD44v6 shRNA to express v6 shRNA, a folded back stem-loop that is processed into siRNA (Fig. 1) that blocks translation of CD44v6 and CD44v6-containing variants by cleaving their mRNAs. Because the CD44v6 shRNA is against variant exon 6, which is abundant in malignant colon cells, the CD44 standard form should remain unaffected.
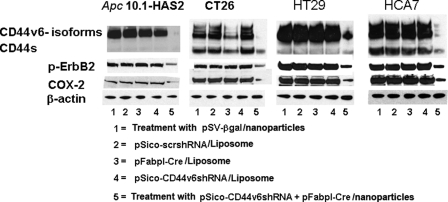
The (pSico-CD44v6 shRNA Plus pFabpl-Cre)/Nanoparticles Silence Expression of CD44v6 and CD44v6-containing Variants in Cancer Cells in Vitro—Validation of these constructs for silencing the expressions of CD44v6 and CD44v6-containing variants is shown in Fig. 10. Apc 10.1-HAS2 clone cells and CT26, HT29, and HCA7 colon cancer cells were treated with Tf-nanoparticles that contained both pSico-CD44v6 shRNA plus pFabpl-Cre (Fig. 10, lanes 5) or with the pSV-β-gal (Fig. 10, lanes 1), as well as with control plasmids transfected with liposomes (Fig. 10, lanes 2-4). The Western blot analyses show that only the nanoparticles with the two plasmids (Fig. 10, lanes 5) silenced CD44v6 and CD44v6-containing variants. The strong suppression of virtually all variants suggests that the v6 exon is likely to be present in all of the expressed variants, similar to the results for the CD44 variants in HeLa cells (54). It also unexpectedly inhibited CD44s to a smaller extent in the CT26 and HCA7 cell lines. As expected, the pSico-CD44v6 shRNA showed little or no effect when transfected with liposomes (Fig. 10, lanes 4), which shows that it has no activity in the absence of the pFabpl-Cre.
FIGURE 10.
(pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles knock down CD44v6, COX-2, and p-ErbB2 in colon pre-neoplastic and cancer cells. Apc 10.1-HAS2 clone cells and CT26, HT29, and HCA7 colon cancer cells were transfected with liposomes and either pSico-scrshRNA, or pFabpl-Cre, or pSico-CD44v6 shRNA, or were treated with nanoparticles containing either pSV-β-gal or the combination of pSico-CD44v6 shRNA plus pFabpl-Cre and cultured for 72 h. Lysates were prepared and probed in Western blots with antibodies to COX-2, human CD44, p-ErbB2, and β-actin. Total ErbB2 remained unchanged in all the treatment groups (data not shown).
Our previous studies (30, 31) showed that endogenous interaction of hyaluronan with CD44 variants regulates COX-2 and phosphorylated p-ErbB2 expression in HCA7 cells. The Western blots in Fig. 10 show that silencing CD44v6-containing variants selectively down-regulates Cox-2 and p-ErbB2 both in the Apc 10.1-HAS2 clone and the three cancer cells (lanes 5). These results provide strong evidence for a hyaluronan-CD44v6 signaling pathway through ErbB2/Cox-2 that can be effectively abrogated by silencing CD44v6 and CD44v6-containing variants.
High expression of COX-2 is implicated in colon tumorigenesis (26, 30, 31). Therefore, we determined whether the expression of a luciferase reporter gene under COX-2 promoter (pCOX-2-luc) is affected by depleting CD44v6 expression in the Apc 10.1-HAS2 clone cells and the CT26, HT29, and HCA7 colon cancer cells by the use of the (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles. All the cultures in the experiments of Fig. 10 were co-transfected for 12 h with pCOX-2-luc and pSV-β-gal (for normalization). They were then treated for 72 h with the same set of reagents as for the experiments in Fig. 11. The luciferase activity and β-galactosidase activity were measured for lysates of each culture, and the results are expressed as ratios of relative light unit to β-galactosidase activity. Fig. 11 shows similar results to those observed in Fig. 10. Only the nanoparticles with combined treatment with the pSico-CD44v6 shRNA and pFabpl-Cre inhibited luciferase activity in each cell culture system. Overall these results demonstrate the possibility that the nanoparticles with the combined plasmids can be used for enhanced transfection efficiency to give the desired endosomal release of CD44v6 shRNA in tumors in vivo.
FIGURE 11.
(pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles suppress COX-2 reporter gene expression in vitro. Apc 10.1-HAS2 clone cells and CT26, HT29, and HCA7 colon cancer cells were co-transfected for 12 h with the pCOX-2 luciferase and the pSV-β-gal reporter plasmids. They were then further transfected for 72 h with liposomes plus either pSico-scrshRNA, or pFabpl-Cre, or pSico-CD44v6 shRNA, or were treated with nanoparticles that contained either pSV-β-gal or the combined pSico-CD44v6 shRNA plus pFabpl-Cre. Cells were extracted with reporter lysis buffer followed by measurement of luciferase and β-galactosidase activity. The graph shows one of three experiments with nearly equivalent results and gives the average ± S.D. of three measurements for each culture.
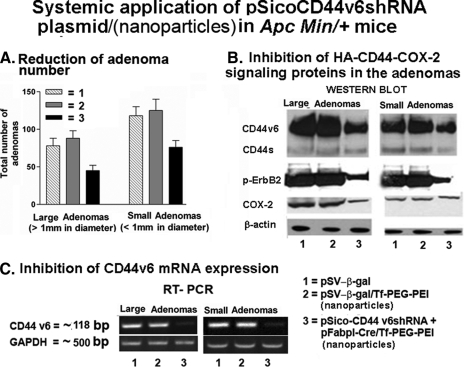
Silencing CD44v6 and CD44v6-containing Variants in Vivo Reduces Tumor Numbers in Apc Min/+ Mice—Apc Min/+ mice carrying intestinal adenomas were treated intraperitoneally(100 μg/100 μl) every other day for 10 days with the pSV-β-gal alone (100 μg) or with pSV-β-gal/nanoparticles (100 μg) or (pSico-CD44v6 shRNA (75 μg) plus pFabpl-Cre (25 μg))/nanoparticles as described under “Experimental Procedures.” In a more recent study we replaced intraperitoneal injection by intravenous injection for the nanoparticle delivery. Tumor numbers were counted in the intestine and then isolated from the mice intestines (Fig. 12A). Lysates were prepared and assayed for Western blot analyses (Fig. 12B) and mRNA analyses (Fig. 12C). In general, the number of tumors (large and small combined) in the intestine before a 10-day treatment regime is approximately between 190 and 230 (80-90 large and 110-120 small). The size of pSico-CD44v6 shRNA + pFabpl-Cre-bearing nanoparticle was much greater (∼7-fold) compared with that of pSVβ-gal (Table 1). Assume the mean size 35 nm of pSVβ-gal/nanoparticle carry one plasmid. The preparation of nanoparticles with combined plasmid is a mixture of various sizes of nanoparticles ranging from 70 nm (two plasmids) to 340 nm (approximately 10 plasmids). The in vitro transfection efficiency of the combined plasmid (pSico-CD44v6 shRNA + pFabpl-Cre) was high (equivalent 80-90%; Figs. 10 and 11). Similar results were obtained when the plasmid combination was delivered into the tumors of Apc Min/+ mice where they produce an amount of shRNA equivalent to an appreciable amount of the tumor accumulated siRNA, which in turn inhibited expression and activity of key signaling proteins in HA/CD44v6 pathway (Fig. 12 and 13). Compared with the other two control treatments, treatment with the nanoparticles that contained the combined plasmids: 1) reduced large (>1 mm diameter) tumor numbers by 40-50% and small (<1 mm) tumor numbers by 25-35% compared with the control treatments; 2) inhibited protein expression of CD44v6 variants (40-50%), phosphorylated p-ErbB2 (50-60%) and Cox-2 (50-60%) compared with controls; and 3) nearly completely silenced CD44v6 mRNA expression compared with controls (Fig. 12C). Total ErbB2 was the same in all treatments (data not shown). It is important to mention that other growth factors, receptor tyrosine kinases, functional state of the CD44 molecules, and that of their intracellular complexes are also important determinants of the HA/CD44-dependent oncogenic functions.
FIGURE 12.
Reduction in number of adenomas, protein expression, and RT-PCR analysis in Apc Min/+ mice adenomas after 10 days of plasmid-nanoparticles treatment. A, inhibition of adenoma growth. Thirty Apc Min/+ mice were randomly divided into three groups. Group 1 received pSV-β-galactosidase (100 μg/100 μl, intraperitoneally) alone. Group 2 received pSV-β-gal nanoparticles (100 μg/100 μl, intraperitoneally) targeted to the Tf-R. Group 3 received pSico-CD44v6 shRNA (75 μg) plus pFabpl-Cre (25 μg)/nanoparticles intraperitoneally every other day. 10 days after the beginning of treatment (i.e. 1 day after the last injection of the plasmid), the animals were sacrificed, and the large (>1 mm) and small (<1 mm) adenomas were counted. Large and small adenomas were excised 10 days after the pSico-CD44v6 shRNA plus pFabpl-Cre-/nanoparticles treatment. The control experiment with pSico-CD44v6 shRNA/nanoparticles or pFabpl-Cre/nanoparticles gave similar results as pSV-β-gal/nanoparticles as described under “Experimental Procedures.” After counting the tumors in groups 1-3, mucosa and enterocytes were removed from segments of intestine by lightly scraping the mucosal surface with the edge of a microscope slide and tumors were excised and pooled. The tumor and adjacent normal tissues were subsequently used for Western blot and RT-PCR analysis. B, Western blot analysis. Tissue extracts were processed for Western blot analysis of CD44, pErbB2, TErbB2, Cox-2, and β-actin. Total ErbB2 remained unchanged in all the treatment groups (data not shown). C, RT-PCR. Total RNAs were extracted from the above set of experiments and analyzed by RT-PCR for mouse CD44v6 and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) expression.
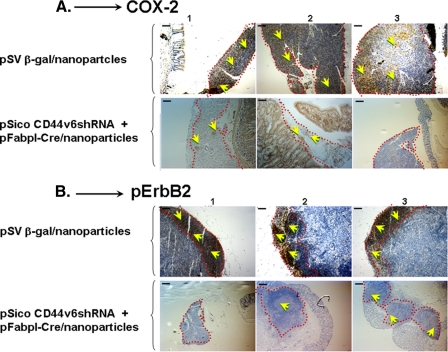
FIGURE 13.
Analysis of Cox-2 and pErbB2 in the paraffin sections of intestines having tumor tissues from various treatment groups of three different Apc Min/+ mice were demonstrated from the experiment as described in Fig. 12 legend. Panels 1-3 refer to different animals. For immunohistochemical analysis, antigens were retrieved from tissue sections from treated and untreated tumors and then treated with antibodies against Cox-2 (A) and pErbB2 (B) following standard avidin-biotin-peroxidase complex method. After probing for pErbB2 and Cox-2, the sections were counter-stained with hematoxylin. The size-matched tumors (outlined with red dotted lines) from various treatments were compared. The upper panels in each case are adenomas from mice treated with pSV-β-gal nanoparticles (controls) that stain heavily for Cox-2 and p-ErbB2 (HRP stain, indicated by yellow arrows) as expected from their high expression in these cells. In contrast, sections in the lower panels in each case are adenomas from mice treated with the nanoparticles that contain pSico-CD44v6 shRNA plus pFabpl-Cre plasmids show smaller sparse or negligible HRP staining for Cox-2 and pErbB2. Importantly, it should be noted that the normal intestine villi treated with (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles stained heavily for Cox-2 and pErbB2, demonstrating the specificity of the nanoparticle for the adenoma (to target CD44v6 mRNA) versus normal intestine. CD44v6 shRNA plus pFabpl-Cre plasmid/nanoparticle treatment did not show any significant change in the HA content in tumor sections (data not shown). Scale bars, 200 μm.
Fig. 13 shows sections of adenomas (outlined by red dotted lines) from individual mice stained for Cox-2 (Fig. 13A) and pErbB2 (Fig. 13B). The upper panels in Fig. 13 in each case are adenomas treated with pSVβ-gal/nanoparticles (controls) that stain heavily for Cox-2 and pErbB2 (HRP stain) as expected from their high expression in Apc 10.1-HAS2 cells as seen in Fig. 10. In contrast, sections in the lower rows of Fig. 13, A and B, adenomas treated with the nanoparticles that contain pSico-CD44v6 shRNA plus pFabpl-Cre plasmids, show smaller, sparse, or negligible HRP staining for Cox-2 and pErbB2 compared with hematoxylin stain. The feeble HRP staining is designated by the yellow arrows in the lower rows of Fig. 13, A and B, indicating the ability of pSico-CD44v6 shRNA + pFabpl-Cre/nanoparticles to target mRNA of CD44v6 and CD44v6-containing variants in the tumors. Although Fig. 13 demonstrates results of three representative Apc Min/+ mice, the rest of the animals of the respective groups showed very similar results. Moreover, immunohistochemical results validate our results in Fig. 12B. In addition, gross examinations of the liver, kidney, lung, and spleen did not show any abnormalities in the animals. Overall, the results demonstrate that the (pSico-CD44v6 shRNA plus pFabpl-Cre plasmids)/nanoparticles have entered the adenoma cells and effectively silenced CD44 variants mRNA with disruption of the downstream HA-CD44 variant signaling pathway.
DISCUSSION
Our previous studies involving perturbation of HA/CD44 signaling helped to understand how epithelial cell-derived HA, and its interaction with CD44, can influence malignant properties. However, these studies did not address the tumor cell response to cell-specific perturbation of the HA-CD44 interaction at the genetic level in vitro and in vivo. In this regard, our results (Fig. 12, A-C) on tumor cell-specific delivery of CD44v6 shRNA demonstrated inhibition of HA/CD44v6 signaling in contrast to attenuation by perturbation of the HA-CD44 interaction using antagonists such as HA oligomers and soluble CD44, and also showed consequent inhibition of in vivo tumor growth. This result indicates that there are differences between attenuation and conditional deletion of mouse CD44v6 and CD44v6-containing variants isoform. Because increased HA induces CD44v6 isoform expression (Fig. 4) and Tf-R expression (Fig. 6), a major goal of this study is to target HA-CD44v6 signaling pathways by CD44v6 shRNA without damaging nontumor tissues, some of whose functions are also CD44s-dependent as seen in Fig. 12B. Thus, besides CD44 variants, we have also chosen Tf-R to actively target tumor cells by Tf-decorated nanoparticles to deliver plasmids that result in cell-specific release of CD44v6 shRNA. Tf-R is present in crypt stem cells of intestine (64). It is likely that the nanoparticles will interact with TF-R and be endocytosed by the crypt stem cells that express the Fabpl gene (65). The Fabpl promoter active in the intestine/colon will produce recombinase from the pFabpl-Cre plasmid and release CD44v6 shRNA in the crypt stem cells. Because the CD44v6 and CD44v6-containing variants are produced in tumors, but not in normal cells, the release of CD44v6 shRNA will therefore have no deleterious effect on crypt stem cells.
We examined the level of Tf-R in the cellular models and tumor cells from Apc Min/+ mice. Our results in Figs. 6 and 8 demonstrate the high expression of Tf-R in colon tumor cells and in the Apc 10.1-HAS2 stable clone as well as in Apc Min/+ mice tumor tissues. These results justify the use of Tf on nanoparticles. In this context, it should also be noted that high expression of Tf-R in hyaluronan up-regulated cells is not merely coincidental. Because increased HAS2 gene expression influences malignant properties in normal epithelial cells (30, 31, 33-35, 66), and because malignant cancer cells express high levels of Tf-R (42, 43), this suggests that increased expression of HAS2 can elevate expression of Tf-R in transformed Apc-HAS2 transfectants. In addition, based on the examination of human transferrin (67) and hyaluronan synthase-2 (68) promoter regions, it is possible that Sp1 transcription factor could activate transcriptions of both HAS2 and Tf-R genes. In addition, Sp3 transcription factors also activate transcription of the HAS2 gene (68). Sp1 knock-out embryos died around E11 of gestation (69), a time point precisely after the death of HAS2 knock-out embryos E9.5-10 (70). This suggests that HA may stimulate Sp transcription factors binding to the HAS2 promoter to induce HAS2 activation. Because Sp1 is also a potential candidate in controlling the tissue-specific expression of Tf-R, and because increased HA induces Tf-R expression in tumor cells or transformed cells (Fig. 6), it is also possible that increased HA regulates Sp transcription factors for the transformed phenotype of epithelial cells and modulates the expression of Tf-R in vivo.
We have demonstrated that “post-transcriptional targeting” by shRNA is possible within tumor cells. Using the intestine/colon-specific promoter-driven Cre (pFabpl-Cre), we confirmed the release of CD44v6 shRNA from a conditionally silenced pSico-CD44v6 shRNA that perturbs the expression of CD44v6 and CD44v6-containing isoforms and reduces adenoma growth in Apc Min/+ mice by inhibiting the HA/CD44v6/Cox-2-induced cell survival pathway. This was accomplished in several steps. These include the following.
We first determined the specificity of the intestine/colon pFabpl-Cre for the Apc 10.1 cells and HCA7 cells. To address this, we delivered pFabpl-Cre and reporter plasmid (pSV-EGFP-β-gal) within the Apc 10.1 cells and HCA7 cells. As seen from Fig. 5, the pFabpl-Cre eliminates the EGFP expression cassette from pSV-EGFP-β-gal with simultaneous expression of β-galactosidase in Apc 10.1 cells and HCA7 cells, whereas prostate-specific pProbasin-Cre has no effect.
For shRNA to be a useful therapeutic agent, the tumor cell-specific plasmid delivery strategy consists of three components as follows: 1) Tf-PEG-PEI; 2) pSico-CD44v6 shRNA; and 3) pFabpl-Cre (52) that produces Cre recombinase selectively in the intestine/colon. These three components when mixed together form nanoparticles whose exterior contains the Tf and interior contains the two plasmids, pSico-CD44v6 shRNA and pFabpl-Cre. The latter provides excellent specificity for the release of CD44v6 shRNA in the intestine/colon. Because the CD44v6 shRNA is against variant exon 6 (v6) that silences the CD44v6 and CD44v6-v9 mRNA, which are abundant in malignant colon cells, it has limited effect on the CD44 standard form (Figs. 3 and 10). Therefore, we determined whether the nanoparticles approach can inhibit HA-CD44v6 interaction and associated anti-apoptotic cell survival pathways necessary for tumor growth in Apc Min/+ mice. However, transfection efficiency of the nanoparticles is crucial for shRNA gene delivery. The efficacy of transfection can be elevated by actively targeting Tf-R on the tumor cell surface (42), which was demonstrated in this study in cellular models and in vivo.
We first examined the size of the nanoparticles, which is a critical parameter for the circulation through the blood, uptake by the target cells, and removal by cells in liver and spleen. Our findings of the size of the nanoparticles (Table 1) are consistent with the findings from others (58, 59). Next, to study the role of CD44v6 shRNA expression vector in colon tumorigenesis, we exploited the transfection of pSV-β-gal/nanoparticles in cellular models as well as in Apc Min/+ mice. The data of Figs. 7 and 9 show that the pSV-β-gal/nanoparticles with high Tf concentrations allow transfection comparable with that of liposome transfection for the following three reasons. 1) These nanoparticles are shielded from nonspecific interaction. 2) The pSV-β-gal/nanoparticles bind selectively to the target cell Tf-R. 3) Binding of the Tf to Tf-R is followed by endocytosis and endosomal release of the reporter plasmid pSV-β-gal, which is translated and expressed in the adenomas (Fig. 9) and cancer cells (Fig. 7), a crucial step in the plasmid delivery process.
For tissue-specific gene delivery of Tf-mediated CD44v6 shRNA in tumor cells, it is crucial to have high levels of Tf-R expression in the tumor cells as has been observed in Fig. 9. Next, we tested whether the HAS2 overexpression with greatly increased HA production in Apc 10.1-HAS2 clones effectively induces CD44v6 expression similar to several different metastatic colon cancer cells, HCA7, CT26, and HT29 cells (Figs. 4 and 10). We then delivered the (pSico-CD44v6 shRNA plus pFabpl-Cre)/nanoparticles systemically in Apc Min/+ mice. The results in Figs. 9 and 12 clearly demonstrate that plasmid DNA and reporter gene expression and the consequent inhibitions of the HA/CD44v6-Cox-2 signaling were in the tumors of all 10 mice tested for these experiments. Moreover, the procedure is well endured by the Apc Min/+ mice with no sign of toxicity. Retardation in tumor development was found in all of the 10 mice by injecting the nanoparticles complex every other day for 10 days satisfying the requirement for tissue-specific gene delivery.
Of particular interest is the use of HA/CD44 antagonists in animal models of tumor. Thus, we extended our studies that highlight the potential importance of the mechanism of constitutive HA-CD44 interaction antagonists that can suppress tumor growth by inhibiting HA-CD44v6-Cox-2 signaling axis, and other malignant properties as a therapeutic target for spontaneous tumors in Apc Min/+ mice. Because HA and CD44 are potential candidates for cancer treatment (13, 71), in the present work we chose CD44 variants as prototype targets for the following reasons. 1) HA-CD44v6 interaction in cancer cells regulates activation of receptor tyrosine kinases that lead to cell survival and proliferation, and to COX-2 expression and function in colon cancer cells (30, 31). 2) CD44v6 (Fig. 12, B and C, and Fig. 13) and Cox-2 are overexpressed in the tumors of Apc Min/+ mice (48, 49). 3) Because the Apc Min/+ mouse model is also relevant to the study of the early stages of human colorectal carcinogenesis (48, 72), our results in the present study validate the use of Apc Min/+ mice as a successful in vivo model to test the effect of CD44 shRNA for the inhibition of tumor growth. For therapeutic gene delivery vectors, optimization of the intracellular steps of the transfection and endosomal release are necessary in future studies. Targeting Tf-R using the dual plasmid/nanoparticle system for a longer period will increase the probability of killing more cancer cells as well as significant inactivation of phospho-ErbB2 and COX-2 protein. Our present study addresses a time frame appropriate to show proof of principle for our novel targeting nanoparticles.
These results highlight the potential importance of the mechanism of HA/CD44 variant interaction-induced anti-apoptotic signaling and other malignant properties as a therapeutic target in colon carcinoma. Understanding how selective alterations of HA/CD44v6 signaling pathway contribute to intestine/colon tumor development and progression will likely provide the novel approaches for therapeutic intervention. Using our experimentally verified Tf-coated nanoparticle delivery of plasmids in the tumor cells, our current studies are directed toward further examining the effect of CD44 variant shRNA on inhibition of growth of various tumors, including prostate and breast, and HA-CD44 variant-associated oncogene signaling.
Acknowledgments
We thank Dr. Bryan P. Toole (Medical University of South Carolina) and Dr. Franklin G. Berger (University of South Carolina) for their support and interest in this work. We also thank Dr. Esmaiel Jabbari (University of South Carolina) for chemical synthesis and size determination of nanoparticles. We thank Margarett H. Romano, Department of Pathology and Laboratory of Medicine, for performing immunohistochemistry.
This work was supported, in whole or in part, by National Institutes of Health Grants P20 RR016434 (to S. M., S. G., and R. R. M.), HL RO1 33756, Mitral-07 CVD 04 (to R. R. M.), 1 P30AR050953 (to V. C. H.), and P20RR017698 (to S. M.), RO1 CA073839 and RO1 CA082867 (to S. M. and S. G.). This work was also supported by Hollings Cancer Center Department of Defense Grant GC-3319-05-44598 (to S. M. and R. R. M.) and Medical University of South Carolina URC Projects 2204000-24330 (to S. M.) and 2204000-24329 (to S. G.).
Footnotes
The abbreviations used are: HA, hyaluronan; Cox-2, cyclooxygenase-2; shRNA, short hairpin RNA; siRNA, silencing RNA; Tf, transferring; Tf-R, Tf receptor; PEG, polyethylene glycol; PEI, polyethyleneimine; Min, multiple intestinal neoplasia; CD44v, CD44 variant isoforms; EGFP, enhanced green fluorescent protein; RT, reverse transcription; PBS, phosphate-buffered saline; HRP, horseradish peroxidase; X-gal, 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside; FAP, familial adenomatous polyposis; CHAPS, 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonic acid; Apc, adenomatous polyposis coli.
S. Misra, unpublished data.
References
- 1.Sprenger, C. C., Plymate, S. R., and Reed, M. J. (2008) Br. J. Cancer 98 250-255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Markwald, R. R., Fitzharris, T. P., Bank, H., and Bernanke, D. H. (1978) Dev. Biol. 62 292-316 [DOI] [PubMed] [Google Scholar]
- 3.Lee, J. Y., and Spicer, A. P. (2000) Curr. Opin. Cell Biol. 12 581-586 [DOI] [PubMed] [Google Scholar]
- 4.Toole, B. P. (2004) Nat. Rev. Cancer 4 528-539 [DOI] [PubMed] [Google Scholar]
- 5.Hascall, V. C., Majors, A. K., De La Motte, C. A., Evanko, S. P., Wang, A., Drazba, J. A., Strong, S. A., and Wight, T. N. (2004) Biochim. Biophys. Acta 1673 3-12 [DOI] [PubMed] [Google Scholar]
- 6.Adamia, S., Maxwell, C. A., and Pilarski, L. M. (2005) Curr. Drug Targets Cardiovasc. Haematol. Disord. 5 3-14 [DOI] [PubMed] [Google Scholar]
- 7.Tammi, R. H., Kultti, A., Kosma, V. M., Pirinen, R., Auvinen, P., and Tammi, M. I. (2008) Semin. Cancer Biol. 18 288-295 [DOI] [PubMed] [Google Scholar]
- 8.Naor, D., Nedvetzki, S., Golan, I., Melnik, L., and Faitelson, Y. (2002) Crit. Rev. Clin. Lab. Sci. 39 527-579 [DOI] [PubMed] [Google Scholar]
- 9.Lesley, J., and Hyman, R. (1992) Eur. J. Immunol. 22 2719-2723 [DOI] [PubMed] [Google Scholar]
- 10.Lesley, J., He, Q., Miyake, K., Hamann, A., Hyman, R., and Kincade, P. W. (1992) J. Exp. Med. 175 257-266 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sleeman, J., Rudy, W., Hofmann, M., Moll, J., Herrlich, P., and Ponta, H. (1996) J. Cell Biol. 135 1139-1150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.van Weering, D. H., Baas, P. D., and Bos, J. L. (1993) PCR Methods Appl. 3 100-106 [DOI] [PubMed] [Google Scholar]
- 13.Ponta, H., Sherman, L., and Herrlich, P. A. (2003) Nat. Rev. Mol. Cell Biol. 4 33-45 [DOI] [PubMed] [Google Scholar]
- 14.Naor, D., Sionov, R. V., Zahalka, M., Rochman, M., Holzmann, B., and Ish-Shalom, D. (1998) Curr. Top. Microbiol. Immunol. 231 143-166 [DOI] [PubMed] [Google Scholar]
- 15.Ochiai, S., Nakanishi, Y., Mizuno, K., Hashimoto, S., Inutsuka, S., Kawasaki, M., Yatsunami, J., and Hara, N. (1997) Nihon Kyobu Shikkan Gakkai Zasshi 35 1179-1185 [PubMed] [Google Scholar]
- 16.Kurozumi, K., Nishida, T., Nakao, K., Nakahara, M., and Tsujimoto, M. (1998) World J. Surg. 22 853-858 [DOI] [PubMed] [Google Scholar]
- 17.Ayhan, A., Tok, E. C., and Bildirici, I. (2001) Gynecol. Oncol. 80 355-358 [DOI] [PubMed] [Google Scholar]
- 18.Ishida, T. (2000) Surg. Today 30 28-32 [DOI] [PubMed] [Google Scholar]
- 19.Yoshida, K., Bolodeoku, J., Sugino, T., Goodison, S., Matsumura, Y., Warren, B. F., Toge, T., Tahara, E., and Tarin, D. (1995) Cancer Res. 55 4273-4277 [PubMed] [Google Scholar]
- 20.Omara-Opyene, A. L., Qiu, J., Shah, G. V., and Iczkowski, K. A. (2004) Lab. Investig. 84 894-907 [DOI] [PubMed] [Google Scholar]
- 21.Miyake, H., Hara, I., Okamoto, I., Gohji, K., Yamanaka, K., Arakawa, S., Saya, H., and Kamidono, S. (1998) J. Urol. 160 1562-1566 [PubMed] [Google Scholar]
- 22.Kim, H., Yang, X. L., Rosada, C., Hamilton, S. R., and August, J. T. (1994) Arch. Biochem. Biophys. 310 504-507 [DOI] [PubMed] [Google Scholar]
- 23.Takahashi, K., Stamenkovic, I., Cutler, M., Saya, H., and Tanabe, K. K. (1995) Oncogene 11 2223-2232 [PubMed] [Google Scholar]
- 24.Ropponen, K. M., Eskelinen, M. J., Lipponen, P. K., Alhava, E., and Kosma, V. M. (1998) Scand. J. Gastroenterol. 33 301-309 [DOI] [PubMed] [Google Scholar]
- 25.Gupta, R. A., and Dubois, R. N. (2001) Nat. Rev. Cancer 1 11-21 [DOI] [PubMed] [Google Scholar]
- 26.Simmons, D. L., Botting, R. M., and Hla, T. (2004) Pharmacol. Rev. 56 387-437 [DOI] [PubMed] [Google Scholar]
- 27.Chen, Q., Shinohara, N., Abe, T., Harabayashi, T., and Nonomura, K. (2004) J. Urol. 172 2153-2157 [DOI] [PubMed] [Google Scholar]
- 28.Dohadwala, M., Luo, J., Zhu, L., Lin, Y., Dougherty, G. J., Sharma, S., Huang, M., Pold, M., Batra, R. K., and Dubinett, S. M. (2001) J. Biol. Chem. 276 20809-20812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tsujii, M., Kawano, S., and DuBois, R. N. (1997) Proc. Natl. Acad. Sci. U. S. A. 94 3336-3340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Misra, S., Hascall, V. C., Berger, F. G., Markwald, R. R., and Ghatak, S. (2008) Connect. Tissue Res. 49 219-224 [DOI] [PubMed] [Google Scholar]
- 31.Misra, S., Obeid, L. M., Hannun, Y. A., Minamisawa, S., Berger, F. G., Markwald, R. R., Toole, B. P., and Ghatak, S. (2008) J. Biol. Chem. 283 14335-14344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yu, Q., and Toole, B. P. (1996) J. Biol. Chem. 271 20603-20607 [DOI] [PubMed] [Google Scholar]
- 33.Ghatak, S., Misra, S., and Toole, B. P. (2005) J. Biol. Chem. 280 8875-8883 [DOI] [PubMed] [Google Scholar]
- 34.Misra, S., Toole, B. P., and Ghatak, S. (2006) J. Biol. Chem. 281 34936-34941 [DOI] [PubMed] [Google Scholar]
- 35.Misra, S., Ghatak, S., and Toole, B. P. (2005) J. Biol. Chem. 280 20310-20315 [DOI] [PubMed] [Google Scholar]
- 36.Misra, S., Ghatak, S., Zoltan-Jones, A., and Toole, B. P. (2003) J. Biol. Chem. 278 25285-25288 [DOI] [PubMed] [Google Scholar]
- 37.Ghatak, S., Misra, S., and Toole, B. P. (2002) J. Biol. Chem. 277 38013-38020 [DOI] [PubMed] [Google Scholar]
- 38.Heidel, J. D., Hu, S., Liu, X. F., Triche, T. J., and Davis, M. E. (2004) Nat. Biotechnol. 22 1579-1582 [DOI] [PubMed] [Google Scholar]
- 39.Paul, C. P., Good, P. D., Winer, I., and Engelke, D. R. (2002) Nat. Biotechnol. 20 505-508 [DOI] [PubMed] [Google Scholar]
- 40.Raper, S. E., Chirmule, N., Lee, F. S., Wivel, N. A., Bagg, A., Gao, G. P., Wilson, J. M., and Batshaw, M. L. (2003) Mol. Genet. Metab. 80 148-158 [DOI] [PubMed] [Google Scholar]
- 41.Kursa, M., Walker, G. F., Roessler, V., Ogris, M., Roedl, W., Kircheis, R., and Wagner, E. (2003) Bioconjugate Chem. 14 222-231 [DOI] [PubMed] [Google Scholar]
- 42.Bellocq, N. C., Pun, S. H., Jensen, G. S., and Davis, M. E. (2003) Bioconjugate Chem. 14 1122-1132 [DOI] [PubMed] [Google Scholar]
- 43.Qian, Z. M., Li, H., Sun, H., and Ho, K. (2002) Pharmacol. Rev. 54 561-587 [DOI] [PubMed] [Google Scholar]
- 44.Miyoshi, Y., Ando, H., Nagase, H., Nishisho, I., Horii, A., Miki, Y., Mori, T., Utsunomiya, J., Baba, S., and Petersen, G. (1992) Proc. Natl. Acad. Sci. U. S. A. 89 4452-4456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fodde, R., and Smits, R. (2001) Trends Mol. Med. 7 369-373 [DOI] [PubMed] [Google Scholar]
- 46.Sasai, H., Masaki, M., and Wakitani, K. (2000) Carcinogenesis 21 953-958 [DOI] [PubMed] [Google Scholar]
- 47.Moser, A. R., Pitot, H. C., and Dove, W. F. (1990) Science 247 322-324 [DOI] [PubMed] [Google Scholar]
- 48.Wielenga, V. J., van der Neut, R., Offerhaus, G. J., and Pals, S. T. (2000) Adv. Cancer Res. 77 169-187 [DOI] [PubMed] [Google Scholar]
- 49.Oshima, M., Dinchuk, J. E., Kargman, S. L., Oshima, H., Hancock, B., Kwong, E., Trzaskos, J. M., Evans, J. F., and Taketo, M. M. (1996) Cell 87 803-809 [DOI] [PubMed] [Google Scholar]
- 50.De Giovanni, C., Landuzzi, L., Nicoletti, G., Astolfi, A., Croci, S., Micaroni, M., Nanni, P., and Lollini, P. L. (2004) Int. J. Cancer 109 200-206 [DOI] [PubMed] [Google Scholar]
- 51.Kaczmarczyk, S. J., and Green, J. E. (2001) Nucleic Acids Res. 29 E56-E56 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Saam, J. R., and Gordon, J. I. (1999) J. Biol. Chem. 274 38071-38082 [DOI] [PubMed] [Google Scholar]
- 53.Ventura, A., Meissner, A., Dillon, C. P., McManus, M., Sharp, P. A., Van Parijs, L., Jaenisch, R., and Jacks, T. (2004) Proc. Natl. Acad. Sci. U. S. A. 101 10380-10385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Cheng, C., Yaffe, M. B., and Sharp, P. A. (2006) Genes Dev. 20 1715-1720 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Saporite-Irwin, S., Geist, R., and Gutman, D. (1997) BioTechniques 23 424-427 [DOI] [PubMed] [Google Scholar]
- 56.Boeckle, S., von Gersdorff, K., van der Piepen, S., Culmsee, C., Wagner, E., and Ogris, M. (2004) J. Gene Med. 6 1102-1111 [DOI] [PubMed] [Google Scholar]
- 57.Snyder, S. L., and Sobocinski, P. Z. (1975) Anal. Biochem. 64 284-288 [DOI] [PubMed] [Google Scholar]
- 58.Kircheis, R., Wightman, L., Schreiber, A., Robitza, B., Rossler, V., Kursa, M., and Wagner, E. (2001) Gene Ther. 8 28-40 [DOI] [PubMed] [Google Scholar]
- 59.Kircheis, R., Wightman, L., and Wagner, E. (2001) Adv. Drug Deliv. Rev. 53 341-358 [DOI] [PubMed] [Google Scholar]
- 60.Lowry, O. H., Rosebrough, N. J., Farr, A. L., and Randall, R. J. (1951) J. Biol. Chem. 193 265-275 [PubMed] [Google Scholar]
- 61.Yu, Q., and Toole, B. P. (1995) BioTechniques 19 122-124, 126-129 [PubMed] [Google Scholar]
- 62.Tucker, J. M., Davis, C., Kitchens, M. E., Bunni, M. A., Priest, D. G., Spencer, H. T., and Berger, F. G. (2002) Cancer Lett. 187 153-162 [DOI] [PubMed] [Google Scholar]
- 63.Choi, S. H., Takahashi, K., Eto, H., Yoon, S. S., and Tanabe, K. K. (2000) Int. J. Cancer 85 523-526 [DOI] [PubMed] [Google Scholar]
- 64.Roy, C. N., and Enns, C. A. (2000) Blood 96 4020-4027 [PubMed] [Google Scholar]
- 65.Moser, A. R., Dove, W. F., Roth, K. A., and Gordon, J. I. (1992) J. Cell Biol. 116 1517-1526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zoltan-Jones, A., Huang, L., Ghatak, S., and Toole, B. P. (2003) J. Biol. Chem. 278 45801-45810 [DOI] [PubMed] [Google Scholar]
- 67.Holloway, K., Sade, H., Romero, I. A., and Male, D. (2007) J. Mol. Biol. 365 1271-1284 [DOI] [PubMed] [Google Scholar]
- 68.Monslow, J., Williams, J. D., Fraser, D. J., Michael, D. R., Foka, P., Kift-Morgan, A. P., Luo, D. D., Fielding, C. A., Craig, K. J., Topley, N., Jones, S. A., Ramji, D. P., and Bowen, T. (2006) J. Biol. Chem. 281 18043-18050 [DOI] [PubMed] [Google Scholar]
- 69.Marin, M., Karis, A., Visser, P., Grosveld, F., and Philipsen, S. (1997) Cell 89 619-628 [DOI] [PubMed] [Google Scholar]
- 70.Camenisch, T. D., Spicer, A. P., Brehm-Gibson, T., Biesterfeldt, J., Augustine, M. L., Calabro, A., Jr., Kubalak, S., Klewer, S. E., and McDonald, J. A. (2000) J. Clin. Investig. 106 349-360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Nakazawa, H., Yoshihara, S., Kudo, D., Morohashi, H., Kakizaki, I., Kon, A., Takagaki, K., and Sasaki, M. (2006) Cancer Chemother. Pharmacol. 57 165-170 [DOI] [PubMed] [Google Scholar]
- 72.Boivin, G. P., Washington, K., Yang, K., Ward, J. M., Pretlow, T. P., Russell, R., Besselsen, D. G., Godfrey, V. L., Doetschman, T., Dove, W. F., Pitot, H. C., Halberg, R. B., Itzkowitz, S. H., Groden, J., and Coffey, R. J. (2003) Gastroenterology 124 762-777 [DOI] [PubMed] [Google Scholar]