Abstract
Background
Ehrlichia chaffeensis is a rickettsial agent responsible for an emerging tick-borne illness, human monocytic ehrlichiosis. Recently, we reported that E. chaffeensis protein expression is influenced by macrophage and tick cell environments. We also demonstrated that host response differs considerably for macrophage and tick cell-derived bacteria with delayed clearance of the pathogen originating from tick cells.
Results
In this study, we mapped differences in the promoter regions of two genes of p28-Omp locus, genes 14 and 19, whose expression is influenced by macrophage and tick cell environments. Primer extension and quantitative RT-PCR analysis were performed to map transcription start sites and to demonstrate that E. chaffeensis regulates transcription in a host cell-specific manner. Promoter regions of genes 14 and 19 were evaluated to map differences in gene expression and to locate RNA polymerase binding sites.
Conclusion
RNA analysis and promoter deletion analysis aided in identifying differences in transcription, DNA sequences that influenced promoter activity and RNA polymerase binding regions. This is the first description of a transcriptional machinery of E. chaffeensis. In the absence of available genetic manipulation systems, the promoter analysis described in this study can serve as a novel molecular tool for mapping the molecular basis for gene expression differences in E. chaffeensis and other related pathogens belonging to the Anaplasmataceae family.
Background
Ehrlichia chaffeensis, an obligate, intracellular, tick-borne bacterium that belongs to the family Anaplasmataceae, is responsible for an emerging disease in humans called human monocytic ehrlichiosis (HME) [1,2]. The transmitting vector of E. chaffeensis, Amblyomma americanum, acquires the pathogen during a blood meal from an infected host [2]. Host cell adaptation and establishment of persistent infection in tick and vertebrate hosts are critical for successful completion of the E. chaffeensis lifecycle and, similarly, for other tick-transmitted rickettsiales of the genera Ehrlichia and Anaplasma [3-7]. It is necessary for the tick-transmitted pathogens to have evolved strategies that support host cell adaptation and to establish persistent infections. There may be many ways by which the pathogens persist; strategies may include altering the host response [8,9], varying expressed proteins relative to time post-infection and differential host-specific protein expression [10-19].
Recently, we reported that Ehrlichia species alter the expression of many proteins in a host cell-specific manner [18-21]. Differentially expressed proteins include outer membrane proteins made from p28-Omp multigene locus having 22 tandomly arranged paralogous genes of E. chaffeensis [18-20]. The major expression from this locus is limited to a subset of genes and is also influenced by vertebrate and tick cell environment. P28-Omp 14 protein is the major expressed protein when E. chaffeensis is grown in tick cells, whereas p28-Omp 19 is expressed predominantly by the organism in macrophages. We also reported that the pathogen originating from tick cells persists longer in a vertebrate host and the host response is significantly different for tick cell-derived bacteria compared with bacteria originating from macrophages [9].
Little is known about the promoter structures and transcriptional regulation of E. chaffeensis genes and their contributions to alter the gene expression in response to tick and vertebrate host cell environments. Promoter analysis under in vivo conditions is not possible at this time because of a lack of methods to transform E. chaffeensis. In the current study, we report the first description of mapping promoter regions of two host-specific differentially expressed genes of E. chaffeensis.
Results
Primer extension analysis of p28-Omp genes 14 and 19
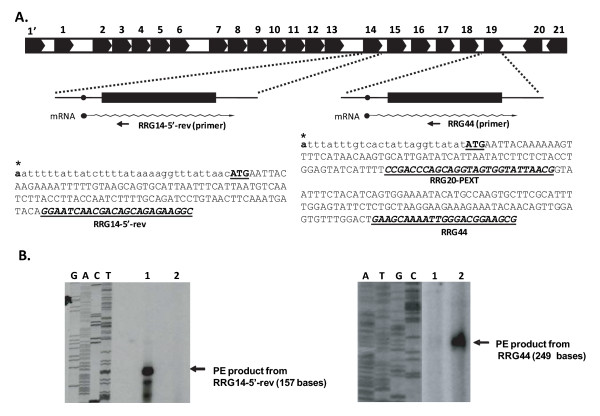
Host-specific differential protein expression from numerous E. chaffeensis genes, including from p28-Omp multi-gene locus, has been reported previously [18-20]. To evaluate the gene expression at transcription level, primer extension analysis was performed for p28-Omp genes 14 and 19 with macrophage and tick cell-derived E. chaffeensis RNA (Figure 1A and 1B). The primer extended products for genes 14 and 19 were detected in tick cell- and macrophage-derived E. chaffeensis RNA, respectively (Figure 1). The analysis also aided in identifying the transcription start sites for genes 14 and 19 located at 34 and 26 nucleotides upstream to the initiation codons, respectively (Figure 1). The nucleotide at the transcription start sites for both the genes is adenosine.
Figure 1.
Primer extension (PE) analysis of p28-Omp genes 14 and 19. Panel A has a cartoon spanning all 22 genes [37]. This panel also has an expansion of cartoons for genes 14 and 19 with predicted transcripts, the primers used for the PE analysis and sequences of the primer extended products with transcription start sites identified with asterisks. PE analysis products resolved on a sequencing gel are shown in panel B. Blots on the left and right represent the data for transcripts of genes 14 and 19, respectively. A sequence ladder for the gene 14 analysis was prepared by using the same primer used for the PE analysis but with a DNA template spanning the gene 14 sequence. For gene 19, PE analysis was performed with RRG 44 primer, and the sequencing ladder was generated by using RRG20-PEXT primer with a gene 19 DNA template. (Lane 1, E. chaffeensis RNA from tick cells; lane 2, E. chaffeensis RNA from macrophages).
Transcriptional analysis by quantitative RT-PCR at different times post-infection
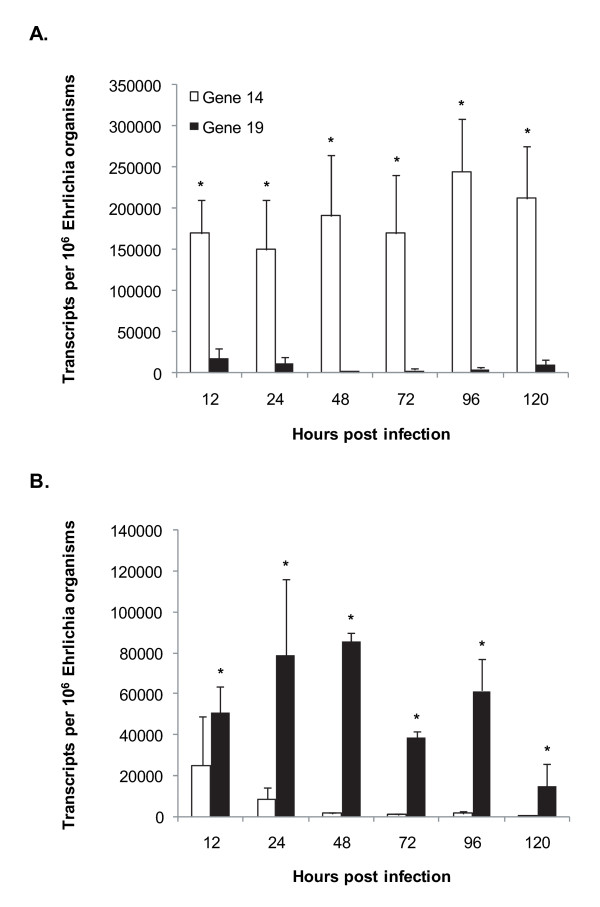
Our previous studies suggested that both p28-Omp genes 14 and 19 are transcriptionally active in E. chaffeensis originating from vertebrate macrophages and tick cells but the expression levels are different [9,19]. The quantitative gene expression differences for genes 14 and 19 were determined by TaqMan-based real-time RT-PCR analysis (quantitative RT-PCR) (Figure 2). Consistent with the previous observations, transcripts for genes 14 and 19 were detected in RNA isolated from both host cell backgrounds. In tick cell-derived E. chaffeensis, p28-Omp gene 14 expression remained higher than expression of p28-Omp gene 19 (Figure 2A). The gene 14 expression in E. chaffeensis also remained high for all time points analyzed post-inoculation in tick cells. In macrophage-derived E. chaffeensis, expression levels were reversed with significantly higher expression for gene 19 (Figure 2B).
Figure 2.
Quantitative RT-PCR analysis. TaqMan-based quantitative RT-PCR analysis was performed with RNA isolated from tick cell (A) and macrophage (B) cultures harvested at different times postinfection. Transcript numbers were estimated and presented per million E. chaffeensis organisms. Data are presented with SE values calculated from three independent experiments (P ≤ 0.05).
P28-Omp 14 and 19 promoter regions sequence analysis
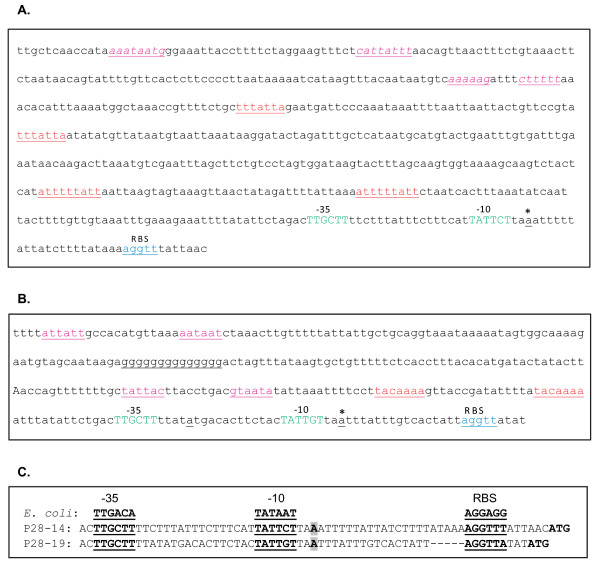
The entire non-coding sequences upstream to genes 14 and 19 were evaluated to identify sequences similar to the consensus E. coli RNA polymerase binding site sequences, -10 and -35, and ribosome binding site sequences (RBS) (Figure 3). Consensus -10 and -35 elements were identified and are located few bases upstream to the transcription start sites mapped by primer extension analysis (Figure 3). Similarly, putative RBS sequences [22] were identified 7 and 4 nucleotides upstream to the initiation codon of genes 14 and 19, respectively. Genes 14 and 19 sequences upstream to the predicted -10 and -35 sequences differed considerably in their lengths and homology (Figure 3A and 3B). The gene 14 upstream sequence is 581 bp in length, which is 273 bp longer than the gene 19 upstream sequence (308 bp). The sequences included several gene-specific direct repeats and palindrome sequences. In addition, a unique 14 nucleotide-long 'G' rich sequence was detected in the gene 19 sequence. The consensus -35 sequence was identical for both the genes, whereas the -10 and RBS sequences differed by one nucleotide each (Figure 3C). Relative distances of the consensus -10 and -35 sequences from transcription start sites also remained the same for both the genes (Figure 3C).
Figure 3.
P28-Omp genes 14 and 19 promoter region sequence analysis. Upstream sequences of genes 14 (panel A) and 19 (panel B) were evaluated for the presence of direct repeats (red text), palindromic sequences (pink text) and for the presence of unique sequences (G-rich region), consensus -35 and -10 regions (green text) and ribosome binding sites (blue text). Panel C shows the comparison of -10, -35 and ribosome binding sites of genes 14 and 19 with the E. coli consensus sequences. Transcription start sites for the genes mapped by primer extension analysis are identified with bold and grey color highlighted text or with an asterisk. Dashes were introduced in the p28-Omp gene 19 sequence to create alignment with the gene 14 sequence.
Evaluation of promoter activities of the sequences upstream to the coding regions of the p28-Omp genes 14 and 19
The transcription analysis assessed by direct RNA mapping and TaqMan-based RT-PCR methods revealed quantitative differences in gene expression for p28-Omp genes 14 and 19, which is influenced by invertebrate and vertebrate host cell environments. It is unclear how the host cell environments influence the Ehrlichia gene expression. Promoter analysis of these differentially expressed genes will be valuable for gaining insights about how differential expression is achieved by E. chaffeensis in vertebrate and tick host environments. Promoter characterization in vivo for E. chaffeensis is not feasible at this time because genetic manipulation systems are yet to be established. Alternatively, characterization of E. chaffeensis promoters may be performed in E. coli or with E. coli RNA polymerase as reported for several C. trachomatis genes [23-30].
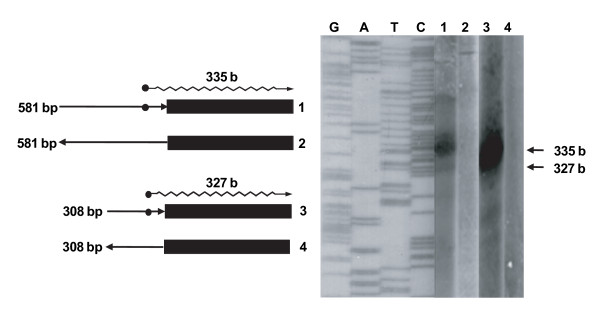
To validate the use of E. coli for mapping the promoters of E. chaffeensis genes,in vitro transcription assays were performed for p28-Omp 14 and 19 promoter regions with E. coli RNA polymerase by following methods reported for Chlamydia species [28-30]. Predicted in vitro transcripts, as estimated from transcription start sites mapped by primer extension described previously, were detected only when p28-Omp 14 and 19 complete upstream sequences were ligated to a segment of lacZ coding sequence (Figure 4). In vitro transcripts were absent in the reactions that contained the complete gene 14 and 19 promoter regions ligated in reverse orientation (Figure 4).
Figure 4.
In vitro transcription analysis. In vitro transcription analysis was performed for the complete upstream sequences of genes 14 and 19 in forward and reverse orientations ligated to a partial lacZ gene segment (301 bp) (solid black boxes). The orientation of ligated promoter regions is shown by arrowhead lines (right arrowhead line, forward orientation; left arrowhead line, reverse orientation). Wiggled arrowhead lines show predicted transcripts of 335 bases for gene 14 and 327 bases for gene 19. Sequence segments and the predicted transcripts for genes 14 and 19 are shown as cartoons on the left, and the observed transcripts are shown on the right of the panels. Puc18 plasmid DNA was used as the template to generate a sequence ladder with an M13 forward primer. Numbers 1 and 2 refer to the constructs for in vitro transcription for gene 14, and 3 and 4 refer to in vitro transcription templates for gene 19.
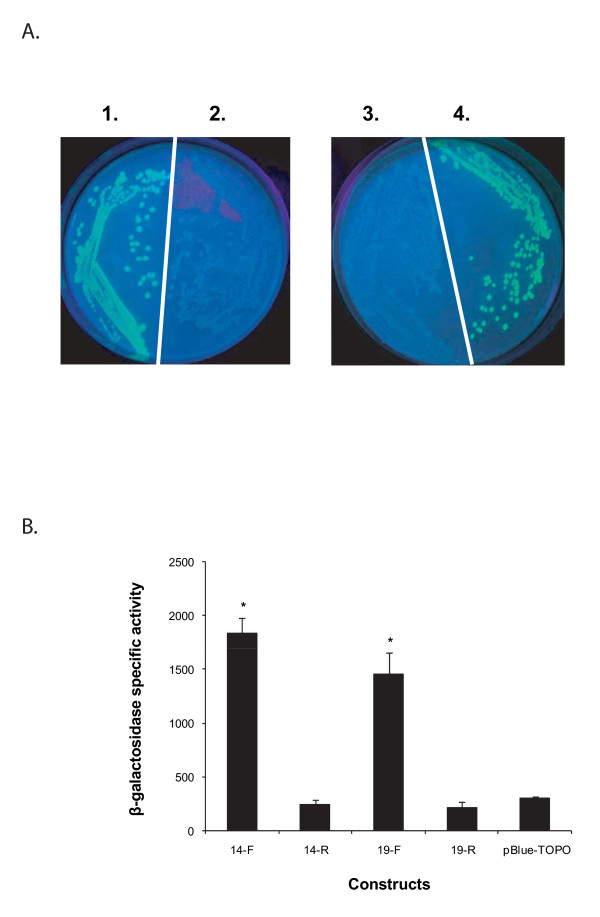
Upstream sequences for p28-Omp genes 14 or 19 were subsequently evaluated in E. coli. Transformants of E. coli containing promoter regions of genes 14 and 19 cloned in front of the promoterless green fluorescent protein (GFP) coding sequence in the pPROBE-NT plasmid were positive for green fluorescence as visualized by the presence of green color colonies (Figure 5A). E. coli transformed with pPROBE-NT plasmids alone were negative for the green fluorescence. The GFP expression was verified by Western blot analysis with GFP-specific polyclonal sera (not shown). Promoter activities for upstream sequences of genes 14 and 19 were further confirmed by another independent method (i.e., by assessing the β-galactosidase activity after inserting the sequences in front of the promoterless lacZ gene in pBlue-TOPO plasmid). The E. coli transformants with plasmids having gene 14 or 19 sequences cloned in correct orientation had significantly more β-galactosidase activity (P ≤ 0.001) than the baseline activity observed for constructs with no promoter sequences or when the sequences were inserted in reverse orientation (Figure 5B).
Figure 5.
(A) Green fluorescent protein (GFP) constructs evaluated for the promoter activity of p28-Omp genes 14 and 19. The pPROBE-NT plasmids containing the promoterless GFP gene (2 and 3) and upstream sequences of genes 14 and 19 in front of the GFP gene (1 and 4, respectively) and a construct containing no promoter sequence were evaluated for GFP expression in E. coli. (B) LacZ constructs evaluated for the promoter activity of p28-Omp genes 14 and 19. The pBlue-TOPO vector containing promoterless lacZ gene (pBlue-TOPO) and upstream sequences of genes 14 and 19 inserted in forward (14-F and 19-F) and reverse orientations (14-R and 19-R) were evaluated for β-galactosidase activity in E. coli. Data are presented with SD values calculated from four independent experiments (P ≤ 0.001).
Promoter deletion analysis
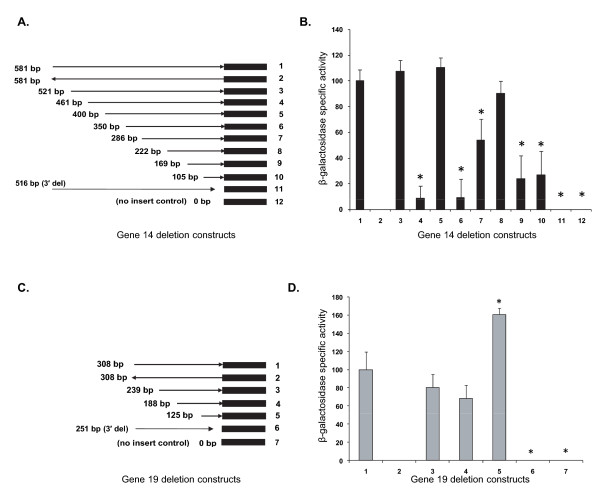
Deletion analyses were performed to assess whether the promoter activities are influenced by the sequences upstream to the transcription start sites of genes 14 and 19; β-galactosidase activity for several pBlue-TOPO plasmid constructs with segments deleted from the 5' end for both the genes were evaluated (Figure 6). Deletions to the sequences ranged from 60 to 476 bp for p28-Omp gene 14 and 69 to 183 bp for gene 19. All deletion constructs for gene 14, except for deletions having 461 and 350 bp segments, had significantly higher β-galactosidase activity compared with negative controls lacking no insert and the insert in the reverse orientation. The first 60 bp deletion from the 5' end resulted in no significant change in β-galactosidase activity compared with that observed for the full-length insert, whereas a deletion of an additional 60 bp caused a decline of about 90% of the enzyme activity. The β-galactosidase activity was restored completely by an additional 61 bp deletion. Further deletion of another 50 bp also resulted in another near-complete loss of activity. Subsequent deletions of 64 bp each caused a stepwise restoration of the enzyme activity to 54 and 91%, respectively. Deletion of another 53 bp caused another drop in β-galactosidase activity to 24%, which remained unaffected with an additional deletion of a 64 bp fragment (Figure 6A and 6B). Similar deletion analysis performed for the gene 19 upstream sequence also resulted in altered β-galactosidase activity compared with the full-length sequence (Figure 6, panels C and D). The 5' end deletions of 69 and 120 bp for this gene resulted in a 20 and 30% decline, respectively, in enzyme activity. These declines, however, were not statistically significant. Deletion of an additional 63 bp caused an increase of about 60% more β-galactosidase activity. To confirm that the RNA polymerase binding regions are located within the sequences spanning up to the consensus -35 sequences, 3' end deletion constructs lacking sequences up to the -35 region for genes 14 and 19 (65 and 57 bp, respectively) were prepared and assessed for β-galactosidase activity. These deletions led to the complete loss of β-galactosidase activity (Figure 6A–B lane 11 and 6C–D lane 6).
Figure 6.
Deletion analysis of promoter regions of genes 14 and 19. β-galactosidase activity of extracts prepared from E. coli cultures of bacteria transformed with various deletion constructs was determined. Panels A and C have cartoons depicting deletion constructs and their orientations for genes 14 and 19, respectively. (Solid black boxes represent lacZ gene, and right and left arrowhead lines show orientation of the promoter regions ligated in front of the lacZ coding sequence. Lengths of the promoter regions in base pairs are indicated on the left. Panels B and D contain the β-galactosidase activity analysis data. (β-galactosidase activity was expressed as percent activity relative to the activity observed for full length promoter segments.) Data are presented with SD values calculated from four independent experiments (P ≤ 0.001).
Location of -10 and -35 regions
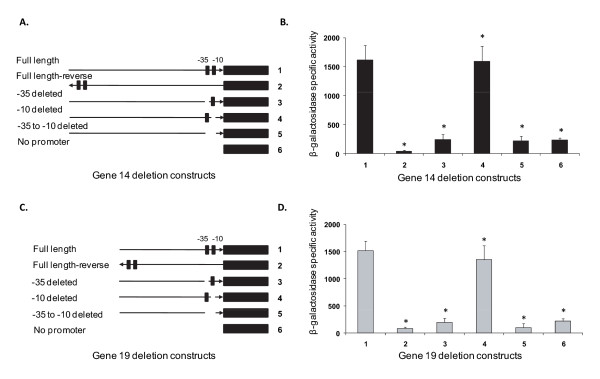
To determine whether the consensus -35 and -10 represented true RNA polymerase binding site regions, constructs lacking either the predicted -35 or -10 alone or the regions spanning from -35 to -10 were generated, and the effect of the loss of these sequences on promoter activity was evaluated by measuring β-galactosidase activity. Deletion of the predicted -35 regions alone or in combination with the -10 for both the genes resulted in decline of β-galactosidase activity to the background levels observed for negative controls. Deletion of the consensus -10 region alone for both the genes, however, resulted in no significant change to the promoter activity (Figure 7). The impact of the deletions of -35 and -10 are very similar for both genes' promoters.
Figure 7.
Deletion analysis spanning the -35 and -10 regions of genes 14 and 19. β-galactosidase activity of extracts prepared from E. coli cultures of bacteria transformed with -35 or -10 deletions or deletions spanning from -35 to 10 were determined. Panels A and C have cartoons depicting deletion constructs and their orientations for genes 14 and 19, respectively. Panels B and D contained the β-galactosidase activity analysis data. Data are presented with SD values calculated from four independent experiments (P ≤ 0.001).
Discussion
Differences in protein expression influenced by vertebrate and tick cell environment are now well documented for E. chaffeensis [18-20] and other tick-transmitted bacteria [12,13,15,16]. We recently reported novel data describing differences in immune response in the murine host against E. chaffeensis originating from tick cells compared with that observed for the bacteria originating from macrophages [9]. Importantly, the murine host takes longer to clear the pathogen originating from tick cells, and the delayed clearance has been associated with altered macrophage, B-cell and cytokine responses. These studies suggest that tick cell-specific altered pathogen protein expression offers a selective advantage to E. chaffeensis for its continued survival when it enters into a vertebrate host from the tick cell environment. To date, no studies have assessed the molecular mechanisms used by E. chaffeensis to achieve differential gene expression.
Primer extension analysis reported in this study confirmed our previous observations of Northern blot analysis that transcripts of p28-Omp genes 14 and 19 are differentially expressed and as monocistronic messages [19]. The primer extension analysis also aided in defining transcription start sites. Adenine, the base found at the transcription start site for genes 14 and 19 of E. chaffeensis, appears to be the most common base at which transcription is initiated from rickettsiales genes, including pathogens of the genera Rickettsia and Anaplasma [31-34]. Our previous studies and those of other investigators also support that genes 14 and 19 are transcriptionally active independent of E. chaffeensis originating from macrophages or tick cells [9,19,21,35-38]. In the current study, quantitative RT-PCR analysis confirmed the previous observations about the presence of messages for genes 14 and 19 in both host cell backgrounds. In addition, the analysis aided in mapping quantitative differences in transcription of differentially expressed genes. The quantitative RT-PCR analysis demonstrates that although genes 14 and 19 are transcriptionally active, levels of transcription are influenced in response to the macrophage and tick cell environments. Gene 19 is higher in its expression in macrophages, and the opposite is true for gene 14 expression.
Promoter regions of genes 14 and 19 differed considerably; the differences include variations in length of the upstream sequences, presence of several gene-specific direct repeats, palindrome sequences and presence of a G-rich region found in gene 19. Importance of palindrome and direct repeat sequences in regulating transcription is well established for many prokaryotes and for a rickettsial pathogen [34,39-42]. For example, the presence of a palindrome sequence in the citrate synthase gene of Rickettsia prowazekii with its possible role in transcriptional regulation is reported by Cai and Winkler [42]. Similarly, transcription factors such as zinc finger proteins that influence gene expression via interacting with G-rich sequences are established for both prokaryotes and eukaryotes [43-49]. The E. chaffeensis genome contains two homologs of zinc finger proteins (Genbank #s ABD44730 and ABD45416) [50]. It is of interest to investigate whether one or both of these putative E. chaffeensis zinc finger proteins act as transcription regulators for p28-Omp gene 19.
Mapping the functions of E. chaffeensis genes in vivo cannot be performed because genetic manipulation systems are yet to be established. To overcome this limitation, we assessed the utility of E. coli RNA polymerase as a surrogate to characterize E. chaffeensis gene promoters as reported for several C. trachomatis genes [23-30]. In vitro transcription assays performed with E. coli RNA polymerase identified the same transcription start sites for p28-Omp genes 14 and 19 as observed in E. chaffeensis. This observation validates the use of E. coli RNA polymerase.
Molecular characterization of promoter sequences located upstream to the transcription start sites of genes 14 and 19 is critical in determining how E. chaffeensis regulates gene expression. In E. coli, expression of reporter gene products, GFP and β-galactosidase, is evident when sequences upstream to the coding regions of p28-Omp genes 14 and 19 were placed in front of promoterless GFP or β-galactosidase genes, respectively. These data are also consistent with previous reports that the E. coli RNA polymerase can complement the functions of rickettsial RNA polymerases of the genera Anaplasma, Ehrlichia and Rickettsia [31,32,37], including recognizing the transcription start sites [32].
Sequential deletions in the gene 14 upstream sequences from the 5' end, whereby some of the direct repeats and palindrome sequences were deleted, resulted in variations in the promoter activity that fluctuated from complete or partial loss of activity compared with that observed for the full-length upstream sequence. Additional deletions caused the restoration of 100% activity, and subsequent additional deletions again led to a decline in promoter activity. Similarly, deletion analysis in the gene 19 promoter region caused loss or gain of promoter activities relative to the inclusion of full-length upstream sequence as a promoter. These data suggest that promoter regions of genes 14 and 19 contain sequence domains that influence binding affinity of RNA polymerase to the respective promoters. Altered promoter activities observed in deletion analysis experiments may have resulted from the deletions of upstream sequences involved in altering DNA topology and making RNA polymerase less or more accessible to its binding domains. Influence of 5' sequences altering the DNA topology for RNA polymerase binding has been well established for promoters of several bacterial organisms such as Bacillus subtilis, C. tracomatis, E. coli, and Klebsiella pneumoniae [23,51-56]. Previous reports also suggest that the inverted and direct repeats contribute to the DNA curvatures, thus affecting RNA polymerase binding to the -35 and -10 regions [23,39]. Although less likely, the presence of E. coli regulators that are homologues of E. chaffeensis may also bind and influence the promoter activity. For example, homologues of R. prowazekii repressors/enhancers in E. coli have been reported for the 16S rRNA gene [32]. Variations in the promoter activity of E. chaffeensis genes observed in E. coli for the deletion constructs may not represent what may occur in the pathogen. Defining the importance of the putative regulatory domains of p28-Omp genes identified in this study requires further analysis in E. chaffeensis or using E. chaffeensis RNA polymerase.
Deletion of the consensus -35 region alone or in combination with the -10 region, but not of the -10 region alone, reduced the promoter activity to background levels for both genes 14 and 19. These data suggest that, independent of the gene assessed, the -35 regions identified contribute to the RNA polymerase binding. It is unclear why deletions of the predicted -10 regions for both the genes had little effect in altering the promoter functions. Greater tolerance to the loss of the -10 regions compared to -35 regions is reported for other prokaryotes [26,57-59]. It is, however, possible that the -10 regions we predicted are not accurate and may be present at a different location. Alternatively, the -10 regions may be less important in E. chaffeensis. This hypothesis is too premature at this time; more detailed mapping of the -10 regions is needed.
In the absence of genetic manipulation methods, an in vitro transcription system can serve as a useful molecular tool for mapping the molecular basis for differences in E. chaffeensis gene expression. For example, extensive studies have already reported using in vitro transcription systems to map regulation of gene expression for another intra-phagosomal bacterium, C. trachomatis, for which genetic manipulation systems are yet to be established [28-30]. In the current study, we also presented the first evidence for a similar in vitro transcription protocol to drive expression from two E. chaffeensis promoter sequences. More detailed investigations may also be performed by using the in vitro transcription protocol with E. coli or E. chaffeensis RNA polymerase, similar to studies carried out for C. trachomatis and R. prowazekii [23-30,32].
Conclusion
In this study, we performed detailed RNA analysis to demonstrate that E. chaffeensis regulates transcription by sensing differences in host cell environments. Experimental evidence presented in this study also demonstrates that gene expression differences are achieved by altering changes in RNA polymerase activity influenced by the sequences located upstream to the transcription start sites. More detailed investigations are needed to map the mechanisms controlling gene expression in E. chaffeensis in different host cell environments.
Methods
In vitro cultivation of E. chaffeensis
E. chaffeensis Arkansas isolate was cultured in vitro in the canine macrophage cell line (DH82) and in the tick cell line (ISE6) as described previously [1,60].
Nucleic acids isolation
About 20 ml of 90–100% infected E. chaffeensis confluent monolayers of DH82 or ISE6 cell cultures recovered from a T-150 flask were used for isolation of total RNA. Total RNA was isolated with the Tri-reagent method by following the manufacturer's instructions (Sigma-Aldrich, St. Louis, MO). The RNA pellet recovered was resuspended in 100 μl of nuclease-free water containing 40 units of RNase inhibitor, RNasin, (Ambion Inc., Austin, TX) and stored at -70°C until use. Quality and concentration of RNA were assessed by spectrophotometry with an ND-1000 spectrophotometer (Nanodrop Technologies, Wilmington, DE) and by calculating the ratio between the optical densities at 260 nm and 280 nm.
Genomic DNA was isolated from 2 ml of 90–100% infected E. chaffeensis confluent monolayer by the sodium dodecyl sulfate (SDS), proteinase K, phenol, chloroform, isoamyl alcohol method [61]. Final purified DNA was resuspended in 100 μl of TE buffer (pH 8.0); concentration was assessed by spectrophotometry with an ND-1000 spectrophotometer and stored at -20°C. Quality of DNA and RNA samples was further confirmed by resolving about 1–5 μg each on a 0.9% agarose gel or 1.5% formaldehyde agarose gel, respectively [61].
Oligonucleotides
Oligonucleotides used for the experiments described in this study are custom synthesized from Integrated DNA Technologies (Coralville, IA) and are listed in Table 1.
Table 1.
List of oligonucleotides used for this study
| Primers | Sequence | Orientation | Amplicon size (bp) |
Annealing temperature (°C) |
|---|---|---|---|---|
| PROMOTER ANALYSIS | ||||
| Gene 14-upstream sequence primers | ||||
| For cloning into pPROBE-NT | ||||
| RRG 183* | 5' GACTCTAGAttgctcaacccataaaataatg | Forward | 596 | 50 |
| RRG 184 | 5' AGTGAGCTCtttataaaagataataaaaatttaag | Reverse | ||
| For cloning into pBlue-TOPO | ||||
| RRG 217 | 5' attgctcaaccataaaataatggga | Forward | 581 | 48 |
| RRG 218 | 5' gttaataaaccttttataaaag | Reverse | ||
| RRG 267 | 5' cagttaactttctgtaaacttc | Forward | 521 | 48 |
| RRG 218** | Reverse | |||
| RRG 268 | 5' atcataagtttacaataatgtc | Forward | 461 | 48 |
| RRG 218 | Reverse | |||
| RRG 269 | 5' cgttttctgctttattagaatg | Forward | 400 | 48 |
| RRG 218 | Reverse | |||
| RRG 270 | 5' gttccgtatttattaatatatg | Forward | 350 | 48 |
| RRG 218 | Reverse | |||
| RRG 271 | 5' catgtactgaatttgtgatttg | Forward | 286 | 48 |
| RRG 218 | Reverse | |||
| RRG 272 | 5' ggataagtactttagcaagtgg | Forward | 222 | 48 |
| RRG 218 | Reverse | |||
| RRG 273 | 5' taagtagtaaagttaactatag | Forward | 169 | 48 |
| RRG 218 | Reverse | |||
| RRG 274 | 5' acttttgttgtaaatttgaaag | Forward | 105 | 48 |
| RRG 218 | Reverse | |||
| RRG 217 | Forward | 516 | 50 | |
| IG14-35 del R | 5' (PO4)-gtctagaatataaaatttctttc | Reverse | ||
| IG14-10 del F | 5' (PO4)-taaatttttattatcttttataaaaggtttattaac | Forward | 8366 | 56 |
| IG14-10 del R | 5' (PO4)-atgaaagaaataaagaaaagcaagtctag | Reverse | ||
| IG14-35 del F | 5' (PO4)-ttctttatttctttcattattc | Forward | 8366 | 48 |
| IG14-35 del R | Reverse | |||
| IG14-10 del F | Forward | 8343 | 51 | |
| IG14-35 del R | Reverse | |||
| Gene 19-upstream sequence primers | ||||
| For cloning into pPROBE-NT | ||||
| RRG 185 | 5' GACTCTAGActtttaattttattattgccacatg | Forward | 334 | 61 |
| RRG 186 | 5' AGTGAGCTCaatagtgacaaataaattaacaatag | Reverse | ||
| For cloning into pBlue-TOPO | ||||
| RRG 185 | Forward | 308 | 60 | |
| RRG 445 | 5' atataacctaatagtgacaaataaattaac | Reverse | ||
| RRG 275 | 5' gtggcaaaagaatgtagcaataag | Forward | 239 | 50 |
| RRG 445 | Reverse | |||
| RRG 276 | 5' gtgctgtttttctcacctttacac | Forward | 188 | 63 |
| RRG 445 | Reverse | |||
| RRG 277 | 5' ctgacgtaatatattaaattttcc | Forward | 125 | 55 |
| RRG 445 | Reverse | |||
| RRG 185 | Forward | 267 | 50 | |
| IG19-35 del R | 5' (PO4)-gtcagaatataaatttttgtataaaatatcg | Reverse | ||
| IG19-10 del F | 5' (PO4)-taatttatttgtcactattaggttat | Forward | 8112 | 56 |
| IG19-10 del R | 5' (PO4)-gtagaagtgtcatataaaagcaag | Reverse | ||
| IG19-35 del F | 5' (PO4)-ttatatgacacttctactattgttaatttatttg | Forward | 8112 | 61.5 |
| IG19-35 del R | Reverse | |||
| IG19-10 del F | Forward | 8088 | 58 | |
| IG19-35 del R | Reverse | |||
| PRIMER EXTENSION ANALYSIS | ||||
| Gene 14 | ||||
| RRG 14-5'rev | 5' gccttctctgctgtcgttgattcc | NA | 52 | |
| Gene 19 | ||||
| RRG 20-PEXT | 5' cgttaataccactacctgctgggtcg | NA | 58 | |
| RRG 44 | 5' cgcttccgtcccaattttgcttc | NA | 58 | |
| IN VITRO TRANSCRIPTION ASSAY | ||||
| Gene 14 upstream full-length+lac Z segment | ||||
| RRG 217 | 5' attgctcaaccataaaataatggga | Forward | 882 | 50 |
| RRG 226 | 5' cgccattcgccattag | Reverse | ||
| RRG 218 | 5' gttaataaaccttttataaaag | Forward | 882 | 50 |
| RRG 226 | Reverse | |||
| Gene 19 upstream full-length+lac Z segment | ||||
| RRG 217 | 5' attgctcaaccataaaataatggga | Forward | 601 | 50 |
| RRG 226 | Reverse | |||
| RRG 445 | 5' atataacctaatagtgacaaataaattaac | Forward | 601 | 50 |
| RRG 226 | Reverse | |||
| IN VITRO TRANSCRIPTION COUPLED TRANSLATION ASSAY | ||||
| RRG 185 | 5' gactctagacttttaattttattattgccacatg | Forward | 848 | 58 |
| RRG 247 | 5' tccggctcgtatgttgtgtg | Reverse | ||
* Text in capital letters refers to sequences inserted for creating restriction enzyme sites. ** Primer sequences were presented only once when a primer was described for the first time.
Primer extension analysis
Primer extension analysis was performed by using a Primer Extension System AMV Reverse Transcriptase kit (Promega, Madison, WI). Briefly, oligonucleotides complementary to the transcripts of p28-Omp genes 14 and 19 were end labeled with [γ-32p] ATP using T4 polynucleotide kinase (Promega, Madison, WI) at 37°C for 10 min. The kinase reaction was stopped by heat inactivation at 90°C for 2 min. The end labeled primers (one ρ mole each) were annealed to E. chaffeensis RNA (~10 μg) by incubating at 58°C for 20 min in 11 μl reactions containing AMV primer extension buffer. E. chaffeensis RNA used for this experiment was isolated from cultures when the infection reached to 80–100%. One micro liter of AMV reverse transcriptase (1 unit) was added, and the reaction was incubated at 42°C for 30 min. The reaction products were concentrated by ethanol precipitation and electrophorosed on a 6% polyacrylamide gel containing 7 M urea, and the gel was transferred to a Whatman paper, dried and exposed to an X-ray film. The primer extended products were detected after developing the film with a Konica film processor (Wayne, NJ).
Quantitative RT-PCR analysis
Quantitative differences in transcripts for p28-Omp genes 14 and 19 were assessed with a TaqMan-based diplex RT-PCR assay using gene-specific primers and probes as we reported earlier [19]. The analysis was performed on total RNA isolated for E. chaffeensis infected DH82 and ISE6 cells at 12, 24, 48, 72, 96 and 120 h post infection. Quantitative data relative to the number of Ehrlichia organisms were calculated [9,19].
Bioinformatics analysis
Sequences upstream from the protein coding regions of E. chaffeensis p28-Omp 14 and 19 were obtained from the GenBank data base and aligned by using the genetic computer group (GCG) programs PileUp and Pretty [62] to search for sequence homologies. Direct repeats and palindrome sequences in the upstream sequences were identified with the GCG programs Repeat and StemLoop, respectively. E. coli σ70 promoter consensus sequences (-10 and -35) [63] were used to locate similar elements manually in p28-Omp genes 14 and 19 sequences upstream to the transcription start sites.
Promoter constructs
Promoter constructs for p28-Omp genes 14 and 19 were made with two independent promoterless reporter genes containing plasmid vectors pPROBE-NT [64] and pBlue-TOPO (Invitrogen Technologies, Carlsbad, CA). The pPROBE-NT vector contains a GFP gene as the reporter gene, whereas a lacZ gene is the reporter gene in the pBlue-TOPO vector. To generate a p28-Omp gene14 promoter region construct, the entire non-coding sequences located between coding sequences of p28-Omp genes 13 and 14 were amplified by using E. chaffeensis genomic DNA as a template and the sequence-specific oligonucleotides (Table 1). A similar strategy was used to prepare the gene 19 promoter constructs by amplifying the DNA segment located between the coding regions of p28-Omp genes 18 and 19. The PCR products were ligated into the promoterless pBlue-TOPO and pPROBE-NT vectors and transformed into E. coli strain, Top10 (Invitrogen Technologies, Carlsbad, CA) and DH5α strain, respectively [61]. One clone each in forward and reverse orientations was selected for the genes 14 and 19 in the pBlue-TOPO plasmid. For the pPROBE-NT constructs, only forward orientation inserts containing plasmids were selected. In addition, nonrecombinant plasmids transformed in E. coli were selected to serve as negative controls.
Promoter deletion constructs
Various deletion fragments of the promoter regions lacking parts of the 5' or 3' end segments of genes 14 and 19 were also generated by PCR and cloning strategy in the pBlue-TOPO plasmid. Deletion constructs of gene 14 and 19 promoters that are lacking the predicted -35 or -10 alone or the regions spanning from -35 to -10 were also generated by PCR cloning strategy but by using a Phusion site-directed mutagenesis kit as per the manufacturer's recommendations (New England Biolabs, MA). Primers used for the deletion analysis experiments are included in Table 1. Presence of correct inserts for the clones was always verified by restriction enzyme and sequence analysis.
Assessment of promoter activity in vitro
Promoter region and reporter gene segments were amplified by PCR using pBlue-TOPO promoter constructs as the templates. Amplicons were then used for in vitro transcription reactions. The entire gene 14 upstream, 5' end non-coding region in forward or reverse orientations along with a 301 bp lacZ gene fragment were amplified from the constructs in pBlue-TOPO (described previously). A similar strategy was followed to generate gene 19 promoter region templates for use in the in vitro transcription analysis. PCR products were purified with the QIAquick PCR Purification Kit (Quiagen, Valencia, CA).
In vitro transcription analysis was performed by following protocol described previously [65] with minor modifications. Briefly, assays were performed in a 10 μl reaction containing 50 mM Tris-acetate (pH 8.0), 50 mM potassium acetate, 8.1 mM magnesium acetate, 27 mM ammonium acetate, 2 mM dithiothreitol, 400 μM ATP, 400 μM GTP, 400 μM UTP, 1.2 μM CTP, 0.21 μM [α-32P] CTP, 18 U of RNasin, 5% glycerol, 100 ng of purified PCR templates and 0.03 U of E. coli RNA polymerase holoenzyme (Epicentre, Madison, WI). The reaction was incubated at 37°C for 15 min and then terminated by adding 4 μl of stop solution (95% formamide, 20 mM EDTA, 0.05% bromophenol blue, 0.05% xylene cyanol). Four micro liters of reaction contents each were resolved in a 6% polyacrylamide gel containing 7 M urea [66]. The gel was transferred to a Whatman paper, dried and exposed to an X-ray film; the in vitro transcripts were detected after developing the film with a Konica film processor (Wayne, NJ).
Assessment of promoter activity in E. coli
The pPROBE-NT constructs containing promoter regions of genes 14 and 19 were assessed for promoter activities by observing green florescence emitted from colonies on agar plates. The promoter activity was further confirmed by performing Western blot analysis using a GFP polyclonal antibody (Rockland Immunochemicals, Inc., Gilbertsville, PA) on protein extracts made from E. coli containing the recombinant plasmids. The pBlue-TOPO promoter constructs were also evaluated for promoter activity by measuring β-galactosidase activity. To accomplish this, E. coli colonies containing the recombinant plasmids were grown to an optical density of 0.4 (at 600 nm); soluble protein preparations from the cell lysates were prepared and assessed for the lacZ expression by using a β-gal assay kit as per the manufacturer's instructions (Invitrogen Technologies, Carlsbad, CA,). About 2.5 or 5 μg of protein preparations were assessed for the β-galactosidase activity using Ortho-Nitrophenyl-β-D-Galactopyranoside (ONPG) as the substrate. The analysis included protein preparations made from no-insert controls as well as E. coli cultures containing constructs with promoter segments in the reverse orientation. The experiments were repeated four independent times with independently isolated protein preparations; samples were also assayed in triplicate each time. Specific activity of β-galactosidase was calculated using the formula outlined in the β-gal assay kit protocol.
Statistical Analysis
Statistical analysis of RT-PCR experiments for measuring the quantitative differences in the gene expression of p28-Omp genes14 and 19 was performed by using the unpaired Student t-test. For promoter deletion analysis experiments, statistical analysis was performed by using repeated measures of ANOVA, and the Bonferroni method was used to adjust for multiple comparisons. GraphPad InStat Software (La Jolla, CA) was used to perform these analyses. A P value of less than 0.05 was considered significant.
Authors' contributions
LP carried out the RNA mapping studies, promoter deletion analysis, in vitro transcription experiments, statistical analysis, and also drafted the manuscript. CC carried out the cell culture experiments, participated in in vitro transcription experiments and compiling references and manuscript editing. RRG conceived of the study and participated in its design and coordination, was instrumental in obtaining financial support, and helped in data analysis and drafting the manuscript to its final form. All authors read and approved the final manuscript.
Contributor Information
Lalitha Peddireddi, Email: lpeddire@vet.k-state.edu.
Chuanmin Cheng, Email: cmcheng@vet.k-state.edu.
Roman R Ganta, Email: rganta@vet.k-state.edu.
Acknowledgements
This study was supported by the Public Health Service grants AI070908 and AI055052 from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD. We thank Dr. Steven Lindow, Department of Plant and Microbiology, University of California, Berkeley, CA, for the kind gift of the pPROBE-NT plasmid. This manuscript is a contribution from the Kansas Agricultural Experiment Station, no. 08-364-J.
References
- Dawson JE, Anderson BE, Fishbein DB, Sanchez CY, Goldsmith CY, Wilson KH. Isolation and characterization of an Ehrlichia sp. from a patient diagnosed with human ehrlichiosis. J Clin Microbiol. 1991;29:2741–2745. doi: 10.1128/jcm.29.12.2741-2745.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paddock CD, Childs JE. Ehrlichia chaffeensis: a prototypical emerging pathogen. Clin Microbiol Rev. 2003;16:37–64. doi: 10.1128/CMR.16.1.37-64.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrew HR, Norval RA. The carrier status of sheep, cattle and African buffalo recovered from heartwater. Vet Parasitol. 1989;34:261–266. doi: 10.1016/0304-4017(89)90056-3. [DOI] [PubMed] [Google Scholar]
- Dumler JS, Sutker WL, Walker DH. Persistent Infection with Ehrlichia chaffeensis. Clin Infect Dis. 1993;17:903–905. doi: 10.1093/clinids/17.5.903. [DOI] [PubMed] [Google Scholar]
- Davidson WR, Lockhart JM, Stallknecht DE, Howerth EW, Dawson JE, Rechav Y. Persistent Ehrlichia chaffeensis infection in white-tailed deer. J Wildl Dis. 2001;37:538–546. doi: 10.7589/0090-3558-37.3.538. [DOI] [PubMed] [Google Scholar]
- French DM, Brown WC, Palmer GH. Emergence of Anaplasma marginale antigenic variants during persistent rickettsemia. Infect Immun. 1999;67:5834–5840. doi: 10.1128/iai.67.11.5834-5840.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuen S, Engvall EO, Artursson K. Persistence of Ehrlichia phagocytophila infection in lambs in relation to clinical parameters and antibody responses. Vet Rec. 1998;143:553–555. doi: 10.1136/vr.143.20.553. [DOI] [PubMed] [Google Scholar]
- Zeidner NS, Dolan MC, Massung R, Piesman J, Fish D. Coinfection with Borrelia burgdorferi and the agent of human granulocytic ehrlichiosis suppresses IL-2 and IFN gamma production and promotes an IL-4 response in C3H/HeJ mice. Parasite Immunol. 2000;22:581–588. doi: 10.1046/j.1365-3024.2000.00339.x. [DOI] [PubMed] [Google Scholar]
- Ganta RR, Cheng C, Miller EC, McGuire BL, Peddireddi L, Sirigireddy KR. Differential clearance and immune responses to tick cell-derived versus macrophage culture-derived Ehrlichia chaffeensis in mice. Infect Immun. 2007;75:135–145. doi: 10.1128/IAI.01127-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbet AF, Lundgren A, Yi J, Rurangirwa FR, Palmer GH. Antigenic variation of Anaplasma marginale by expression of MSP2 mosaics. Infect Immun. 2000;68:6133–6138. doi: 10.1128/IAI.68.11.6133-6138.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brayton KA, Meeus PF, Barbet AF, Palmer GH. Simultaneous variation of the immunodominant outer membrane proteins, MSP2 and MSP3, during anaplasma marginale persistence in vivo. Infect Immun. 2003;71:6627–6632. doi: 10.1128/IAI.71.11.6627-6632.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia-Garcia JC, de la FJ, Blouin EF, Johnson TJ, Halbur T, Onet VC. Differential expression of the msp1alpha gene of Anaplasma marginale occurs in bovine erythrocytes and tick cells. Vet Microbiol. 2004;98:261–272. doi: 10.1016/j.vetmic.2003.10.021. [DOI] [PubMed] [Google Scholar]
- De SA, Telford SR, Brunet LR, Barthold SW, Fikrig E. Borrelia burgdorferi OspA is an arthropod-specific transmission-blocking Lyme disease vaccine. J Exp Med. 1996;183:271–275. doi: 10.1084/jem.183.1.271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwan TG, Piesman J, Golde WT, Dolan MC, Rosa PA. Induction of an outer surface protein on Borrelia burgdorferi during tick feeding. Proc Natl Acad Sci USA. 1995;92:2909–2913. doi: 10.1073/pnas.92.7.2909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jauron SD, Nelson CM, Fingerle V, Ravyn MD, Goodman JL, Johnson RC. Host cell-specific expression of a p44 epitope by the human granulocytic ehrlichiosis agent. J Infect Dis. 2001;184:1445–1450. doi: 10.1086/324428. [DOI] [PubMed] [Google Scholar]
- Lohr CV, Brayton KA, Shkap V, Molad T, Barbet AF, Brown WC. Expression of Anaplasma marginale major surface protein 2 operon-associated proteins during mammalian and arthropod infection. Infect Immun. 2002;70:6005–6012. doi: 10.1128/IAI.70.11.6005-6012.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rurangirwa FR, Stiller D, French DM, Palmer GH. Restriction of major surface protein 2 (MSP2) variants during tick transmission of the ehrlichia Anaplasma marginale. Proc Natl Acad Sci USA. 1999;96:3171–3176. doi: 10.1073/pnas.96.6.3171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singu V, Liu H, Cheng C, Ganta RR. Ehrlichia chaffeensis expresses macrophage- and tick cell-specific 28-kilodalton outer membrane proteins. Infect Immun. 2005;73:79–87. doi: 10.1128/IAI.73.1.79-87.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singu V, Peddireddi L, Sirigireddy KR, Cheng C, Munderloh UG, Ganta RR. Unique Macrophage and Tick Cell-specific Protein Expression from the p28/p30 Omp Multigene Locus in Ehrlichia Species. Cell Microbiol. 2006;8:1475–1487. doi: 10.1111/j.1462-5822.2006.00727.x. [DOI] [PubMed] [Google Scholar]
- Seo GM, Cheng C, Tomich J, Ganta RR. Total, membrane, and immunogenic proteomes of macrophage- and tick cell-derived Ehrlichia chaffeensis evaluated by LC-MS/MS and MALDI-TOF methods. Infect Immun. 2008;76:4823–32. doi: 10.1128/IAI.00484-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ganta RR, Peddireddi L, Seo GM, Dedonder SE, Cheng C, Chapes SK. Molecular characterization of Ehrlichia interactions with tick cells and macrophages. Front Biosci. 2009;14:3259–73. doi: 10.2741/3449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steitz JA, Jakes K. How ribosomes select initiator regions in mRNA: base pair formation between the 3' terminus of 16S rRNA and the mRNA during initiation of protein synthesis in Escherichia coli. Proc Natl Acad Sci USA. 1975;72:4734–4738. doi: 10.1073/pnas.72.12.4734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mathews SA, Stephens RS. DNA structure and novel amino and carboxyl termini of the Chlamydia sigma 70 analogue modulate promoter recognition. Microbiology. 1999;145:1671–1681. doi: 10.1099/13500872-145-7-1671. [DOI] [PubMed] [Google Scholar]
- Koo IC, Walthers D, Hefty PS, Kenney LJ, Stephens RS. ChxR is a transcriptional activator in Chlamydia. Proc Natl Acad Sci USA. 2006;103:750–755. doi: 10.1073/pnas.0509690103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson AC, Tan M. Stress response gene regulation in Chlamydia is dependent on HrcA-CIRCE interactions. J Bacteriol. 2004;186:3384–3391. doi: 10.1128/JB.186.11.3384-3391.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thouvenot B, Charpentier B, Branlant C. The strong efficiency of the Escherichia coli gapA P1 promoter depends on a complex combination of functional determinants. Biochem J. 2004;383:371–382. doi: 10.1042/BJ20040792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu HH, Tan M. Sigma28 RNA polymerase regulates hctB, a late developmental gene in Chlamydia. Mol Microbiol. 2003;50:577–584. doi: 10.1046/j.1365-2958.2003.03708.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akers JC, Tan M. Molecular mechanism of tryptophan-dependent transcriptional regulation in Chlamydia trachomatis. J Bacteriol. 2006;188:4236–4243. doi: 10.1128/JB.01660-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu HH, Di Russo EG, Rounds MA, Tan M. Mutational analysis of the promoter recognized by Chlamydia and Escherichia coli sigma(28) RNA polymerase. J Bacteriol. 2006;188:5524–5531. doi: 10.1128/JB.00480-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaumburg CS, Tan M. Mutational analysis of the Chlamydia trachomatis dnaK promoter defines the optimal -35 promoter element. Nucleic Acids Res. 2003;31:551–555. doi: 10.1093/nar/gkg150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbet AF, Agnes JT, Moreland AL, Lundgren AM, Alleman AR, Noh SM. Identification of functional promoters in the msp2 expression loci of Anaplasma marginale and Anaplasma phagocytophilum. Gene. 2005;353:89–97. doi: 10.1016/j.gene.2005.03.036. [DOI] [PubMed] [Google Scholar]
- Pang H, Winkler HH. Transcriptional analysis of the 16s rRNA gene in Rickettsia prowazekii. J Bacteriol. 1996;178:1750–1755. doi: 10.1128/jb.178.6.1750-1755.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw EI, Marks GL, Winkler HH, Wood DO. Transcriptional characterization of the Rickettsia prowazekii major macromolecular synthesis operon. J Bacteriol. 1997;179:6448–6452. doi: 10.1128/jb.179.20.6448-6452.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radulovic S, Rahman MS, Beier MS, Azad AF. Molecular and functional analysis of the Rickettsia typhi groESL operon. Gene. 2002;298:41–48. doi: 10.1016/S0378-1119(02)00922-8. [DOI] [PubMed] [Google Scholar]
- Cheng C, Paddock CD, Reddy GR. Molecular Heterogeneity of Ehrlichia chaffeensis Isolates Determined by Sequence Analysis of the 28-Kilodalton Outer Membrane Protein Genes and Other Regions of the Genome. Infect Immun. 2003;71:187–195. doi: 10.1128/IAI.71.1.187-195.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohashi N, Rikihisa Y, Unver A. Analysis of Transcriptionally Active Gene Clusters of Major Outer Membrane Protein Multigene Family in Ehrlichia canis and E. chaffeensis. Infect Immun. 2001;69:2083–2091. doi: 10.1128/IAI.69.4.2083-2091.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long SW, Zhang XF, Qi H, Standaert S, Walker DH, Yu XJ. Antigenic variation of Ehrlichia chaffeensis resulting from differential expression of the 28-kilodalton protein gene family. Infect Immun. 2002;70:1824–1831. doi: 10.1128/IAI.70.4.1824-1831.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu X, McBride JW, Zhang X, Walker DH. Characterization of the complete transcriptionally active Ehrlichia chaffeensis 28 kDa outer membrane protein multigene family. Gene. 2000;248:59–68. doi: 10.1016/S0378-1119(00)00147-5. [DOI] [PubMed] [Google Scholar]
- Ni X, Westpheling J. Direct repeat sequences in the Streptomyces chitinase-63 promoter direct both glucose repression and chitin induction. Proc Natl Acad Sci USA. 1997;94:13116–13121. doi: 10.1073/pnas.94.24.13116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen-Kupiec R, Blank C, Leigh JA. Transcriptional regulation in Archaea: in vivo demonstration of a repressor binding site in a methanogen. Proc Natl Acad Sci. 1997;94:1316–1320. doi: 10.1073/pnas.94.4.1316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma D, Cook DN, O'Brien DA, Hearst JE. Analysis of the promoter and regulatory sequences of an oxygen-regulated bch operon in Rhodobacter capsulatus by site-directed mutagenesis. J Bacteriol. 1993;175:2037–2045. doi: 10.1128/jb.175.7.2037-2045.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai J, Winkler HH. Transcriptional regulation in the obligate intracytoplasmic bacterium Rickettsia prowazekii. J Bacteriol. 1996;178:5543–5545. doi: 10.1128/jb.178.18.5543-5545.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamper JT, Kamper U, Rogers LM, Kolattukudy PE. Identification of regulatory elements in the cutinase promoter from Fusarium solani f. sp. pisi (Nectria haematococca) J Biol Chem. 1994;269:9195–9204. [PubMed] [Google Scholar]
- Passantino R, Antona V, Barbieri G, Rubino P, Melchionna R, Cossu G. Negative regulation of beta enolase gene transcription in embryonic muscle is dependent upon a zinc finger factor that binds to the G-rich box within the muscle-specific enhancer. J Biol Chem. 1998;273:484–494. doi: 10.1074/jbc.273.1.484. [DOI] [PubMed] [Google Scholar]
- Lin CJ, Tam RC. Transcriptional regulation of CD28 expression by CD28GR, a novel promoter element located in exon 1 of the CD28 gene. J Immunol. 2001;166:6134–6143. doi: 10.4049/jimmunol.166.10.6134. [DOI] [PubMed] [Google Scholar]
- Detillieux KA, Meyers AF, Meij JT, Cattini PA. An A/G-rich motif in the rat fibroblast growth factor-2 gene confers enhancer activity on a heterologous promoter in neonatal rat cardiac myocytes. Mol Cell Biochem. 1998;188:169–176. doi: 10.1023/A:1006886307083. [DOI] [PubMed] [Google Scholar]
- Stolt P, Stoker NG. Mutational analysis of the regulatory region of the Mycobacterium plasmid pAL5000. Nucleic Acids Res. 1997;25:3840–3846. doi: 10.1093/nar/25.19.3840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van BA, Scherer S, van AL, Verbrugh H. Short-sequence DNA repeats in prokaryotic genomes. Microbiol Mol Biol Rev. 1998;62:275–293. doi: 10.1128/mmbr.62.2.275-293.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou AY, Archdeacon J, Kado CI. Agrobacterium transcriptional regulator Ros is a prokaryotic zinc finger protein that regulates the plant oncogene ipt. Proc Natl Acad Sci USA. 1998;95:5293–5298. doi: 10.1073/pnas.95.9.5293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hotopp JC, Lin M, Madupu R, Crabtree J, Angiuoli SV, Eisen J. Comparative genomics of emerging human ehrlichiosis agents. PLoS Genet. 2006;2:e21. doi: 10.1371/journal.pgen.0020021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nickerson CA, Achberger EC. Role of curved DNA in binding of Escherichia coli RNA polymerase to promoters. J Bacteriol. 1995;177:5756–5761. doi: 10.1128/jb.177.20.5756-5761.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espinosa-Urgel M, Tormo A. Sigma s-dependent promoters in Escherichia coli are located in DNA regions with intrinsic curvature. Nucleic Acids Res. 1993;21:3667–3670. doi: 10.1093/nar/21.16.3667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McAllister CF, Achberger EC. Effect of polyadenine-containing curved DNA on promoter utilization in Bacillus subtilis. J Biol Chem. 1988;263:11743–11749. [PubMed] [Google Scholar]
- Plaskon RR, Wartell RM. Sequence distributions associated with DNA curvature are found upstream of strong E. coli promoters. Nucleic Acids Res. 1987;15:785–796. doi: 10.1093/nar/15.2.785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molina-Lopez JA, Govantes F, Santero E. Geometry of the process of transcription activation at the sigma 54-dependent nifH promoter of Klebsiella pneumoniae. J Biol Chem. 1994;269:25419–25425. [PubMed] [Google Scholar]
- Perez-Martin J, Timmis KN, de LV. Co-regulation by bent DNA. Functional substitutions of the integration host factor site at sigma 54-dependent promoter Pu of the upper-TOL operon by intrinsically curved sequences. J Biol Chem. 1994;269:22657–22662. [PubMed] [Google Scholar]
- Hook-Barnard I, Johnson XB, Hinton DM. Escherichia coli RNA polymerase recognition of a sigma70-dependent promoter requiring a -35 DNA element and an extended -10 TGn motif. J Bacteriol. 2006;188:8352–8359. doi: 10.1128/JB.00853-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sohaskey CD, Zuckert WR, Barbour AG. The extended promoters for two outer membrane lipoprotein genes of Borrelia spp. uniquely include a T-rich region. Mol Microbiol. 1999;33:41–51. doi: 10.1046/j.1365-2958.1999.01443.x. [DOI] [PubMed] [Google Scholar]
- Hayashi K, Shiina T, Ishii N, Iwai K, Ishizaki Y, Morikawa K. A role of the -35 element in the initiation of transcription at psbA promoter in tobacco plastids. Plant Cell Physiol. 2003;44:334–341. doi: 10.1093/pcp/pcg041. [DOI] [PubMed] [Google Scholar]
- Munderloh UG, Liu Y, Wang M, Chen C, Kurtti TJ. Establishment, maintenance and description of cell lines from the tick Ixodes scapularis. J Parasitol. 1994;80:533–543. doi: 10.2307/3283188. [DOI] [PubMed] [Google Scholar]
- Sambrook J, Russell DW. Molecular Cloning: A Laboratory Manual. 2. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press; 2000. [Google Scholar]
- Devereux J, Haeberli P, Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984;12:387–395. doi: 10.1093/nar/12.1Part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaumburg CS, Tan M. A positive cis-acting DNA element is required for high-level transcription in Chlamydia. J Bacteriol. 2000;182:5167–5171. doi: 10.1128/JB.182.18.5167-5171.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller WG, Leveau JH, Lindow SE. Improved gfp and inaZ broad-host-range promoter-probe vectors. Mol Plant Microbe Interact. 2000;13:1243–1250. doi: 10.1094/MPMI.2000.13.11.1243. [DOI] [PubMed] [Google Scholar]
- Wilson AC, Tan M. Stress response gene regulation in Chlamydia is dependent on HrcA-CIRCE interactions. J Bacteriol. 2004;186:3384–3391. doi: 10.1128/JB.186.11.3384-3391.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carle GF, Olson MV. Separation of chromosomal DNA molecules form yeast by orthogonal-field-alteration gel electrophoresis. Nucleic Acids Res. 1984;12:5647–5664. doi: 10.1093/nar/12.14.5647. [DOI] [PMC free article] [PubMed] [Google Scholar]