Abstract
Identification of genes expressed in response to prion infection may elucidate biomarkers for disease, identify factors involved in agent replication, mechanisms of neuropathology and therapeutic targets. Although several groups have sought to identify gene expression changes specific to prion disease, expression profiles rife with cell population changes have consistently been identified. Cuprizone, a neurotoxicant, qualitatively mimics the cell population changes observed in prion disease, resulting in both spongiform change and astrocytosis. The use of cuprizone-treated animals as an experimental control during comparative expression profiling allows for the identification of transcripts whose expression increases during prion disease and remains unchanged during cuprizone-triggered neuropathology. In this study, expression profiles from the brains of mice preclinically and clinically infected with Rocky Mountain Laboratory (RML) mouse-adapted scrapie agent and age-matched controls were profiled using Affymetrix gene arrays. In total, 164 genes were differentially regulated during prion infection. Eighty-three of these transcripts have been previously undescribed as differentially regulated during prion disease. A 0.4% cuprizone diet was utilized as a control for comparative expression profiling. Cuprizone treatment induced spongiosis and astrocyte proliferation as indicated by glial fibrillary acidic protein (Gfap) transcriptional activation and immunohistochemistry. Gene expression profiles from brain tissue obtained from cuprizone-treated mice identified 307 differentially regulated transcript changes. After comparative analysis, 17 transcripts unaffected by cuprizone treatment but increasing in expression from preclinical to clinical prion infection were identified. Here we describe the novel use of the prion disease mimetic, cuprizone, to control for cell population changes in the brain during prion infection.
Key words: RML infection, cuprizone, microarray, gene expression, comparative profiling
Introduction
Transmissible spongiform encephalopathies (TSEs) or prion diseases are infectious, neurodegenerative conditions that are inevitably fatal. Human TSEs include Creutzfeldt-Jakob disease, Gerstmann-Sträussler-Scheinker syndrome, fatal familial insomnia and kuru. Animal TSEs include scrapie in sheep, bovine spongiform encephalopathy (“mad cow disease”) in cattle, chronic wasting disease in cervids, and transmissible mink encephalopathy. Common pathological features shared by TSEs include spongiform vacuolation, gliosis and neuronal loss.1 These characteristic changes may also be accompanied by the accumulation of amyloid plaques in the neuropil of infected brains.
Several groups have identified individual genes that are highly expressed during prion disease, typically using subtractive hybridization of cDNA libraries or microarray analysis. RNA expression profiles have been developed for many different TSE strains at different stages during infection.2–17 Many of these RNA expression profiles were generated from whole brain homogenates from clinically affected animals. Early in the disease process, low-grade microglia and astrocytes are activated. As a result, these cells proliferate, progressively amplifying during infection,18,19 changing the cell population of the brain. This neuroinflammation is a generalized reaction that occurs in many brain diseases including Alzheimer disease,20 Parkinson disease21 and multiple sclerosis,22 among others. Not surprisingly, most of the RNAs differentially regulated during TSE infection are related to activated astrocytes and microglia. RNA changes resulting from inflammation mask transcript changes associated with prion disease-specific neuropathology. Greenwood et al. (2005) addressed this problem by developing RNA expression profiles from neuronal cell lines that are persistently infected with TSE agent,23 identifying a number of neuronal transcripts associated with prion disease (clusterin—associated with the clearing of cellular debris and apoptosis; metallothionein 1—inhibits oxidative damage and apoptosis) and several transcripts not previously identified in other studies. The ability to focus on neuronal changes in a brain infection would be an important progression of this line of research.
We hypothesized that gene profiling of animals treated with the prion disease mimetic, cuprizone, would allow us to identify transcripts associated with cell population changes, providing greater focus to those transcripts related to prion pathogenesis. Cuprizone (bis-cyclohexanone-oxaldihydraxone) is a copper chelator which, when fed to mice, produces demyelination in the central nervous system.24 Demyelination occurs soon after cuprizone dosing, exclusively affecting oligodendrocytes.25 Pattison and Jebbett observed that mice, fed a diet supplemented with 0.4% cuprizone, developed histopathological lesions in the brain very similar to those induced by murine TSE agent.26 Both cuprizone treatment and TSEs exhibit extracellular vacuolation, astrocytosis and astrocyte hypertrophy in the white matter of the cerebellum, midbrain and cerebellar peduncles. Additionally, cuprizone treatment, like prion disease, induces microglial activation and recruitment27 without loss of the blood-brain barrier.28 As cuprizone treatment causes neuroinflammation similar to prion disease, it provides a useful model system for identifying transcripts associated with broad neuroinflammation in the absence of the prion-specific PrPTSE. Here we report the use of cuprizone as a control for transcriptional events associated with neuroinflammation, facilitating the identification of several novel transcripts upregulated during a prion infection.
Results
Identification of previously undescribed transcripts differentially regulated during prion disease.
Microarray analysis was performed to detect differences in gene expression in the mouse brain at three time points during prion infection. C57BL/6 mice were intraperitoneally infected with RML-infected mouse brain homogenate and sacrificed at 108 dpi, 158 dpi and at the presentation of clinical disease (198 dpi), with age-matched, mock-inoculated controls at each time point. At 108 and 158 dpi, no clinical symptoms were present, however, protease-resistant PrP was detected at both 158 and 198 dpi (Fig. 2I, L and O).
Figure 2.
Spongiosis and astrocytosis in mice treated with cuprizone, infected with RML or controls. (A, D, G, J and M) Hematoxylin and eosin staining shows spongiosis in the white matter of the cerebellum after eight weeks of cuprizone treatment and after 158 and 198 days of RML infection. (B, E, H, K and N) Anti-GFAP immunohistochemistry shows star-shaped astrocytosis in the frontal cortex after cuprizone treatment and after 158 and 198 days of RML infection. (C, F, I, L and O) PrPTSE staining in the medulla. Red, positive staining can be seen at 158 and 198 dpi and is not observed after cuprizone treatment. Scale bars: (A, D, G, J, M, F, L and O) = 50 µm; (B, E, H, K, N, C and I) = 100 µm.
RNA samples were obtained from brain samples at all three time points and differentially regulated genes were identified using microarray analysis. The RMA method was used to background correct, normalize and summarize gene expression probe set data (representing individual genes) to identify differentially-expressed genes at each time point during prion disease. Differentially regulated genes were filtered on expression (10.0–99.0th expression percentile) to control for extreme or minimal signal intensity. Following statistical analysis (described in Materials and Methods), 164 genes were identified as being differentially regulated during prion disease. Many of these changes have been identified previously2–17 and include several transcripts known to be associated with proliferating astrocytes and microglia, such as cystatin F (leukocystatin), lymphocyte antigen 86, cathepsin S, CD53 antigen, CD68 antigen and glial fibrillary acidic protein. Supplemental Table 2 includes all 164 differentially regulated transcripts identified during our prion time course analysis. Our analysis, however, detected 83 transcripts that have not previously been identified as differentially expressed during prion disease, likely due to our lowering of the threshold of detection from the usual ≥2-fold to ≥1.5-fold. It should be noted that all transcripts identified as differentially expressed were deemed statistically significant changes. These transcripts were analyzed using the Gene Ontology (GO) classification system. The majority of transcripts identified have not yet been classified, however, 27 were found to be categorized in a variety of biological processes (Table 1). Genes related to the immune response and response to stimulus GO-terms were strongly represented, including genes encoding for guanylate binding proteins (Gbp3, Gbp6) and interferon-induced proteins (Ifi35, Ifitm1, Rsad2). Genes involved in protein modification and intracellular cellular signaling were also upregulated. Several genes downregulated during prion disease belong to the ubiquitin cycle and cellular developmental processes.
Table 1.
Previously unidentified, differentially regulated gene expression changes in RML-infected brain tissue
| Probe set ID | Gene symbol | Gene title | 108 dpi | 158 dpi | 198 dpi |
| Immune Response | |||||
| 1431591_s_at | Isg15 | ISG15 ubiquitin-like modifier | 1.2 | 2.3 | 2.1 |
| 1418392_a_at | Gbp3 | guanylate binding protein 3 | 1.2 | 1.8 | 1.7 |
| 1426397_at | Tgfbr2 | transforming growth factor, beta receptor II | 1.1 | 1.2 | 1.7 |
| 1424617_at | Ifi35 | interferon-induced protein 35 | 1.1 | 1.6 | 1.6 |
| 1451318_a_at | Lyn | Yamaguchi sarcoma viral (v-yes-1) oncogene homolog | 1.4 | 1.4 | 1.6 |
| 1425156_at | Gbp6 | guanylate binding protein 6 | 1.1 | 1.7 | 1.5 |
| 1418641_at | Lcp2 | lymphocyte cytosolic protein 2 | 1.0 | 1.6 | 1.4 |
| Intracellular Signaling Cascade | |||||
| 1460287_at | Timp2 | tissue inhibitor of metalloproteinase 2 | 1.3 | 1.6 | 2.6 |
| 1460188_at | Ptpn6 | protein tyrosine phosphatase, non-receptor type 6 | 1.6 | 1.9 | 2.4 |
| 1455132_at | A430107D22Rik | RIKEN cDNA A430107D22 gene | 1.5 | 1.5 | 1.4 |
| 1452398_at | Plce1 | phospholipase C, epsilon 1 | 1.1 | 1.3 | 1.6 |
| 1438936_s_at | Ang | angiogenin, ribonuclease, RNase A family, 5 | 1.0 | 1.4 | 1.9 |
| 1428479_at | Nfatc1 | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 | 1.6 | 1.3 | 1.4 |
| 1416675_s_at | Plcd1 | phospholipase C, delta 1 | 1.6 | 1.2 | 1.2 |
| Protein Modification Process | |||||
| 1418191_at | Usp18 | ubiquitin specific peptidase 18 | 1.5 | 3.6 | 3.9 |
| 1451584_at | Havcr2 | hepatitis A virus cellular receptor 2 | 1.3 | 1.8 | 2.1 |
| 1416340_a_at | Man2b1 | mannosidase 2, alpha B1 | 1.5 | 1.4 | 1.7 |
| 1449342_at | Ptplb | protein tyrosine phosphatase-like (proline instead of catalytic arginine), member b | −1.1 | 1.4 | 1.5 |
| 1433768_at | Palld | palladin, cytoskeletal associated protein | 1.3 | 1.1 | 1.5 |
| Response to Stimulus | |||||
| 1423547_at | Lyz2 | lysozyme 2 | 1.8 | 3.4 | 5.5 |
| 1421009_at | Rsad2 | radical S-adenosyl methionine domain containing 2 | 1.3 | 2.1 | 2.2 |
| 1424254_at | Ifitm1 | interferon induced transmembrane protein 1 | −1.1 | 1.7 | 1.1 |
| Ubiquitin Cycle | |||||
| 1433664_at | Ube2q2 | ubiquitin-conjugating enzyme E2Q (putative) 2 | −1.3 | −1.3 | −1.5 |
| 1419562_at | Birc6 | baculoviral IAP repeat-containing 6 | −1.8 | −1.2 | −1.3 |
| Cellular Developmental Process | |||||
| 1437947_x_at | Vdac1 | voltage-dependent anion channel 1 | −1.2 | −1.7 | −1.3 |
| 1453134_at | Pik3ca | phosphatidylinositol 3-kinase, catalytic, alpha polypeptide | −1.7 | −1.4 | −1.3 |
| 1449044_at | Eef1e1 | eukaryotic translation elongation factor 1 epsilon 1 | −1.7 | 1.0 | −1.1 |
| Other | |||||
| 1448891_at | Fcrls | Fc receptor-like S, scavenger receptor | 2.2 | 2.0 | 2.8 |
| 1421457_a_at | Samsn1 | SAM domain, SH3 domain and nuclear localization signals, 1 | 1.3 | 1.9 | 2.3 |
| 1451655_at | Slfn8 | schlafen 8 | 1.1 | 1.8 | 2.3 |
| 1418825_at | Irgm | immunity-related GTPase family, M | 1.2 | 2.0 | 2.3 |
| 1420591_at | Gpr84 | G protein-coupled receptor 84 | 1.4 | 1.7 | 2.2 |
| 1424754_at | Ms4a7 | membrane-spanning 4-domains, subfamily A, member 7 | −1.0 | 1.3 | 2.1 |
| 1443698_at | Xaf1 | XIAP associated factor 1 | 1.1 | 1.8 | 2.1 |
| 1436346_at | Cd109 | CD109 antigen | 1.1 | −1.1 | 2.1 |
| 1449227_at | Ch25h | cholesterol 25-hydroxylase | 1.1 | 1.6 | 2.0 |
| 1451564_at | Parp14 | poly (ADP-ribose) polymerase family, member 14 | 1.4 | 1.9 | 2.0 |
| 1421408_at | Igsf6 | immunoglobulin superfamily, member 6 | 1.2 | 1.3 | 2.0 |
| 1454757_s_at | D12Ertd647e | DNA segment, Chr 12, ERATO Doi 647, expressed | 1.2 | 1.7 | 1.9 |
| 1451537_at | Chi3l1 | chitinase 3-like 1 | 1.1 | 1.7 | 1.8 |
| 1419879_s_at | Trim25 | tripartite motif-containing 25 | 1.2 | 1.3 | 1.8 |
| 1456133_x_at | Itgb5 | integrin beta 5 | 1.5 | 1.3 | 1.8 |
| 1421374_a_at | Fxyd1 | FXYD domain-containing ion transport regulator 1 | −1.6 | 1.4 | 1.8 |
| 1448561_at | Ncf2 | neutrophil cytosolic factor 2 | 1.4 | 1.6 | 1.8 |
| 1451132_at | Pbxip1 | pre-B-cell leukemia transcription factor interacting protein 1 | 1.3 | 1.3 | 1.8 |
| 1426454_at | Arhgdib | Rho, GDP dissociation inhibitor (GDI) beta | 1.1 | 1.7 | 1.7 |
| 1449591_at | Casp4 | caspase 4, apoptosis-related cysteine peptidase | 1.0 | 1.5 | 1.7 |
| 1434945_at | Lpcat2 | lysophosphatidylcholine acyltransferase 2 | 1.2 | 1.3 | 1.7 |
| 1450234_at | Ms4a6c | membrane-spanning 4-domains, subfamily A, member 6C | 1.3 | 1.7 | 1.7 |
| 1460351_at | S100a11 | S100 calcium binding protein A11 (calgizzarin) | −1.6 | 1.4 | 1.7 |
| 1422571_at | Thbs2 | thrombospondin 2 | 1.2 | 1.3 | 1.6 |
| 1429525_s_at | Myo1f | myosin IF | 1.2 | 1.5 | 1.6 |
| 1418099_at | Tnfrsf1b | tumor necrosis factor receptor superfamily, member 1b | 1.2 | 1.4 | 1.6 |
| 1458019_at | Mus musculus Transcribed locus | 1.0 | 1.4 | 1.6 | |
| 1424829_at | A830007P12Rik | RIKEN cDNA A830007P12 gene | 1.3 | 1.2 | 1.5 |
| 1454748_at | Naprt1 | nicotinate phosphoribosyltransferase domain containing 1 | 1.6 | 1.3 | 1.5 |
| 1451139_at | Slc39a4 | solute carrier family 39 (zinc transporter), member 4 | 1.3 | 1.1 | 1.5 |
| 1437886_at | Klhl6 | kelch-like 6 (Drosophila) | 1.2 | 1.3 | 1.5 |
| 1420624_a_at | Vamp8 | vesicle-associated membrane protein 8 | −1.5 | 1.2 | 1.3 |
| 1421221_at | Bco2 | beta-carotene oxygenase 2 | 1.3 | 1.5 | 1.3 |
| 1449960_at | Nptx2 | neuronal pentraxin 2 | −1.2 | 1.5 | 1.3 |
| 1418451_at | Gng2 | guanine nucleotide binding protein (G protein), gamma 2 | −1.5 | 1.1 | 1.2 |
| 1424058_at | Prrc1 | proline-rich coiled-coil 1 | 1.5 | 1.1 | 1.2 |
| 1427903_at | Phpt1 | phosphohistidine phosphatase 1 | −1.7 | 1.1 | 1.1 |
| 1419812_s_at | Ccdc56 | coiled-coil domain containing 56 | −1.6 | 1.1 | 1.0 |
| 1426187_a_at | Hax1 | HCLS1 associated X-1 | −1.6 | 1.1 | 1.0 |
| 1417653_at | Pvalb | parvalbumin | −2.6 | 1.0 | 1.0 |
| 1435522_a_at | 2310016E02Rik | RIKEN cDNA 2310016E02 gene | −2.0 | 1.1 | 1.0 |
| 1428464_at | Ndufa3 | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 3 | −2.1 | 1.1 | 1.0 |
| 1416979_at | Pomp | proteasome maturation protein | −1.6 | 1.1 | 1.0 |
| 1424122_s_at | Commd1 | COMM domain containing 1 | −1.7 | 1.2 | −1.0 |
| 1416277_a_at | Rplp1 | ribosomal protein, large, P1 | −1.6 | 1.1 | −1.0 |
| 1450818_a_at | Ndufa7 | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 7 (B14.5a) | −1.5 | 1.1 | −1.0 |
| 1452790_x_at | Ndufa3 | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 3 | −2.0 | 1.0 | −1.0 |
| 1427102_at | Slfn4 | schlafen 4 | 1.1 | 1.8 | −1.0 |
| 1434391_at | Hnrnpu | Heterogeneous nuclear ribonucleoprotein U | −1.7 | 1.1 | −1.0 |
| 1418083_at | 0610009B22Rik | RIKEN cDNA 0610009B22 gene | −1.5 | −1.0 | −1.1 |
| 1442887_at | AI853363 | expressed sequence AI853363 | −1.7 | −1.0 | −1.2 |
| 1422884_at | Snrpd3 | small nuclear ribonucleoprotein D3 | −1.7 | 1.5 | −1.4 |
| 1428570_at | Ccnc | cyclin C | −1.5 | −1.4 | −1.4 |
| 1450770_at | 3632451O06Rik | RIKEN cDNA 3632451O06 gene | −1.3 | −1.4 | −1.5 |
| 1417121_at | Gabra6 | gamma-aminobutyric acid (GABA-A) receptor, subunit alpha 6 | −1.1 | −1.3 | −1.6 |
| 1433452_at | B630019K06Rik | RIKEN cDNA B630019K06 gene | −1.1 | −1.5 | −1.6 |
Transcripts were subjected to the Gene Ontology classification system for discovery of biological processes affected by prion disease. Fold change vs. control at three time points are shown here.
Neuroinflammatory changes induced by prion disease and cuprizone treatment.
Oral dosing of mice with the copper chelator cuprizone was used to generate chronic astrocytosis similar to that seen during prion disease. To define the impact of cuprizone treatment, we initiated a time course study of the prion disease mimetic, cuprizone. Cuprizone-treated mice were age-matched to the mice used for our prion studies, then orally dosed for 3, 4, 6, 7, 8 or 10 weeks with 0.4% cuprizone, with untreated age-matched mice used as controls. To determine the most appropriate cuprizone treatment (i.e., treatment time that mimics prion pathology most closely), quantitative PCR (qPCR) was used to measure astrocyte-specific glial fibrillary acidic protein (Gfap) expression levels during prion time courses and cuprizone treatment (Fig. 1). As expected during prion disease, Gfap expression increased over the course of infection. With cuprizone treatment, Gfap expression peaked after seven weeks of treatment and reached a plateau after eight weeks. During clinical disease prion infection (198 dpi), Gfap abundance was approximately 12-fold above the Gfap levels measured in uninfected animals and 5-times higher than the Gfap measured in the cuprizone-treated animals (Fig. 1). At 158 dpi, Gfap levels were comparable to those observed in the 8 and 10 week cuprizone treated brains suggesting an equivalent amount of astrocytosis. The eight week cuprizone treatment was selected for gene expression analysis.
Figure 1.
Expression of Gfap during cuprizone treatment and prion disease. Relative Gfap mRNA levels from 3, 4, 6, 7, 8 and 10 weeks of cuprizone treatment and 108, 158 and 198 dpi RML infection were generated by the Pfaffl method after qPCR, as indicated by fold change versus age-matched controls. After 8 and 10 weeks of cuprizone intoxication, Gfap levels most closely resemble those observed at 158 dpi of prion disease. Tbp was used as an endogenous control for normalization.
Spongiform vacuolation and astrocytosis were compared in the cuprizone-treated and prion-infected brains. Following H&E staining of brain slices from cuprizone-treated mice, vacuoles were observed in the white matter and granular layer of the cerebellum as well as in the pons and frontal cortex (Fig. 2A and D). During prion disease, vacuoles were first observed in the cerebellum and midbrain at 158 dpi with spongiosis becoming more prevalent in the cerebellum, midbrain and frontal cortex at clinical disease (Fig. 2J and M). Robust astrocytosis was observed, immunohistochemically, using an antibody for the detection of astrocyte-specific GFAP, in the frontal cortex, thalamus, corpus callosum and interposed nucleus (Fig. 2E) in cuprizone-treated animals. In prion-infected animals, basal astrocyte levels were observed at 108 dpi, with astrocyte levels rising throughout the brain at 158 and 198 dpi (Fig. 2H, K and N), consistent with the expression of Gfap observed by qPCR.
PrPTSE was not observed in the brains of either mock-infected or cuprizone-treated mice when assayed for immunohistochemically (Fig. 2C and F). Although PrPTSE was not observed in brain samples from infected animals at 108 dpi, punctate staining was observed, at 158 dpi, in the pons and medulla and, at clinical stage, in the pons, medulla, midbrain, thalamus and frontal cortex (Fig. 2I, L and O).
Cuprizone treatment induces differential gene expression.
Microarray analysis identified 319 genes differentially expressed in mouse brain after eight weeks of cuprizone treatment. Of these, 307 transcripts were grouped according to their functional annotations (Fig. 3). The majority of upregulated transcripts were associated with two biological processes; (1) immune system (including inflammatory response) and (2) metabolic processes. Immune system genes that were upregulated include chemokine (C-C motif) ligands, Fc receptors, and components of the complement system. Metabolic process genes included proteolytic cathepsins (C, D, H, L, S and Z), and lysosymes 1 and 2. Other upregulated genes were related to the regulation of apoptosis, cell communication, cellular adhesion and localization. Of the genes downregulated during cuprizone treatment, several genes play a role in axon ensheathement (proteolipid protein (myelin)1 and claudin 11), likely due to the demyelinating effects of cuprizone.
Figure 3.
Functional analysis of transcripts differentially regulated in the brains of mice treated with cuprizone. The DAVID annotational database was utilized to determine which gene ontology biological processes were affected by cuprizone treatment (p ≤ 0.05). Upregulated transcripts are in green; downregulated transcripts are in blue.
In a simple comparison of the gene expression profiles of mice infected with prion disease and those treated with cuprizone, 54% (88 of 164) of the transcripts differentially expressed during prion disease were also differentially expressed following cuprizone treatment. These transcripts include 28 genes associated with the immune system, eight genes involved with cell death, and five genes associated in lipid metabolism. It is clear that similar molecular mechanisms exist during prion disease and cuprizone treatment, presumably due to the activation of astrocytes and microglia in both, as well as the death of specific cell types (e.g., neurons during prion disease, oligodendrocytes during cuprizone treatment), which lead to spongiform vacuoles upon histological examination. Conversely, 231 of the genes upregulated by cuprizone treatment were not induced by prion disease, suggesting that stress response pathways activated during cuprizone treatment may be compromised during prion infection.
Comparative analysis of prion-infected gene expression profiles and gene expression profiles associated with cuprizone treatment.
To identify transcripts related to prion disease and unchanged during cuprizone-induced neuroinflammation, probe sets from the prion disease time course experiment were re-analyzed (see Materials and Methods). One cluster emerged containing transcripts that increased in expression over the time course of prion disease (Fig. 4A). The transcripts in this cluster were statistically assessed, producing 333 transcripts which were increasing in expression over the course of prion disease. In parallel, probe sets from the cuprizone data set were also filtered on expression and a transcript with no expression change between control and cuprizone treatment was identified (TATA box binding protein—Tbp). Transcripts with 100-85% similarity were recognized, using the Euclidean similarity metric (Fig. 4B), producing 22,729 transcripts that were unchanged during cuprizone treatment. Analysis of both the cluster containing transcripts increasing in expression over the course of prion disease and the unchanging cuprizone data set identified 17 transcripts that were present in both data sets. These represent transcripts upregulated during prion disease but remained unchanging following cuprizone treatment (Fig. 4C). In total, 14 of the 17 transcripts were annotated. Two of the genes upregulated during prion disease are involved in the neuropeptide signaling, triggered by a peptide neurotransmitter binding to a cell surface receptor. These transcripts and their log2 transformed expression ratios are shown in Figure 5. Full gene names and fold change versus control values are available in Supplemental Table 3.
Figure 4.
Comparative analysis between transcripts increasing in expression during prion disease and unchanging during cuprizone treatment. (A) After SOM clustering analysis, one cluster emerged containing transcripts that increased in expression over the time course of prion infection (shown here, 333 total transcripts). (B) The cuprizone data set was mined for one transcript whose expression was unchanged as compared to control (Tbp). Using the Euclidean similarity metric, transcripts with 100-85% similar expression patterns to Tbp were identified (22,729 total transcripts). (C) Venn diagram illustrating the comparative gene expression analysis of both data sets described above. In total, 17 probe sets belonged to both data sets and represent transcripts whose expression increased during prion disease but did not change with cuprizone treatment. This overlap signifies transcripts whose expression is related to prion disease and not associated to the changing cell population that occurs during neuroinflammation.
Figure 5.
Annotated transcripts upregulated during prion disease and unrelated to cuprizone-induced neuroinflammation. Of the seventeen transcripts that increased in expression over the course of prion disease and remained unchanged during cuprizone treatment, fourteen were found to be annotated. The log2 transformed expression ratios versus control are shown here for ease in visualization, allowing gene expression to be centered on zero. The shaded region represents non-significant, unchanging transcript levels. Unchanging was classified as |0.0-0.3| log2 transformed expression ratio versus control.
Validation of microarray data.
To independently validate the gene expression data generated by our microarray experiments, qPCR was used to measures the expression of three genes identified as upregulated: radical S-adenosyl methionine domain containing 2 (Rsad2) which was previously unreported as upregulated during prion disease, as well as apolipoprotein D (ApoD) and suppressor of cytokine signaling 3 (Socs3) which were upregulated in prion disease and not during cuprizone treatment. Both ApoD8,11,13,16 and Socs3,14 have been previously described in prion disease gene array studies, and ApoD has been a useful gene for validation via qPCR.11 QPCR analyses used the same total RNA samples as used for initial microarray studies. For all genes examined, expression levels found by microarray analysis were confirmed, although, in general, we observed higher relative fold-change levels with qPCR than microarray analysis. This has been described as a common phenomenon, though trends remain consistent across assays.33 The general expression pattern of each transcript was confirmed, validating the expression results we obtained from microarray analysis (Fig. 6).
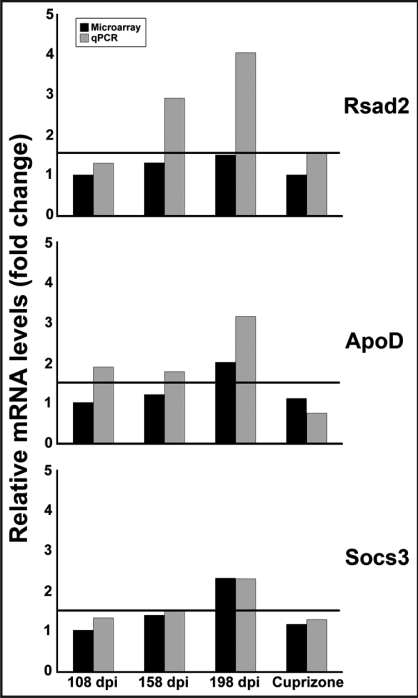
Figure 6.
Validation of selected microarray results by qPCR. The expression of Rsad2, ApoD and Socs3 was analyzed from RML-infected brain homogenate obtained at two preclinical time points (108 and 158 dpi) and at clinical disease (198 dpi) as well as after eight weeks of cuprizone treatment. Pooled RNA samples from eight infected/treated and eight controls were studied at each time point. The chart above shows relative mRNA levels, as indicated by fold change versus age-matched controls, generated by the Pfaffl method after qPCR, with a threshold line at 1.5-fold. Tbp was used as an endogenous control for normalization. Samples used for qPCR were from the same samples used for microarray analysis. Microarray data was taken from Table 1 and Supplemental Table 2.
Discussion
We identified novel transcripts differentially regulated during prion disease, transcripts differentially regulated after eight weeks of treatment with 0.4% cuprizone as well as transcripts upregulated during mouse-adapted prion disease that were unchanged during cuprizone-induced neuroinflammation.
Several studies have been undertaken to identify gene expression changes associated with prion disease.2–17 We identified many prion disease-induced transcripts that had been reported previously including Gfap, several cathepsins, components of the complement system, many Fc receptors, Cst7, Serpina3n, Lgals3, CD84, CD68 and CD52. Another 83 previously unidentified differentially regulated genes were identified during preclinical and clinical RML prion infection. This increase in the number of gene expression changes is likely due to the use of the Affymetrix Mouse Genome 430 2.0 Array, a more comprehensive mouse genome expression microarray than previously used and the inclusion of transcripts with a differential expression of ≥1.5 fold. Many of the transcripts that had been newly described as downregulated in response to prion infection (Table 1) were not present in our differentially regulated cuprizone dataset. The comparative analysis of the transcripts increasing in expression over the course of prion disease and unchanging with cuprizone treatment removed all of the downregulated transcripts from the analysis. Future investigation is necessary to determine if these gene changes are unique to prion disease and what their downregulation means for understanding prion pathology.
The upregulation of several transcripts related to the immune response GO-term suggests the response of immune cells in the brain to prion infection. At 158 dpi, many immune response related transcripts are upregulated ≥1.5-fold. When C57Bl/6 mice are inoculated i.p. with the RML prion agent, T-cells do not infiltrate the brain until the breakdown of the blood-brain barrier, 185 dpi34 and prion disease does not illicit an antibody-based immune response.35 Therefore, the upregulated transcripts related to the immune response GO-term are not expected to be associated with activated T-cells. It is likely that these transcripts are expressed by astrocytes and microglia, which are proliferating during preclinical infection. The expression of two of these genes, lsg15 and Lyn, is upregulated during astrocytosis in other systems; Isg15 is expressed by astrocytes following viral infection36 and Lyn is reported to be expressed by microglia after stimulation with beta-amyloid.37
We report the first gene expression analysis of brain tissue from mice treated with 0.4% cuprizone treatment. The use of 0.2% cuprizone as a demyelinating agent has been described previously and gene expression studies have been undertaken, but the neuropathological similarities to prion disease have only been described when the intake of cuprizone is ≥0.4%.26 The gene expression profile in response to cuprizone treatment described here is distinct and more robust than previous studies.38–41 Fifty-six transcripts reported as upregulated during cuprizone treatment are related to the immune response GO-term, many of which are also upregulated during prion disease, including Clec7a, Ccl3, Ccl6, Gp49a, parts of the classical complement system, several murine MHC components, and activated microglial markers like CD68 and Ly86. In addition, many transcripts involved in the regulation of apoptosis were also upregulated during cuprizone treatment, likely due to the demyelination process that selectively targets oligodendrocytes. More than seventy transcripts upregulated during cuprizone treatment were annotated as involved in metabolic processes, including several cathepsins, Hexa, Hexb and lysozymes. The upregulation of these transcripts is likely due to the breaking down of both cuprizone and the cellular remnants of oligodendrocytes undergoing apoptosis.
The use of cuprizone as a control for comparative gene expression analysis is a novel technique facilitating the identification of prion disease-specific transcript changes that are not related to the robust neuroinflammation observed during prion disease. By identifying transcripts whose expression increases during prion disease but remain unchanged with cuprizone treatment, transcript changes related to the changing cell population in the brain during neuroinflammation (i.e., proliferation of astrocytes and microglia) could be removed. In previous gene expression studies, known markers for astrocytes and microglia were among the most upregulated transcripts identified. While these changes are important for understanding the state of the brain during prion disease, they may mask important gene expression changes related to prion pathogenesis. Similarities are clearly present between prion disease and its mimetic, cuprizone, making it a useful control for comparative gene expression analysis. In total, 14 annotated transcripts were identified as increasing in expression during prion disease and not related to the neuroinflammatory profile associated with cuprizone treatment. To note, nine of these transcripts (Hsbp1, Socs3, Ifi44, D12Ertd647e, Casp4, Agrp, Plce1, Ptpbl, Ddx58) were upregulated preclinically and may provide insights into the progressive impairment of the CNS during prion infection.
We identified signaling pathways that may be important to prion pathogenesis. Transcripts upregulated during prion disease but not during cuprizone treatment fell into two signaling pathways. The products of Plce1 and Agrp are both involved in the neuropeptide signaling pathway. PLCE1 hydrolyses various polyphosphoinositides that activate the MAP kinase pathway, stimulating cell growth and differentiation.42 PLCE1 may be involved in stimulating the proliferation of glial cells in the brain during prion infection. AGRP is a neuropeptide secreted by the hypothalamus that controls body weight.43,44 AGRP also plays a role in adipocytokine signaling, another signaling pathway found to be represented in our data by AGRP, SLC2A4 and SOCS3. Adipocytes, or fat cells, secrete numerous cytokines that are important for metabolic homeostasis and play a role in inflammation.45 Slc2a4 (also known as Glut4) codes for a glucose transporter found in muscle and adipose tissue whose transporter action is dependent upon insulin.46 In adipocytes, SOCS3 works to suppress the expression of cytokines by inhibiting the JAK/STAT pathway in a negative feedback loop.47 SOCS3, however, plays a role in numerous signaling cascades and its expression may not be related to adipocytokine signaling alone. The role of these pathways in prion infection as well as confirmation with protein data is currently being explored.
In summary, eighty-three previously undescribed, annotated transcripts have been identified that are differentially regulated in the mouse brain during RML prion disease. We have also reported 307 annotated transcripts that are differentially regulated in the brain after eight weeks of 0.4% cuprizone treatment. We have described a comparative analysis technique that has allowed us to identify transcripts that were upregulated over the course of prion disease but remained unchanged during cuprizone treatment. With this novel analysis tool, we identified 14 annotated transcripts that change in expression during prion disease and do not appear to be related to the cell population changes occurring during disease. These transcripts may provide stepping stones for further investigation as to their roles in neurodegeneration and dysfunction during prion disease.
Materials and Methods
Animals and treatments.
C57Bl/6 mice were intraperitoneally (i.p.) inoculated with 50 ul of a 10% brain homogenate (in PBS) prepared from a mouse clinically infected with the RML strain of TSE agent. Control mice were mock-inoculated with a 10% brain homogenate prepared from uninfected C57Bl/6 mice. Mice were sacrificed at two preclinical time points (108 and 158 days post-inoculation [dpi]) and at the appearance of clinical symptoms (198 dpi). For the cuprizone control, C57Bl/6 mice were fed chow supplemented with 0.4% cuprizone for 3, 4, 6, 7, 8 or 10 weeks, a dose at which no mortality was observed. These mice were age-matched to the mice used in our prion experiments in such a way that at the termination of each cuprizone treatment, the animals age would coincide with the onset of clinical disease in the prion infected mice. Control mice were fed a normal diet. Upon sacrifice, all brains were removed and bi-sected sagittally. One half was used for RNA isolation and gene expression analysis while the other half was embedded in Optimal Cutting Temperature medium (Tissue Tek O.C.T. Compound, Sakura Finetek, Torrance, CA), frozen in liquid nitrogen and stored at −80°C until sagittal sectioning.
RNA purification and array hybridization.
Total RNA was purified from brain tissue (eight mice per treatment) using the Qiagen RNeasy Mini Kit protocol (Qiagen, Valencia, CA). cDNA was synthesized using the ImProm-II Reverse Transcription kit (Promega, Madison, WI) and an anchored T7-oligo(dT) V primer. Second strand synthesis was performed using DNA polymerase I (Invitrogen, Carlsbad, CA). Biotinylated cRNA was produced by in vitro transcription, incorporating biotin 14-CTP (Invitrogen, Carlsbad, CA) and biotin 16-UTP (Roche Applied Science, Indianapolis, IN) following manufacturer's instructions (Ambion, Austin, TX). Samples were hybridized onto Mouse Genome 430 2.0 Array (Affymetrix, Santa Clara, CA) at 45°C for 16 hours with a constant rotation at 60 rpm. The arrays were washed and stained using an Affymetrix fluidics station and scanned on an Affymetrix scanner. Each treatment array was performed in triplicate.
Data analysis.
Raw microarray data was uploaded into the GeneSpring GX 9.0 Analysis Platform (Agilent, Santa Clara, CA). Microarray data was normalized and background corrected using Robust Multi-Array Average (RMA) analysis.29 For identification of previously undescribed transcripts differentially regulated during prion disease, probe sets from the prion infection time course were filtered on expression (10.0–99.0th expression percentile) and statistically assessed using one-way ANOVA analysis (post hoc Tukey HSD and Benjamini-Hochberg correction; p ≤ 0.05) and further filtered by ≥1.5 fold expression. The cuprizone treatment data set was also filtered on expression (10.0–99.0th expression percentile) and subjected to unpaired t-test analysis (Benjamini-Hochberg correction; p ≤ 0.05) and further filtering by ≥1.5 fold expression against control.
For comparative analysis, the data sets from the prion infection time course were re-analyzed by filtering on expression (10.0–99.0th expression percentile) and clustering using 3 × 4 Self-Organizing Map (SOM) analysis. One cluster contained transcripts whose expression increased during the course of prion disease; these transcripts were statistically assessed using one-way ANOVA analysis (post hoc Tukey HSD and Benjamini-Hochberg correction; p ≤ 0.05) and further filtered by ≥1.5 fold expression. Transcripts from the cuprizone data set with no expression change between control and cuprizone treatment were identified using the Euclidean similarity metric (a coefficient of correlation 0.85–1.0). To categorize transcripts specific to prion disease (i.e., not associated with cuprizone-induced neuroinflammation), the prion time course dataset was compared to the unchanging cuprizone dataset. Probe sets present in both analyses are transcripts upregulated during prion disease and unchanged with cuprizone treatment. Annotational information and gene ontology biological functions were determined using the DAVID functional annotation database.30
Staining and immunohistochemistry.
Ten micron thick sagittal cryo-sections were cut and placed on glass slides for staining or immunohistochemistry (IHC). Slides were stained with hematoxylin and eosin (H&E) for histological detection of vacuolation. For IHC, slides were prepared as previously described31 with minor variations. Slides were incubated with primary antibody of either anti-GFAP (Promega Co., Madison, WI) or anti-prion protein, [SAF-83] (Cayman Chemical, Ann Arbor, Michigan), diluted 1:500 or 1:1,000, respectively, overnight at 4°C. After washing, the sections were incubated with biotinylated secondary antibody anti-rabbit IgG (1:200) for 1 h at room temperature, washed again and the antibody complex was detected using 3,3′-diaminobenzidine (DAB) (Sigma-Aldrich, St. Louis, MO) for GFAP or 3-amino-9-ethyl-carbazole (AEC) (Vector Laboratories, Burlingame, CA) for prion protein, according to manufacturer's protocol.
Quantitative RT-PCR.
cDNA was created as described above from total RNA isolated from mouse brain homogenates for additional PCR amplification using iQ Sybr Green Supermix (BioRad, Hercules, CA). The PCR amplification cycle varied with each gene being amplified (Suppl. Table 1). Relative expression values were determined using the iCycler software package and Pfaffl analysis using Tbp as the housekeeping gene for normalization.32
Acknowledgements
We would like to thank Debbie McKenzie (University of Alberta) for helpful discussion and assistance in manuscript preparation and David Westaway (University of Alberta) for critical reading of the manuscript. This research was supported by NIH grant RO1 NS060034 (to Judd M. Aiken).
Abbreviations
- RML
rocky mountain lab
- TSE
transmissible spongiform encephalopathy
- i.p.
intraperitoneal
- DPI
days post-inoculation
- RMA
robust multi-array average
- qPCR
quantitative PCR
- Gfap
glial fibrillary acidic protein
- Tbp
TATA box binding protein
- GO
gene ontology
Footnotes
Previously published online as a Prion E-publication: http://www.landesbioscience.com/journals/prion/article/9059
Supplementary Material
References
- 1.Collinge J. Prion diseases of humans and animals: Their causes and molecular basis. Annu Rev Neurosci. 2001;24:519–550. doi: 10.1146/annurev.neuro.24.1.519. [DOI] [PubMed] [Google Scholar]
- 2.Duguid JR, Rohwer RG, Seed B. Isolation of cDNAs of scrapie-modulated RNAs by subtractive hybridization of a cDNA library. Proc Natl Acad Sci USA. 1988;85:5738–5742. doi: 10.1073/pnas.85.15.5738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Duguid JR, Bohmont CW, Liu NG, Tourtellotte WW. Changes in brain gene expression shared by scrapie and alzheimer disease. Proc Natl Acad Sci USA. 1989;86:7260–7264. doi: 10.1073/pnas.86.18.7260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Duguid JR, Dinauer MC. Library subtraction of in vitro cDNA libraries to identify differentially expressed genes in scrapie infection. Nucleic Acids Res. 1990;18:2789–2792. doi: 10.1093/nar/18.9.2789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Diedrich JF, Minnigan H, Carp RI, Whitaker JN, Race R, Frey W, 2nd, Haase AT. Neuropathological changes in scrapie and alzheimer's disease are associated with increased expression of apolipoprotein E and cathepsin D in astrocytes. J Virol. 1991;65:4759–4768. doi: 10.1128/jvi.65.9.4759-4768.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Duguid J, Trzepacz C. Major histocompatibility complex genes have an increased brain expression after scrapie infection. Proc Natl Acad Sci USA. 1993;90:114–117. doi: 10.1073/pnas.90.1.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Diedrich JF, Carp RI, Haase AT. Increased expression of heat shock protein, transferrin and beta 2-microglobulin in astrocytes during scrapie. Microb Pathog. 1993;15:1–6. doi: 10.1006/mpat.1993.1051. [DOI] [PubMed] [Google Scholar]
- 8.Dandoy-Dron F, Guillo F, Benboudjema L, Deslys JP, Lasmezas C, Dormont D, et al. Gene expression in scrapie. cloning of a new scrapie-responsive gene and the identification of increased levels of seven other mRNA transcripts. J Biol Chem. 1998;273:7691–7697. doi: 10.1074/jbc.273.13.7691. [DOI] [PubMed] [Google Scholar]
- 9.Baker CA, Manuelidis L. Unique inflammatory RNA profiles of microglia in creutzfeldtjakob disease. Proc Natl Acad Sci USA. 2003;100:675–679. doi: 10.1073/pnas.0237313100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Booth S, Bowman C, Baumgartner R, Dolenko B, Sorensen G, Robertson C, et al. Molecular classification of scrapie strains in mice using gene expression profiling. Biochem Biophys Res Commun. 2004;325:1339–1345. doi: 10.1016/j.bbrc.2004.10.150. [DOI] [PubMed] [Google Scholar]
- 11.Booth S, Bowman C, Baumgartner R, Sorensen G, Robertson C, Coulthart M, et al. Identification of central nervous system genes involved in the host response to the scrapie agent during preclinical and clinical infection. J Gen Virol. 2004;85:3459–3471. doi: 10.1099/vir.0.80110-0. [DOI] [PubMed] [Google Scholar]
- 12.Brown AR, Webb J, Rebus S, Williams A, Fazakerley JK. Identification of upregulated genes by array analysis in scrapie-infected mouse brains. Neuropathol Appl Neurobiol. 2004;30:555–567. doi: 10.1111/j.1365-2990.2004.00565.x. [DOI] [PubMed] [Google Scholar]
- 13.Riemer C, Neidhold S, Burwinkel M, Schwarz A, Schultz J, Kratzschmar J, et al. Gene expression profiling of scrapie-infected brain tissue. Biochem Biophys Res Commun. 2004;323:556–564. doi: 10.1016/j.bbrc.2004.08.124. [DOI] [PubMed] [Google Scholar]
- 14.Xiang W, Windl O, Wunsch G, Dugas M, Kohlmann A, Dierkes N, et al. Identification of differentially expressed genes in scrapie-infected mouse brains by using global gene expression technology. J Virol. 2004;78:11051–11060. doi: 10.1128/JVI.78.20.11051-11060.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Brown AR, Rebus S, McKimmie CS, Robertson K, Williams A, Fazakerley JK. Gene expression profiling of the preclinical scrapie-infected hippocampus. Biochem Biophys Res Commun. 2005;334:86–95. doi: 10.1016/j.bbrc.2005.06.060. [DOI] [PubMed] [Google Scholar]
- 16.Skinner PJ, Abbassi H, Chesebro B, Race RE, Reilly C, Haase AT. Gene expression alterations in brains of mice infected with three strains of scrapie. BMC Genomics. 2006;7:114. doi: 10.1186/1471-2164-7-114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xiang W, Hummel M, Mitteregger G, Pace C, Windl O, Mansmann U, Kretzschmar HA. Transcriptome analysis reveals altered cholesterol metabolism during the neurodegeneration in mouse scrapie model. J Neurochem. 2007;102:834–847. doi: 10.1111/j.1471-4159.2007.04566.x. [DOI] [PubMed] [Google Scholar]
- 18.Williams AE, Lawson LJ, Perry VH, Fraser H. Characterization of the microglial response in murine scrapie. Neuropathol Appl Neurobiol. 1994;20:47–55. doi: 10.1111/j.1365-2990.1994.tb00956.x. [DOI] [PubMed] [Google Scholar]
- 19.DeArmond SJ, Kristensson K, Bowler RP. PrPSc causes nerve cell death and stimulates astrocyte proliferation: A paradox. Prog Brain Res. 1992;94:437–446. doi: 10.1016/s0079-6123(08)61771-9. [DOI] [PubMed] [Google Scholar]
- 20.Eikelenboom P, Bate C, Van Gool WA, Hoozemans JJ, Rozemuller JM, Veerhuis R, Williams A. Neuroinflammation in alzheimer's disease and prion disease. Glia. 2002;40:232–239. doi: 10.1002/glia.10146. [DOI] [PubMed] [Google Scholar]
- 21.McGeer PL, Itagaki S, Boyes BE, McGeer EG. Reactive microglia are positive for HLADR in the substantia nigra of parkinson's and alzheimer's disease brains. Neurology. 1988;38:1285–1291. doi: 10.1212/wnl.38.8.1285. [DOI] [PubMed] [Google Scholar]
- 22.Martino G, Adorini L, Rieckmann P, Hillert J, Kallmann B, Comi G, Filippi M. Inflammation in multiple sclerosis: The good, the bad and the complex. Lancet Neurol. 2002;1:499–509. doi: 10.1016/s1474-4422(02)00223-5. [DOI] [PubMed] [Google Scholar]
- 23.Greenwood AD, Horsch M, Stengel A, Vorberg I, Lutzny G, Maas E, et al. Cell line dependent RNA expression profiles of prion-infected mouse neuronal cells. J Mol Biol. 2005;349:487–500. doi: 10.1016/j.jmb.2005.03.076. [DOI] [PubMed] [Google Scholar]
- 24.Blakemore WF. Observations on oligodendrocyte degeneration, the resolution of status spongiosus and remyelination in cuprizone intoxication in mice. J Neurocytol. 1972;1:413–426. doi: 10.1007/BF01102943. [DOI] [PubMed] [Google Scholar]
- 25.Ludwin SK. Central nervous system demyelination and remyelination in the mouse: An ultrastructural study of cuprizone toxicity. Lab Invest. 1978;39:597–612. [PubMed] [Google Scholar]
- 26.Pattison IH, Jebbett JN. Histopathological similarities between scrapie and cuprizone toxicity in mice. Nature. 1971;230:115–117. doi: 10.1038/230115a0. [DOI] [PubMed] [Google Scholar]
- 27.Hiremath MM, Saito Y, Knapp GW, Ting JP, Suzuki K, Matsushima GK. Microglial/macrophage accumulation during cuprizone-induced demyelination in C57BL/6 mice. J Neuroimmunol. 1998;92:38–49. doi: 10.1016/s0165-5728(98)00168-4. [DOI] [PubMed] [Google Scholar]
- 28.Bakker DA, Ludwin SK. Blood-brain barrier permeability during cuprizone-induced demyelination. implications for the pathogenesis of immune-mediated demyelinating diseases. J Neurol Sci. 1987;78:125–137. doi: 10.1016/0022-510x(87)90055-4. [DOI] [PubMed] [Google Scholar]
- 29.Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP. Exploration, normalization and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4:249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 30.Dennis G, Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA. DAVID: Database for annotation, visualization and integrated discovery. Genome Biol. 2003;4:3. [PubMed] [Google Scholar]
- 31.Hoefert VB, Aiken JM, McKenzie D, Johnson CJ. Labeling of the scrapie-associated prion protein in vitro and in vivo. Neurosci Lett. 2004;371:176–180. doi: 10.1016/j.neulet.2004.08.065. [DOI] [PubMed] [Google Scholar]
- 32.Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:45. doi: 10.1093/nar/29.9.e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lee CK, Weindruch R, Prolla TA. Gene-expression profile of the ageing brain in mice. Nat Genet. 2000;25:294–297. doi: 10.1038/77046. [DOI] [PubMed] [Google Scholar]
- 34.Lewicki H, Tishon A, Homann D, Mazarguil H, Laval F, Asensio VC, et al. T cells infiltrate the brain in murine and human transmissible spongiform encephalopathies. J Virol. 2003;77:3799–3808. doi: 10.1128/JVI.77.6.3799-3808.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Porter DD, Porter HG, Cox NA. Failure to demonstrate a humoral immune response to scrapie infection in mice. J Immunol. 1973;111:1407–1410. [PubMed] [Google Scholar]
- 36.Roberts ES, Zandonatti MA, Watry DD, Madden LJ, Henriksen SJ, Taffe MA, Fox HS. Induction of pathogenic sets of genes in macrophages and neurons in NeuroAIDS. Am J Pathol. 2003;162:2041–2057. doi: 10.1016/S0002-9440(10)64336-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Moore KJ, El Khoury J, Medeiros LA, Terada K, Geula C, Luster AD, Freeman MW. A CD36-initiated signaling cascade mediates inflammatory effects of beta-amyloid. J Biol Chem. 2002;277:47373–47379. doi: 10.1074/jbc.M208788200. [DOI] [PubMed] [Google Scholar]
- 38.Morell P, Barrett CV, Mason JL, Toews AD, Hostettler JD, Knapp GW, Matsushima GK. Gene expression in brain during cuprizone-induced demyelination and remyelination. Mol Cell Neurosci. 1998;12:220–227. doi: 10.1006/mcne.1998.0715. [DOI] [PubMed] [Google Scholar]
- 39.Arnett HA, Wang Y, Matsushima GK, Suzuki K, Ting JP. Functional genomic analysis of remyelination reveals importance of inflammation in oligodendrocyte regeneration. J Neurosci. 2003;23:9824–9832. doi: 10.1523/JNEUROSCI.23-30-09824.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Jurevics H, Largent C, Hostettler J, Sammond DW, Matsushima GK, Kleindienst A, et al. Alterations in metabolism and gene expression in brain regions during cuprizone-induced demyelination and remyelination. J Neurochem. 2002;82:126–136. doi: 10.1046/j.1471-4159.2002.00954.x. [DOI] [PubMed] [Google Scholar]
- 41.Bedard A, Tremblay P, Chernomoretz A, Vallieres L. Identification of genes preferentially expressed by microglia and upregulated during cuprizone-induced inflammation. Glia. 2007;55:777–789. doi: 10.1002/glia.20477. [DOI] [PubMed] [Google Scholar]
- 42.Lopez I, Mak EC, Ding J, Hamm HE, Lomasney JW. A novel bifunctional phospholipase c that is regulated by galpha 12 and stimulates the Ras/mitogen-activated protein kinase pathway. J Biol Chem. 2001;276:2758–2765. doi: 10.1074/jbc.M008119200. [DOI] [PubMed] [Google Scholar]
- 43.Shutter JR, Graham M, Kinsey AC, Scully S, Luthy R, Stark KL. Hypothalamic expression of ART, a novel gene related to agouti, is upregulated in obese and diabetic mutant mice. Genes Dev. 1997;11:593–602. doi: 10.1101/gad.11.5.593. [DOI] [PubMed] [Google Scholar]
- 44.Ollmann MM, Wilson BD, Yang YK, Kerns JA, Chen Y, Gantz I, Barsh GS. Antagonism of central melanocortin receptors in vitro and in vivo by agouti-related protein. Science. 1997;278:135–138. doi: 10.1126/science.278.5335.135. [DOI] [PubMed] [Google Scholar]
- 45.Guzik TJ, Mangalat D, Korbut R. Adipocytokines—novel link between inflammation and vascular function? J Physiol Pharmacol. 2006;57:505–528. [PubMed] [Google Scholar]
- 46.Watson RT, Khan AH, Furukawa M, Hou JC, Li L, Kanzaki M, et al. Entry of newly synthesized GLUT4 into the insulin-responsive storage compartment is GGA dependent. EMBO J. 2004;23:2059–2070. doi: 10.1038/sj.emboj.7600159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Emanuelli B, Peraldi P, Filloux C, Sawka-Verhelle D, Hilton D, Van Obberghen E. SOCS-3 is an insulin-induced negative regulator of insulin signaling. J Biol Chem. 2000;275:15985–15991. doi: 10.1074/jbc.275.21.15985. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.