Abstract
The cerebellum provides an excellent system for understanding how afferent and target neurons coordinate sequential intercellular signals and cell-autonomous genetic programs in development. Mutations in the orphan nuclear receptor RORα block Purkinje cell differentiation with a secondary loss of afferent granule cells. We show that early transcriptional targets of RORα include both mitogenic signals for afferent progenitors and signal transduction genes required to process their subsequent synaptic input. RORα acts through recruitment of gene-specific sets of transcriptional cofactors, including β-catenin, p300, and Tip60, but appears independent of CBP. One target promoter is Sonic hedgehog and recombinant Sonic hedgehog restores granule precursor proliferation in RORα-deficient cerebellum. Our results suggest a link between RORα and β-catenin pathways, confirm that a nuclear receptor employs distinct coactivator complexes at different target genes, and provide a logic for early RORα expression in coordinating expression of genes required for reciprocal signals in cerebellar development.
Cellular communication during brain development remains a crucial aspect of neuroscience that is not fully understood. The development of a circuit typically requires a series of reciprocal signals between cell types to coordinate cell number, migration, cytodifferentiation, axon pathfinding, synaptogenesis, pruning and cell type specific genetic programs that respond to these signals. In cerebellum, Purkinje neurons are the sole output of a stereotyped local circuit and organize this circuit in development. Purkinje cells must therefore negotiate signaling interactions with multiple afferent cell populations as they differentiate.
The cerebellum develops from a plate of cells that form a proliferative ventricular zone along the dorsal neural tube in mid-gestation (reviewed in (Goldowitz and Hamre, 1998; Hatten and Heintz, 1995; Wang and Zoghbi, 2001). Purkinje cell precursors leave the mitotic cycle and the ventricular zone during embryonic days 11–13 (E11–13) in mouse and begin to express RORα by E12.5. Basket and stellate inhibitory interneurons arise from this ventricular zone later. Migratory cells from the rhombic lip form a second germinal zone (the external granule layer, or EGL) by E12.5 that will give rise to glutamatergic granule cells. Signals from Purkinje cells are required for the proliferation, differentiation and maintenance of afferent neurons, particularly cerebellar granule cells that extend parallel fibers and brain stem olivary neurons that extend climbing fibers to Purkinje cell dendrites. Purkinje neurons in turn become dependent on signals from these cells. However, the genetic circuits that coordinate these activities are not understood.
Staggerer is a classical mutation of RORα that blocks Purkinje cell differentiation, resulting in congenital ataxia and cerebellar hypoplasia (Sidman et al., 1962). Elegant developmental studies in staggerer mice and staggerer <–> wild-type chimeras indicated that the immature synaptic arrangements, immature cell morphology, and retention of embryonic cell surface properties and other molecular markers are intrinsic to mutant Purkinje cells, while subsequent loss of granule cells is a secondary and noncell-autonomous consequence (Crepel et al., 1980; Hatten and Messer, 1978; Herrup and Mullen, 1979; Landis and Sidman, 1978; Sotelo and Changeux, 1974; Trenkner, 1979). In particular, staggerer Purkinje cells are competent to receive innervation from olivary climbing fibers, their first afferents in development, but not from granule cell parallel fibers shortly thereafter (Landis and Reese, 1977), suggesting a differential synaptic competence of these immature cells. Positional cloning demonstrated that staggerer is a null mutation of Rora, the gene encoding RORα (Hamilton et al., 1996). Independent alleles of Rora created by gene targeting show identical phenotypes (Dussault et al., 1998; Steinmayr et al., 1998). Within the cerebellum, Rora RNA is expressed at high levels in Purkinje cells and at much lower levels in basket and stellate cells (Hamilton et al., 1996; Nakagawa et al., 1997). Although consensus in vitro binding sites have been described (Giguere et al., 1995; Giguere et al., 1994), few endogenous targets have been demonstrated.
Here, we present a systematic analysis of the genetic program controlled by RORα during cerebellar development. Our results indicate transcription-level coordination of outgoing signals from Purkinje cells with activation of cell-autonomous machinery to receive subsequent signals from target cells. Granule precursors (which do not express RORα) express reduced levels of proliferation markers as early as E15.5 in staggerer, indicating the importance of embryonic Purkinje-to-granule mitogenic signaling. Sonic hedgehog (Shh) is a strong candidate for this signal as staggerer Purkinje cells express reduced levels of Shh. RORα binds the Shh promoter in vivo and is required for recruitment of transcriptional cofactors β-catenin and p300 to sites in the Shh promoter. Further, recombinant SHH is sufficient to stimulate proliferation of granule cell precursors in staggerer cerebellar slice cultures. RORα also regulates several genes required in Purkinje cells to process incoming excitatory synaptic input from granule cells, including a group of functionally interacting genes required for calcium second messenger signaling during granule-to-Purkinje synaptic signaling. RORα binds in vivo to promoters for each of five putative direct target genes tested, including Shh, Slc1a6, Itpr1, Pcp4, and Pcp2. Interestingly, RORα recruits distinct combinations of functionally important coactivators on each target promoter, indicating a critical role of promoter context in combinatorial control of gene expression by an orphan nuclear receptor. Together, these data link RORα-dependent transcriptional strategies to synaptic pathways for cell-intrinsic and extrinsic signaling in cerebellar development.
Results
RORα has a small and specific initial effect on global gene expression
To define the genetic programs controlled by RORα in developing Purkinje cells, we profiled RNA expression from staggerer and wild-type cerebellum every two days during perinatal development. RORα expression begins by E12.5 (Figure 1A), but the staggerer cerebellum is morphologically normal through E17.5 (Vogel et al., 2000). Thinning of the EGL is seen at birth and gross hypoplasia and cytological abnormalities are evident by P4. RNA samples prepared from sex-matched littermates of each genotype were converted into labeled cRNA for hybridization to Affymetrix Mu11k arrays. Approximately two-thirds of probe sets on the array are called “present” in at least two of 24 hybridizations performed. The concordance for 158,148 replicate data points (replicate samples with same genotype and age for each probe set) is 0.95; for probe sets with positive expression values Pearson’s r is 0.96 with a bias of 0.999.
Figure 1. RORα has a systematic effect on developmental RNA expression profile in cerebellum.
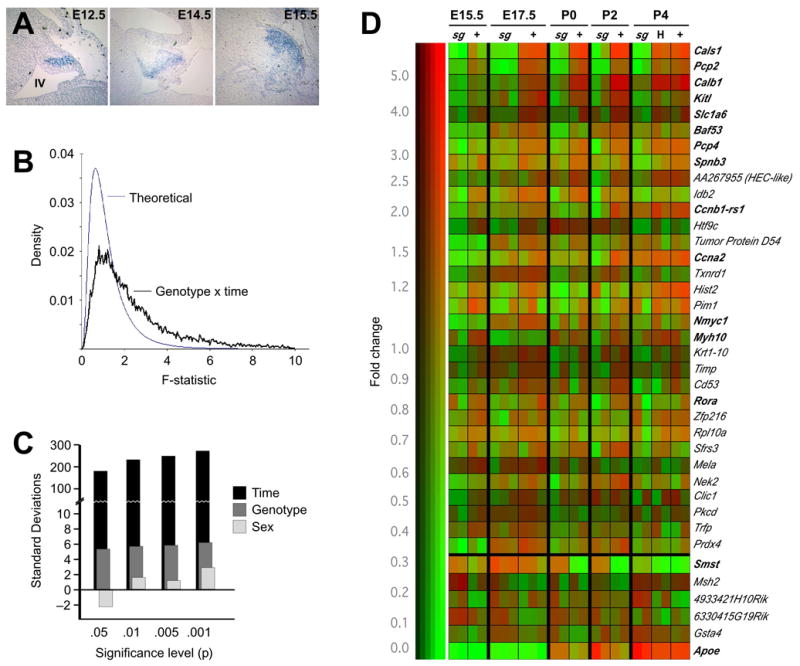
(A) In situ hybridization shows expression of Rora RNA in developing Purkinje cells as early as E12.5 and expressing cell populate the presumptive cortex by E15.5. Caudal is clockwise from top, position of the fourth ventricle is indicated in the first panel.
(B) Distribution of F statistics generated by standard two-factor ANOVA indicate significant effects of genotype and time (but not gender) in our multidimensional data set compared to the distribution generated under the null hypothesis of no effect.
(C) The number of standard deviations by which the data depart from expectation is plotted. The number of probe sets reaching given significance levels (p-values) for effect of time, Rora genotype, or gender is compared to the distribution of numbers expected by chance under the null hypothesis.
(D) Clustergram of genes identified in our analysis indicates the strength of genotype dependence and that the majority of differentially expressed genes do not have a strong heterozygote effect. Color indicates relative expression level among experiments and brightness indicates absolute signal strength. Probe sets with a p-value less than 0.05 for genotype effect by ANOVA, a minimum average difference of 500 according to MAS and a minimum 1.25-fold change are shown. Genes are arranged in order by the product of F-statistic for genotype and approximate fold change. For upregulated genes, only those with consistent patterns across the full perinatal window are shown.
We looked for systematic effects of age, staggerer genotype, gender, and interactions among these variables. Using a 2-factor analysis of variance (ANOVA; (Neter et al., 1985) we compared the distribution of F statistics for genotype and developmental time to the distribution under the null hypothesis of no effect (Figure 1B). The 95th percentile of the distribution of F for genotype and time together in our data set is approximately 117 standard deviations away from that expected by chance alone. Significant departures from expectation at other significance levels, and for each variable in isolation, indicate systematic effects of RORα genotype and developmental time on expression profiles (Figure 1C). By contrast, we see essentially no systematic impact of gender and among individual genes only the inactive X chromosome-specific transcript Xist emerges as a consistent and significant gender effect.
Expression of known target and control genes confirm the accuracy of the array data. In adult staggerer cerebellum, Itpr1 (Nakagawa et al., 1996) and Pcp2 (Hamilton et al., 1996) are not detected, while Calb1 (Nakagawa et al., 1996), Pcp4/Pep19 (Sangameswaran et al., 1989) and the mutant Rora transcript (Hamilton et al., 1996) are expressed at reduced levels when compared to wild-type expression. We confirmed these reported differences by semi-quantitative RT-PCR (not shown). Our array data identify each of these genes as significantly reduced during the perinatal window (Table 1 and Figure 1D). By contrast, several other Purkinje cell markers (Wnt3, Neurod2, Tead2 and Homer2) and housekeeping genes (Gapd, Eno2, Cd98, and repetitive elements such as MLV) do not show significant genotype effects.
Table 1.
Probe sets with ANOVA-based experiment-wide p-values for genotype effect ≤0.05 and fold change ≥1.25 at both E15.5 and E17.5
| Probe ID | Gene | ANOVA F(genotype) | p-value | change E15 & E17 | F x fold-change |
|---|---|---|---|---|---|
|
Decreased expression | |||||
| X61397_s_at | Cals1 | 157.378 | 0.0000 | 4.25 | 668.86 |
| M21532_s_at | Pcp2 | 116.929 | 0.0000 | 2.00 | 233.86 |
| M21531_s_at | Calb1 | 73.531 | 0.0000 | 2.25 | 165.44 |
| U44725_s_at | Kitl | 48.147 | 0.0000 | 3.00 | 144.44 |
| D83262_at | Slc1a6 (EAAT4) | 18.462 | 0.0015 | 3.25 | 60.00 |
| AA415606_at | Baf53a | 43.954 | 0.0000 | 1.25 | 54.94 |
| X17320_s_at | Pcp4 | 33.003 | 0.0002 | 1.25 | 41.25 |
| AF026489_at | Spnb3 | 29.314 | 0.0003 | 1.25 | 36.64 |
| ESTs, Weakly similar to retinoblastoma-associated protein | |||||
| AA267955_s_at | HEC | 25.684 | 0.0005 | 1.25 | 32.10 |
| Msa.1693.0_s_at | Idb2 | 20.554 | 0.0011 | 1.50 | 30.83 |
| AA426917_s_at | Ccnb1-rs1 | 22.250 | 0.0008 | 1.25 | 27.81 |
| X56044_s_at | Htf9c | 5.674 | 0.0385 | 4.50 | 25.53 |
| Msa.17592.0_s_at | pigpen protein | 19.534 | 0.0013 | 1.25 | 24.42 |
| Z26580_s_at | Ccna2 | 16.353 | 0.0024 | 1.25 | 20.44 |
| AA408677_rc_s_at | Txnrd1 | 14.784 | 0.0032 | 1.25 | 18.48 |
| Z30940_f_at | Hist2 | 12.012 | 0.0061 | 1.50 | 18.02 |
| Msa.1076.0_at | Pim1 | 13.469 | 0.0043 | 1.25 | 16.84 |
| X03919_s_at | Nmyc1 | 10.722 | 0.0084 | 1.50 | 16.08 |
| Msa.38014.0_s_at | Myh10 | 7.948 | 0.0182 | 1.75 | 13.91 |
| D78354_at | Plscr1 | 8.952 | 0.0135 | 1.50 | 13.43 |
| V00830_f_at | Krt1-10 | 8.721 | 0.0145 | 1.50 | 13.08 |
| V00755_s_at | Timp | 8.657 | 0.0147 | 1.50 | 12.99 |
| Msa.31660.0_s_at | Cd53 | 8.149 | 0.0171 | 1.50 | 12.22 |
| Msa.2058.0_s_at | Rora | 9.772 | 0.0108 | 1.25 | 12.21 |
| Msa.18074.0_f_at | Zfp216 | 6.892 | 0.0254 | 1.75 | 12.06 |
| Msa.29968.0_s_at | Mm.27526, arginyl-tRNA synthetase | 7.575 | 0.0204 | 1.50 | 11.36 |
| Msa.1847.0_f_at | Rpl10a | 8.785 | 0.0142 | 1.25 | 10.98 |
| Msa.16618.0_s_at | Sfrs3 | 8.264 | 0.0165 | 1.25 | 10.33 |
| AA450768_s_at | Mm.200828, ESTs | 6.845 | 0.0258 | 1.50 | 10.27 |
| Msa.54.0_f_at | Mela | 5.042 | 0.0486 | 2.00 | 10.08 |
| U95610_s_at | Nek2 | 7.627 | 0.0200 | 1.25 | 9.53 |
| Msa.19334.0_f_at | Clic1 | 5.884 | 0.0357 | 1.50 | 8.83 |
| X60304_at | Pkcd | 6.485 | 0.0290 | 1.25 | 8.11 |
| AA104750_at | Trfp | 5.616 | 0.0393 | 1.25 | 7.02 |
| U96746_s_at | Prdx4 | 4.990 | 0.0495 | 1.25 | 6.24 |
|
| |||||
|
Increased expression | |||||
| X51468_f_at | Smst | 58.140 | 0.0000 | 1.25 | 72.68 |
Reasoning that direct targets of RORα should be enriched among the earliest expression differences, we combined statistical and filtering methods to identify these genes. Probe sets with a p-value ≤ 0.05 by ANOVA for a genotype effect over the entire data set were filtered for empirical criteria of at least 1.25-fold enriched in wild-type relative to staggerer at both E15.5 and E17.5 and a minimum expression level (set to the point at which 75% of genes are called absent by Affymetrix Microarray Suite (MAS) in wild-type samples at that those times; this eliminates ~40% of probe sets). These criteria predict 32 genes down-regulated in embryonic staggerer cerebellum (Table 1). (Several genes were potentially up-regulated by these criteria; however, a majority of these genes show unusually poor reproducibility or complex expression patterns between genotypes over time and so are not considered further in this analysis). We have similarly sorted significant genes for effects at later and broader time windows (see supplemental data), which reveals an increasing divergence between genotypes over time. Similar results were obtained using the error correcting model in SAM (Tusher et al., 2001). We selected several early genes for further study based on statistical significance and consistent changes in pattern, with no changes in sign of the differences between genotypes. RORα-dependence of gene expression level and cellular pattern of expression were confirmed by quantitative RT-PCR (Q-PCR) and in situ hybridization. Specific genes are discussed below according to their expression patterns and likely developmental role.
Proliferation markers implicate early Purkinje cell mitogenic signaling to the EGL
Surprisingly, a large fraction (~ 1/4) of the earliest significant expression differences between staggerer and littermate controls are cell cycle and proliferation-related genes. The pattern of expression differences among these genes predicts early and progressive decrease in cell proliferation in the perinatal staggerer cerebellum (Table 1, Figure 1D, Figure 2A, and supplementary data). During this period of development, granule cell progenitors of the EGL are the only significant source of mitotic cells in the cerebellum. Several cyclins (Ccna1, Ccnb1-rs, Ccnb2, Ccnd1), a topoisomerase (Top2a), and a dUTPase sequence were identified by several modest-stringency filters of the complete data set. Closer statistical examination of the microarray data reveals additional proliferation genes and an interesting pattern of progressive cell cycle marker loss in staggerer cerebellum. By E15.5, modest but reproducible changes in the B-cyclins, thymidilate synthase (Tyms), Pcna, and Nmyc and its binding partner Baf53 (Park et al., 2002) are seen in staggerer samples. (An Nmyc-down-regulated gene, Ndr2, is up-regulated in the postnatal staggerer cerebellum, supplementary data). By E17.5, expression levels of Ccna1 and Ccnd1 (3 of 3 comparisons) are also reduced. Additional cell cycle markers decrease in early postnatal staggerer cerebellum. Two cyclin inhibitors, Cdkn1a and Cdkn2d, show modest increases in expression, consistent with reduced mitogenic signaling to the granule cell precursors in the EGL (not shown). In situ hybridization of Ccna2 and dUTPase selectively labels the proliferating pool in the EGL (Figure 2B).
Figure 2. Progressive loss of proliferation markers in external granule layer.
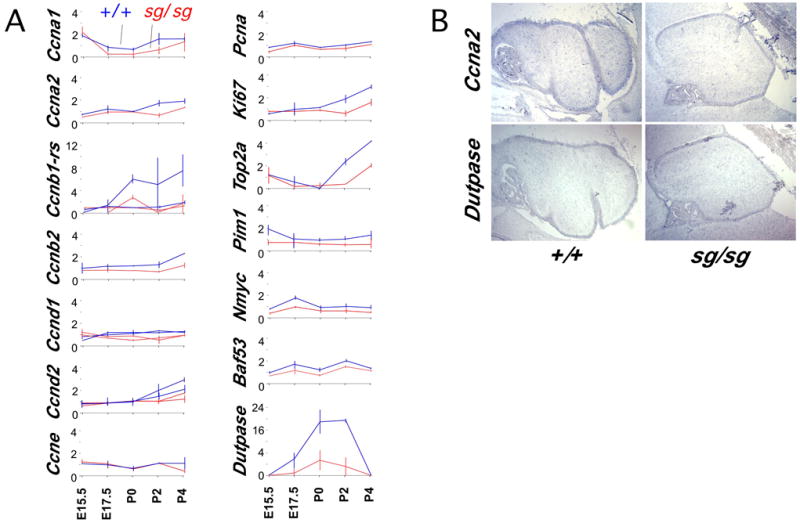
(A) Normalized expression data from Affymetrix Mu11K microarrays are plotted as line graphs. Staggerer samples are in red, littermate controls in blue. Average difference values for each probe set are normalized to make the average of all 24 hybridizations equal to 1. Horizontal lines indicate range of values among replicate samples. Multiple lines in a given plot represent overlay of normalized data from independent probe sets on the array. During this perinatal window, only the cells in the external granule layer should contribute in large number to pool of dividing cells in the cerebellum.
(B) Paired serial sections from control and staggerer specimens were mounted together on single slides and processed for in situ hybridization. In situ hybridization shows Ccna2 and dUTPase RNA expression is restricted to the EGL within the cerebellum at P2. Note the thinning of the staggerer EGL by this time.
Candidate mitogens affected by RORα
Unbiased analysis of the array data also identified expression differences in growth factor genes Gdf10, Kit-ligand (Kitl) and Smst, which might suggest them as candidate mitogens for the granule cell precursors (Figure 1D). All three genes are expressed in the Purkinje cell layer and their expression is validated with Q-PCR at P0 (Figure 3). However, Gdf10 expression is not in Purkinje neurons (Zhao et al., 1999). We find that Gdf10 is expressed in later-migrating cells from the ventricular zone (compare E15.5 expression in Figure 3 to Pcp4 in Figure 6A). C-kit receptors for KITL are only expressed on inhibitory and glial cells in cerebellum (Kim et al., 2003; Zhang and Fedoroff, 1997), and Kitl-deficient mice have no obvious cerebellar defect. Smst expression is increased in staggerer, the only consistently up-regulated gene we have found in staggerer Purkinje cells (Table 1, Figure 3). In situ hybridization shows a dramatic increase in the number of Smst expressing cells. Although SMST can exert stage-specific effects on the proliferation and differentiation of granule cells (Yacubova and Komura, 2002), its elevated expression in vivo is linked to stress responses (Matsui et al., 1993; Zupanc, 1999; Zupanc and Clint, 2001). Thus, although several signaling genes are affected by loss of RORα, none detected by array data seems likely to explain the observed loss of proliferation in the EGL.
Figure 3. RORα regulates candidate mitogenic factors in Purkinje cells.
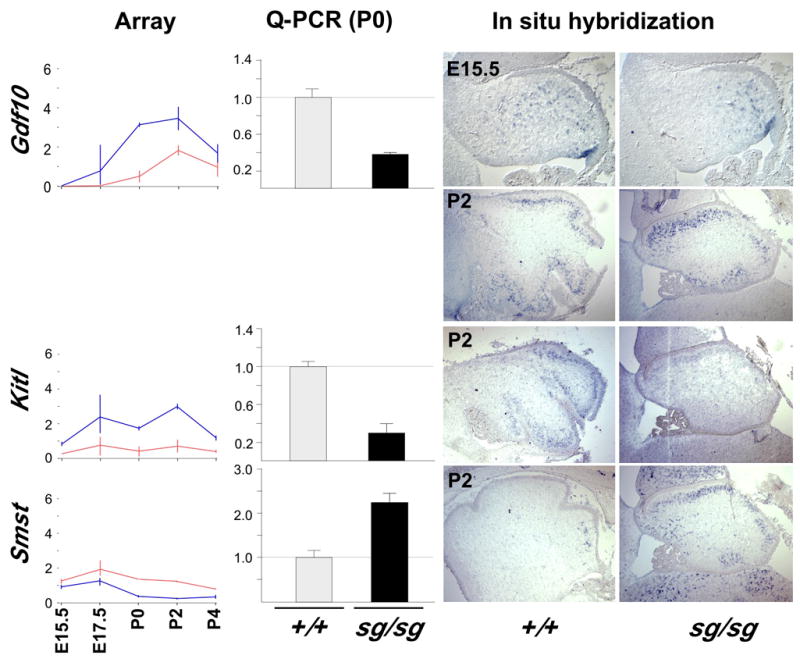
Line plots of normalized array data from Kitl, Smst, and Gdf10. Q-PCR confirms reduced expression of Gdf10, but increased expression of Smst RNA in staggerer by P0. Difference by genotype has a p-value ≤0.02 for Gdf10 and p≤0.0023 for Smst by 2-sided t-test. In situ hybridization at postnatal day 2 (as well as E15.5 for Gdf10) indicates the relevant cell population and qualitatively confirms altered expression level for Kitl, Smst, and Gdf10, including dramatic increase in Smst-positive cells by P2. Although Gdf10 expression is roughly in the Purkinje cell layer at P2, expression at E15.5 is not consistent with Purkinje cells (compare with Pcp4 pattern in Figure 6), suggesting expression in Bergmann glia or inhibitory interneurons.
Figure 6. RORα regulates Purkinje cell genes involved in calcium signaling.
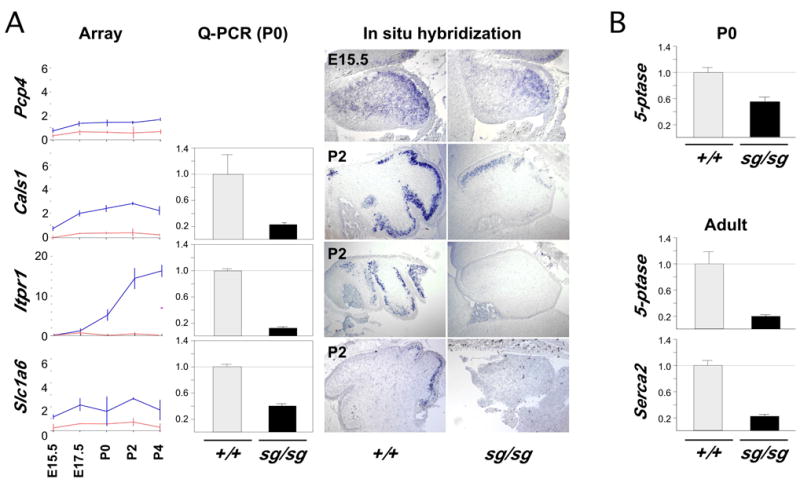
(A) Line plots include normalized array data from Pcp4, Slc1A6, Itpr1, and Cals1. In situ hybridization at postnatal day 2 (E15.5 for Pcp4) indicates the relevant cell population and qualitatively confirms decreased expression. Quantitative real-time PCR at P0 demonstrates reduced expression for Cals1, Slc1A6 and Itpr1. Difference by genotype was significant for all three genes (p≤0.01).
(B) SCA1 target genes that are relevant to calcium signaling are decreased in staggerer as well. Q-PCR showed 5-phosphatase levels decreased at P0, when Serca2/Atp2a2 is not reliably detected. Both genes are decreased in adult. Difference by genotype is significant for 5-phosphatase in both P0 (p ≤0.05) and adult (p ≤0.025) as is Serca2 (p≤0.03).
Early loss of Shh signaling in staggerer cerebellum
Purkinje cell-derived Sonic hedgehog (SHH) provides a potent mitogenic signal to the EGL in postnatal cerebellum (Dahmane and Ruiz-i-Altaba, 1999; Kenney and Rowitch, 2000; Wallace, 1999; Wechsler-Reya and Scott, 1999; Zhao et al., 2002), but Shh was not reliably detected in our microarray data. We therefore monitored Shh expression by Q-PCR (Figure 4). Shh RNA expression is reduced two to three-fold at E15.5, E17.5 and P0, suggesting SHH as a strong candidate for mediating the early effects of Purkinje cells on EGL proliferation.
Figure 4. Shh signaling disrupted in staggerer.
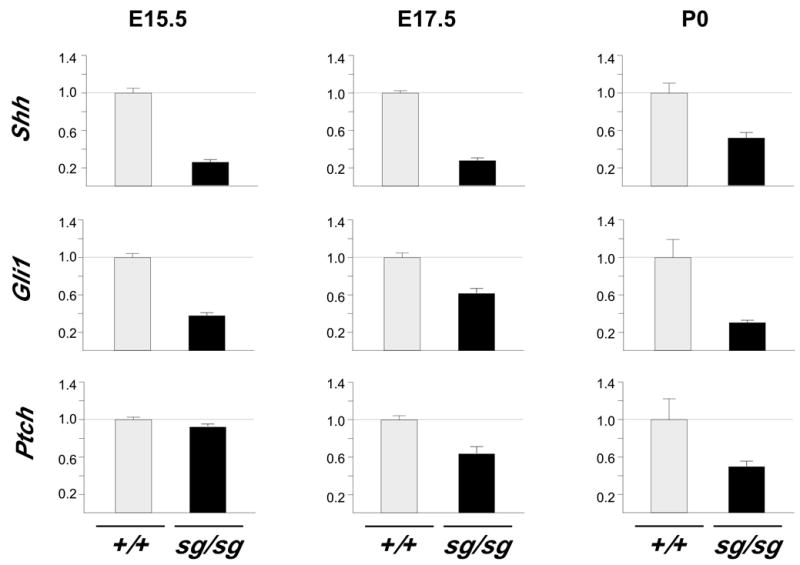
Q-PCR demonstrates reduced expression of Shh, Gli1, and Ptch RNA in staggerer cerebellum. Shh and Gli1 are significantly altered (p ≤0.05) at E15.5, while Ptch is not. All three genes are significantly different at both E17.5 and P0 (p ≤0.05).
To ask whether this decrease in Sonic hedgehog expression is sufficient to reduce signaling to granule cell precursors, we examined expression status of its known direct target genes in mutant and control cerebellum. Nmyc, an immediate early response to SHH signaling (Kenney et al., 2003) was identified as a significant expression difference in our initial microarray analysis (Figure 1D), but other direct targets such as Gli and Ptch were not reliably detected in that experiment. Quantification of Gli1 and Ptch using a commercial TaqMan assay demonstrated that expression of Gli1 is reduced at E15.5 and expression of both genes is reduced at E17.5 and P0 (Figure 4). All three of these diagnostic SHH targets are reduced in staggerer prior to significant loss of cell number and in greater magnitude than loss of cell number at birth. In contrast, other EGL markers present in the array data, including Zippro1 (RU49), Zic1, Zic2, and Zic3, are reliably detected but not statistically different at these times (supplementary data).
Exogenous SHH reverses granule cell proliferation deficit in staggerer
To ask whether SHH is sufficient to overcome the loss of granule cell precursor proliferation in the context of other potential signaling changes in staggerer, we cultured cerebellar slices from mutant and wild-type animals with or without recombinant SHH (rSHH) and labeled newly synthesized DNA by bromodeoxyuridine (BrdU) incorporation. First, sections from P4 mutant cerebellum were cultured for two days prior to labeling, to exhaust endogenous mitogens. BrdU detection by a fluorescein-conjugated antibody indicates that rSHH is sufficient to stimulate proliferation in the staggerer EGL (Figure 5A). Next, sections from P0 staggerer and control cerebellum were labeled beginning on the first day of culture and BrdU incorporation was quantified by fluorescence imaging (Figure 5B,C). Mutant and control slices in the absence of rSHH show approximately four-fold difference in BrdU labeling. However, addition of rSHH is sufficient to stimulate BrdU incorporation in the EGL of mutant slices even beyond the level of untreated nonmutant controls. Incorporation in treated staggerer sections at both 1 μg/ml and 0.5 μg/ml (not shown) is less than in control sections, consistent with the idea that the loss of endogenous SHH in staggerer is in a physiologically dose-responsive range. Differences between genotypes and between treatment groups confirm that the untreated staggerer EGL has reduced proliferation compared to littermate controls and that this reduction can be overcome by exogenous SHH.
Figure 5. Exogenous SHH stimulates staggerer granule cell precursor proliferation.
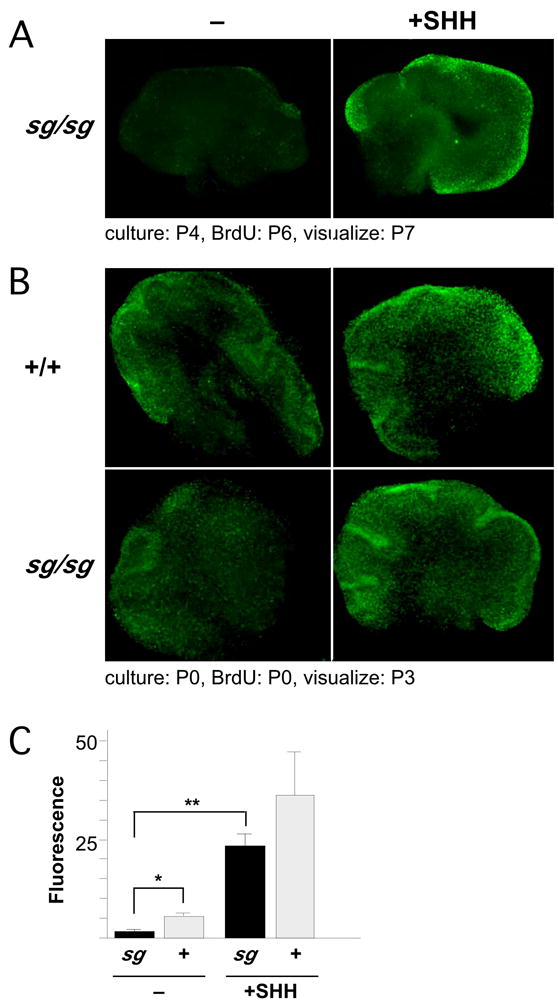
(A) rSHH is sufficient to induce proliferation in sg granule precursors. 300 μm sections from a P4 staggerer were cultured in the absence (left) or presence (right) of rSHH. After 2 days, dividing cells were labeled by BrdU incorporation for 24 hours, then fixed and visualized with a FITC-conjugated antibody under epifluorescence.
(B) Diminished proliferation of neonatal sg granule precursors is overcome by rSHH. 300 μm sections of wild-type and staggerer cerebellum at P0 were cultured immediately in media containing BrdU in the presence or absence of exogenous SHH. After 3 days in culture, sections were fixed, incubated with FITC-conjugated anti-BrdU antibody, and visualized by epifluorescence.
(C) Fluorescence density was measured across each section in using MetaMorph software. Average values from multiple sections ± s.e.m. are shown. * p ≤0.05; ** p ≤0.01.
RORα regulates expression of genes for calcium-mediated signal transduction
The Purkinje cell-expressed genes that show the most marked expression differences indicate a surprisingly small number of functional classes. The largest fraction of these genes is required for calcium second messenger signaling and glutamatergic signaling, a key source of signal-induced calcium flux in Purkinje cells. We validated several of these key expression differences by Q-PCR and by in situ hybridization, using matched littermate pairs (Figure 6A).
The calcium signal transduction genes that show RORα-dependent expression include Pcp4 (a calmodulin inhibitor), Itpr1 (IP3 receptor and calmodulin target), Cals1 (an Itpr1 binding partner), and Calb1 (a calcium buffer). The time course of the array data show that expression of these genes is significantly reduced by E17.5, before loss of Purkinje cell number (Vogel et al., 2000), suggesting that these expression differences reflect an altered regulatory mechanism rather than secondary pathology. Quantitative PCR data confirm the magnitude of reduced expression at birth. In situ hybridization shows specific expression of each of these genes in the Purkinje cells during development and indicates reduced expression levels in these cells in staggerer mutants.
A second functionally related set of RORα-responsive genes is required for glutamatergic signaling. Slc1a6 (which encodes EAAT4, the major glutamate transporter of Purkinje cells), and Spnb3 (brain-specific β-spectrin III, which anchors EAAT4 to the cytoskeleton; (Jackson et al., 2001)), also emerge from ANOVA and filtering analysis of the earliest significant changes in staggerer. Interestingly, Grm1, which encodes the later-expressed major metabotropic receptor at parallel fiber synapses, was not expressed in adult staggerer cerebellum in a previous RT-PCR screen for RORα-responsive genes (B.A.H., unpublished), though its expression is not in our array experiments. As with the calcium signaling genes, Q-PCR and in situ hybridization data confirm the timing, site and magnitude of diminished Slc1a6 expression in staggerer.
Intriguingly, Itpr1 and Slc1a6 were also identified among six genes down-regulated prior to onset of behavioral or pathologic symptoms in a mouse model of the SCA1 polyglutamine repeat disorder. Loss of expression was proposed to be mediated by sequestration of transcriptional coactivators used by nuclear receptors, including p300 (Lin et al., 2000). The other four genes identified in SCA1 were not represented or did not report expression in our array data. We used Q-PCR to test the expression levels of two of the remaining SCA1-downregulated genes with potential relevance to calcium signaling, Atp2a2 (a calcium-transporting ATPase also called Serca2), and an IP 5-phosphatase. Both genes show consistent and markedly reduced expression in staggerer compared to wild-type littermates when they are detectably expressed (Figure 6B).
RORα binds promoters of early-responding genes in vivo and is required for recruitment of coactivators
The discovery of several RORα-responsive genes in developing Purkinje cells permitted us to test the hypothesis that the earliest-responding genes are enriched for direct binding targets of RORα and to test whether loss of RORα affects recruitment of coactivators to their promoters by chromatin immunoprecipitations (Ch-IP). As we and others have previously found evidence for cross-talk between RORα and thyroid receptor pathways (Hamilton et al., 1996; Koibuchi and Chin, 1998; Kuno-Murata et al., 2000), we examined in vivo promoter-specific binding by RORα, TRβ (the major thyroid receptor in Purkinje cells (Strait et al., 1991), and selected nuclear receptor cofactors by Ch-IP from rapidly dissected cerebellum (Figure 7A).
Figure 7. RORα and coactivator binding and activation of target genes.
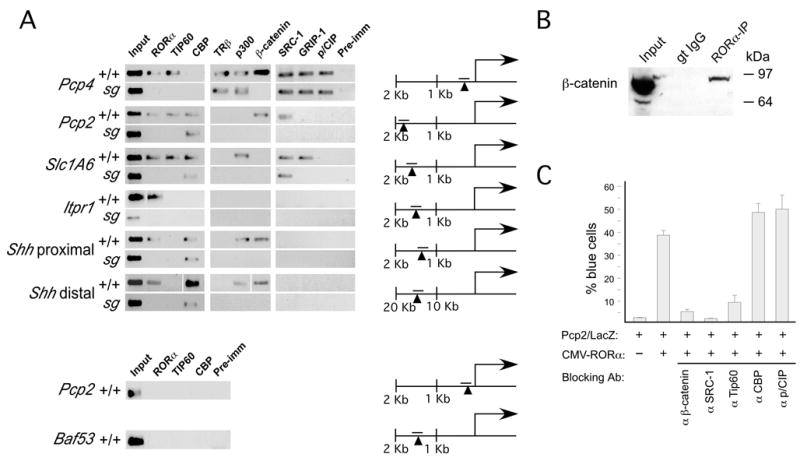
ChIP experiments on freshly dissected cerebella demonstrate in vivo binding of RORα to Pcp4, Pcp2, Slc1A6, Shh (at two sites), and Itpr1 promoters, but not to Baf53a and a second site in the Pcp2 promoter. RORα recruits specific sets of coactivators to each target promoter, including Tip60, β-catenin, and p300. None of the examined coactivators were bound to the Itpr1 promoter. Location of the PCR product relative to the known or presumed start of transcription is indicated for each gene; arrowhead indicates putative RORα binding site. (For the Shh distal site, a space in the image indicates different gels).
(B) Co-immunoprecipitation assays were performed on freshly dissected cerebella from P0 mice. β-catenin was pulled down with an RORα antibody but not a control IgG from the same host species. β-catenin was detected with two separate antibodies.
(C) Microinjection of blocking antibodies against β-catenin, SRC-1, and Tip60, but not CBP, p/CIP, and p/CAF blocked activation of the Pcp2/LacZ reporter in the presence of a CMV-RORα expression construct.
We assayed binding of RORα and TRβ at promoters for six RORα-responsive genes defined by expression data. Itpr1, Pcp2, Pcp4, Shh, and Slc1a6 are all Purkinje cell-selective genes within the cerebellum. RORα antibodies immunoprecipitated all five Purkinje cell promoters in non-mutant animals at P0. As a control for antibody specificity, parallel experiments show no binding in littermate staggerer mutants. In contrast, TRβ binds only at Pcp4, where it is independent of staggerer genotype. RORα is not bound to negative control promoter fragments at Baf53 (a myc protein binding partner involved in cell proliferation (Park et al., 2002) and an apparent indirect target of RORα in granule precursors) nor to an unoccupied site in the proximal Pcp2 promoter.
We next examined whether RORα is required for recruitment of coactivator complexes at the six in vivo RORα-binding sites we identified. By comparing coactivator recruitment in wild-type and staggerer cerebellum, we directly assayed the RORα-dependence of recruitment for each cofactor. By comparing across promoters, we could assess whether the set of recruited cofactors indicates a uniform complex or promoter-specific sets of cofactors. At the Pcp2 promoter we find RORα-dependent recruitment of Tip60, β-catenin, and SRC-1, while the Pcp4 promoter exhibited RORα-dependent recruitment of just Tip60 and β-catenin. Two distinct sites in the Shh promoter demonstrated identical RORα-dependent recruitment pattern, including β-catenin and p300, but not Tip60. The Slc1a6 promoter exhibited RORα-dependent recruitment of Tip60, p300, and GRIP-1, but not β-catenin. Surprisingly, the Itpr1 promoter failed to recruit any of the tested cofactors even though RORα is present on the promoter and required for its activation, suggesting other cofactors may be used at this promoter.
RORα has previously been shown to interact with several coactivators in biochemical assays, but not β-catenin. To test whether β-catenin interacts with RORα in vivo, we performed co-immunoprecipitation on protein lysates from freshly dissected tissue with anti-RORα or control serum and examined the precipitated materials by western blot. Immunoprecipitation with anti-RORα specifically co-precipitates significant levels of β-catenin (Figure 7B).
To determine whether the cofactors identified by ChIP exhibit a functional role in RORα dependent gene activation, we used a single cell nuclear microinjection assay (Figure 7C). To model RORα-dependent activation on a target promoter, a reporter construct containing 2 kb of the Pcp2 promoter fused to LacZ (Vandaele et al., 1991) was microinjected with or without an RORα expression plasmid into CV-1 cells. Co-injection of RORα resulted in a dramatic increase in activation of the reporter, consistent with its role as an activator of the Pcp2 gene. Co-injection of purified specific blocking antibodies to β-catenin, Tip60, or SRC-1 (Baek et al., 2002; Jepsen et al., 2000), all of which bind to the Pcp2 promoter in an RORα-dependent manner in situ, showed a dramatic decrease in reporter activation in the presence of the RORα expression construct. However, injection of blocking antibodies to CBP, which is bound to the Pcp2 promoter in an RORα independent fashion, and pCIP, which is not bound at all, did not significantly affect activity (controls in supplementary figure 1). These results are consistent with the functional requirement for RORα-recruited coactivators in induction of its target genes in cerebellar development.
Discussion
It is important to understand the mechanisms by which nuclear receptors mediate events in brain development and the potential unique function of orphan receptors versus liganded receptors. Our findings demonstrate an important role for RORα in the transcriptional coordination of sequential signaling pathways during cerebellar development and suggest a preliminary model for the reciprocal nature of these pathways (Figure 8A) and a mechanistic model for RORα-regulated expression of the component genes (Figure 8B). Identification of the earliest RORα-dependent genes suggests that RORα coordinates expression of both an outgoing Shh signal to granule cell precursors and a series of genes required to receive and interpret excitatory input from the mature granules that result. These are the earliest known effects of RORα and precede morphological abnormalities in the RORα-deficient cerebellum. RORα directly binds to the promoters of all five of the early target genes we tested. Intriguingly, the coactivators recruited in an RORα-dependent fashion exhibit target gene specificity, revealing additional patterns of coactivator usage for a nuclear receptor. Further, RORα is required to recruit β-catenin to the Shh promoter, suggesting the possibility that RORα mediates an unexpected link between the Wnt/β-catenin pathway and Shh expression. This analysis provides new insights into the molecular mechanisms of RORα and sheds new light on the genetic architecture of Purkinje cell differentiation.
Figure 8. A model for reciprocal signaling in Purkinje and granule cell differentiation.
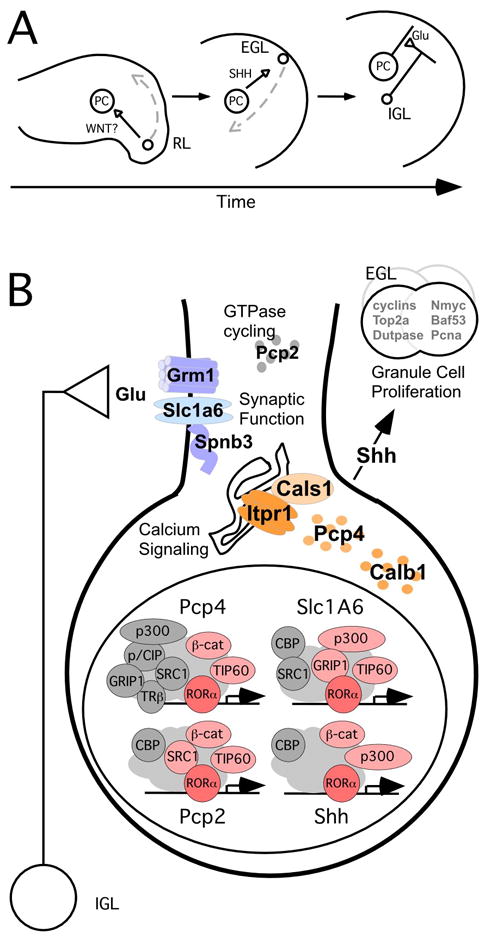
(A) Speculative model for sequential intercellular signaling events surrounding RORα-stimulated gene expression. Wnt1 is expressed by migratory granule cell precursors (GCPs) at the rhombic lip (RL). WNT expression may influence the nuclear accumulation of β-catenin in Purkinje cells (PC). Nuclear β-catenin acts as a cofactor on RORα-regulated promoters, including Shh. Purkinje cell SHH stimulates proliferation of GCPs in the external granule cell layer (EGL). Granule cells in the internal granule cell layer (IGL) make glutamatergic synapses on Purkinje cells, where RORα also regulates expression of signal transduction molecules to receive and process this input.
(B) Model for RORα-dependent gene expression in Purkinje cells. In Purkinje cell nuclei, RORα (red) recruits promoter-specific sets of coactivators (light red) to target genes in the context of additional, independent factors (gray). The products of known direct and potentially direct target genes are involved in receiving signals from afferent cells (blue), processing those signals via calcium release (orange), and the stimulation of proliferation of GCPs in the EGL (SHH).
RORα controls mitogenic potential in the EGL through Sonic hedgehog
Diminished expression of several EGL-expressed cell cycle and proliferation marker genes in the array data led us to identify Sonic hedgehog as a direct transcriptional target of RORα. It has been shown that Purkinje cell-derived SHH is both necessary for normal levels of granule cell genesis and sufficient to induce granule precursor proliferation in postnatal explant cultures in a dose-responsive manner (Wechsler-Reya and Scott, 1999), by stimulation of Nmyc, its binding partner Baf53, and various cyclins (Kenney et al., 2003; Kenney and Rowitch, 2000). Our work extends this to a much earlier developmental period and, importantly, demonstrates that Shh expression is regulated in Purkinje cell differentiation by RORα.
By three criteria, SHH appears to be the limiting factor in Purkinje-to-granule mitogenic signaling in staggerer. Shh expression level is reduced in staggerer mutant cerebellum prior to reduction in cell number. Reduced expression of SHH signaling targets, Nmyc, Ptch, and Gli1, also precedes the decrease in cell number, indicating a reduction in mitogenic signaling rather than a change in cell composition. Although granule cell number in the staggerer cerebellum may be slightly decreased near the end of embryonic development and is ~20% decreased at P0 (Yoon, 1972), we see a two to three-fold decrease in Shh and Gli1 at E15.5 and in Ptch at E17.5. Other EGL markers, including the pro-proliferation Zic family of transcription factors are not significantly altered at this time, further indicating that the reduction in Shh pathway expression is indicative of impaired signaling rather than cell number. Proliferation in the EGL accelerates after birth, consistent with the much larger magnitude difference in DNA labeling we see in postnatal slice cultures. Importantly, recombinant SHH is sufficient to drive proliferation in the EGL of staggerer slice cultures in a dose-responsive manner. Our data further defines the Shh pathway in granule cell genesis by demonstrating that RORα is an important transcriptional regulator of Shh in cerebellar development.
RORα regulates genes for calcium-mediated signal transduction
Differentiating Purkinje neurons must integrate a wide variety of extracellular cues. One of the most important signals received by Purkinje cells is excitatory input from granule cells through calcium-mobilizing receptors at the parallel fiber synapse. Strikingly, a large fraction of the genes we find regulated by RORα in the embryonic staggerer cerebellum are related to calcium-mediated signaling (e.g. Calb1, Pcp4, Itpr1, and Cals1). Moreover, these gene products may act as a functional unit: ITPR1 binds and co-localizes with the Cals1-encoded carbonic anhydrase related protein (CARP) in Purkinje cell soma and dendrites (Hirota et al., 2003) and ITPR1-mediated calcium release is inhibited by calmodulin, which in turn is inhibited by interaction with PCP4 protein (Slemmon et al., 1996). Thus, RORα appears to coordinately regulate transcript levels for interacting proteins involved in signal-dependent calcium release.
We also find reduced expression of genes required for excitatory neurotransmission at glutamatergic synapses, a major source of calcium mobilizing signals in Purkinje cells. These genes include Slc1a6, and later Grm1, which is required for elimination of supernumerary climbing fiber synapses (Kano et al., 1997). Pcp2, which contains a GoLoco G-protein modulatory domain and localizes to Purkinje cell dendrites (Luo and Denker, 1999; Zhang et al., 2002), could potentially function in this pathway. Interestingly, we also see modest but highly significant reduction in the major spectrin isoform gene, Spnb3. SPNB3 links EAAT4 to the cytoskeleton through a direct interaction (Jackson et al., 2001) and has the potential to coordinate larger cell surface complexes. Cell surface changes are particularly interesting in the context of altered cell surface properties and differential synaptic competence: staggerer Purkinje cells are able to receive climbing fibers, but neither stabilize nor mature postsynaptic responses to parallel fibers (Landis and Sidman, 1978), both of which are glutamatergic inputs. This could be due to a failure to translate recognition events into cytoskeletal rearrangement to form an appropriate postsynaptic site. Genetically staggerer Purkinje cells within chimeric animals also do not appear to receive parallel fiber input (Herrup and Mullen, 1979), implying that this is a cell-autonomous phenotype distinct from the effect of staggerer on granule cell genesis.
RORα-dependent coactivator recruitment exhibits promoter specificity
While recent work has shed considerable light on ligand-dependent and independent activities of ligand-activated nuclear receptors, the presumptive ligand-independent orphan receptors such as the RORs are less well understood. While RORα can interact in vitro with cofactors used by other nuclear receptors, (Atkins et al., 1999; Delerive et al., 2002; Lau et al., 1999), we have now identified specific DNA factors that are indeed recruited to the promoters of RORα target genes in an RORα-dependent fashion. We have documented that factors recruited by RORα including Tip60, SRC-1, and β-catenin have a functional role in RORα-dependent transcriptional activation. Intriguingly, CBP is not required for RORα-dependent activation and RORα is not required for CBP recruitment, a coactivator requirement common to virtually all ligand dependent nuclear receptors examined to date. This may emphasize the importance of Tip60 histone acetyltransferase (HAT) activity in RORα-induced gene activation. Recruitment of Tip60 has recently been shown to be a required coactivator for specific NF-κB gene targets (Baek et al., 2002), and is recruited to each of three RORα-responsive promoters in an RORα-dependent manner. Thus, our results demonstrate unique and distinct RORα-dependent recruitment of coactivators to target gene promoters, and functional activity of several recruited cofactors. This finding is likely to be prototypic for other nuclear receptors, particularly orphan receptors.
Our results link RORα to β-catenin in transcriptional activation, and are the first to indicate recruitment of β-catenin by an apparently unliganded nuclear receptor. β-catenin binds to RORα in vivo and is recruited to RORα-responsive promoters in an RORα-dependent manner. Although perturbations in Purkinje neurons ultimately will be required to prove the physiological importance of this interaction, we show that β-catenin is required for RORα-dependent activation of the Pcp2 promoter in a CV-1 cell culture model. Because nuclear β-catenin is often associated with Wnt pathway signaling, these results may also suggest a link between early Wnt signaling in the embryonic cerebellum and RORα-dependent activation of genes in the Purkinje cell lineage. Wnt signaling pathways are involved in multiple stages of cerebellar development. In particular, Wnt1 plays a role in the maintenance of the midbrain-hindbrain region during early development of the cerebellar anlage (Brault et al., 2001), and is also expressed by migratory granule cell progenitors by E12.5 (Shimamura et al., 1994). Mature granule cells also express Wnt7a, which refines afferent mossy fiber synapses (Hall et al., 2000). Alternatively, RORα could be acting on a Wnt-independent pool of nuclear β-catenin.
However, while RORα is required for recruitment of β–catenin and p300, it does not recruit CBP to the Shh promoter. Intriguingly, we find the same RORα-dependent complex at a second, remote site several kb upstream of Shh. This may indicate redundant use of the coregulatory apparatus in both promoter and enhancer, highly similar RORα-dependent complexes formed at independent sites, or communication between proximal and distal sites by looping, similar to that proposed for formation of the androgen receptor complex on the prostate specific antigen gene promoter and enhancer (Shang et al., 2002).
RORα coordinates outgoing and incoming signaling pathways
By examining a subset of RORα-responsive promoters, we are able to produce a snapshot of RORα’s role in promoting transcriptional activation, through promoter-specific recruitment of coactivating factors, as part of the developmental program of RORα-dependent gene expression in cerebellar development. The identity of direct RORα target genes in the early cerebellum suggests that RORα coordinates the activation of outgoing mitogenic signals to afferent precursors with the activation of signal transduction machinery required to receive their subsequent input. Taken together, our data link RORα to signaling through β-catenin, confirm that a nuclear receptor employs distinct coactivator complexes in activation of different target genes, and provide a logic for early RORα expression in coordinating signaling to afferent cells with preparing Purkinje cells to receive calcium-mediated signals in reply.
Experimental Procedures
Mice
Mice were originally obtained from the Jackson Laboratory and maintained locally. C57BL/6J–Rorasg + +/+ Myo5ad Bmp4se mice were backcrossed to C57BL/6J to remove the Myo5ad Bmp4se alleles. The colony was subsequently maintained by heterozygote matings. For prenatal timepoints heterozygous parents were bred in timed matings, with noon of the day after mating designated as E0.5. Concordance with developmental stage was confirmed by gross morphologic criteria (Kaufman, 1992). Genotyping was performed using a 3-primer PCR assay that produces alternate products from intact and staggerer intragenic deletion alleles and a second 3-primer assay for the closely related X and Y chromosome genes Zfx and Zfy; PCR products were resolved by electrophoresis through 2–3% agarose. Primers are Rora.sg, CTAGTCGGGGCTGAAACAGA; Rora.wt, GTGTTGAGCTGTTGGCCC; Rora.both, GGTTATAAAAGCCTGCTTCCG and Zfx, CAGAACACACTATTGAACAAAACG; Zfy, GTCAAATAGGTGCAATATCATCTT; ZfxZfy, CTCCATTCATACGAAAGACTATCC.
Microarray analysis
Cerebella from 203 mice were dissected in cold PBS under a microscope while blind to genotype. Each cerebellum was transferred immediately to 0.5 ml Trizol reagent, homogenized (Brinkmann Polytron 7mm generator at half power), and stored at −80°C. After genotyping, RNA fractions were prepared from individual sex-matched littermate pairs or pools including animals from several litters. Matched sets of 5 to 10 μg of total RNA was used for cDNA synthesis. Labeled cRNA target synthesis and hybridization to Affymetrix Mu11K probe arrays was performed according to manufacturer’s protocol.
Expression profiles were extracted using both Affymetrix software to generate spreadsheets and pairwise comparisons. GeneSpring 3.2.8 was used to visualize more complex patterns. Two to three replicates were used per genotype per time point. Differences reported here met the criteria of being significant in the Affymetrix analysis in at least two replicates at one time point on the end of the developmental series or at multiple time points within the developmental series with non-overlapping ranges in absolute difference values.
We used four computational approaches to identify candidate expression differences: rule-based, pair-wise comparisons in the Affymetrix Microarray Suite (MAS 4.0, Affymetrix), data filtering in GeneSpring (Silicon Genetics), permutation-based statistics in SAM (Tusher et al., 2001) and a standard 2-factor analysis of variance (ANOVA, (Neter et al., 1985). Lists of genes identified independently by each method are provided in supplementary data. All methods identify a consistent core of highly significant gene expression differences. Specific genes described here were selected for statistical significance, consistent or monotonic changes in magnitude during development, and no changes in sign for significant differences between genotypes.
Quantitative Real-Time PCR
RNA was prepared from P0 cerebellum of 117 individual mice and reverse transcribed using Superscript (Invitrogen) enzyme and primed with random hexamers. Real-time PCR assays were performed at the Center for Aids Research Genomics Core (Veterans Medical Research Foundation, La Jolla, CA). Forward primers used for QPCR were tagged with Z-sequence (ACAGAACCTGACCGTACA) for use with the Uniprimer fluorescence system (Intergen). Gli1 and Ptch Taqman assays were obtained from Applied Biosystems. Amplification and signal detection were performed on a Prism 7700 and analysis was performed using Sequence Detection System software (Applied Biosystems). All assays were done in duplicate and normalized to either 18S RNA or Gapdh. Primer sequences are listed in supporting information online.
In situ hybridization
In situ hybridization to RNA was performed by standard methods (Wilkinson and Nieto, 1993). Briefly, mice were perfused with PBS followed by 4% paraformaldehyde in PBS, post-fixed overnight in 4% paraformaldehyde in PBS at 4°C and embedded in paraffin. Dewaxed sections were hybridized overnight with digoxygenin-labeled cRNA and then washed, treated with RNAse A, washed again and incubated with alkaline phosphatase-conjugated anti-digoxygenin FAB fragments (Roche) that had been blocked with total embryonic head extract.
Co-Immunoprecipitation Assay
Co-IP was performed as described (Ezhevsky et al. 2001). Cerebella from 52 P0 mice were rapidly dissected on ice. Immunoprecipitation was performed with anti-RORα antibodies obtained from Santa Cruz Biotechnology. Membranes were probed with anti-β-catenin antibodies obtained from Santa Cruz Biotechnology (polyclonal) and from BD Biosciences (monoclonal).
Chromatin Immunoprecipitation Assays
Ch-IP was performed as described (Baek et al., 2002; Jepsen et al., 2000). Cerebella from 294 P0 mice were rapidly dissected on ice and individually processed immediately for crosslinking. Antibodies were obtained from Santa Cruz Biotechnology; all except for RORα have been reported previously in this assay (Baek et al., 2002; Jepsen et al., 2000). Precipitated fractions were assayed by PCR for the presence of each promoter using 3 μl of a 50 μl DNA extraction and 25 cycles of amplification. Primer sequences are listed in supporting information online.
Microinjection experiments
CV-1 cells were seeded at subconfluent density on glass coverslips, and rendered quiescent prior to injection by overnight incubation in serum free medium. All plasmids were diluted in rhodamine-conjugated dextran (Molecular Probes) to a final concentration of 0.1 μg/ml DNA as described (Lavinsky et al., 1998). Following microinjection, cells were incubated overnight to allow expression of the reporter gene, and subsequently stained for β-galactosidase expression. At least 250 cells were injected in every case, and all assays were performed at least in duplicate. Reporter gene expression was expressed as the percentage of rhodamine-containing injected cells that show any degree of staining for β-galactosidase. Rescue experiments were conducted as described (Torchia et al., 1997). All antibodies demonstrating a negative phenotype with the PCP2/LacZ reporter were tested in control experiments in the same cells with a reporter responsive to retinoic acid (Kamei et al., 1996).
Cerebellar cultures
300 μm sections from 12 rapidly dissected P0 and 7 P4 cerebella were transferred into Millicel culture inserts (Millipore) containing Neural Basal Media (Gibco) supplemented with N2, B27, L-glutamine, Pen/Strep, gentamycin (Gibco), and BrdU (Sigma). Slices were treated with 3 μg/ml recombinant Shh protein (BD Biosciences) for 48 hours and then pulsed with BrdU for an additional 18 hours. Sections were washed with PBS, fixed with 4 % PFA, followed by addition of 2 N HCl, neutralized with 0.1 N Borate, and permeabilized with 0.4% Triton. BrdU was detected with a biotinylated sheep anti-BrdU antibody (Biodesign) and FITC-streptavidin (Jackson Immunoresearch). Sections were mounted in ProLong (Molecular Probes) to preserve fluorescence prior to microscopy.
Supplementary Material
Supplemental Figure 1: Microinjection controls
Supplemental Table 1: Average difference values for each hybridization
Supplemental Table 2: Computed F-statistics by 2-factor ANOVA, with corresponding p-values and t-tests for effects of genotype, developmental time, and gender.
Supplemental Table 3: Genotype-dependent expression differences predicted by various methods.
Supplemental Table 4: Primer sequences for ChIP assays and QPCR.
Acknowledgments
We thank Linda Feng and Matt Hazle of the UCSD Cancer Center/VAMC GeneChip core facility for assistance with GeneChip hybridization, Yihua Zhao for mining UniGene data for the Mu11K gene list, Christine Plotkin of the Center for AIDS Research and Andrew Li for assistance with real-time PCR, and Tiffany Poon, Dorothy Concepcion, and Leslie Keeler for additional technical support. We thank Jacques Corbeil, Joseph Gleeson, Karl Herrup and John Welch for helpful discussions. This work was supported in part by start-up funds, a Basil O’Connor Scholar Award from the March of Dimes to B.A.H. and grants from the National Institutes of Health to N.J.S, M.G.R. and B.A.H. D. G. is supported in part by training grant T32 GM008666. S.H.B. is supported in part by the California Breast Cancer Research Program (BCRP Fellowship). M.G.R. is an investigator of the Howard Hughes Medical Institute. B.A.H. is a Pew Scholar in the Biomedical Sciences.
References
- Atkins GB, Hu X, Guenther MG, Rachez C, Freedman LP, Lazar MA. Coactivators for the Orphan Nuclear Receptor ROR{alpha} Mol Endocrinol. 1999;13:1550–1557. doi: 10.1210/mend.13.9.0343. [DOI] [PubMed] [Google Scholar]
- Baek SH, Ohgi KA, Rose DW, Koo EH, Glass CK, Rosenfeld MG. Exchange of N-CoR Corepressor and Tip60 Coactivator Complexes Links Gene Expression by NF-κB and β-Amyloid Precursor Protein. Cell. 2002;110:55–67. doi: 10.1016/s0092-8674(02)00809-7. [DOI] [PubMed] [Google Scholar]
- Brault V, Moore R, Kutsch S, Ishibashi M, Rowitch D, McMahon A, Sommer L, Boussadia O, Kemler R. Inactivation of the β-catenin gene by Wnt1-Cre-mediated deletion results in dramatic brain malformation and failure of craniofacial development. Development. 2001;128:1253–1264. doi: 10.1242/dev.128.8.1253. [DOI] [PubMed] [Google Scholar]
- Crepel F, Delhaye-Bouchaud N, Guastavino JM, Sampaio I. Multiple innervation of cerebellar Purkinje cells by climbing fibres in staggerer mutant mouse. Nature. 1980;283:483–484. doi: 10.1038/283483a0. [DOI] [PubMed] [Google Scholar]
- Dahmane N, Ruiz-i-Altaba A. Sonic hedgehog regulates the growth and patterning of the cerebellum. Development. 1999;126:3089–100. doi: 10.1242/dev.126.14.3089. [DOI] [PubMed] [Google Scholar]
- Delerive P, Chin WW, Suen CS. Identification of reverb alpha as a ROR alpha target gene. J Biol Chem. 2002;277:35013 – 35018. doi: 10.1074/jbc.M202979200. [DOI] [PubMed] [Google Scholar]
- Dussault I, Fawcett D, Matthyssen A, Bader JA, Giguere V. Orphan nuclear receptor ROR alpha-deficient mice display the cerebellar defects of staggerer. Mechanisms of Development. 1998;70:147–53. doi: 10.1016/s0925-4773(97)00187-1. [DOI] [PubMed] [Google Scholar]
- Giguere V, McBroom LD, Flock G. Determinants of target gene specificity for RORα1: monomeric DNA binding by an orphan nuclear receptor. Mol Cell Biol. 1995;15:2517–26. doi: 10.1128/mcb.15.5.2517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giguere V, Tini M, Flock G, Ong E, Evans RM, Otulakowski G. Isoform-specific amino-terminal domains dictate DNA-binding properties of RORα, a novel family of orphan hormone nuclear receptors. Genes Dev. 1994;8:538–53. doi: 10.1101/gad.8.5.538. [DOI] [PubMed] [Google Scholar]
- Goldowitz D, Hamre K. The cells and molecules that make a cerebellum. Trends Neurosci. 1998;21:375–82. doi: 10.1016/s0166-2236(98)01313-7. [DOI] [PubMed] [Google Scholar]
- Hall AC, Lucas FR, Salinas PC. Axonal remodeling and synaptic differentiation in the cerebellum is regulated by WNT-7a signaling. Cell. 2000;100:525–35. doi: 10.1016/s0092-8674(00)80689-3. [DOI] [PubMed] [Google Scholar]
- Hamilton BA, Frankel WN, Kerrebrock AW, Hawkins TL, FitzHugh W, Kusumi K, Russell LB, Mueller KL, van Berkel V, Birren B, et al. Disruption of the nuclear hormone receptor RORα in staggerer mice. Nature. 1996;379:736–739. doi: 10.1038/379736a0. [DOI] [PubMed] [Google Scholar]
- Hatten ME, Heintz N. Mechanisms of neural patterning and specification in the developing cerebellum. Ann Rev Neurosci. 1995;18:385–408. doi: 10.1146/annurev.ne.18.030195.002125. [DOI] [PubMed] [Google Scholar]
- Hatten ME, Messer A. Postnatal cerebellar cells from staggerer mutant mice express embryonic cell surface characteristic. Nature. 1978;276:504–506. doi: 10.1038/276504a0. [DOI] [PubMed] [Google Scholar]
- Herrup K, Mullen RJ. Staggerer chimeras: intrinsic nature of Purkinje cell defects and implications for normal cerebellar development. Brain Res. 1979;178:443–57. doi: 10.1016/0006-8993(79)90705-4. [DOI] [PubMed] [Google Scholar]
- Hirota J, Ando H, Hamada K, Mikoshiba K. Carbonic anhydrase related protein is a novel binding protein for Inositol1,4,5-Trisphosphate receptor type 1. Biochem J. 2003;372:435–41. doi: 10.1042/BJ20030110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson M, Song W, Liu MY, Jin L, Dykes-Hoberg M, Lin CI, Bowers WJ, Federoff HJ, Sternweis PC, Rothstein JD. Modulation of the neuronal glutamate transporter EAAT4 by two interacting proteins. Nature. 2001;410:89–93. doi: 10.1038/35065091. [DOI] [PubMed] [Google Scholar]
- Jepsen K, Hermanson O, Onami TM, Gleiberman AS, Lunyak V, McEvilly RJ, Kurokawa R, Kumar V, Liu F, Seto E, et al. Combinatorial roles of the nuclear receptor corepressor in transcription and development. Cell. 2000;102:753–763. doi: 10.1016/s0092-8674(00)00064-7. [DOI] [PubMed] [Google Scholar]
- Kamei Y, Xu L, Heinzel T, Thorchia J, Kurokawa R, Gloss B, Lin SC, Heyman RA, Rose DW, Glass CK, Rosenfeld MG. A CBP integrator complex mediates transcriptional activation and AP-1 inhibition by nuclear receptors. Cell. 1996;85:403–414. doi: 10.1016/s0092-8674(00)81118-6. [DOI] [PubMed] [Google Scholar]
- Kano M, Hashimoto K, Kurihara H, Watanabe M, Inoue Y, Aiba A, Tonegawa S. Persistent Multiple Climbing Fiber Innervationof Cerebellar Purkinje Cells in Mice Lacking mGluR1. Neuron. 1997;18:71–79. doi: 10.1016/s0896-6273(01)80047-7. [DOI] [PubMed] [Google Scholar]
- Kaufman M. The Atlas of Mouse Development. San Diego, CA: Academic Press; 1992. [Google Scholar]
- Kenney AM, Cole MD, Rowitch DH. Nmyc upregulation by sonic hedgehog signaling promotes proliferation in developing cerebellar granule neuron precursors. Development. 2003;130:15–28. doi: 10.1242/dev.00182. [DOI] [PubMed] [Google Scholar]
- Kenney AM, Rowitch DH. Sonic hedgehog promotes G(1) cyclin expression and sustained cell cycle progression in mammalian neuronal precursors. Mol Cell Biol. 2000;20:9055–67. doi: 10.1128/mcb.20.23.9055-9067.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Im JO, Won YJ, Yoon SY, Lee EJ, Lee JH, Hong HN. Upregulation of c-Kit receptor and stem cell factor in cerebellar inhibitory synapses in response to kainic acid. J Neurosci Res. 2003;71:72–78. doi: 10.1002/jnr.10466. [DOI] [PubMed] [Google Scholar]
- Koibuchi N, Chin WW. ROR alpha gene expression in the perinatal rat cerebellum: ontogeny and thyroid hormone regulation. Endocrinology. 1998;139:2335–41. doi: 10.1210/endo.139.5.6013. [DOI] [PubMed] [Google Scholar]
- Kuno-Murata M, Koibuchi N, Fukuda H, Murata M, Chin WW. Augmentation of Thyroid Hormone Receptor-Mediated Transcription by Ca2+/Calmodulin-Dependent Protein Kinase Type IV. Endocrinology. 2000;141:2275–2278. doi: 10.1210/endo.141.6.7612. [DOI] [PubMed] [Google Scholar]
- Landis DM, Reese TS. Structure of the Purkinje cell membrane in staggerer and weaver mutant mice. J Comp Neurol. 1977;171:247–60. doi: 10.1002/cne.901710208. [DOI] [PubMed] [Google Scholar]
- Landis DM, Sidman RL. Electron microscopic analysis of postnatal histogenesis in the cerebellar cortex of staggerer mutant mice. J Comp Neurol. 1978;179:831–63. doi: 10.1002/cne.901790408. [DOI] [PubMed] [Google Scholar]
- Lau P, Bailey P, Dowhan D, Muscat G. Exogenous expression of a dominant negative RORalpha1 vector in muscle cells impairs differentiation: RORalpha1 directly interacts with p300 and myoD. Nucl Acids Res. 1999;27:411–420. doi: 10.1093/nar/27.2.411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lavinsky RM, Jepsen K, Heinzel T, Torchia J, Mullen TM, Del-Rio AL, Ricote M, Ngo S, Gemsch J, Hilsenbeck SG, et al. Diverse signaling pathways modulate nuclear receptor recruitment of N-CoR and SMRT complexes. Proc Natl Acad Sci U S A. 1998;95:2920–2925. doi: 10.1073/pnas.95.6.2920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin X, Antalffy B, Kang D, Orr HT, Zoghbi HY. Polyglutamine expansion down-regulates specific neuronal genes before pathologic changes in SCA1. Nat Neurosci. 2000;3:157–63. doi: 10.1038/72101. [DOI] [PubMed] [Google Scholar]
- Luo Y, Denker BM. Interaction of Heterotrimeric G Protein Galpha o with Purkinje Cell Protein-2: Evidence for a novel Nucleotide Exchange Factor. J Biol Chem. 1999;274:10685–10688. doi: 10.1074/jbc.274.16.10685. [DOI] [PubMed] [Google Scholar]
- Matsui K, Masui A, Kato N, Adachi K. Levels of somatostatin and cholecystokinin in the brain of ataxic mutant mice. Life Sci. 1993;53:333–40. doi: 10.1016/0024-3205(93)90752-o. [DOI] [PubMed] [Google Scholar]
- Nakagawa S, Watanabe M, Inoue Y. Regional variation in expression of calbindin and inositol 1,4,5-trisphosphate receptor type 1 mRNAs in the cerebellum of the staggerer mutant mouse. European Journal of Neuroscience. 1996;8:1401–7. doi: 10.1111/j.1460-9568.1996.tb01602.x. [DOI] [PubMed] [Google Scholar]
- Nakagawa S, Watanabe M, Inoue Y. Prominent expression of nuclear hormone receptor ROR alpha in Purkinje cells from early development. Neuroscience Research. 1997;28:177–84. doi: 10.1016/s0168-0102(97)00042-4. [DOI] [PubMed] [Google Scholar]
- Neter J, Wasserman W, Kutner MH. Applied Linear Statistical Methods. 2. Homewood, IL: Richard D. Irwin & Co; 1985. [Google Scholar]
- Park J, Wood MA, Cole MD. BAF53 Forms Distinct Nuclear Complexes and Functions as a Critical c-Myc-Interacting Nuclear Cofactor for Oncogenic Transformation. Mol Cell Biol. 2002;22:1307–1316. doi: 10.1128/mcb.22.5.1307-1316.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sangameswaran L, Hempstead J, Morgan JI. Molecular cloning of a neuron-specific transcript and its regulation during normal and aberrant cerebellar development. Proc Natl Acad Sci U S A. 1989;86:5651–5. doi: 10.1073/pnas.86.14.5651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shang Y, Myers M, Brown M. Formation of the androgen receptor transcription complex. Molecular Cell. 2002;9:601–610. doi: 10.1016/s1097-2765(02)00471-9. [DOI] [PubMed] [Google Scholar]
- Shimamura K, Hirano S, McMahon A, Takeichi M. Wnt-1-dependent regulation of local E-cadherin and alpha N-catenin expression in the embryonic mouse brain. Development. 1994;120:2225–2234. doi: 10.1242/dev.120.8.2225. [DOI] [PubMed] [Google Scholar]
- Sidman RL, Lane PW, Dickie MM. Staggerer, a new mutation in the mouse affecting the cerebellum. Science. 1962;137:610–612. doi: 10.1126/science.137.3530.610. [DOI] [PubMed] [Google Scholar]
- Slemmon JR, Morgan JI, Fullerton SM, Danho W, Hilbush BS, Wengenack TM. Camstatins Are Peptide Antagonists of Calmodulin Based Upon a Conserved Structural Motif in PEP-19, Neurogranin, and Neuromodulin. J Biol Chem. 1996;271:15911–15917. doi: 10.1074/jbc.271.27.15911. [DOI] [PubMed] [Google Scholar]
- Sotelo C, Changeux JP. Transsynaptic degeneration ‘en cascade’ in the cerebellar cortex of staggerer mutant mice. Brain Res. 1974;67:519–526. doi: 10.1016/0006-8993(74)90499-5. [DOI] [PubMed] [Google Scholar]
- Steinmayr M, Andre E, Conquet F, Rondi-Reig L, Delhaye-Bouchaud N, Auclair N, Daniel H, Crepel F, Mariani J, Sotelo C, Becker-Andre M. staggerer phenotype in retinoid-related orphan receptor alpha-deficient mice. Proceedings of the National Academy of Sciences of the United States of America. 1998;95:3960–5. doi: 10.1073/pnas.95.7.3960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strait KA, Schwartz HL, Seybold VS, Ling NC, Oppenheimer JH. Immunofluorescence localization of thyroid hormone receptor protein β1 and variant α2 in selected tissues: Cerebellar Purkinje cells as a model for β1 receptor-mediated developmental effects of thyroid hormone in brain. Proc Nat’l Acad Sci, USA. 1991;88:3887–3891. doi: 10.1073/pnas.88.9.3887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torchia J, Rose DW, Inostroza J, Kamei Y, Glass CK, Rosenfeld MG. p/CIP: A novel factor related to nuclear receptor activation is a heteromeric partner of CBP/p300 and required for CBP coactivation function. Nature. 1997;387:677–685. doi: 10.1038/42652. [DOI] [PubMed] [Google Scholar]
- Trenkner E. Postnatal cerebellar cells of staggerer mutant mice express immature components on their surface. Nature. 1979;277:566–7. doi: 10.1038/277566a0. [DOI] [PubMed] [Google Scholar]
- Tusher VG, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A. 2001;98:5116–21. doi: 10.1073/pnas.091062498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vandaele S, Nordquist DT, Feddersen RM, Tretjakoff I, Peterson AC, Orr HT. Purkinje cell protein-2 regulatory regions and transgene expression in cerebellar compartments. Genes Dev. 1991;5:1136–48. doi: 10.1101/gad.5.7.1136. [DOI] [PubMed] [Google Scholar]
- Vogel MW, Sinclair M, Qiu D, Fan H. Purkinje cell fate in staggerer mutants: agenesis versus cell death. Journal of Neurobiology. 2000;42:323–37. doi: 10.1002/(sici)1097-4695(20000215)42:3<323::aid-neu4>3.0.co;2-2. [DOI] [PubMed] [Google Scholar]
- Wallace VA. Purkinje-cell-derived Sonic hedgehog regulates granule neuron precursor cell proliferation in the developing mouse cerebellum. Curr Biol. 1999;9:445–8. doi: 10.1016/s0960-9822(99)80195-x. [DOI] [PubMed] [Google Scholar]
- Wang VY, Zoghbi HY. Genetic regulation of cerebellar development. Nat Rev Neurosci. 2001;2:484–91. doi: 10.1038/35081558. [DOI] [PubMed] [Google Scholar]
- Wechsler-Reya RJ, Scott MP. Control of neuronal precursor proliferation in the cerebellum by Sonic Hedgehog. Neuron. 1999;22:103–14. doi: 10.1016/s0896-6273(00)80682-0. [DOI] [PubMed] [Google Scholar]
- Wilkinson DG, Nieto A. Detection of messenger RNA by in situ hybridization to tissue sections and whole mounts. Methods Enzymol. 1993;225:361–373. doi: 10.1016/0076-6879(93)25025-w. [DOI] [PubMed] [Google Scholar]
- Yacubova E, Komura H. Stage-specific control of neuronal migration by somatostatin. Nature. 2002;415:77–81. doi: 10.1038/415077a. [DOI] [PubMed] [Google Scholar]
- Yoon CH. Developmental mechanism for changes in cerebellum of “staggerer” mouse, a neurological mutant of genetic origin. Neurology. 1972;22:743–754. doi: 10.1212/wnl.22.7.743. [DOI] [PubMed] [Google Scholar]
- Zhang SC, Fedoroff S. Cellular localization of stem cell factor and c-kit receptor in the mouse nervous system. J Neurosci Res. 1997;47:1–15. [PubMed] [Google Scholar]
- Zhang X, Zhang H, Oberdick J. Conservation of the developmentally regulated dendritic localization of a Purkinje cell-specific mRNA that encodes a G-protein modulator: comparison of rodent and human Pcp2(L7) gene structure and expression. Molecular Brain Research. 2002;105:1–10. doi: 10.1016/s0169-328x(02)00379-0. [DOI] [PubMed] [Google Scholar]
- Zhao Q, Kho A, Kenney AM, Yuk Di DI, Kohane I, Rowitch DH. Identification of genes expressed with temporal-spatial restriction to developing cerebellar neuron precursors by a functional genomic approach. Proc Natl Acad Sci U S A. 2002;99:5704–9. doi: 10.1073/pnas.082092399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao R, Lawler AM, Lee SJ. Characterization of GDF-10 Expression Patterns and Null Mice. Developmental Biology. 1999;212:68–79. doi: 10.1006/dbio.1999.9326. [DOI] [PubMed] [Google Scholar]
- Zupanc GK. Up-regulation of somatostatin after lesions in the cerebellum of the teleost fish Apteronotus leptorhynchus. Neurosci Lett. 1999;268:135–8. doi: 10.1016/s0304-3940(99)00299-2. [DOI] [PubMed] [Google Scholar]
- Zupanc GK, Clint SC. Radial glia-mediated up-regulation of somatostatin in the regenerating adult fish brain. Neurosci Lett. 2001;309:149–52. doi: 10.1016/s0304-3940(01)02061-4. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figure 1: Microinjection controls
Supplemental Table 1: Average difference values for each hybridization
Supplemental Table 2: Computed F-statistics by 2-factor ANOVA, with corresponding p-values and t-tests for effects of genotype, developmental time, and gender.
Supplemental Table 3: Genotype-dependent expression differences predicted by various methods.
Supplemental Table 4: Primer sequences for ChIP assays and QPCR.


