Abstract
Background
The role of the RNA polymerase sigma factor RpoN in regulation of gene expression in Geobacter sulfurreducens was investigated to better understand transcriptional regulatory networks as part of an effort to develop regulatory modules for genome-scale in silico models, which can predict the physiological responses of Geobacter species during groundwater bioremediation or electricity production.
Results
An rpoN deletion mutant could not be obtained under all conditions tested. In order to investigate the regulon of the G. sulfurreducens RpoN, an RpoN over-expression strain was made in which an extra copy of the rpoN gene was under the control of a taclac promoter. Combining both the microarray transcriptome analysis and the computational prediction revealed that the G. sulfurreducens RpoN controls genes involved in a wide range of cellular functions. Most importantly, RpoN controls the expression of the dcuB gene encoding the fumarate/succinate exchanger, which is essential for cell growth with fumarate as the terminal electron acceptor in G. sulfurreducens. RpoN also controls genes, which encode enzymes for both pathways of ammonia assimilation that is predicted to be essential under all growth conditions in G. sulfurreducens. Other genes that were identified as part of the RpoN regulon using either the computational prediction or the microarray transcriptome analysis included genes involved in flagella biosynthesis, pili biosynthesis and genes involved in central metabolism enzymes and cytochromes involved in extracellular electron transfer to Fe(III), which are known to be important for growth in subsurface environment or electricity production in microbial fuel cells. The consensus sequence for the predicted RpoN-regulated promoter elements is TTGGCACGGTTTTTGCT.
Conclusion
The G. sulfurreducens RpoN is an essential sigma factor and a global regulator involved in a complex transcriptional network controlling a variety of cellular processes.
Background
RpoN (σ54 or sigma 54) is a subunit of the RNA polymerase and plays a critical role in the regulation of gene expression by recognizing specific promoter elements and initiating transcription. RpoN-dependent promoters do not have conserved -35 and -10 elements typically found in the promoters recognized by sigma factors in the σ70 family. Instead, a GG dinucleotide around position -24 and a GC dinucleotide around position -12 with respect to the transcription initiation site are highly conserved among RpoN-dependent promoters [1]. The consensus of 186 RpoN-dependent promoter elements from 47 bacterial species was reported to be mrNrYTGGCACG...4bp...TTGCWNNw [2]. In contrast to σ70 family sigma factors, RpoN is able to bind to a promoter without the core RNA polymerase (RNAP) [1]. RNAP containing RpoN (RNAP/RpoN) can form a stable closed complex with the promoter. In addition, RNAP/RpoN requires a transcription factor, the enhancer-binding protein (EBP), for initiation of transcription. Some of the EBP family members are response regulators in two-component regulatory systems [3]. RpoN was first identified in Escherichia coli and was reported to regulate the transcription initiation of nitrogen assimilation genes [4]. Since then, RpoN homologs have been identified in bacteria from different phylogenetic origins and are involved in regulation of genes related to a diverse functional categories [5], including genes for pili and flagella biosynthesis and quorum sensing in Pseudomonas aeruginosa [6-8], C4-dicarboxylate transport in Mesorhizobium ciceri [9] and carbon metabolism in Gram positive bacteria, Bacillus subtilis and Listeria monocytogenes [10-13].
Geobacter species are important agents in the bioremediation of subsurface environments contaminated with organic or metal contaminants [14]. They also appear to be the primary contributors to current production in microbial fuel cells harvesting electricity from the environment [15]. Some physiological responses of Geobacter species can be predicted with constraint-based genome-scale metabolic models that determine the optimal flux of metabolites for a given environmental condition [16]. The ability to predictively model the physiological responses of environmentally relevant microorganisms to a wide diversity of environmental conditions is a major goal of environmental biotechnology [14,17]. However, the current version of these models lacks regulatory modules that could increase their predictive value. Progress has been made using a combination of bioinformatic tools and molecular biological methods to identify regulatory components, such as operon structures, promoter elements, and transcription factors and their binding sites, as well as global transcriptomic and proteomic expression patterns that provide the basis for building regulatory modules in G. sulfurreducens [18-21]. Sigma factors are key to constructing bacterial transcriptional regulatory networks. In G. sulfurreducens, homologs of RpoD, RpoS, RpoH, RpoE and FliA of the σ70 family have been identified and physiological roles of G. sulfurreducens RpoS [22,23], RpoH [24] and RpoE (G. Reguera et al, unpublished) have been elucidated.
An ortholog of the rpoN gene (GSU1887) is present in the G. sulfurreducens genome [19]. It encodes a protein, which shares a considerable degree of similarity to RpoN sigma factors from other bacteria. We report here that an rpoN deletion mutant could not be isolated under conditions tested in this study. In order to identify the components of the RpoN regulon in G. sulfurreducens, genome-wide microarray transcriptional profiling of an RpoN over-expression strain and genome-wide prediction of RpoN-regulated promoters were employed. We discuss below our findings that RpoN-dependent genes carry out important functions that may contribute to the reasons why no viable rpoN deletion mutants could be obtained.
Results
The Geobacter sulfurreducens rpoN gene cluster
An rpoN ortholog (GSU1887), which encodes the RpoN sigma factor, is present in the G. sulfurreducens genome. Phylogenetic analysis showed that G. sulfurreducens RpoN is very similar to other experimentally characterized RpoN sigma factors, such as the Escherichia coli RpoN [19]. The G. sulfurreducens RpoN polypeptide displays characteristic structures of the members of the RpoN family, including an N-terminal glutamine-rich region (the first 50 amino acids), a C-terminal X-link, a helix-turn-helix (HTH) DNA-binding motif, and an RpoN box (ARRTVTKYRE) [25].
Analysis of the chromosomal region surrounding the G. sulfurreducens rpoN gene revealed that its downstream genes encode a homolog of a ribosomal subunit interface protein (GSU1886) whose N-terminal domain is homologous to the RpoN modulation protein found in Klebsiella pneumoniae [26] and a homolog of Hpr (Ser) kinase/phosphorylase (GSU1885). The upstream genes encode an ATP binding protein (GSU1888) and two conserved proteins (GSU1889 and GSU1890) with unknown function (Figure 1a).
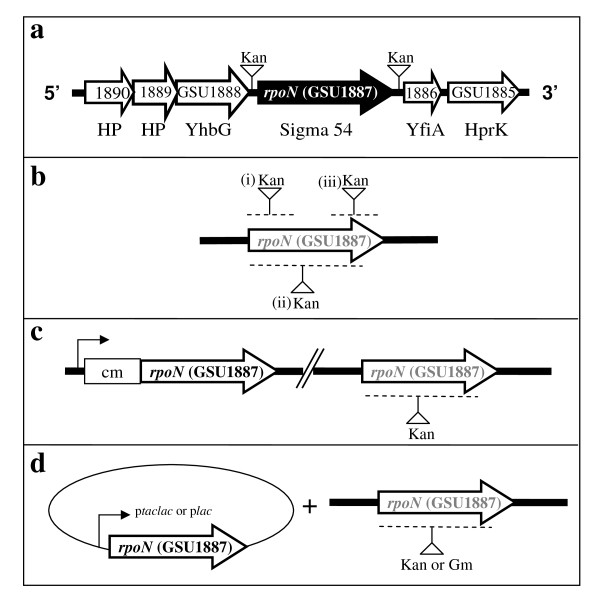
Figure 1.
The rpoN gene cluster and the mutation schemes. (a) Genes surrounding rpoN are shown as open arrows. HP: conserved hypothetical protein with unknown function; YhbG: ABC transporter, ATP binding protein; YfiA: ribosomal subunit interface-associated sigma-54 modulation protein; HprK: Hpr(Ser) kinase/phosphorylase. Insertion of a kanamycin resistance cassette upstream or downstream of the intergenic region of the rpoN gene resulted in viable mutants (a). (b) Scheme showing attempts of construction of deletion of (i) the 5'-end, (ii) the whole, or (iii) the 3'-end of the rpoN coding region. (c) An extra copy of the rpoN gene was inserted on the chromosome and was under the control of the chloramphenicol resistance cassette promoter. (d) An extra copy of the rpoN gene was introduced in trans under the control of a lac promoter (constitutively expressed) or a taclac promoter (IPTG-inducible). The position of insertion of the antibiotic resistance cassette (kanamycin, Kan or gentamycin, Gm) is indicated with an inverted triangle and a vertical bar. The regions which were attempted to replace with the antibiotic resistance cassette insertion are indicated by dashed line.
In order to understand the physiological role of RpoN and of genes it controls in G. sulfurreducens, construction of a deletion mutant of the rpoN gene was attempted. However, an rpoN mutant could not be isolated under different growth conditions using media with different electron acceptors (fumarate or Fe(III) citrate) and amendments (glutamine or glutamate) (Table 1). Different mutagenesis strategies were attempted, including deletion of the whole coding region of the rpoN gene, only the 5'-end of the rpoN coding region, or only the 3'-end of the rpoN coding region (Figure 1b). No viable mutants were derived from above attempts. We also constructed an rpoN diploid strain (DLCN43) in which an extra copy of the rpoN gene was integrated into the chromosome and controlled by the promoter from a chloramphenicol resistance cassette (Figure 1c). Attempts of deleting and replacing the original rpoN gene from DLCN43 also failed. Furthermore, an extra copy of the rpoN gene under the control of a constitutively active lac promoter, or under the control of an IPTG (isopropyl-β-D-thiogalactoside)-inducible taclac promoter was introduced to the wild type strain in trans (Figure 1d). Attempts of deleting the chromosomal rpoN gene yielded viable isolates. However, further analysis of these isolates indicated that none of them had deletion of the chromosomal rpoN gene. In contrast, insertion of a kanamycin resistance cassette at the intergenic regions immediately upstream or downstream of the rpoN gene yielded viable mutants, suggesting that the inability to isolate an rpoN-deletion mutant was not due to the polar effects of the kanamycin resistance cassette. The conditions and mutagenesis methods used in this work for our attempts to isolate a null rpoN mutant are summarized in Table 1, and mutation scheme is illustrated in Figure 1.
Table 1.
List of mutagenesis and selection media for attempts to generate a null rpoN mutant.
| Mutagenesis | Selection medium (electron donor/acceptor) |
|---|---|
| 1. Deletion and replacement of the rpoN gene by double-crossover (Figure 1b) | Acetate/fumarate |
| Acetate/Fe(III) citrate | |
| Acetate/Fe(III) citrate amended with glutamine | |
| Acetate/Fe(III) citrate amended with glutamate | |
| Acetate/Fe(III) citrate amended with both glutamine and glutamate | |
| 2. Deletion and replacement of the 5'-end of the rpoN gene by double-crossover (Figure 1b) | Acetate/fumarate |
| Acetate/Fe(III) citrate | |
| 3. Deletion and replacement of the 3'-end of the rpoN gene by double-crossover (Figure 1b) | Acetate/fumarate |
| Acetate/Fe(III) citrate | |
| 4. Deletion and replacement of the upstream intergenic region of the rpoN gene by double-crossover (Figure 1a) | Acetate/fumarate |
| 5. Deletion and replacement of the downstream intergenic region of the rpoN gene by double-crossover (Figure 1a) | Acetate/fumarate |
| 6. Integration of a linear DNA fragment on the chromosome, providing another copy of the rpoN gene on another location of the chromosome (Figure 1c) | Acetate/fumarate |
| Acetate/Fe(III) citrate | |
| 7. Integration of a linear DNA fragment on the chromosome, providing another copy of the rpoN gene in trans, which is either constitutively expressed by a lac promoter, or is IPTG inducible (Figure 1d) | Acetate/fumarate |
| Acetate/Fe(III) citrate | |
| Acetate/Fe(III) citrate amended with both glutamine and glutamate | |
RpoN expression patterns and over-expression of RpoN
In order to understand RpoN expression under different growth conditions, Western blot analysis was performed using anti-sera against RpoN with cell extracts prepared from cultures grown with a variety of electron donors/acceptors. RpoN levels were similar in cultures grown in the presence of ammonium or when nitrogen fixation was required in ammonia-free medium, and RpoN was constitutively expressed under all other conditions tested (Figure 2a).
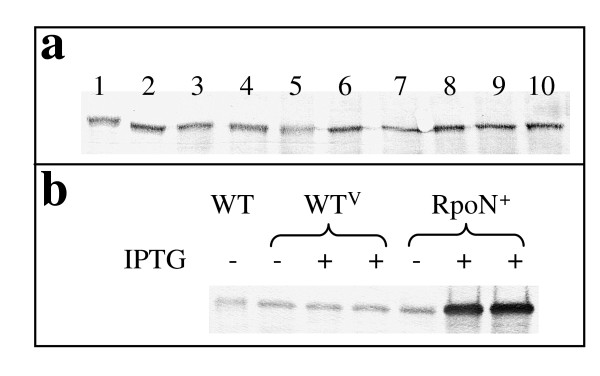
Figure 2.
RpoN expression. (a) RpoN expression under different growth conditions. 1: NBAF; 2: NBH2F; 3: NBLF; 4: ammonium-free NBAF; 5: FWAFC; 6: FWH2FC; 7: FWLFC; 8: FWAF; 9: FWH2F; 10: FWLF. Media abbreviations were detailed in Methods. (b) RpoN over-expression. Total protein (5 μg) was separated by 10% SDS-PAGE and analyzed by Western blot analysis with the RpoN-specific antiserum. Two biological samples were shown for IPTG-induced WTV and RpoN+ strains. IPTG was added at final concentration 1 mM.
Because an rpoN deletion mutant could not be obtained, a strain in which the rpoN gene was over-expressed under the control of the IPTG-inducible taclac promoter was generated to gain insights into the function of RpoN. The strain capable of over-expressing RpoN (DL1/pCDrpoN) and the control strain harboring the empty vector pCD341 (DL1/pCD341) were designated RpoN+ and WTV, respectively, for simplification. The over-expression of the RpoN protein in RpoN+ after induction was confirmed using Western blot analysis (Figure 2b). The abundance of the RpoN protein in the RpoN+ strain was 5.4 times more than that of the wild type with IPTG induction, but it stayed at a similar level as that of the wild type in the absence of IPTG. The WTV strain, with or without the addition of IPTG, had levels of RpoN similar to those in the wild type strain.
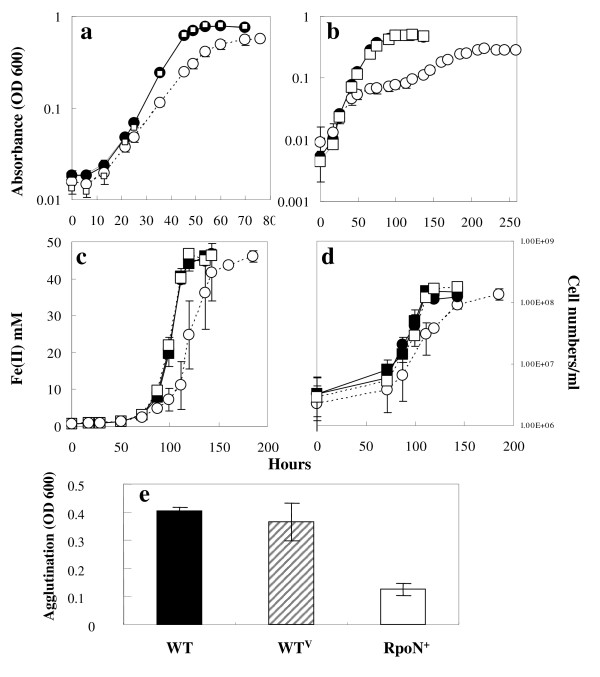
When induced with IPTG, the RpoN+ strain grew slower and had a longer lag phase than the WTV strain with either fumarate or Fe(III) citrate as the electron acceptor and acetate as the electron donor (Figure 3a, c &3d). The effect of over-expressing RpoN on growth was more pronounced when cells were grown in media lacking ammonia (Figure 3b). The doubling times for the WTV and the RpoN+ strains were 7.5 and 9.7 hours, respectively, in the presence of IPTG in the NBAF medium (Figure 3a). Under nitrogen fixation conditions, the doubling times for the WTV and the RpoN+ strains were 12.5 and 51.3 hours, respectively, in the presence of IPTG in the ammonium-free NBAF medium (Figure 3b). For FWAFC media, the doubling times were 9.6 and 14.9 hours with IPTG for WTV and RpoN+, respectively (Figure 3d). The RpoN+ strain grew similarly with the WTV strain in the absence of IPTG in all media. These results suggest that over-expression of RpoN inhibited growth under various conditions.
Figure 3.
Characterization of the RpoN over-expression strain. Cell growth with fumarate as an electron acceptor was monitored by absorbance at 600 nm (a)(b). (a) acetate as the electron donor and fumarate as the electron acceptor (NBAF medium); (b): ammonia-free NBAF. Growth with Fe(III) as an electron acceptor was monitored by Fe(II) production (c) as well as cell numbers (d). Filled square: the WTV strain without IPTG; Empty square: the WTV strain with IPTG. Filled circle: the RpoN+ strain without IPTG; empty circle: the RpoN+ strain with IPTG. (a)-(d): Data are means ± standard deviations of triplicates. The production of pili was measured by agglutination assays (e). Data are means ± standard deviation of triplicates from two independent experiments (e).
The RpoN regulon identified by genome-scale transcriptome analysis and prediction of RpoN-dependent promoters
To elucidate the function of RpoN in Geobacter species, the transcriptome of the RpoN+ and the WTV strains was compared in order to identify those genes whose transcription is regulated by RpoN. Due to different growth rates between the two strains in the NBAF medium (Figure 3), total RNA was isolated independently from three sets of WTV and RpoN+ cultures during the exponential growth phase, about OD600 = 0.3–0.35 and 0.2–0.25 for the WTV and RpoN+ strains, respectively.
With a 1.5 fold-change cutoff and a relatively strong cutoff for false discovery rate (FDR) of p < 0.0005, the RpoN+ strain was found to have 138 genes with increased transcript levels and 59 genes with decreased transcript levels when compared to the WTV strain. The fold change of the transcription level of the rpoN gene itself was 6.61 higher in the RpoN+ strain than in the WTV strain, which is in agreement with protein over-expression results obtained by the Western blot analysis. A complete list of the differentially expressed genes and their expression ratio is provided as additional files (see Additional files 1 &2). The 196 genes with significant changes in expression level in the RpoN+ strain were assigned to 16 functional groups. Most of them encoded hypothetical proteins or proteins with unknown function (63 genes). The top three categories of genes with annotated functions consisted of genes associated with protein synthesis (33), energy metabolism (21 genes), and transport and binding (12).
In addition to observing the RpoN regulon by using gene expression microarrays, we employed computational analysis using the PromScan software [27] to identify RpoN recognition sequence elements within the G. sulfurreducens genome. The search identified 798 putative RpoN recognition sequence elements with scores ≥ 80 (data not shown). Among these sequence elements, 467 elements were located upstream of protein-coding genes, in the same strand orientation as their potential target genes. Of these possible promoter elements that could potentially regulate transcription of protein-coding genes, 110 were located in noncoding regions. We considered these 110 sequence elements to be the most likely RpoN-regulated promoters. Their sequences and genome locations are provided in Additional file 3, while their predicted target genes and operons are provided in Additional file 4. The consensus sequence of these 110 predicted G. sulfurreducens RpoN-regulated promoter elements listed in Additional file 3 was TTGGCACGGTTTTTGCT, where the -24 GG and the -12 GC dinucleotides are in bold. The highest scoring RpoN-regulated promoter was located upstream of the flagella biosynthesis operon containing the fliA gene encoding the RNA polymerase sigma factor FliA (σ28) (see Additional file 4).
Genome locations of the 110 RpoN-dependent sequence elements identified using the PromScan analysis were cross-examined with the list of genes identified by the transcriptome analysis with a fold change cutoff of 1.25. The results of this comparison are listed in Additional file 5, and selected operons encoding genes related to nitrogen assimilation, appendages and solute transport are listed in Table 2. These combined data showed that RpoN regulatory elements can be found in not only up-regulated genes, but also down-regulated genes. The increased transcript levels observed for some genes could result from increased RpoN availability for transcription initiation that is the limiting factor at a normal rpoN transcript level. The decreased transcript levels for some genes could be due to the fact that RpoN alone can bind to the -24/-12 elements without the core RNAP, and therefore the promoter regions may not be accessible by other sigma factors, or less core RNAP may be available in the excess of RpoN [1,28].
Table 2.
Genes containing RpoN-dependent promoters identified by the PromScan analysis and the transcriptome analysis.
| Operons§ | Genes | Annotations | Fold changes* | PromScan score |
|---|---|---|---|---|
| Amino acid biosynthesis and Nitrogen assimilation | ||||
| Glutamine synthase (GS) | GSU1835 (glnA)† | glutamine synthetase (GS) | +1.39 | 80 |
| GSU1836 (glnB)† | nitrogen regulatory protein PII | +1.77 | ||
| Glutamate synthase (GOGAT) | GSU1235 | hypothetical protein | N. D. | 88 |
| GSU1236 | hypothetical protein | N. D. | ||
| GSU1237 | pyridine nucleotide-disulphide oxidoreductase family protein | -1.44 | ||
| GSU1238 | iron-sulfur cluster-binding protein | N. D. | ||
| GSU1239 (gltB)† | glutamate synthase-related protein (GOGAT) | N. D. | ||
| Nitrogen assimilation | GSU2802 | NAD(+) – dinitrogen-reductase ADP-D-ribosyltransferase | N. D. | 86 |
| GSU2803 | dinitrogenase iron-molybdenum cofactor family protein | -1.21 | ||
| GSU2804 | ferredoxin family protein | -1.24 | ||
| GSU2805 (nifX) | nitrogenase molybdenum-iron cofactor biosynthesis protein NifX | -1.27 | ||
| GSU2806 (nifEN)† | nitrogenase molybdenum-iron cofactor biosynthesis protein NifEN | -1.34 | ||
| Appendages and Motility | ||||
| Flagella biogenesis | GSU3050 (flgA) | flagella basal body P-ring formation protein FlgA | -1.34 | 93 |
| GSU3051 (flgG) | flagellar basal-body rod protein FlgG | N. D. | ||
| GSU3052 (flgG) | flagellar basal-body rod protein FlgG | -1.19 | ||
| GSU3053 (fliA)† | RNA polymerase sigma factor for flagellar Operon/gene/gene | -1.50 | ||
| GSU3054 | ParA family protein | -1.21 | ||
| GSU3055 (flhF) | flagellar biosynthetic protein FlhF | -1.34 | ||
| GSU3056 (flhA) | flagellar biosynthetic protein FlhA | -1.42 | ||
| Flagella basal body | GSU0407 (flgB) | Flagellar basal-body rod protein FlgB | -1.38 | 92 |
| GSU0408 (flgC) | Flagellar basal-body rod protein FlgC | -1.31 | ||
| Flagella biogenesis | GSU0420 (fliL)† | flagellar protein FliL | -1.47 | 86 |
| GSU0421 (fliM) | flagellar motor switch protein FliM | -1.35 | ||
| GSU0422 (fliN) | flagellar motor switch protein FliN | -1.93 | ||
| GSU0423 (fliP) | flagellar biosynthetic protein FliP | N. D. | ||
| GSU0424 (fliQ) | flagellar biosynthetic protein FliQ | -1.32 | ||
| GSU0425 (fliR) | flagellar biosynthesis protein FliR | -1.72 | ||
| GSU0426 (flhB) | flagellar biosynthetic protein FlhB | -1.11 | ||
| Flagella biogenesis | GSU3040 | hypothetical protein | N. D. | 86 |
| GSU3041 | carbon storage regulator | -1.04 | ||
| GSU3042 (flgL) | flagellar hook-associated protein FlgL | -1.05 | ||
| GSU3043 (flgK) | flagellar hook-associated protein FlgK | -1.05 | ||
| GSU3044 | hypothetical protein | -1.21 | ||
| GSU3045 (flgM) | negative regulator of flagellin synthesis FlgM | -1.13 | ||
| GSU3046 (flgJ)† | flagellar protein FlgJ-like protein | -1.41 | ||
| Metabolisms | ||||
| Formate dehydrogenase | GSU0777(fdnG)† | formate dehydrogenase, major subunit, selenocysteine-containing | N. D. | 90 |
| GSU0778 | formate dehydrogenase, iron-sulfur subunit | N. D. | ||
| GSU0779 | formate dehydrogenase, b-type cytochrome subunit | -1.19 | ||
| GSU0780 | formate dehydrogenase accessory protein FdhD | N. D. | ||
| GSU0781 | twin-arginine translocation protein, TatA/E family | -1.41 | ||
| Solute transporter | ||||
| Fumarate/succinate exchanger | GSU2750 | hypothetical protein | +1.30 | 83 |
| GSU2751 (dcuB)† | C4-dicarboxylate transporter (DcuB) | +1.63 | ||
§Operon predictions in G. sulfurreducens are described in [18].
†Genes were discussed in the text and/or RpoN regulatory elements have been confirmed in their promoter regions by primer extension analyses
*Not all genes listed in the table matched the criteria for ≥ 1.25 fold-change cutoffs by the gene expression microarray analysis as described in Methods, but these genes are listed because one or more genes from the same operon matched the cutoff criteria.
N.D.: Not detected.
Both microarray transcriptome analysis and computational prediction of RpoN promoter elements concluded that the dcuB gene, which encodes the fumarate/succinate exchanger (C4-dicarboxylate transporter), is RpoN-dependent. Both analyses also indicated that genes encoding components for nitrogen assimilation, such as glutamine synthetase (GS) (GSU1835), are RpoN-dependent (Table 2). Furthermore, our computational analysis identified an RpoN promoter upstream of the glutamate synthetase (GSU1239, GOGAT) operon. The microarray analysis indicated that the gene encoding glutamate dehydrogenase (GDH) (GSU1305) is RpoN-dependent.
In silico modeling analysis was utilized to understand the role of GDH, GS and GOGAT enzymes in G. sulfurreducens metabolism. The growth of G. sulfurreducens was simulated under different growth conditions, varying electron donors, electron acceptors, and nitrogen sources. In silico modeling analysis suggested that missing both GS and GOGAT enzymes or GDH, GS and GOGAT is lethal under all growth conditions.
Other genes identified by both the transcriptome analysis and the computational promoter prediction include those encoding flagella biosynthesis, formate dehydrogenase, alcohol dehydrogenase, and acetyl-CoA carboxylase (Table 2 and Additional file 5).
An RpoN-dependent regulatory element, located upstream of the pilA gene encoding pilin, the building block for nanowires [29], was identified by the PromScan software in this study and by 5'-RACE analysis in an earlier study [30]. Therefore, the effect of RpoN-overexpression was tested on the pili formation via a cell agglutination assay at 25°C [31]. In the presence of IPTG, the RpoN+ strain displayed less agglutination at 25°C than WTV and WT (Figure 3e), which strongly suggests that RpoN is involved in pili biogenesis.
Genes that were differentially expressed in the RpoN+ strain but for which no RpoN-dependent regulatory elements could be found include those encoding components of stress response/molecular chaperones, central metabolism, extracellular electron transfer, and genes encoding regulatory proteins (see Additional files 1 &2), suggesting their possible indirect regulation by RpoN or a possibility that their RpoN promoters may be too divergent from promoters in other bacteria to be detected using computational approaches. Alternatively, the physiological states caused by the overexpression of RpoN in the RpoN+ strain, which were different from those in the WTV strain, such as slower growth, might affect gene regulation.
Evaluation of predicted RpoN-dependent promoters via primer extension analysis
In order to further validate the results of the microarray transcriptome analysis and of the computational promoter prediction and to examine promoters regulating differentially expressed genes, primer extension assays were carried out on 12 selected operons or singleton genes, including seven genes with increased expression and five genes with decreased expression in the microarray analysis (Table 3). These selections included 1) genes involved in cellular functions that were reported in other bacteria to be RpoN-dependent, such as flagella biosynthesis and nitrogen assimilation; or 2) genes involved in physiological functions that were not previously reported to be RpoN-dependent in other bacteria, but were predicted to have conserved RpoN-dependent -24/-12 promoter elements in their regulatory regions, or 3) genes encoding enzymes that are essential for cell growth. The trend of changes in transcript levels observed in the primer extension assays was similar to that reported in the microarray analysis (Figure 4, Table 3 and Additional file 6), with an exception of GSU1836 (glnB), which encodes a PII nitrogen regulatory protein.
Table 3.
Summary of 12 genes whose 5' ends of mRNA were analyzed by primer extension assays.
| Gene # | ID | Promoter elements validated | In agreement with microarray data | In agreement with PromScan |
|---|---|---|---|---|
| Up-regulated according to microarray analysis | ||||
| GSU0364 | ppcB | RpoD | Yes | Yes |
| GSU1836 | glnB | RpoN | No | Yes |
| GSU2005 | ABC transporter | RpoD | Yes | No |
| GSU2302 | Trehalose phosphatase | RpoD | Yes | Yes |
| GSU2490 | Oxalate-formate antiporter | RpoD | Yes | No |
| GSU2751 | dcuB | RpoN | Yes | Yes |
| GSU3206 | dksA | RpoD | Yes | No |
| Down-regulated according to microarray analysis | ||||
| GSU0420 | fliL | RpoN | Yes | Yes |
| GSU0777 | fdnG | RpoN | Yes | Yes |
| GSU0939 | PII | RpoD | Yes | No |
| GSU2806 | nifEN | RpoN | Yes | Yes |
| GSU3046 | flgJ | RpoN | Yes | Yes |
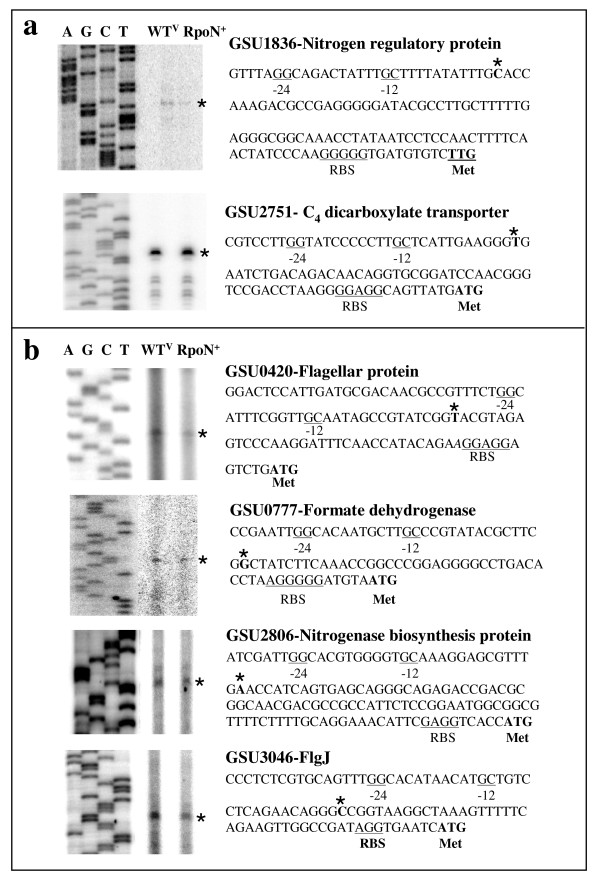
Figure 4.
RpoN-dependent gene expression. Representative genes, (a) GSU1836 and GSU2751 (up-regulated in the RpoN+ strain), and (b) GSU0420, GSU0777, GSU2806 and GSU3046 (down-regulated in the RpoN+ strain) identified by the microarray analysis were further analyzed by primer extension assays. The results of the primer extension assays and their promoter regions are shown. The 5' ends of mRNA are indicated by asterisks. The putative -24/-12 elements and RBS are underlined. Translation start codons are shown in bold and are indicated by Met.
Eight genes out of the 12 selected genes had promoter elements identified by the primer extension assays to match those identified by the computational predictions, and six of these eight genes were found to be regulated by promoter sequences highly similar to other bacterial RpoN promoter elements (Table 3). These six RpoN-regulated genes were GSU0420 (fliL), GSU1836 (glnB), GSU2806 (nifEN), GSU3046 (flgJ), GSU2751 (dcuB) and GSU0777 (fdnG) (Figure 4 &5 and Table 3). Alignment of these six G. sulfurreducens RpoN promoter elements and of the promoter element upstream of the pilA gene identified in the previous study [30] was shown (Figure 5).
Figure 5.
G. sulfurreducens RpoN-dependent promoter elements. (a) Alignment of G. sulfurreducens RpoN-dependent promoters identified by primer extension assays in this study. Conserved nucleotides which are the same to the consensus sequences from (b) are labeled in red. (b) Sequence logo of 110 G. sulfurreducens RpoN-regulated promoters predicted by PromScan in non-coding regions upstream of target protein-coding genes.
While conserved RpoN-regulated promoters were predicted for other four genes/operons (GSU0939, GSU2005, GSU2490 and GSU3206) by the computational tools (see Additional file 4), no -24/-12 RpoN-dependent promoter elements could be identified by primer extension assays for these genes/operons (see Additional file 6). It is possible that the predicted RpoN-regulated promoter elements for these four genes/operons are activated only under certain conditions, such as nitrogen-fixing conditions, which were different from the conditions for the primer extension assays.
In conclusion, the primer extension analyses showed that RpoN-regulated promoters are located upstream of genes for a fumarate/succinate exchanger (GSU2751, dcuB), glutamine synthetase (GSU1836-1835 operon, glnB-glnA), flagella biosynthesis proteins (GSU0420–0426, and GSU3040–3046), nitrogen assimilation enzymes (GSU2802–2806), and a formate dehydrogenase (GSU0777–0781). In addition, an RpoN-regulated promoter was also identified for the pilA gene (GSU1496) by 5'-RACE analysis [30].
Discussion
The G. sulfurreducens RpoN regulon
The G. sulfurreducens RpoN regulon was identified using the microarray transcriptome analysis combined with the computational analysis. Both methods indicated that RpoN plays an important role in influencing the expression of a number of genes that are important for growth in subsurface environment and in microbial fuel cells. Most evidently, both methods demonstrated that the G. sulfurreducens RpoN controls the expression of a fumarate/succinate exchanger (DcuB), which is essential under fumarate respiration [32]. Thus, no viable rpoN deletion mutant was isolated when fumarate was the terminal electron acceptor. This is the first report to our knowledge identifying a -24/-12 RpoN-dependent promoter element in the dcuB regulatory region. In other bacteria, such as E. coli, the dcuB gene is transcribed by RNA polymerase in complex with the RpoD sigma factor, and its expression is under a hierarchical control involving FNR, CRP, and a two-component regulatory system [33].
Glutamate and glutamine, the essential biomass components and major intracellular nitrogen donors, are the products of ammonia assimilation and are synthesized in bacteria via two pathways [34]. The G. sulfurreducens RpoN controls the expression of enzymes involved in both pathways for ammonia assimilation, namely, 1) the GDH-dependent pathway, in which glutamate is synthesized by reductive amination of 2-ketoglutarate and 2) the GS-GOGAT pathway, in which GS converts glutamate and ammonia to glutamine and GOGAT transfers the amide group from glutamine to 2-ketoglutarate. Therefore, deleting the rpoN gene would result in deficiencies in ammonia assimilation and thus, cell death. In E. coli, the GS-GOGAT pathway is used under energy-rich and nitrogen-limiting conditions and the expression of both enzymes is under the control of RpoN, whereas the GDH pathway is employed under energy-limiting and excess ammonium conditions, and the transcription of GDH is controlled by RNAP/RpoD [34-36]. In fact, the expression of other known bacterial GDHs is controlled by RNAP/RpoD, including those in Neisseria meningitides, Klebsiella aerogenes, Psychrobacter sp, Streptococcus pneumoniae and Pseudomonas aeruginosa [37-42].
Acetate is a primary electron donor for Geobacter species in soils and sediments [43], especially during groundwater uranium bioremediation [44]. The microarray study suggests that RpoN positively regulates the expression of genes essential for acetate oxidation coupled to metal reduction such as fumarase (GSU0994) and acetyl-CoA transferases (GSU0174 and GSU0490) [45]. The inability to isolate an rpoN null mutant could also be due to the poor expression of these essential TCA cycle enzymes in the absence of RpoN. The ability to recover such a mutant with Fe(III) as the electron acceptor would also be limited due to low expression of the cytochromes essential for Fe(III) reduction, such as the omcB gene [46], which was up-regulated in the RpoN+ strain (see Additional file 1).
Flagella are considered to play an important role in Fe(III) oxide reduction by Geobacter species [47]. The regulatory regions from both operons containing GSU0420 and GSU3046 encoding flagella proteins contain sequences homologous to RpoN-dependent promoter sequences (Figure 4 and 5). As presented in Table 2 and Additional file 4, the operon containing the fliA gene (σ28, GSU3053) along with other flagella biosynthesis genes (flg-1, flg-2, flh-A, and others) was predicted to have the highest scoring RpoN-regulated promoter, indicating that this promoter was highly conserved and that σ28 gene expression in G. sulfurreducens is likely regulated by RpoN. Interestingly, these results are similar to Campylobacter spp and Vibrio spp, in which RpoN regulates the σ28 gene expression (see reviews and references within [48-50]). In other bacterial species, e.g. Salmonella enterica serovar Typhimurium, the regulation of fliA is controlled by the FlhCD transcriptional regulator [50]. FlhCD is absent from the G. sulfurreducens genome, which also appears to lack FlhCD binding sites [19]. It is interesting to note that C. jejuni and V. cholerae use pili and flagella to achieve virulence [50], whereas in Geobacter species, flagella and pili are implicated in extracellular Fe(III) reduction [29,47] and higher power production in microbial fuel cells [51].
Unique features of the G. sulfurreducens RpoN sigma factor
In most bacteria that have been studied, the rpoN gene deletion resulted in viable mutants or mutants requiring certain nutrient addendum [8,52,53]. However, despite the high amino acid sequence similarity of the G. sulfurreducens RpoN to other bacterial homologs, an rpoN null mutant was not obtained after multiple attempts. The only other case in which the rpoN gene is essential was reported in Myxococcus xanthus, another delta-proteobacterium, for reasons that have yet to be elucidated [54].
This work also suggested that the expression of RpoN in G. sulfurreducens is under a tight control in a complex manner for several reasons: (1) over-expression of RpoN inhibited growth under various growth conditions, (2) over-expression of RpoN induced up-regulation of genes involved in stress responses (see Additional file 1), and (3) our inability to isolate a viable rpoN deletion mutant even in the presence of another copy of the rpoN gene, in trans or in cis, which was under the control of artificial promoters such as the lac promoter or the promoter from the chloramphenicol resistance gene. Furthermore, the G. sulfurreducens genome contains 28 genes encoding transcription factors from the EBP family, whose members are required for transcription initiation directed by RNAP/RpoN. This number of the EBPs is much higher than that found in most bacteria. For instance, E. coli, which has a larger genome (4.6 Mbp) than G. sulfurreducens (3.8 Mbp), has only 12 EBPs [4,55]. Approximately half of the EBPs in G. sulfurreducens belong to two-component regulatory systems. This further suggests that cellular responses to various environmental conditions are directed by gene expression regulated by RpoN. It has been reported for other bacteria that EBP-encoding genes are often located adjacent or close to their target promoters ([3] and references within). This was also found to be the case for some of the EBPs in G. sulfurreducens. It has been shown that PilR, a member of the EBP family, regulates the pilA gene, which has an RpoN-dependent promoter and is located immediately downstream of the pilR gene [30]. Genes encoding EBPs are also located upstream of the dcuB and fdnG genes, which were shown to contain an RpoN-dependent promoter (Figure 4). In addition, a gene encoding an EBP is located upstream of GSU3364, which was predicted to contain an RpoN-dependent promoter (see Additional file 4). However, it appears that G. sulfurreducens EBPs are not always located adjacent or close to their target promoters for the cases of fliL, glnB (GSU1836), nifEN, and flgJ, which were shown to contain an RpoN-dependent promoter in their regulatory regions (Figure 4).
Data analysis-combining computational prediction with microarray analysis
Due to the absence of an RpoN deletion mutant, our analyses were restricted to the use of an RpoN over-expressing strain. Activation of RpoN-dependent transcription requires the presence of EBPs activated by modification such as phosphorylation, and therefore, an increase in the amount of RpoN alone would not directly lead to increasing expression of every RpoN-regulated gene. This is exactly what we have observed with our microarray analysis. With a relatively strong cutoff for false discovery rate of p < 0.0005, the majority (~85%) of differentially expressed genes had fold changes between 1.50 and 2. When the results from the computational prediction are compared with the results from the transcriptome analysis, the use of an arbitrary cutoff level may not be able to detect all RpoN-regulated genes in the RpoN over-expressing strain. For instance, from the transcriptome analysis, the fold change of the nifEN gene encoding a subunit of nitrogenase complex was -1.34 (Table 2), while an RpoN-dependent promoter for this gene was identified in G. sulfurreducens (Figure 4) and in other species [34]. Therefore, it is likely that the fold change in expression of the nifEN gene and other genes did not reach the 1.5 threshold due to complexity of gene regulation in the RpoN over-expressing strain. In order to provide a broader list of potential RpoN targets, we therefore provided a list of genes predicted to be under control of RpoN-regulated promoters (see Additional file 4), with predicted sequences. To increase the possibility of capturing even weaker effects of RpoN on gene expression, the threshold level for expression changes was set at 1.25 for comparison of the microarray transcriptome analysis and the computer prediction (Table 2 and Additional file 5). By lowering the cutoff of fold changes and by using additional validation by the primer extension assays, we were able to confirm RpoN regulatory elements located upstream of operons/genes identified by both methods (Table 2 &3), suggesting that this strategy is feasible.
The results from the transcriptome analysis and the primer extension assays showed that RpoN-dependent promoters can be found upstream of both up- and down-regulated genes in the RpoN+ strain. For example, increased transcript levels observed for some genes, such as dcuB, could result from increased RpoN availability for transcription initiation, which is the limiting factor at a normal rpoN transcript level. Therefore, such genes were up-regulated with the increase in the RpoN level. In contrast, excess RpoN may inhibit transcription due to lack of proportional increase in EBPs and/or core RNAP, thereby making excess RpoN an inhibitor or repressor instead of an activator. This might be explained by the degree of the conservation of the RpoN-recognition sequences. When the RpoN-recognition sequences of glnB, dcuB, fliL, fdnG, nifEN, and flgJ, for which their RpoN-recognition sequences were identified by the primer extension assays (Figure 4), were analyzed, it was found that the RpoN-recognition sequences of the genes repressed in the overexpressing strain (fliL, fdnG, nifEN, and flgJ) are slightly more similar to the consensus of the RpoN-recognition sequences (10 or 11 nucleotides conserved out of 13 consensus nucleotides) than those for the up-regulated genes (glnB and dcuB, 9 nucleotides conserved out of 13 consensus nucleotides) (Figure 5a). This slight difference could make significant difference in transcriptional activation and repression. Therefore, RpoN might bind more tightly to the promoters that are more similar to the consensus and inhibit their transcription in the absence of a proportionally increased amount of an EBP for these promoters. Furthermore, the decreased transcript levels for some genes could also be due to "sigma factor antagonism" in which other sigma factor(s) could not access the promoter regions that were engaged by RpoN [28], or because less core RNAP may be available for other sigma factors in the excess of RpoN. It is likely that more than one sigma factor and/or transcriptional regulators can affect gene expression in the same cell. For example, it has been proposed that the transcription of RpoN-dependent promoters is affected by ppGpp and its cofactor DksA through a mechanism in which more core RNAP are available for RNAP/RpoN holoenzyme formation due to the short half life of RNAP/RpoD induced by binding to ppGpp and DksA [56,57]. Alternatively, the physiological states caused by the overexpression of RpoN in the RpoN+ strain, which were different from those in the WTV strain, such as slower growth, and/or growth conditions, which affect gene regulation, such as fumarate respiration, resulted in positive effects on some genes and negative effects on others. For instance, the dcuB gene was up-regulated in the RpoN+ strain, because the active EBP was likely present for the activation of the dcuB gene under the conditions for the microarray transcriptome analysis, during which cells needed to grow on fumarate, and thus the fumarate/succinate exchanger encoded by the dcuB gene was required for growth. In contrast, it is unlikely that the genes for nitrogen fixation such as nifEN were essential for growth in the presence of ammonia and thus the active EBP for these genes was scarce, if present, under the conditions for the microarray transcriptome analysis, resulting in the inhibition of these genes in the RpoN+ strain. It is also possible that RpoN may affect regulation of different promoters under a different set of conditions or different stages of cell growth.
Conclusion
The results presented here demonstrate that G. sulfurreducens has an RpoN ortholog, which exhibits typical structural characteristics shared by the RpoN family. However, unlike most of other bacterial rpoN genes, the G. sulfurreducens rpoN was indispensable for growth under all conditions tested. By combining data from the computational prediction with the microarray analysis of the RpoN over-expression strain, the regulon of G. sulfurreducens RpoN was identified, which includes a number of genes that are important for growth in subsurface environments and microbial fuel cells. The G. sulfurreducens RpoN regulates the expression of the dcuB gene encoding a fumarate/succinate exchanger, which is essential for fumarate respiration. The G. sulfurreducens RpoN controls both pathways of glutamate/glutamine syntheses, including the GDH (glutamate dehydrogenase) pathway and the GS/GOGAT (glutamine synthase/glutamate synthase) pathway. Thus, deletion of the rpoN gene would hinder cells' ability for ammonia assimilation, and therefore this mutant would not be viable. This study provides information on transcriptional regulatory networks in G. sulfurreducens, which would increase the predictive value of the regulatory modules in the genome-wide in silico models. Further studies for the RpoN transcriptional network in global regulation in G. sulfurreducens are currently underway to fine-tune the regulatory modules in the models.
Methods
Bacterial strains and culturing conditions
Escherichia coli strain JM109 [endA-1 recA-1 gyrA-96 thi hsdR-17(rk-, mk+) relA-1 supE-44 Δ(lac-proAB)(F' traD-36 proAB lacIqZΔM15)] [58], or TOP10 [F-mcrA Δ(mrr-hsdRMS-mcrBC) Φ 80lacZΔM15 ΔlacX-74 recA-1 araD-139 Δ(ara-leu)7697 galU galK rpsL (StrR) endA-1 nupG] [59] was cultured in LB medium at 37°C with shaking. Targeted gene disruption experiments were performed on G. sulfurreducens strain DL1 [60,61] to produce strains DLCN29 (a kanamycin cassette insertion between GSU1888 and 1887 (rpoN)), DLCN32 (a kanamycin cassette insertion between GSU1887 and GSU1886) (Figure 1) and an rpoN diploid strain DLCN43. G. sulfurreducens strains were routinely cultured anaerobically in NB acetate-fumarate (NBAF) or freshwater acetate-Fe(III) citrate (FWAFC) medium at 30°C as previously described [61]. NB and FW are the two basic mineral solutions, and differ mainly in buffering capacity and trace element contents. Acetate (15 mM) and fumarate (40 mM) were the electron donor and electron acceptor, respectively, for the general propagation unless otherwise stated. Both can be substituted with either lactate (20 mM) or hydrogen as an electron donor, or Fe(III) citrate (55 mM) as an electron acceptor when necessary (for a complete media composition please see references [61,62]).
DNA manipulations
Genomic DNA was extracted with the Qiagen Genomic-tip 100/G. Plasmid DNA and PCR products were purified with the Qiagen mini plasmid purification kits and PCR purification kits, respectively (Qiagen). DNA cloning and other manipulations were carried out according to the methods outlined by Sambrook et al. [63]. Restriction enzymes and other DNA-modifying enzymes were from New England Biolabs. Probes for Southern blot analyses were labeled with [α-32P]dCTP using the NEBlot kit (New England BioLabs). [α-32P]dCTP was from PerkinElmer Life and Analytical Sciences. Qiagen Taq DNA polymerase, unless otherwise stated, was used for all PCR amplifications.
Single-step gene replacement
Sequences were deleted with single-step gene replacement as previously described [64]. To disrupt the intergenic regions either upstream (between rpoN (GSU1887) and GSU1888) or downstream (between rpoN and GSU1886) of the rpoN gene, a linear DNA fragment was generated by recombinant PCR [64,65] from three primary PCR products. For disruption of the intergenic region upstream of the rpoN gene, a 2.1 kb linear DNA fragment was composed of three PCR products: (1) the 3' end of GSU1888 (0.5 kb, amplified with primers rpoNU-1 and rpoNU-2); (2) 5' end of the rpoN gene (0.5 kb, amplified with primers rpoNU-5 and rpoNU-6); and (3) a kanamycin resistant cassette (KanR) (1.1 kb, amplified with primers rpoNU-3 and rpoNU-4). For disruption of the intergenic region downstream of the rpoN gene, three primary PCR reactions were performed to amplify a 2.1 kb linear DNA fragment: (1) the 3' end of rpoN [0.5 kb, position to position, amplified with primers rpoND-1 and rpoND-2); (2) 5' end of the GSU1886 gene (0.5 kb, amplified with primers rpoND-5 and rpoND-6); and (3) a KanR cassette (1.1 kb, amplified with primers rpoND-3 and rpoND-4). Recombinant PCR was performed with these three PCR products as templates with distal primer pairs, rpoNU-1/rpoNU-6 and rpoND-1/rpoND-6 for upstream or downstream intergenic region mutation respectively. PCR conditions were as previously described, except that the annealing temperature was 58°C [64]. All primer sequences used in this work are listed in Additional file 7.
Electroporation, mutant isolation and genotype confirmation were performed as previously described [61,64]. One of each of the mutants, designated DLCN29 and DLCN32, was chosen as the representative strain.
Construction of an rpoN diploid strain of G. sulfurreducens (DLCN43)
A 2.5 kb linear DNA fragment containing the chloramphenicol resistance cassette (CmR) followed by the coding region of the rpoN gene was constructed using cross-over PCR [65]. The chloramphenicol resistance cassette was amplified with Cm-rpoNF1 (Cla I site) and Cm-rpoNR2 using pACYC184 as the template. The rpoN gene was amplified with C-rpoNF3 and C-rpoNR4. The two PCR products were joined together by cross-over PCR as described in [64,65]. The resulted PCR product (Cm-rpoN) was Klenow filled-in and ligated to the Sma I-cut pLA01 (as described below), resulted in plasmid pLA03.
The plasmid pLA01 is a derivative of pCR2.1-TOPO that the 5'-end of the periplasmic c-type cytochrome gene (ppcA) which was amplified with primer pair: ppcAF1 and ppcAR2 was cloned into pCR2.1-TOPO using TOPO TA cloning kit (Invitrogen). Therefore, plasmid pLA03 contains the 5'-end ppcA followed by Cm-rpoN: the CmR resistance cassette and the rpoN gene, and the 3'-end of ppcA. The plasmid pLA03 was linearized and electroporated into G. sulfurreducens DL1 and CmR transformants were selected. The insertion of the Cm-rpoN construction within the ppcA gene was verified by PCR and the resultant strain was named DLCN43.
In order to interrupt any of the two copies of the rpoN gene in DLCN43, a linear PCR fragment containing the rpoN gene disrupted by the kanamycin resistance cassette was constructed with cross-over PCR. The 5' region of rpoN was amplified with primer pair: RpoNKmII-1 and RpoNKmII-2. The 3' region of rpoN was amplified with RpoNKmII-5 and RpoNKmII-6. The KanR cassette was amplified with RpoNKmII-3 and RpoNKmII-4. The recombinant PCR was carried out as described in the previous section and the resultant recombinant PCR product was electroporated into the strain DLCN43. A total of 15 KanR transformants were isolated. However, all 15 transformants had the KanR insertion within the Cm-rpoN locus.
Over-expression of rpoN in trans under the control of a lac or an IPTG-inducible taclac promoter
The complete rpoN coding sequence was amplified with primer sets RpoNfor-XbaI and RpoNrev-EcoRI for insertion to pJMG (lac promoter, gentamycin resistant) [66,67] or RpoNfor-EcoRI and RpoNrev-HindIII for insertion to pCD341 (taclac promoter, kanamycin resistant) [68] using Phusion High-Fidelity DNA polymerase (New England Biolabs) under the following conditions: 98°C, 30 s followed by 30 cycles of 98°C,20 s; 58°C, 20 s; 72°C, 60 s; and a final extension at 72°C for 10 min. The PCR product of the rpoN coding sequence was digested with restriction enzyme sets of Xba I and EcoR I or EcoR I and Hind III and inserted into the Xba I and EcoR I sites of the vector pJMG or the EcoR I and Hind III sites of the vector pCD341 via ligation; the resulting plasmids were designated pJMG rpoN or pCD rpoN, respectively. The rpoN gene in pJMGrpoN or pCDrpoN was then sequenced to screen for PCR artifacts.
Following electroporation of strain DL1 with pJMGrpoN or pCDrpoN, a gentamycin-resistant transformant or a kanamycin-resistant transformant, was isolated and designated DL1/pJMGrpoN or DL1/pCDrpoN (RpoN+ for simplification), respectively. The presence of the plasmid in the DL1 strain was confirmed by plasmid purification and PCR.
The over-expression of rpoN for the strain containing pCDrpoN was achieved by adding 1 mM IPTG, a non-degradable analog of lactose to the medium. In the absence of lactose, transcription from the taclac promoter is inhibited by the lacZ repressor [69]. Upon addition of lactose or IPTG, the lacZ repressor is inactivated, therefore inducing transcription of the rpoN operon.
Primer extension analyses
Total RNA was isolated from mid-exponential-phase cultures with RNeasy Midi kits (Qiagen) followed by treatment with RNase-free DNase (Ambion). Primer extension experiments were performed at 42°C using AMV reverse transcriptase (Roche) with primers GSU0364-06, GSU0420-04, GSU0777-04, GSU0938-06, GSU1836-04, GSU2005-02, GSU2302-04, GSU2490-02, GSU2751-02, GSU2806–08, GSU3046-02, and GSU3206-06, respectively for the corresponding promoter regions. The sequencing ladders presented in Figure 3 and Additional file 6 were also generated with these same primers using Thermo Sequenase Cycle sequencing kit (USB).
DNA microarray hybridization and statistical analysis
DNA microarray hybridization was carried out as previously described [70]. Briefly, total RNA was extracted from three sets of identically treated batch cultures of the wild type harboring an empty vector (DL1/pCD341 or WTV for simplification) and the RpoN overexpressing strains (RpoN+). Ten micrograms of RNA from the wild type and the RpoN+ strain samples were chemically labeled with Cy3 or Cy5 fluorescent dyes respectively, using the MicroMax ASAP RNA Labeling Kit (Perkin Elmer), according to manufacturer's instructions. Labeled RNA was fragmented in a 20 μl volume at 70°C for 30 min using Ambion's Fragmentation Reagent and competitively hybridized to 12 K Arrays (Combimatrix) according to manufacturer's protocol. The arrays were scanned using a GenePix 4000B scanner (Molecular Devices), and analyzed using GenePix and Acuity 4.0 software. LIMMA mixed model analysis (R-package LIMMA [71]) was applied to the normalized Log2 expression ratios to identify differentially expressed genes. The P-value was then corrected for multiple comparisons according to Benjamini and Hochberg's procedure [72] to control the false discovery rate (FDR). Genes whose expression was significantly changed are listed in Additional files 1 &2 according to their fold changes (≤ -1.5 for down-regulation and ≥ +1.5 for up-regulation) and the P-values (≤ 0.0005). A gene was considered differentially expressed if at least half of its probes had a P ≤ 0.0005 and a fold change ≤ -1.5 or ≥ +1.5.
Gene expression microarray data (raw data and statistically processed data files) for the G. sulfurreducens over-expressing RpoN strain are available from the NCBI GEO (Gene Expression Omnibus) database http://www.ncbi.nlm.nih.gov/geo/, with accession GSE8022.
Computational analysis of RpoN-regulated promoters and their target operons
RpoN-regulated promoters were predicted in the genome of G. sulfurreducens using the PromScan software [27]. This software assigned scores representing the Kullback-Leibler distance for predicted RpoN sites in the G. sulfurreducens genome, based on 186 known RpoN promoter sites from 47 bacterial species [2]. The predicted sequence elements were ranked according to their PromScan scores, and sequence elements with scores equal to or exceeding the default cutoff of 80 were selected for further consideration.
The operon organization of the G. sulfurreducens genome was predicted using a commercial version of the FGENESB software (V. Solovyev, A. Salamov, and P. Kosarev, unpublished; Softberry, Inc; 2003–2008). The reference June 1, 2004 version of operon annotation used in this study has been described previously [18]. For all RpoN-regulated promoter elements predicted by PromScan, we compared their genome location and strand orientation relative to operons and singleton ORFs. Those sequence elements that were located upstream of and in the same direction with protein-coding genes were considered to be possible RpoN-regulated promoter elements. Only those elements that did not overlap with coding genes (according to gene boundaries predicted by the FGENESB software) were selected for further consideration.
To compare the predicted locations of RpoN-regulated promoters with experimental evidence, we identified predicted RpoN promoters located upstream of and in the same orientation with genes with significantly altered expression in the RpoN+ strain. This was achieved by comparing the list of suggested target genes located downstream of RpoN promoters (see Additional file 4) to the list of genes with significantly altered expression in the RpoN+ strain and identifying the genes present in both lists.
Consensus sequences of predicted RpoN promoters was computed using our software, CONSENS by J. Krushkal [73]. Each nucleotide reported in the output consensus sequence represents the most frequent nucleotide. For ambiguous nucleotides co-occurring with equal highest frequencies, degenerate symbols were used according to the IUPAC-IUB ambiguity codes. Sequence logos of the predicted promoter sites were drawn using the WebLogo package v. 3 beta at http://weblogo.berkeley.edu/[74].
In silico analysis of G. sulfurreducens growth
In silico modeling was utilized to analyze the possible phenotypes of G. sulfurreducens mutants in which genes encoding enzymes for ammonia assimilation pathway were deleted. The constraint-based genome-scale metabolic model of G. sulfurreducens [16] was applied in simulating cell growth using flux balance analysis and linear optimization [75] in SimPheny (Genomatica, Inc., CA). Biomass synthesis was selected as the objective function to be maximized in growth simulations. The following external metabolites were allowed to freely enter and leave the network for simulations of anaerobic growth on minimal media: Ca2+, CO2, Fe2+, H+, H2O, K+, Mg2+, Na+, NH4+, PO43-, and SO42-. Acetate was supplied to the metabolic model as electron donor and Fe(III) or fumarate was supplied as electron acceptor for the simulations. All other external metabolites were only allowed to leave the system.
Preparation of antisera against RpoN
The rpoN coding region was amplified with primers pGEXrpoNEcoRIfor and pGEXrpoNXhoIrev, digested with EcoR I and Xho I, and inserted into the EcoR I-Xho I sites of pGEX-4T-1 (GE). Competent E. coli strain JM109 was transformed with the resulting plasmid, pGEXrpoN. The E. coli cell lysates containing the over-expressed GST-tagged RpoN was size-fractioned by sodium dodecyl sulfate-polyacrylamide gel electrophoresis. The band corresponding to RpoN protein was cut, crushed, and used to immunize New Zealand rabbits for antibody production against RpoN as described by Harlow and Lane [76].
Analytical techniques
Protein concentration was determined using the bicinchoninic acid method with bovine serum albumin as a standard [77]. Western blot analyses were carried out by using antiserum against RpoN according to the protocol described by Ausubel et al [78]. Immunoreactive bands were visualized using an alkaline phosphatase-conjugated goat anti-rabbit secondary antibody (Pierce) and 1-step NBT/BCIP plus suppressor (Pierce) according to the manufacturer's instructions. Growth of fumarate cultures was monitored by measuring turbidity at 600 nm in a Genesys 2 spectrophotometer (Spectronic Instruments). Cell density of Fe(III)-grown cultures were determined using epifluorescence microscopy with acridine orange staining [79]. Fe(II) concentrations were determined with the ferrozine assay as previously described [80]. Agglutination assays were preformed as described by Reguera et al [31].
Authors' contributions
CL designed this study, carried out experimental work, drafted the manuscript and interpreted microarray analysis and PromScan data. JK oversaw the computational prediction of RpoN-regulated promoters, interpreted results and drafted the manuscript. TU drafted the manuscript and carried out part of the primer extension assays. MP predicted RpoN-regulated promoter elements and localized them relative to gene expression changes and operon locations. JS performed in silico modeling analysis to predict the effects of GS, GOGAT and/or GDH deletion mutations on cell growth under different conditions. KJ immunized rabbits for antiserum production against RpoN. CN participated in RpoN mutagenesis. GR participated in agglutination analysis. RD and BP carried out microarray hybridization. RMA assisted with interpretation of RpoN promoter analysis and bioinformatics data integration. DRL designed the study and participated in interpretation of the results and drafted the manuscript. All authors participated in editing this manuscript and approved of its final form.
Supplementary Material
List of up-regulated genes in the RpoN over-expressing strain as compared to the wild type strain, based on fold change cutoff 1.5.
List of down-regulated genes in the RpoN over-expressing strain as compared to the wild type strain, based on fold change cutoff 1.5.
List of 110 predicted RpoN-regulated promoters located in the noncoding regions, upstream of and in the same orientation with protein-coding genes.
List of 110 predicted RpoN-regulated promoters and their target downstream genes.
List of operons/genes containing RpoN-dependent promoters identified by the PromScan analysis and also identified using transcriptome analysis.
RpoN-dependent gene expression. Representative genes, (a) GSU0364, GSU2005, GSU2302, GSU2490 and GSU3206 (up-regulated in the RpoN+ strain), and (b) GSU0938 (down-regulated in the RpoN+ strain) identified by the microarray analysis were analyzed by primer extension assays. The results of the primer extension assays and their promoter regions are shown. The 5' ends of mRNA are indicated by asterisks. RBS sites are underlined. Translation start codons are in bold and are indicated by Met.
Primers used in this work.
Contributor Information
Ching Leang, Email: leang@microbio.umass.edu.
Julia Krushkal, Email: jkrushka@utmem.edu.
Toshiyuki Ueki, Email: tueki@microbio.umass.edu.
Marko Puljic, Email: neuropercolation@yahoo.com.
Jun Sun, Email: jsun@genomatica.com.
Katy Juárez, Email: katy@ibt.unam.mx.
Cinthia Núñez, Email: cinthia@ibt.unam.mx.
Gemma Reguera, Email: reguera@msu.edu.
Raymond DiDonato, Email: rdidonat@hotmail.com.
Bradley Postier, Email: bpostier@biology2.wustl.edu.
Ronald M Adkins, Email: radkins1@utmem.edu.
Derek R Lovley, Email: dlovley@microbio.umass.edu.
Acknowledgements
This research was supported by the Office of Science (BER), U. S. Department of Energy, Cooperative Agreement No. DE-FC02-02ER63446.
We are grateful for the excellent technical support from Betsy Blunt and Lorrie A. Perpetua. We thank Anna Liu for help with Limma analysis for the RpoN+ microarray data set. We also thank Yanhua Qu (University of Tennessee, Memphis) for technical assistance with the online supplement and Jeanete Peeples (University of Tennessee, Memphis) for editorial assistance.
References
- Buck M, Cannon W. Specific binding of the transcription factor sigma-54 to promoter DNA. Nature. 1992;10(6385):422–424. doi: 10.1038/358422a0. [DOI] [PubMed] [Google Scholar]
- Barrios H, Valderrama B, Morett E. Compilation and analysis of sigma54-dependent promoter sequences. Nucleic Acids Res. 1999;10(22):4305–4313. doi: 10.1093/nar/27.22.4305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studholme DJ, Dixon R. Domain architectures of sigma54-dependent transcriptional activators. J Bacteriol. 2003;10(6):1757–1767. doi: 10.1128/JB.185.6.1757-1767.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitzer L, Schneider BL. Metabolic context and possible physiological themes of sigma54-dependent genes in Escherichia coli. Microbiol Mol Biol Rev. 2001;10(3):422–444. doi: 10.1128/MMBR.65.3.422-444.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studholme DJ, Buck M. The biology of enhancer-dependent transcriptional regulation in bacteria: insights from genome sequences. FEMS Microbiol Lett. 2000;10(1):1–9. doi: 10.1111/j.1574-6968.2000.tb09074.x. [DOI] [PubMed] [Google Scholar]
- Ishimoto KS, Lory S. Formation of pilin in Pseudomonas aeruginosa requires the alternative sigma factor (RpoN) of RNA polymerase. Proc Natl Acad Sci USA. 1989;10(6):1954–1957. doi: 10.1073/pnas.86.6.1954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson LS, Webb JS, Rice SA, Kjelleberg S. The alternative sigma factor RpoN regulates the quorum sensing gene rhlI in Pseudomonas aeruginosa. FEMS Microbiol Lett. 2003;10(2):187–195. doi: 10.1016/S0378-1097(03)00097-1. [DOI] [PubMed] [Google Scholar]
- Totten PA, Lara JC, Lory S. The rpoN gene product of Pseudomonas aeruginosa is required for expression of diverse genes, including the flagellin gene. J Bacteriol. 1990;10(1):389–396. doi: 10.1128/jb.172.1.389-396.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautam US, Jajoo A, Singh A, Chakrabartty PK, Das SK. Characterization of an rpoN mutant of Mesorhizobium ciceri. J Appl Microbiol. 2007;10(5):1798–1807. doi: 10.1111/j.1365-2672.2007.03432.x. [DOI] [PubMed] [Google Scholar]
- Ali NO, Bignon J, Rapoport G, Debarbouille M. Regulation of the acetoin catabolic pathway is controlled by sigma L in Bacillus subtilis. J Bacteriol. 2001;10(8):2497–2504. doi: 10.1128/JB.183.8.2497-2504.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arous S, Buchrieser C, Folio P, Glaser P, Namane A, Hebraud M, Hechard Y. Global analysis of gene expression in an rpoN mutant of Listeria monocytogenes. Microbiology. 2004;10(Pt 5):1581–1590. doi: 10.1099/mic.0.26860-0. [DOI] [PubMed] [Google Scholar]
- Debarbouille M, Martin-Verstraete I, Kunst F, Rapoport G. The Bacillus subtilis sigL gene encodes an equivalent of sigma 54 from Gram-negative bacteria. Proc Natl Acad Sci USA. 1991;10(20):9092–9096. doi: 10.1073/pnas.88.20.9092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin I, Debarbouille M, Klier A, Rapoport G. Induction and metabolite regulation of levanase synthesis in Bacillus subtilis. J Bacteriol. 1989;10(4):1885–1892. doi: 10.1128/jb.171.4.1885-1892.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovely DR, Mahadevan R, Nevin K. In: Microbial Biodegradation-Genomics and Molecular Biology. Diaz E, editor. Norfolk, UK: Caister Academic Press; 2008. Systems Biology Approach to Bioremediation with Extracellular Electron Transfer. [Google Scholar]
- Lovley DR. Bug juice: harvesting electricity with microorganisms. Nat Rev Microbiol. 2006;10(7):497–508. doi: 10.1038/nrmicro1442. [DOI] [PubMed] [Google Scholar]
- Mahadevan R, Bond DR, Butler JE, Esteve-Nunez A, Coppi MV, Palsson BO, Schilling CH, Lovley DR. Characterization of metabolism in the Fe(III)-reducing organism Geobacter sulfurreducens by constraint-based modeling. Appl Environ Microbiol. 2006;10(2):1558–1568. doi: 10.1128/AEM.72.2.1558-1568.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovely DR, Nevin K. In: Bioenergy. Wall JD, editor. Washington, DC: ASM Press; 2008. Electricity Production with Electricigens; pp. 295–306. [Google Scholar]
- Krushkal J, Yan B, DiDonato LN, Puljic M, Nevin KP, Woodard TL, Adkins RM, Methe BA, Lovley DR. Genome-wide expression profiling in Geobacter sulfurreducens : identification of Fur and RpoS transcription regulatory sites in a relGsu mutant. Functional & integrative genomics. 2007;10(3):229–255. doi: 10.1007/s10142-007-0048-5. [DOI] [PubMed] [Google Scholar]
- Yan B, Lovley DR, Krushkal J. Genome-wide similarity search for transcription factors and their binding sites in a metal-reducing prokaryote Geobacter sulfurreducens. Biosystems. 2007;10(2):421–441. doi: 10.1016/j.biosystems.2006.10.006. [DOI] [PubMed] [Google Scholar]
- Yan B, Nunez C, Ueki T, Esteve-Nunez A, Puljic M, Adkins RM, Methe BA, Lovley DR, Krushkal J. Computational prediction of RpoS and RpoD regulatory sites in Geobacter sulfurreducens using sequence and gene expression information. Gene. 2006;10:73–95. doi: 10.1016/j.gene.2006.06.025. [DOI] [PubMed] [Google Scholar]
- Yan B, Methe BA, Lovley DR, Krushkal J. Computational prediction of conserved operons and phylogenetic footprinting of transcription regulatory elements in the metal-reducing bacterial family Geobacteraceae. J Theor Biol. 2004;10(1):133–144. doi: 10.1016/j.jtbi.2004.04.022. [DOI] [PubMed] [Google Scholar]
- Nunez C, Adams L, Childers S, Lovley DR. The RpoS Sigma Factor in the Dissimilatory Fe(III)-Reducing Bacterium Geobacter sulfurreducens. J Bacteriol. 2004;10(16):5543–5546. doi: 10.1128/JB.186.16.5543-5546.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunez C, Esteve-Nunez A, Giometti C, Tollaksen S, Khare T, Lin W, Lovley DR, Methe BA. DNA microarray and proteomic analyses of the RpoS regulon in Geobacter sulfurreducens. J Bacteriol. 2006;10(8):2792–2800. doi: 10.1128/JB.188.8.2792-2800.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ueki T, Lovley DR. Heat-shock sigma factor RpoH from Geobacter sulfurreducens. Microbiology. 2007;10(Pt 3):838–846. doi: 10.1099/mic.0.2006/000638-0. [DOI] [PubMed] [Google Scholar]
- Merrick MJ. In a class of its own – the RNA polymerase sigma factor sigma 54 (sigma N) Mol Microbiol. 1993;10(5):903–909. doi: 10.1111/j.1365-2958.1993.tb00961.x. [DOI] [PubMed] [Google Scholar]
- Merrick MJ, Gibbins JR. The nucleotide sequence of the nitrogen-regulation gene ntrA of Klebsiella pneumoniae and comparison with conserved features in bacterial RNA polymerase sigma factors. Nucleic Acids Res. 1985;10(21):7607–7620. doi: 10.1093/nar/13.21.7607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studholme DJ, Buck M, Nixon T. Identification of potential sigmaN-dependent promoters in bacterial genomes. Microbiology. 2000;10(Pt 12):3021–3023. doi: 10.1099/00221287-146-12-3021. [DOI] [PubMed] [Google Scholar]
- Boucher JC, Schurr MJ, Deretic V. Dual regulation of mucoidy in Pseudomonas aeruginosa and sigma factor antagonism. Mol Microbiol. 2000;10(2):341–351. doi: 10.1046/j.1365-2958.2000.01846.x. [DOI] [PubMed] [Google Scholar]
- Reguera G, McCarthy KD, Mehta T, Nicoll JS, Tuominen MT, Lovley DR. Extracellular electron transfer via microbial nanowires. Nature. 2005;10(7045):1098–1101. doi: 10.1038/nature03661. [DOI] [PubMed] [Google Scholar]
- Juarez K, Kim BC, Nevin K, Olvera L, Reguera G, Lovley DR, Methe BA. PilR, a transcriptional regulator for pilin and other genes required for Fe(III) reduction in Geobacter sulfurreducens. J Mol Microbiol Biotechnol. 2009;10(3–4):146–158. doi: 10.1159/000115849. [DOI] [PubMed] [Google Scholar]
- Reguera G, Pollina RB, Nicoll JS, Lovley DR. Possible Nonconductive Role of Geobacter sulfurreducens Pilus Nanowires in Biofilm Formation. J Bacteriol. 2007;10(5):2125–2127. doi: 10.1128/JB.01284-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butler JE, Glaven RH, Esteve-Nunez A, Nunez C, Shelobolina ES, Bond DR, Lovley DR. Genetic characterization of a single bifunctional enzyme for fumarate reduction and succinate oxidation in Geobacter sulfurreducens and engineering of fumarate reduction in Geobacter metallireducens. J Bacteriol. 2006;10(2):450–455. doi: 10.1128/JB.188.2.450-455.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janausch IG, Zientz E, Tran QH, Kroger A, Unden G. C4-dicarboxylate carriers and sensors in bacteria. Biochim Biophys Acta. 2002;10(1–2):39–56. doi: 10.1016/s0005-2728(01)00233-x. [DOI] [PubMed] [Google Scholar]
- Merrick MJ, Edwards RA. Nitrogen control in bacteria. Microbiol Rev. 1995;10(4):604–622. doi: 10.1128/mr.59.4.604-622.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helling RB. Why does Escherichia coli have two primary pathways for synthesis of glutamate? J Bacteriol. 1994;10(15):4664–4668. doi: 10.1128/jb.176.15.4664-4668.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitzer L. Nitrogen assimilation and global regulation in Escherichia coli. Annu Rev Microbiol. 2003;10:155–176. doi: 10.1146/annurev.micro.57.030502.090820. [DOI] [PubMed] [Google Scholar]
- Bender RA. The role of the NAC protein in the nitrogen regulation of Klebsiella aerogenes. Mol Microbiol. 1991;10(11):2575–2580. doi: 10.1111/j.1365-2958.1991.tb01965.x. [DOI] [PubMed] [Google Scholar]
- Bormann ER, Eikmanns BJ, Sahm H. Molecular analysis of the Corynebacterium glutamicum gdh gene encoding glutamate dehydrogenase. Mol Microbiol. 1992;10(3):317–326. doi: 10.1111/j.1365-2958.1992.tb01474.x. [DOI] [PubMed] [Google Scholar]
- Di Fraia R, Wilquet V, Ciardiello MA, Carratore V, Antignani A, Camardella L, Glansdorff N, Di Prisco G. NADP+-dependent glutamate dehydrogenase in the Antarctic psychrotolerant bacterium Psychrobacter sp. TAD1. Characterization, protein and DNA sequence, and relationship to other glutamate dehydrogenases. Eur J Biochem. 2000;10(1):121–131. doi: 10.1046/j.1432-1327.2000.00972.x. [DOI] [PubMed] [Google Scholar]
- Hashim S, Kwon DH, Abdelal A, Lu CD. The arginine regulatory protein mediates repression by arginine of the operons encoding glutamate synthase and anabolic glutamate dehydrogenase in Pseudomonas aeruginosa. J Bacteriol. 2004;10(12):3848–3854. doi: 10.1128/JB.186.12.3848-3854.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kloosterman TG, Hendriksen WT, Bijlsma JJ, Bootsma HJ, van Hijum SA, Kok J, Hermans PW, Kuipers OP. Regulation of glutamine and glutamate metabolism by GlnR and GlnA in Streptococcus pneumoniae. J Biol Chem. 2006;10(35):25097–25109. doi: 10.1074/jbc.M601661200. [DOI] [PubMed] [Google Scholar]
- Pagliarulo C, Salvatore P, De Vitis LR, Colicchio R, Monaco C, Tredici M, Tala A, Bardaro M, Lavitola A, Bruni CB. et al. Regulation and differential expression of gdhA encoding NADP-specific glutamate dehydrogenase in Neisseria meningitidis clinical isolates. Mol Microbiol. 2004;10(6):1757–1772. doi: 10.1111/j.1365-2958.2003.03947.x. [DOI] [PubMed] [Google Scholar]
- Lovley DR, Holmes DE, Nevin KP. Dissimilatory Fe(III) and Mn(IV) reduction. Adv Microb Physiol. 2004;10:219–286. doi: 10.1016/S0065-2911(04)49005-5. [DOI] [PubMed] [Google Scholar]
- Anderson RT, Vrionis HA, Ortiz-Bernad I, Resch CT, Long PE, Dayvault R, Karp K, Marutzky S, Metzler DR, Peacock A. et al. Stimulating the in situ activity of Geobacter species to remove uranium from the groundwater of a uranium-contaminated aquifer. Appl Environ Microbiol. 2003;10(10):5884–5891. doi: 10.1128/AEM.69.10.5884-5891.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Segura D, Mahadevan R, Juarez K, Lovley DR. Computational and Experimental Analysis of Redundancy in the Central Metabolism of Geobacter sulfurreducens. PLoS computational biology. 2008;10(2):e36. doi: 10.1371/journal.pcbi.0040036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leang C, Coppi MV, Lovley DR. OmcB, a c-type polyheme cytochrome, involved in Fe(III) reduction in Geobacter sulfurreducens. J Bacteriol. 2003;10(7):2096–2103. doi: 10.1128/JB.185.7.2096-2103.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Childers SE, Ciufo S, Lovley DR. Geobacter metallireducens accesses insoluble Fe(III) oxide by chemotaxis. Nature. 2002;10(6882):767–769. doi: 10.1038/416767a. [DOI] [PubMed] [Google Scholar]
- McCarter LL. Regulation of flagella. Curr Opin Microbiol. 2006;10(2):180–186. doi: 10.1016/j.mib.2006.02.001. [DOI] [PubMed] [Google Scholar]
- Soutourina OA, Bertin PN. Regulation cascade of flagellar expression in Gram-negative bacteria. FEMS Microbiol Rev. 2003;10(4):505–523. doi: 10.1016/S0168-6445(03)00064-0. [DOI] [PubMed] [Google Scholar]
- Kazmierczak MJ, Wiedmann M, Boor KJ. Alternative sigma factors and their roles in bacterial virulence. Microbiol Mol Biol Rev. 2005;10(4):527–543. doi: 10.1128/MMBR.69.4.527-543.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reguera G, Nevin KP, Nicoll JS, Covalla SF, Woodard TL, Lovley DR. Biofilm and nanowire production leads to increased current in Geobacter sulfurreducens fuel cells. Appl Environ Microbiol. 2006;10(11):7345–7348. doi: 10.1128/AEM.01444-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brun YV, Shapiro L. A temporally controlled sigma-factor is required for polar morphogenesis and normal cell division in Caulobacter. Genes Dev. 1992;10(12A):2395–2408. doi: 10.1101/gad.6.12a.2395. [DOI] [PubMed] [Google Scholar]
- Powell BS, Court DL, Inada T, Nakamura Y, Michotey V, Cui X, Reizer A, Saier MH Jr, Reizer J. Novel proteins of the phosphotransferase system encoded within the rpoN operon of Escherichia coli. Enzyme IIANtr affects growth on organic nitrogen and the conditional lethality of an erats mutant. J Biol Chem. 1995;10(9):4822–4839. doi: 10.1074/jbc.270.9.4822. [DOI] [PubMed] [Google Scholar]
- Keseler IM, Kaiser D. sigma(54), a vital protein for Myxococcus xanthus. Proc Natl Acad Sci USA. 1997;10(5):1979–1984. doi: 10.1073/pnas.94.5.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S, Brocchieri L, Mrazek J, Kaiser D. Distinguishing features of delta-proteobacterial genomes. Proc Natl Acad Sci USA. 2006;10(30):11352–11357. doi: 10.1073/pnas.0604311103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernardo LM, Johansson LU, Solera D, Skarfstad E, Shingler V. The guanosine tetraphosphate (ppGpp) alarmone, DksA and promoter affinity for RNA polymerase in regulation of sigma-dependent transcription. Mol Microbiol. 2006;10(3):749–764. doi: 10.1111/j.1365-2958.2006.05129.x. [DOI] [PubMed] [Google Scholar]
- Szalewska-Palasz A, Johansson LU, Bernardo LM, Skarfstad E, Stec E, Brannstrom K, Shingler V. Properties of RNA polymerase bypass mutants: implications for the role of ppGpp and its co-factor DksA in controlling transcription dependent on sigma54. J Biol Chem. 2007;10(25):18046–18056. doi: 10.1074/jbc.M610181200. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C, Vieira J, Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;10(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Grant SG, Jessee J, Bloom FR, Hanahan D. Differential plasmid rescue from transgenic mouse DNAs into Escherichia coli methylation-restriction mutants. Proc Natl Acad Sci USA. 1990;10(12):4645–4649. doi: 10.1073/pnas.87.12.4645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caccavo F Jr, Lonergan DJ, Lovley DR, Davis M, Stolz JF, McInerney MJ. Geobacter sulfurreducens sp. nov., a hydrogen- and acetate-oxidizing dissimilatory metal-reducing microorganism. Appl Environ Microbiol. 1994;10(10):3752–3759. doi: 10.1128/aem.60.10.3752-3759.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coppi MV, Leang C, Sandler SJ, Lovley DR. Development of a genetic system for Geobacter sulfurreducens. Appl Environ Microbiol. 2001;10(7):3180–3187. doi: 10.1128/AEM.67.7.3180-3187.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coppi MV, O'Neil RA, Lovley DR. Identification of an uptake hydrogenase required for hydrogen-dependent reduction of Fe(III) and other electron acceptors by Geobacter sulfurreducens. J Bacteriol. 2004;10(10):3022–3028. doi: 10.1128/JB.186.10.3022-3028.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual. 2. New York: Cold Spring Harbor Laboratory Press;; 1989. [Google Scholar]
- Lloyd JR, Leang C, Hodges Myerson AL, Coppi MV, Cuifo S, Methe B, Sandler SJ, Lovley DR. Biochemical and genetic characterization of PpcA, a periplasmic c-type cytochrome in Geobacter sulfurreducens. Biochem J. 2003;10(Pt 1):153–161. doi: 10.1042/BJ20020597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murphy KC, Campellone KG, Poteete AR. PCR-mediated gene replacement in Escherichia coli. Gene. 2000;10(1–2):321–330. doi: 10.1016/S0378-1119(00)00071-8. [DOI] [PubMed] [Google Scholar]
- Butler JE, Kaufmann F, Coppi MV, Nunez C, Lovley DR. MacA, a diheme c-type cytochrome involved in Fe(III) reduction by Geobacter sulfurreducens. J Bacteriol. 2004;10(12):4042–4045. doi: 10.1128/JB.186.12.4042-4045.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marx CJ, Lidstrom ME. Development of improved versatile broad-host-range vectors for use in methylotrophs and other Gram-negative bacteria. Microbiology. 2001;10(Pt 8):2065–2075. doi: 10.1099/00221287-147-8-2065. [DOI] [PubMed] [Google Scholar]
- Dehio M, Knorre A, Lanz C, Dehio C. Construction of versatile high-level expression vectors for Bartonella henselae and the use of green fluorescent protein as a new expression marker. Gene. 1998;10(2):223–229. doi: 10.1016/S0378-1119(98)00319-9. [DOI] [PubMed] [Google Scholar]
- Barkley MD, Bourgeois S. In: The operon. Miller JH, Reznikoff WS, editor. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory; 1978. Repressor recognition of operator and effectors; pp. 177–220. [Google Scholar]
- Postier B, Didonato R Jr, Nevin KP, Liu A, Frank B, Lovley D, Methe BA. Benefits of in-situ synthesized microarrays for analysis of gene expression in understudied microorganisms. J Microbiol Methods. 2008;10(1):26–32. doi: 10.1016/j.mimet.2007.07.004. [DOI] [PubMed] [Google Scholar]
- Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Statistical applications in genetics and molecular biology. 2004;10 doi: 10.2202/1544-6115.1027. [DOI] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate – a Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B-Methodological. 1995;10(1):289–300. [Google Scholar]
- Zharkikh AA, Rzhetsky A, Morosov PS, Sitnikova TL, Krushkal JS. VOSTORG: a package of microcomputer programs for sequence analysis and construction of phylogenetic trees. Gene. 1991;10(2):251–254. doi: 10.1016/0378-1119(91)90419-C. [DOI] [PubMed] [Google Scholar]
- Crooks GE, Hon G, Chandonia JM, Brenner SE. WebLogo: a sequence logo generator. Genome research. 2004;10(6):1188–1190. doi: 10.1101/gr.849004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards JS, Palsson BO. Metabolic flux balance analysis and the in silico analysis of Escherichia coli K-12 gene deletions. BMC bioinformatics. 2000;10:1. doi: 10.1186/1471-2105-1-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harlow E, Lane D. Antibodies: a Laboratory Manual. New York: Cold Spring Harbor Laboratory; 1988. [Google Scholar]
- Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC. Measurement of protein using bicinchoninic acid. Anal Biochem. 1985;10(1):76–85. doi: 10.1016/0003-2697(85)90442-7. [DOI] [PubMed] [Google Scholar]
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K, eds. Current protocols in molecular biology. New York: John Wiley & Sons, Inc.; 1999. [Google Scholar]
- Lovley DR, Phillips EJP. Novel mode of microbial energy metabolism: organic carbon oxidation coupled to dissimilatory reduction of iron or manganese. Appl Environ Microbiol. 1988;10(6):1472–1480. doi: 10.1128/aem.54.6.1472-1480.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovley DR, Philips EJ. Organic matter mineralization with the reduction of ferric iron in anaerobic sediments. Appl Environ Microbiol. 1986;10:683–689. doi: 10.1128/aem.51.4.683-689.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
List of up-regulated genes in the RpoN over-expressing strain as compared to the wild type strain, based on fold change cutoff 1.5.
List of down-regulated genes in the RpoN over-expressing strain as compared to the wild type strain, based on fold change cutoff 1.5.
List of 110 predicted RpoN-regulated promoters located in the noncoding regions, upstream of and in the same orientation with protein-coding genes.
List of 110 predicted RpoN-regulated promoters and their target downstream genes.
List of operons/genes containing RpoN-dependent promoters identified by the PromScan analysis and also identified using transcriptome analysis.
RpoN-dependent gene expression. Representative genes, (a) GSU0364, GSU2005, GSU2302, GSU2490 and GSU3206 (up-regulated in the RpoN+ strain), and (b) GSU0938 (down-regulated in the RpoN+ strain) identified by the microarray analysis were analyzed by primer extension assays. The results of the primer extension assays and their promoter regions are shown. The 5' ends of mRNA are indicated by asterisks. RBS sites are underlined. Translation start codons are in bold and are indicated by Met.
Primers used in this work.