Abstract
The adenovirus type 5 (Ad5) early region 1B 55-kDa (E1B-55K) protein is a multifunctional regulator of cell-cycle-independent virus replication that participates in many processes required for maximal virus production. As part of a study of E1B-55K function, we generated the Ad5 mutant H5pm4133, carrying stop codons after the second and seventh codons of the E1B reading frame, thereby eliminating synthesis of the full-length 55K product and its smaller derivatives. Unexpectedly, phenotypic studies revealed that H5pm4133 fully exhibits the characteristics of wild-type (wt) Ad5 in all assays tested. Immunoblot analyses demonstrated that H5pm4133 and wt Ad5 produce very low levels of two distinct polypeptides in the 48- to 49-kDa range, which lack the amino-terminal region but contain segments from the central and carboxy-terminal part of the 55K protein. Genetic and biochemical studies with different Ad5 mutants show that at least one of these isoforms consists of two closely migrating polypeptides of 433 amino acid residues (433R) and 422R, which are produced by translation initiation at two downstream AUG codons of the 55K reading frame. Significantly, a virus mutant producing low levels of the 433R isoform alone replicated to levels comparable to those of wt Ad5, demonstrating that this polypeptide provides essentially all functions of E1B-55K required to promote maximal virus growth in human tumor cells. Altogether, these results extend previous findings that the wt Ad5 E1B region encodes a series of smaller isoforms of E1B-55K and demonstrate that very low levels of at least one of these novel proteins (E1B-433R) are sufficient for a productive infection.
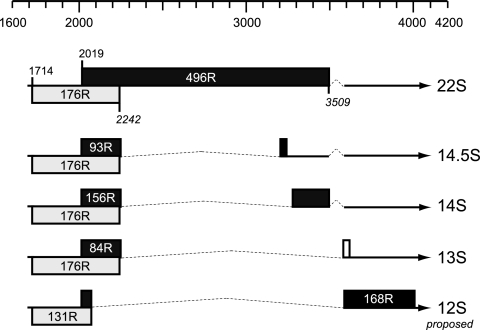
Products of early region 1B (E1B) from subgroup C adenovirus type 5 (Ad5) are required for productive infection of human cells and for complete transformation of primary mammalian cells in cooperation with the E1A gene products (reviewed in reference 6). Most of these activities can be assigned to two different polypeptides of 176 amino acid residues (176R) (E1B 19-kDa protein [E1B-19K]) and 496R (E1B-55K). Both proteins are produced by translation of two overlapping reading frames starting from two separate initiation codons of the 22S transcript (28, 48) (Fig. 1). The 19-kDa product is also translated from three alternatively spliced transcripts of 13S, 14S, and 14.5S, which additionally direct the synthesis of three smaller 55K-related proteins (156R, 93R, and 84R) containing the amino-terminal 79 residues of E1B-55K (12, 45). E1B-156R also shares an identical carboxy terminus with 55K, while the 93R and 84R E1B polypeptides contain unique carboxy termini.
FIG. 1.
Ad5 E1B transcription unit. The positions (nucleotide numbers) on the Ad5 DNA sequence from 1600 to 4200 are indicated on the top. The Ad5 E1B mRNAs are indicated by thin lines and arrows; introns are indicated by dotted lines. The reading frames predicted from DNA sequences and mapping studies are identified by boxes. Included also are the relevant nucleotides for translation start sites. Positions of translation stop sites are shown in italics. The figure was assembled based on material in several publications cited in the text.
It has been well established that E1B-19K and E1B-55K promote efficient viral replication via a number of different mechanisms. In the early phase of the infection, both E1B proteins counteract antiproliferative processes induced by the host cell, including activation of p53-dependent and -independent apoptosis, induction of cell cycle arrest, and stimulation of cellular DNA damage responses (reviewed in references 49 and 50). In the late phase of the infection, E1B-55K additionally controls efficient late viral protein production by stimulating the preferential cytoplasmic accumulation and translation of the viral late mRNAs while simultaneously blocking host cell mRNA nuclear export and host protein synthesis (15, 18). These multiple early and late lytic functions of 55K require interactions with a variety of viral and cellular factors, most importantly with the protein product from early region 4 open reading frame 6 (E4orf6) (reviewed in reference 7 and references therein). Several studies have shown that E4orf6 alters the intracellular distribution of E1B-55K in virus-infected cells, directing the E1B protein to the nuclear matrix compartment (27) and the sites of viral RNA transcription and processing (19, 34). Also, a substantial amount of novel information demonstrates that E4orf6 connects E1B-55K to components of a cellular E3 ubiquitin ligase, thereby allowing the proteasomal degradation of its cellular interaction partners p53, Mre11, and DNA ligase IV (4, 8, 9, 37, 44). It appears that the latter activity also involves nuclear depletion via the CRM1-mediated export pathway and cytoplasmic deposition of Mre11 and other components of the Mre11/Rad50/Nbs1 (MRN) DNA double-strand-break repair complex into aggresomes (23, 29). Consistent with their role in the removal of aggregated proteins from the cytosol, deposition of Mre11 into aggresomes accelerates proteolytic degradation of MRN subunits (29). Finally, considerable evidence suggests that the E1B-55K/E4orf6 complex directly participates in the selective nuclear export of late viral mRNAs through active nucleocytoplasmic shuttling, covalent conjugation of the small ubiquitin-related modifier protein 1 (SUMO1) (23), and significantly through its E3 ubiquitin-protein ligase activity (10, 51).
As opposed to the case with E1B-19K and E1B-55K, no functions have been attributed to the smaller E1B proteins 84R, 93R, and 156R in infection yet. Phenotypic analyses with Ad2/5 virus mutants expressing the 55K protein indicate only that they are dispensable for productive infection in HeLa cells (32, 36). Nevertheless, since E1B-156R shares its amino- and carboxy-terminal regions with E1B-55K and possesses transforming potential, it may share several functions with the 496R product (43). In addition to the three smaller E1B species, the existence of a 168R E1B protein has been proposed for Ad2/5 (Fig. 1), which is translated from a 12S mRNA generated by alternative splicing of the major 22S transcript (2, 28, 45). Finally, work from Barker and Berk has shown that an Ad2/5 mutant produces very low levels of E1B-55K or a 55K-related protein that may lack the amino-terminal region of the adenovirus protein but is sufficient for a productive infection in HeLa cells (5).
In this study, we demonstrate that the Ad2/5 E1B region produces at least four additional isoforms of E1B-55K, including two polypeptides of 433R and 422R. Genetic and biochemical studies with different Ad5 mutants show that both polypeptides are synthesized by initiating translation at the second and third downstream AUG of the E1B-55K reading frame. Interestingly, both isoforms are expressed at very low levels and lack 63 and 74 amino acids from the N terminus of the full-length 496R E1B-55K product. Nevertheless, 433R, and to a lesser extent 422R, comprise all currently known structural and functional domains critical for E1B's early and late lytic functions and provide basically all functions sufficient for a normal productive infection in human tumor cell lines.
MATERIALS AND METHODS
Cell culture.
A549 (DSMZ ACC 107; Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Braunschweig, Germany), H1299 (31), and 293 (20) were grown as monolayer cultures in Dulbecco's modified Eagle's medium (DMEM) supplemented with 5% fetal calf serum, 100 U of penicillin, and 100 μg of streptomycin per ml in a 5%-CO2 atmosphere at 37°C.
Construction of Ad5 recombinants.
Plasmid pE1-1390 contains termination codons (opal and amber) at the third and eighth codons of the 55K reading frame (R3Opa/E8Amb) (Fig. 2A) and was derived from pE1-1235 (21, 23) by site-directed mutagenesis with oligonucleotide primers 1147 and 1148 (Table 1). To eliminate initiation of translation at the second and/or the third AUG of the E1B-55K mRNA, A-to-G transitions were introduced into pE1-1390 at nucleotides (nt) 2208 (pE1-1633), 2241 (pE1-1635), and 2208 plus 2241 (pE1-1889) by site-directed mutagenesis with the complementary oligonucleotide pairs 1285/1286 and 1287/1288, respectively (Table 1). For the construction of pE1-1680, mutations were introduced at nt 2274 plus 2280 into pE1B-1390 by site-directed mutagenesis with oligonucleotide primers 1144 and 1160 (Table 1), resulting in two additional termination codons (ochre and opal) at the 86th and 88th codons (E86Och/R88Opa) of the 55K reading frame. The mutations in these plasmids were verified by DNA sequencing. Next, the 7.7-kb SwaI/BstZ17I fragment from pH5pg4100 (21) was replaced with the corresponding fragments from the plasmids to generate adenoviral plasmids pH5pm4133 (R3Opa/E8Amb), pH5pm4149 (R3Opa/E8Amb/E86Och/R88Opa), pH5pm4177 (R3Opa/E8Amb/M64V), pH5pm4178 (R3Opa/E8Amb/M75V), and pH5pm4179 (R3Opa/E8Amb/M64V/M75V). The recombinant plasmids were partially sequenced to confirm the mutations in the E1B-55K gene (Table 2).
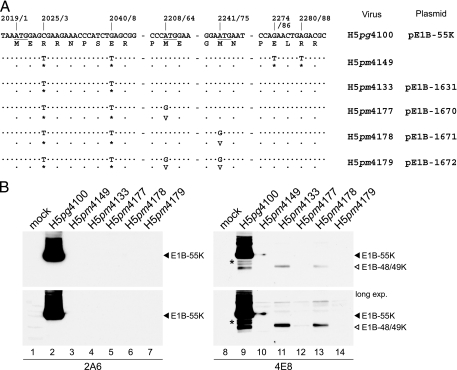
FIG. 2.
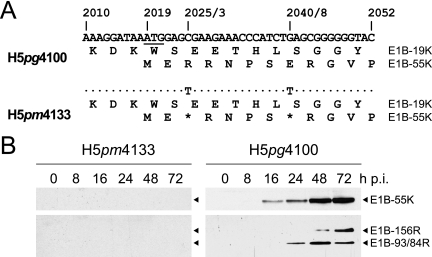
Effect of mutations on expression of E1B-55K and its related products. (A) Mutations in virus H5pm4133. Numbers refer to positions of nucleotides in the wt H5pg4100 DNA sequence, and numbers after the diagonal slash denote the positions of the amino acid residues. The first initiation codon of the 55K reading frame is underlined. Segments of the amino acid sequences of E1B-19K (176R) and E1B-55K (496R) are shown below. The positions of the point mutations in H5pm4133 are indicated, and dots identify identical nucleotides. Asterisks denote the introduced opal and amber codons in the 55K reading frame from H5pm4133. (B) Steady-state expression levels of E1B-55K proteins in mutant and wt virus-infected A549 cells. A549 cells were infected at a multiplicity of 20 focus-forming units per cell. Cells were harvested at the indicated times postinfection (p.i.), and total cell extracts were prepared. Proteins (50-μg samples) from each time point were separated by SDS-12% PAGE and subjected to immunoblotting using anti-E1B-55K mouse MAb 2A6.
TABLE 1.
Primers used in this work
| Primer no. | Primer sequencea |
|---|---|
| 1147 | 5′-GGATAAATGGAGTGAAGAAACCCATCTTAGCGGGGGGTACC-3′ |
| 1148 | 5′-GGTACCCCCCGCTAAGATGGGTTTCTTCACTCCATTTATCC-3′ |
| 1144 | 5′-GGCTGAACTGTATCCATAACTGTGACGCATTTTGACAATTACAG-3′ |
| 1160 | 5′-CTGTAATTGTCAAAATGCGTCACAGTTATGGATACAGTTCAGCC-3′ |
| 1285 | 5′-CGGCAGGAGCAGAGCCCGTGGAACCCGAGAGC-3′ |
| 1286 | 5′-GCTCTCGGGTTCCACGGGCTCTGCTCCTGCCG-3′ |
| 1287 | 5′-GGCCTGGACCCTCGGGAGTGAATGTTGTACAGG-3′ |
| 1288 | 5′-CCTGTACAACATTCACTCCCGAGGGTCCAGGCC-3′ |
| 1476 | 5′-GAGTTTTATAAAGGATAAGTGGAGCGAAGAAACCC-3′ |
| 1477 | 5′-GGGTTTCTTCGCTCCACTTATCCTTTATAAAACTC-3′ |
Underlining indicates mutations introduced by the oligonucleotide primers.
TABLE 2.
Virus mutants used in this work
| Virus | Plasmid | Mutations | Amino acid changes |
|---|---|---|---|
| H5pm4133 | pE1B-1631 | 2025 C → T, 2040 G → T | R3Opa/E8Amb |
| H5pm4149 | NAa | 2025 C → T, 2040 G → T, 2274 G → T, 2280 A → T | R3Opa/E8Amb/E86Och/R88Opa |
| H5pm4177 | pE1B-1670 | 2025 C → T, 2040 G → T, 2208 A → G | R3Opa/E8Amb/M64V |
| H5pm4178 | pE1B-1671 | 2025 C → T, 2040 G → T, 2241 A → G | R3Opa/E8Amb/M75V |
| H5pm4179 | pE1B-1672 | 2025 C → T, 2040 G → T, 2208 A → G, 2241 A → G | R3Opa/E8Amb/M64V/M75V |
NA, not applicable.
For the generation of virus mutants (Table 2), the viral genomes were released from the recombinant plasmids by PacI digestion. Five micrograms of viral DNA was used to transfect 3 × 105 complementing 293 cells by using Lipofectamine. After 5 days, cells were harvested and viruses were released by three cycles of freezing and thawing. The viruses were propagated in 293 monolayer cells and purified by cesium chloride equilibrium density centrifugation as described below. Viral DNAs were isolated from viral particles as described previously (42) and analyzed by HindIII restriction endonuclease digestion. In addition, the viral DNAs were partially sequenced to verify the presence of the E1B-55K mutations.
Antibodies.
Primary antibodies specific for adenovirus proteins used in this study included E1B-55K mouse monoclonal antibody (MAb) 2A6 (41), rat MAb 7C11 (23), rat MAb 4E8 (see below), E2A-72K (DBP) mouse MAb B6-8 (38), L4-100K rat MAb 6B10 (26), E4orf6 rabbit polyclonal antibody 1807 (11), E4orf6 mouse MAb RSA3 (30), and Ad5 rabbit polyclonal serum L133 (23). Primary antibodies specific for cellular proteins included p53 mouse MAb DO-1 (Santa Cruz), ß-actin mouse MAb AC-15 (Sigma-Aldrich, Inc.), and Mre11 rabbit polyclonal antibody pNB 100-142 (Novus Biologicals, Inc.). Secondary antibodies conjugated to horseradish peroxidase for detection of proteins by immunoblotting were donkey anti-rabbit immunoglobulin G (IgG), goat anti-mouse IgG, and sheep anti-rat IgG (Amersham Biosciences). Fluorescent secondary antibodies were affinity purified fluorescein isothiocyanate (FITC)-conjugated donkey anti-mouse IgG and Texas Red-conjugated donkey anti-rat IgG (Dianova). These were used at a 1:100 dilution in all immunofluorescence experiments.
The E1B rat MAb 4E8 was generated against the synthetic peptide TD1 (TEDGQGLKGVKRERGAC), corresponding to amino acids 94 to 110 of the Ad5 E1B-55K protein, and used to immunize LOU/C rats.
Viruses.
H5pg4100 served as the wild-type (wt) Ad5 parent virus in these studies (23). H5pm4149 has been described recently (13) and carries four stop codons in the 55K reading frame (Table 2). The construction of the other E1B mutant viruses used in these studies (Table 2) has been described before. All viruses were propagated in 293 monolayer cultures. Infections were performed in one-fifth of the normal culture volume in DMEM for 2 h at 37°C with gentle rocking every 10 min. The virus suspension was then replaced with normal growth medium. Three to 5 days postinfection, infected cells and supernatants were harvested and freeze-thawed three times to release progeny virions. Virus was harvested by sequential centrifugation in discontinuous and equilibrium cesium chloride gradients exactly as described previously (46). The titers of the viruses used in this study were determined by a fluorescent focus assay exactly as described previously (21).
To measure virus growth, infected cells were harvested at 24 h, 48 h, and 72 h postinfection and lysed by three cycles of freeze-thawing. The cell lysates were serially diluted in DMEM for infection of 293 cells, and virus yield was determined by quantitative E2A-72K immunofluorescence staining at 24 h after infection as described above. Viral DNA replication was determined by quantitative PCR exactly as described previously (39). PCR products were analyzed on a 1% agarose gel and quantitated using the ChemiDoc system and QuantityOne software (Bio-Rad).
Protein analysis.
For protein analysis, cells were resuspended in lysis buffer (50 mM Tris-HCl [pH 8.0], 150 mM NaCl, 5 mM EDTA, 1 mM dithiothreitol, 0.1% sodium dodecyl sulfate [SDS], 1% Nonidet P-40, 0.1% Triton X-100, 0.5% sodium deoxycholate) supplemented with a protease inhibitor cocktail containing 1% (vol/vol) phenylmethylsulfonyl fluoride, 0.1% (vol/vol) aprotinin, 1 μg/ml leupeptin, 1 μg/ml pepstatin, and 1 mM dithiothreitol. After 1 h on ice, the lysates were sonicated and the insoluble debris was pelleted at 15,000 × g at 4°C.
For immunoblotting, equal amounts of total protein were separated by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose membranes (Schleicher & Schüll). Membranes were incubated overnight in phosphate-buffered saline (PBS)-0.1% Tween 20 containing 5% nonfat dry milk and then for 2 h in PBS-1% nonfat dry milk containing the appropriate primary antibody. Membranes were washed three times in PBS-0.1% Tween 20, incubated with a secondary antibody linked to horseradish peroxidase (Jackson ImmunoResearch) in PBS-0.1% Tween 20, and washed three times in PBS-0.1% Tween 20. The bands were visualized by enhanced chemiluminescence as recommended by the manufacturer (Pierce) on X-ray films (CEA RP). Autoradiograms were scanned and cropped using Adobe Photoshop CS4, and figures were prepared using Adobe Illustrator CS4 software.
Indirect immunofluorescence.
For indirect immunofluorescence, cells were grown on glass coverslips and infected as described above. At the indicated times, cells were fixed in ice-cold methanol at −20°C for 15 min and permeabilized in PBS-0.5% Triton X-100 for 30 min at room temperature. After 1 h in Tris-buffered sulfate-BG buffer (20 mM Tris-HCl [pH 7.6], 137 mM NaCl, 3 mM KCl, 1.5 mM MgCl2, 0.05% Tween 20, 0.05% sodium azide, 5 mg/ml glycerol, and 5 mg/ml bovine serum albumin), they were reacted for 1 h with the anti-E1B-55K rat MAb 4E8 or mouse MAb 2A6 diluted to 1:10 in PBS and washed three times in PBS-0.1% Tween 20, followed by incubation with the corresponding FITC- or Texas Red-conjugated secondary antibodies. Coverslips were washes three times in PBS-0.1% Tween 20 and mounted in Glow medium (Energene), and digital images were acquired on a DMRB fluorescence microscope (Leica) with a charge-coupled-device camera (Diagnostic Instruments). Images were cropped using Adobe Photoshop CS4 and assembled with Adobe Illustrator CS4.
Plasmids and transient transfections.
Plasmid pE1B-55K contains the Ad5 E1 region from nt 2019 to 4113 and expresses wt Ad5 E1B-55K and its related products 156R, 93R, and 84R (Fig. 1) under the control of the cytomegalovirus major immediate-early promoter from vector pcDNA3 (Invitrogen). Plasmids pE1B-1631, pE1B-1670, pE1B-1671, and pE1B-1672 were derived from pE1B-55K by site-directed mutagenesis with oligonucleotide primers (Table 1) and contain termination codons (opal and amber) at the third and eighth codons of the 55K reading frame (see Fig. 4A). To eliminate initiation of translation at the second and the third AUG of the E1B-55K mRNA, A-to-G transitions were introduced at nt 2208 (pE1B-1670) and 2241 (pE1B-1671) by site-directed mutagenesis with complementary oligonucleotide pairs as described above (Table 1). For the construction of pE1B-1672, the mutation at nt 2241 was introduced into pE1B-1670 by site-directed mutagenesis with oligonucleotide primers (Table 1). The mutations were confirmed by DNA sequencing. Plasmid pC53-SN3 produces human wt p53 (14), and pRE-LUC contains five p53-binding sites in front of the RGC firefly luciferase reporter (16, 17, 22).
FIG. 4.
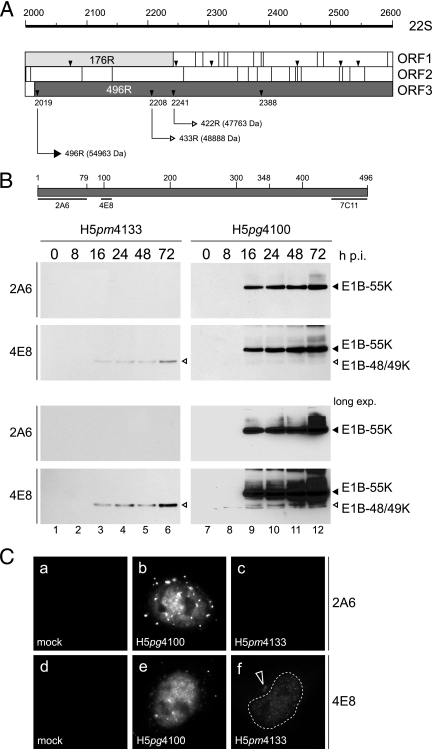
wt H5pg4100 and H5pm4133 express a novel 49-kDa isoform of the E1B-55K product. (A) Schematic drawing of the E1B-55K coding region surrounding the starting methionine at nucleotide position 2019. Numbers refer to positions of nucleotides in the wt E1B 22S mRNA. Open reading frame (ORF) maps spanning the region from nt 2000 to 2600 of the E1B 22S mRNA are shown. The three reading frames (ORF1 to ORF3) of the E1B sense strand are shown below. Gray-shaded regions indicate parts of the 19K and 55K protein coding sequences. Vertical lines bisecting each reading frame indicate stop codons, whereas small triangles denote methionine codons. Numbers below refer to nucleotide positions of the methionine codons, and thin-lined arrows indicate translation of the corresponding E1B 496R, 433R, and 422R polypeptides. Numbers in parentheses designate the theoretical molecular mass calculated from the protein sequences. (B) Top: schematic drawing of the E1B-55K 496R protein. Numbers refer to positions of amino acid residues in the viral polypeptide, and regions of 55K recognized by MAbs 2A6, 4E8, and 7C11 are indicated below by thin lines. Bottom: Immunoblot analysis of total-cell lysates from wt and mutant virus-infected cells. A549 cells were infected at a multiplicity of 50 FFU per cell. Cells were harvested at the indicated times postinfection, and total cell extracts were prepared. Proteins (50-μg samples) from each time point were separated by SDS-12% PAGE and subjected to immunoblotting using anti-E1B-55K mouse MAb 2A6 or anti-E1B-55K rat MAb 4E8. The positions of the E1B proteins are indicated on the right, and their molecular masses were estimated from their electrophoretic mobilities. Long exp., longer exposure. (C) Steady-state localization of E1B-55K and its isoforms in wt and mutant virus-infected cells. A549 cells were mock infected or infected with wt or E1B mutant virus H5pm4133 at a multiplicity of 50 FFU per cell. Cells were fixed at 24 h after infection and labeled with anti-E1B-55K mouse MAb 2A6 (panels a, b, and c) or anti-E1B-55K rat MAb 4E8 (panels d, e, and f) and FITC- and Texas Red-conjugated secondary antibodies. In panel d, the nucleus is indicated by a dotted line. Magnification, ×7,600.
For dual luciferase assays, subconfluent H1299 cells were transfected as described previously (33), using the indicated amounts of reporter and effector plasmids and 1.0 μg of pRL-TK (Promega), which expresses Renilla luciferase under the control of the herpes simplex virus thymidine kinase promoter. Total cell extracts were prepared 48 h after transfection in lysis buffer, and RGC firefly luciferase activity was assayed with 10 μl of extract in an automated luminometer (Lumat LB9510; Berthold). All samples were normalized for transfection efficiency by measuring Renilla luciferase activity.
RESULTS
Construction and phenotypic analysis of the viral mutant H5pm4133.
As part of our studies of E1B-55K function during productive infection, we generated mutant virus H5pm4133, which contains stop codons (opal and amber) after the second and seventh codon of the E1B-55K reading frame (Fig. 2). We expected that these mutations would eliminate the synthesis of E1B-55K and its related products derived from the 13S, 14S, and 14.5S mRNAs without affecting the integrity of the E1B-19K (E1B-176R) protein. As anticipated, no signal was obtained with total extracts from A549 cells infected with H5pm4133, while E1B-55K (496R) and its related forms E1B-156R, E1B-93R, and E1B-84R were detectable in wt H5pg4100-infected cells with MAb 2A6, recognizing an epitope in the amino-terminal region of the adenovirus proteins (43) (Fig. 2; see Fig. 4B).
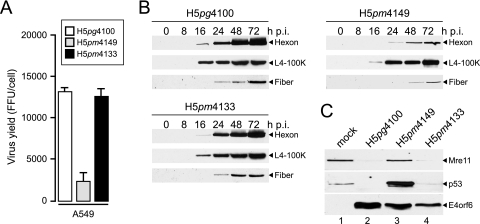
To assess the effect of mutations on overall virus growth properties, total virus yield was determined in A549 cells and compared to that in wt H5pg4100 and H5pm4149-infected cells (Fig. 3A). In line with previous reports, the virus mutant H5pm4149, which lacks E1B-55K and its related proteins (13) (Table 2), was defective for growth in A549 cells compared to the wt virus. Unexpectedly, however, H5pm4133 grew to levels comparable to that of wt H5pg4100. Identical results (not shown) were obtained from analyses using HeLa and H1299 cells. Given these findings, infected-cell extracts were analyzed for late viral protein expression by immunoblotting (Fig. 3B). Extracts were prepared at different times after infection and probed for expression of hexon, L4-100K, and fiber. In agreement with the data obtained in the virus growth experiments, H5pm4149 showed substantially reduced expression of hexon and fiber whereas H5pm4133 produced the viral late gene products at levels similar to those of the wt virus. These results, along with the observation that H5pm4133 was not impaired in the onset of viral DNA synthesis and viral DNA accumulation (data not shown), indicate that this mutant expresses some form of the E1B-55K protein that is sufficient for a productive infection. Further support for this idea came from studies where we examined the proteasomal degradation of p53 and Mre11 by the viral E1B-55K/E4orf6/E3 ubiquitin-ligase complex during infection (37, 44) (Fig. 3C). Immunoblotting of total extracts from infected A549 cells showed that the steady-state concentrations of Mre11 and p53 were significantly reduced in late wt- and H5pm4133 mutant virus-infected cells, whereas no reduction of Mre11 and p53 levels was observed at the same time point in A549 cells infected with H5pm4149.
FIG. 3.
Virus growth and viral late protein synthesis. (A) A549 cells were infected with wt H5pg4100 or E1B mutant viruses at a multiplicity of 50 focus-forming units (FFU) per cell. Viral particles were harvested 72 h postinfection, and virus yield was determined by quantitative E2A-72K immunofluorescence staining on 293 cells. The results represent the averages from three independent experiments. Error bars indicate the standard errors of the means. (B) Viral late protein synthesis. A549 cells were infected with the wt or mutant viruses at a multiplicity of 50 FFU per cell. Total cell extracts were prepared at the indicated times postinfection (p.i.). Proteins (10-μg samples) were separated by SDS-12% PAGE, transferred to nitrocellulose membranes, and probed with the anti-Ad5 rabbit polyclonal serum L133. Bands corresponding to the hexon, L4-100K, and fiber proteins are indicated on the right. (C) Steady-state concentrations of Mre11, p53, and E4orf6 in infected cells. A549 cells were infected with wt H5pg4100 or E1B mutant viruses at a multiplicity of 50 FFU per cell. Whole-cell extracts were prepared at 48 h after infection, and proteins (50-μg samples) were separated on SDS-12% polyacrylamide gels and subjected to immunoblotting using anti-Mre11 rabbit polyclonal antibody pNB-100-142, anti-p53 mouse MAb DO-1, or anti-E4orf6 mouse MAb RSA3.
wt Ad5 and H5pm4133 express several isoforms of the large E1B protein.
The results from these studies were highly reminiscent of data from previous work by Barker and Berk with the Ad2/5 chimeric E1B-55K mutant pm2022 (5). This mutant contains a C-to-T transition in the Ad2 E1B-55K gene, which converts the third codon of the reading frame to an opal stop codon, thereby eliminating synthesis of 55K after only two amino acids. Biochemical studies showed that pm2022 expresses very low levels of several truncated forms of the Ad2 E1B-55K protein in the 46-to-50-kDa range, which are possibly synthesized by reinitiating translation at the second or third downstream AUG of the 22S E1B mRNA (Fig. 4A). More significantly, it appeared that these isoforms are sufficient to proceed through the productive infection in HeLa cells (5).
These observations, together with our results described above, prompted us to investigate whether similar polypeptides are expressed in H5pm4133-infected A549 cells (Fig. 4B). Total-cell lysates were prepared at different times after infection with H5pm4133 and wt H5pg4100, and immunoblots were probed with mouse MAb 2A6 or rat MAb 4E8 (Fig. 4B), which was raised against a synthetic peptide derived from the central region of the viral protein (Fig. 4B). As before, E1B-55K was seen with MAb 2A6 in lysates from wt- but not from H5pm4133-infected cells. The E1B-55K protein was also detected with MAb 4E8 in wt H5pg4100-infected cells (Fig. 4B, lanes 9 to 12). Significantly, in the same immunoblots this MAb reacted with low levels of at least two faster-migrating bands in the 48-to-49-kDa range. When larger amounts of total extracts from wt-infected cells were analyzed by immunoblotting with MAb 4E8, two additional bands in the range of 52 kDa to 53 kDa were detected (Fig. 5B). Identical results were obtained from analyses employing rat MAb 7C11 (see Fig. 7B), recognizing an epitope in the carboxy-terminal region of Ad5 E1B-55K (Fig. 4B), and other wt Ad5 strains, such as H5wt300 and H5dl309. More importantly, a ∼49-kDa band was also present in the extracts from H5pm4133-infected cells (Fig. 4B, lanes 3 to 6), demonstrating that wt and Hpm4133 mutant virus-infected A549 cells contain a 48/49-kDa isoform of the large E1B product which lacks the amino-terminal epitope of MAb 2A6 but includes the central and carboxy-terminal regions of the adenovirus protein. In support of this, no staining of E1B-55K was observed in H5pm4133-infected A549 cells by indirect immunofluorescence with MAb 2A6 (Fig. 4C, panel c). In contrast, a weak and diffuse nuclear staining was evident with MAb 4E8 in the mutant virus-infected cells (Fig. 4C, panel f). In addition, in all of these cells examined (n > 50), MAb 4E8 exhibited a faint cytoplasmic condensation in close proximity to the nuclear membrane, a characteristic of large E1B proteins from subgroup C adenoviruses.
FIG. 5.
The AUGs at positions 64 and 75 initiate translation of two E1B isoforms. (A) Mutations in the 5′ coding region of the E1B-55K mutant viruses (Virus) and plasmid constructs (Plasmid). In wt virus H5pg4100 and plasmid pE1B-55K (indicated on the right), numbers refer to positions of nucleotides in the Ad5 wt DNA sequence and numbers after the diagonal slash denote the positions of the amino acid residues. The first three initiation codons of the 55K reading frame are underlined. The amino acid sequence is shown below. The positions of the point mutations in the virus mutants and plasmid constructs are indicated, and dots identify identical nucleotides and amino acid residues, respectively. Asterisks denote the introduced opal, amber, and ochre codons in the 55K reading frame. (B) Steady-state expression levels of E1B-55K proteins in wt and mutant virus-infected cells. A549 cells were infected with a multiplicity of 50 FFU per cell. Whole-cell extracts were prepared, and samples containing 50 μg of protein were examined by SDS-12% PAGE followed by immunoblotting with antibody 2A6 (left) or 4E8 (right). Lower panels represent longer exposures (long exp.) of the same immunoblots to visualize expression of the 422R E1B isoform in H5pm4177-infected cells. The designations of the viruses and plasmids are indicated on the right. The asterisk in lane 9 indicates two bands in the 52-to-53-kDa range.
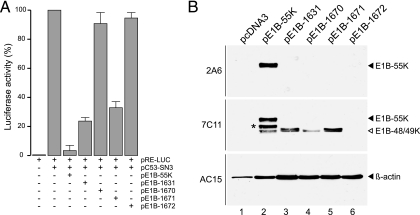
FIG. 7.
Effect of mutations on inhibition of p53-stimulated transcription. (A) Subconfluent H1299 cells were transfected with reporter and effector plasmids as described in Materials and Methods. The mean and standard deviation are presented for three independent experiments, each performed in duplicate. (B) Expression of E1B proteins in transfected H1299 cells. Luciferase samples were immunoblotted by normalizing the amount of extract used according to Renilla luciferase activity and probing the immunoblots with MAbs 2A6, 7C11, and AC-15. The asterisk in lane 2 indicates a slower-migrating band in the 52-to-53-kDa range.
Taken together with earlier results from Barker and Berk, these results show that early regions 1B of subgroup C Ad2 and Ad5 produce several isoforms of the large E1B product in the 48-to-53 kDa-range, all of which lack the amino-terminal epitope of MAb 2A6. The apparent molecular masses of the 48-kDa and 49-kDa species are consistent with translation initiation at the second or third in-frame AUG of the 55K reading frame (Fig. 4A), which would yield polypeptides of 422R (47,763 Da) and 433R (48,888 Da), respectively.
Two downstream AUGs of the E1B-55K reading frame initiate translation of the E1B-49K isoform.
To test this possibility, we generated a set of mutant viruses where the AUGs at positions 64 (H5pm4177), 75 (H5pm4178), and 64 plus 75 (H5pm4179) were converted to valine codons in the opal3/amber8 background (Fig. 5A). A549 cells were infected with wt and mutant viruses, and E1B protein expression was tested by immunoblotting using MAbs 2A6 and 4E8 (Fig. 5B). In total-cell lysates prepared from wt virus-infected cells, MAb 2A6 detected the full-length 496R E1B-55K protein (Fig. 5B, lane 2). Consistent with the data obtained in the immunoblots shown in Fig. 4B, no 2A6-positive signals were seen in the total-cell lysates from infections with H5pm4133 (Fig. 5B, lane 4) and the other E1B virus mutants containing the opal3 and amber8 mutations (Fig. 5B, lanes 3 and 5 to 7). As before (Fig. 4B), a ∼49-kDa form was observed with MAb 4E8 in total-cell lysates from A549 cells infected with H5pm4133 and, significantly, H5pm4178, which lacks the third in-frame AUG at position 75. Extremely low steady-state levels of this band were evident in H5pm4177-infected cells (Fig. 5B, lane 12), whereas no specific signals were reproducibly detected in total-cell lysates from cells infected with H5pm4149 (Fig. 5B, lane 10) and H5pm4179 (Fig. 5B, lane 14) lacking both downstream AUGs. A 4E8-positive band in the 48-to-49-kDa range was also evident in total-cell extracts from wt virus-infected cells (Fig. 5B, lane 9). Finally, in the same extracts, MAb 4E8 reacted with E1B-55K and two bands in the 52-to-53-kDa range (Fig. 5B, lane 9), which were reproducibly not observed in infections with the mutant viruses containing the opal3/amber8 mutations.
Altogether, these data show that the smaller form of ∼49 kDa present in lysates from infections with H5pm4133 consists of two closely migrating proteins of 433R and 422R and confirm that translation of these isoforms initiates primarily at AUG 64 and, probably to a much lesser extent, at AUG 75. Whether the same downstream AUGs are used as the initiating methionines for the 48/49-kDa and 52/53-kDa forms in wt H5pg4100-infected cells is unclear.
The E1B-49K protein is sufficient to promote virus replication.
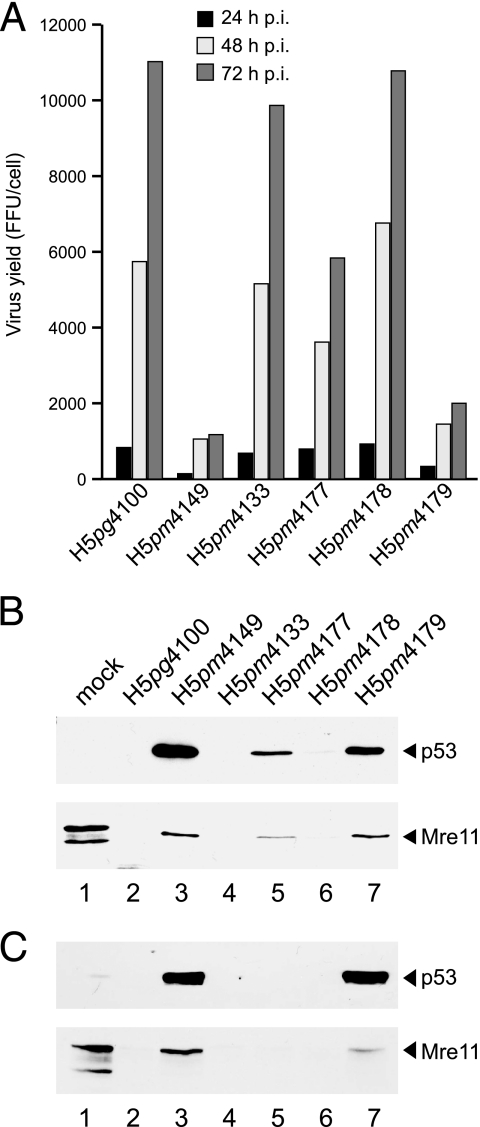
To reveal the effect of mutations on virus growth, we determined the virus yield from wt and E1B mutant virus-infected A549 cells (Fig. 6A). As expected, H5pm4149, lacking all E1B-55K functions, was severely defective for virus growth, while replication of H5pm4133 (R3Opa/E8Amb) and, significantly, H5pm4178 (R3Opa/E8Amb/M75V) expressing the 433R species was comparable to that of wt H5pg4100. In contrast, H5pm4177 (R3Opa/E8Amb/M64V), lacking the AUG at position 64, was moderately defective, whereas H5pm4179 (R3Opa/E8Amb/M64V/M75V), lacking both downstream AUGs, replicated to levels nearly reduced to those of H5pm4149. Thus, the low steady-state concentrations of the 433R E1B isoform and to a much lesser extent of the 422R species are sufficient to proceed through a productive infection. In line with this result, the shorter isoform was less efficient in the proteasomal degradation of p53 and Mre11 via the E4orf6/E3 ubiquitin-ligase complex than the 433R species (Fig. 6B, lanes 5 and 6), which was as effective as the wt 496R E1B protein (Fig. 6B, lane 2). However, the 422R form was as efficient as the 433R product at degrading p53 and Mre11 when steady-state concentrations of 422R were elevated to levels comparable levels to those of 433R by increasing the multiplicity of infection 10-fold (Fig. 6C, lane 5).
FIG. 6.
Effect of amino acid substitutions in E1B-55K on virus growth and proteasomal degradation of p53 and Mre11. (A) A549 cells were infected with wt H5pg4100 or E1B mutant viruses at a multiplicity of 50 FFU per cell. Viral particles were harvested at the indicated time points after infection, and virus yield was determined by quantitative E2A-72K immunofluorescence staining on 293 cells. The results represent the averages from two independent experiments. FFU, focus-forming units. (B) A549 cells were infected as described above, and whole-cell extracts were prepared. Proteins (50-μg samples) from the 48-h-p.i. time point were separated on SDS-12% polyacrylamide gels and subjected to immunoblotting using anti-p53 mouse MAb DO-1 or anti-Mre11 rabbit polyclonal antibody pNB-100-142. (C) A549 cells were infected at a multiplicity of 50 FFU per cell (H5pg4100, H5pm4149, H5pm4133, H5pm4178, and H5pm4179) or 500 FFU per cell (H5pm4177), and steady-state concentrations of p53 and Mre11 were determined by immunoblotting using MAbs DO-1 and pNB-100-142, respectively.
The E1B-49K protein can block p53-stimulated transcription.
To evaluate the ability of the E1B isoforms to repress p53 transactivation, we introduced mutations in plasmid pE1B-55K, containing the wt E1B region, from nt 2019 to 4113 in pcDNA3 (Fig. 5A). As in the E1B virus mutants, the third and eighth codons of the E1B-55K reading frame were changed in all constructs into an opal and amber stop codon (R3Opa/E8Amb), respectively. Additionally, the downstream AUGs at positions 64 (pE1B-1670), 75 (pE1B-1671), and 64 plus 75 (pE1B-1672) were converted to valine codons (Fig. 5A). The E1B expression plasmids were then cotransfected with a luciferase reporter (pRE-LUC) containing five p53-binding sites into human p53-negative H1299 cells (Fig. 7).
Cotransfection of pRE-LUC with a plasmid expressing human wt p53 (pC53-SN3) substantially enhanced expression severalfold (Fig. 7A), whereas inclusion of a third plasmid (pE1B-55K) expressing wt E1B-55K and the smaller 7C11-positive isoforms (Fig. 7B, lane 2) blocked the ability of p53 to enhance expression from the reporter approximately 20-fold. Inhibition of p53-stimulated transcription in the range of 70% to 80% was observed with pE1B-1631 (R3Opa/E8Amb) and pE1B-1671 (R3Opa/E8Amb/M75V), producing the 433R/422R and 433R proteins, respectively. We note that the E1B-433R protein migrated as a doublet in these assays (Fig. 7B, lanes 3 and 5). No negative effect on p53 transactivation was seen in cotransfections with plasmids lacking AUG64 (pE1B-1670) and AUG 64 plus AUG 75 (pE1B-1672). Consistent with this, no E1B protein expression was detected in the immunoblots with MAbs 2A6 and 7C11 in pE1B-1672-transfected cells (Fig. 7B, lane 6). In contrast, MAb 7C11 reacted with very low levels of a single band in total lysates from pE1B-1670-transfected cells (Fig. 7B, lane 4), most probably corresponding to the 422R species. Hence, the observed increase in promoter activity with pE1B-1670 is likely due to the reduced expression levels of the 422R isoform. Interestingly, as before (Fig. 5), MAb 7C11 also detected a slower-migrating band in the 52-to-53-kDa range (Fig. 7B, lane 2), which, analogous to virus-infected cells (Fig. 5), was clearly absent in cell lysates from transfections with the different mutant constructs.
Taken together, these studies show that plasmid-transfected cells express essentially the same set of E1B isoforms as wt and mutant virus-infected cells. Thus, internal translation initiation at AUG 64 and AUG 75 is independent of other viral gene products and importantly from sequence elements upstream of the first AUG of the 22S mRNA (nt position 2019), which are absent in the pcDNA3 vector background. Moreover, the 433R protein can functionally replace the 55K product in the inhibition of p53 transactivation, showing that the amino-terminal 63 amino acids of E1B-55K are dispensable for this activity.
DISCUSSION
Subgroup C Ad2/5 early region 1B encodes at least five different gene products generated by translation of several alternatively spliced mRNAs (Fig. 1). Translation of the major 22S mRNA uses discrete but overlapping reading frames to produce E1B-19K and E1B-55K, the two principal proteins of the E1B region. This report demonstrates that this polycistronic transcript directs the synthesis of several additional proteins, which migrate with apparent molecular masses of 48 kDa to 53 kDa in SDS-polyacrylamide gels. All of these products lack the amino-terminal epitope of MAb 2A6 but contain segments from the central and carboxy-terminal parts of the 496R 55K protein (Fig. 4, 5, and 7). These data extend results from previous work that several smaller isoforms of E1B-55K exist in Ad2/5-infected cells (2, 5, 28, 45, 48) and demonstrate for the first time that at least one of these polypeptides, with a molecular mass of approximately 49 kDa, can support efficient virus replication in human tumor cells. Furthermore, our data provide an explanation for the findings described by Barker and Berk (5) and identify the molecular basis underlying the phenotype of E1B virus mutants containing stop codons in the 5′ coding region of the 55K mRNA, such as the Ad2/5 chimeric E1B virus mutant pm2022 (5).
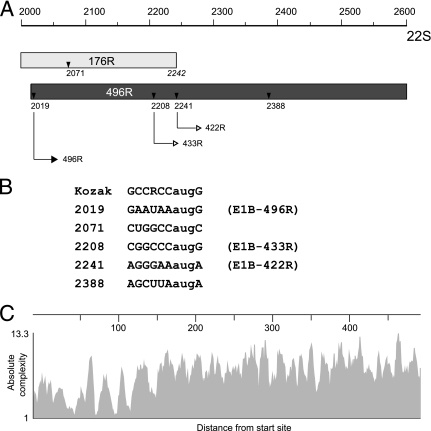
Results from genetic and biochemical studies show that the ∼49-kDa species seen in the opal3/amber8 background consists of two closely migrating polypeptides of 433R and 422R, which differ substantially in their steady-state concentrations and in their ability to support virus growth (Fig. 5, 6, and 7). Both isoforms are produced from internal initiation sites at nt 2208 (AUG 64) and 2241 (AUG 75) (Fig. 8A). Thus, both downstream AUGs of the 55K reading frame can be recognized as initiating codons by one or more processes currently known to bypass the conventional scanning mechanism, including ribosome shunting, internal ribosome entry, leaky scanning, and translation reinitiation (24, 40). Leaky scanning is possible because translation of the full-length 496R protein is likely initiated by this process and because the sequences flanking the AUGs (nt 2019 and 2071) do not conform to the Kozak consensus sequence, whereas the AUG at nt 2208 is in a good Kozak context (Fig. 8B). A similar scenario has been described for the formation of a truncated 40-kDa isoform of the Ad5 IV2a protein (35). Hence, the 40S ribosomal subunits might bypass the AUG codons at nt 2019 and 2071 and initiate instead at AUG 2208 and probably much more rarely at AUG 2241. In fact, reduced initiation at this start codon may explain the apparent differences in steady-state concentrations between the 433R and 422R isoforms (Fig. 5 and 7). Reinitiation at AUG 2208 and/or AUG 2241 in combination with leaky scanning could also occur because the opal3 mutation in combination with the AUG codon at nt 2019 may provide a small upstream open reading frame necessary for translation reinitiation of the 40S subunit at downstream AUGs (24, 25). Other mechanisms, such as internal ribosome entry or ribosome shunting, are also possible, in particular because the corresponding region of the 22S E1B mRNA (nt 2000 to 2260) allows the prediction of a highly structured sequence element (data not shown), typically required for translation initiation by internal ribosome entry or ribosome shunting. Further experiments are now under way to test these hypotheses.
FIG. 8.
Ad5 E1B transcription unit may encode two additional isoforms of the E1B-55K protein. (A) Schematic drawing of the E1B-55K coding region from nt 2000 to 2600. Numbers refer to positions of nucleotides in the wt E1B 22S mRNA. Gray-shaded regions indicate parts of the 19K (176R) and 55K (496R) coding sequences. Small triangles denote methionine codons. Numbers below refer to nucleotide positions of the methionine codons, and thin-lined arrows indicate translation of the corresponding E1B 496R, 433R, and 422R polypeptides. The position of the translation stop site of E1B-176R is shown in italics below the coding sequence. (B) Comparison of sequences flanking the AUG codons at positions 2019, 2071, 2208, 2241, and 2238 with the Kozak consensus sequence. R, adenine or guanine. (C) Absolute complexity (y axis) of aligned E1B sequences from subgroup A to G adenoviruses was calculated using the Vector NTI software program (Invitrogen) in terms of average pairwise alignment scores. Higher values in the y axis show higher sequence conservation. The x axis indicates the distance from the start site. Amino acid sequences of large E1B proteins were derived from Ad subtypes 1, 2, 3, 4, 5, 7, 8, 9, 11, 12, 14, 16, 17, 19, 21, 26, 34, 35, 37, 40, 41, 46, 48, 49, 50, 52, and 53.
At present, it is unclear whether the same downstream AUGs of the 55K reading frame serve as internal translation initiation sites for the production of the isoforms seen in the wt context. In cells infected with H5pg4100, we observe at least four polypeptides in the 48-to-53-kDa range, including a smaller form of 48 to 49 kDa, which may correspond to the 433R/422R isoforms present in the opal3 and amber8 background. Similar to the situation with the mutant virus, formation of these isoforms is independent of other viral proteins, since a similar set of bands is evident in immunoblots from plasmid-based transfection assays (Fig. 7B). Also, all of these proteins lack the epitope of MAb 2A6 and thus lack all or part of the amino-terminal 63 amino acids of the full-length 496R 55K protein (Fig. 5 and 7). Moreover, preliminary data (not shown) indicate that a virus mutant lacking AUG 64 plus AUG 75 (M64V/M75V) produces the 52/53-kDa but not the 48/49-kDa species. We therefore tentatively conclude that AUG 64 and/or 75 are used in the wt context and direct the synthesis of the 48/49-kDa species by one or more processes that allow internal translation initiation as discussed before. Consequently, a different scenario must underlie the formation of the 52/53-kDa proteins. One possibility is that translation of the 52/53-kDa proteins initiates from a non-AUG codon (CUG, ACG, or GUG) (47) located upstream from AUG codon 64. In fact, the 55K reading frame from wt Ad5 contains a single GUG codon at nt 2079, and initiation at this codon would give a product of the expected size of approximately 52 kDa (476R). Assuming that translation can initiate at this non-AUG codon, the 53-kDa species could represent a different posttranslationally modified form of the 52-kDa protein. Nevertheless, it remains unclear why initiation is abrogated in the presence of the opal3 and amber8 codons (Fig. 5 and 7), since the 52/53-kDa forms are absent in infections and transfections with the corresponding mutant viruses or plasmids (Fig. 5 and 7). Hence, another possibility is that these proteins are generated by proteolytic cleavage, thereby eliminating the 2A6 epitope. Although the amino-terminal region of the 55K protein contains numerous potential cleavage sites cleaved by proteases, none of them would yield a product which matches the observed size of both truncated E1B proteins.
Given the very high steady-state concentrations of E1B-55K typically present in wt Ad2/5-infected cells, probably the most striking observation from this study is that virus mutants producing low levels of the 433R isoform replicate nearly as efficiently as the wt virus in human tumor cell lines (Fig. 6). We consider it unlikely that a read-through of both stop codons at low levels and production of full-length E1B-55K at undetectable quantities constitute the mechanism underlying the wt phenotype of these virus mutants. The different growth properties of the mutant viruses containing the opal3 and amber8 codons (Fig. 6A), the observed differences in their ability to facilitate proteasomal degradation of p53 and Mre11 (Fig. 6B), and the data from the reporter gene assays (Fig. 7) are not compatible with a low-level read-through scenario.
Assuming that read-through of two consecutive stop codons is very rare, results from this study further suggest that low concentrations of E1B-55K protein activity are sufficient for a productive infection in tumor cell lines and indicate that the amino-terminal 63 amino acids of the E1B-55K protein are dispensable for virus growth. This is consistent with the fact that the amino-terminal segment (amino acids 1 to 100) is the least conserved region among the large E1B proteins from the seven different human subgroups (Fig. 8C) and the observation that the 433 amino acids of this isoform comprise all currently known structural and functional domains critical for 55K's early and late lytic functions (recently reviewed in reference 7). In support of this, the 433R E1B protein retains the ability to block p53-stimulated transcription in transient reporter gene assays (Fig. 7) and to localize in one or more cytoplasmic bodies in H5pm4133-infected cells, a characteristic of large E1B proteins from subgroup C Ad2/5 (Fig. 4C, panel f). These structures likely correspond to aggresomes, recently implicated in the inactivation and rapid degradation of MRN components by E1B-55K (3, 29). Nonetheless, given the apparent differences between the steady-state levels of the 496R protein and the 433R isoform, it is tempting to speculate that the amino-terminal 63 amino acids contain elements that augment the half-life of the full-length product and thus contribute to the high steady-state concentrations of E1B-55K typically present in wt virus-infected cells. Additionally and/or alternatively, it is possible that this region includes a yet-unknown function required for efficient virus growth, which is compensated by some cellular activity in human tumor cells infected with the opal3/amber8-containing viruses. Therefore, given the central role of E1B-55K in productive infection and tumor-specific adenovirus replication (1), the mutants described here, expressing the 433R isoform alone, may also prove useful in the development of adenovirus-based cancer therapeutics and should be tested in the context of primary human cells.
Acknowledgments
We thank Hannah Götze and Sylvia Allmeier for able technical assistance.
This work was supported by the Erich and Gertrud Roggenbuck-Foundation, Hamburg, Germany.
Footnotes
Published ahead of print on 8 July 2009.
REFERENCES
- 1.Alemany, R. 2007. Cancer selective adenoviruses. Mol. Aspects Med. 28:42-58. [DOI] [PubMed] [Google Scholar]
- 2.Anderson, C. W., R. C. Schmitt, J. E. Smart, and J. B. Lewis. 1984. Early region 1B of adenovirus 2 encodes two coterminal proteins of 495 and 155 amino acid residues. J. Virol. 50:387-396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Araujo, F. D., T. H. Stracker, C. T. Carson, D. V. Lee, and M. D. Weitzman. 2005. Adenovirus type 5 E4orf3 protein targets the Mre11 complex to cytoplasmic aggresomes. J. Virol. 79:11382-11391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Baker, A., K. J. Rohleder, L. A. Hanakahi, and G. Ketner. 2007. The adenovirus E4 34k and E1b 55k oncoproteins target host DNA ligase IV for proteasomal degradation. J. Virol. 13:7034-7040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Barker, D. D., and A. J. Berk. 1987. Adenovirus proteins from both E1B reading frames are required for transformation of rodent cells by viral infection and DNA transfection. Virology 156:107-121. [DOI] [PubMed] [Google Scholar]
- 6.Berk, A. J. 2005. Recent lessons in gene expression, cell cycle control, and cell biology from adenovirus. Oncogene 24:7673-7685. [DOI] [PubMed] [Google Scholar]
- 7.Blackford, A. N., and R. J. Grand. 11 February 2009. Adenovirus E1B-55K: a protein with multiple roles in viral infection and cell transformation. J. Virol. doi: 10.1128/JVI.02417-08. [DOI] [PMC free article] [PubMed]
- 8.Blanchette, P., and P. E. Branton. 2009. Manipulation of the ubiquitin-proteasome pathway by small DNA tumor viruses. Virology 384:317-323. [DOI] [PubMed] [Google Scholar]
- 9.Blanchette, P., C. Y. Cheng, Q. Yan, G. Ketner, D. A. Ornelles, T. Dobner, R. C. Conaway, J. W. Conaway, and P. E. Branton. 2004. Both BC-box motifs of adenovirus protein E4orf6 are required to assemble an E3 ligase complex that degrades p53. Mol. Cell. Biol. 24:9619-9629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Blanchette, P., K. Kindsmüller, P. Groitl, F. Dallaire, T. Speiseder, P. E. Branton, and T. Dobner. 2008. Control of mRNA export by adenovirus E4orf6 and E1B55K proteins during productive infection requires the E4orf6 ubiquitin ligase activity. J. Virol. 82:2642-2651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Boivin, D., M. R. Morrison, R. C. Marcellus, E. Querido, and P. E. Branton. 1999. Analysis of synthesis, stability, phosphorylation, and interacting polypeptides of the 34-kilodalton product of open reading frame 6 of the early region 4 protein of human adenovirus type 5. J. Virol. 73:1245-1253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bos, J. L., L. J. Polder, R. Bernards, P. I. Schrier, P. J. van den Elsen, A. J. van der Eb, and H. van Ormondt. 1981. The 2.2 kb E1b mRNA of human Ad12 and Ad5 codes for two tumor antigens starting at different AUG triplets. Cell 27:121-131. [DOI] [PubMed] [Google Scholar]
- 13.Dallaire, F., P. Blanchette, P. Groitl, T. Dobner, and P. E. Branton. 2009. Identification of integrin α3 as a new substrate of the adenovirus E4orf6/E1B55K E3 ubiquitin ligase complex. J. Virol. 83:5329-5338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dobner, T., N. Horikoshi, S. Rubenwolf, and T. Shenk. 1996. Blockage by adenovirus E4orf6 of transcriptional activation by the p53 tumor suppressor. Science 272:1470-1473. [DOI] [PubMed] [Google Scholar]
- 15.Dobner, T., and J. Kzhyshkowska. 2001. Nuclear export of adenovirus RNA. Curr. Top. Microbiol. Immunol. 259:25-54. [DOI] [PubMed] [Google Scholar]
- 16.Endter, C., B. Hartl, T. Spruss, J. Hauber, and T. Dobner. 2005. Blockage of CRM1-dependent nuclear export of the adenovirus type 5 early region 1B 55-kDa protein augments oncogenic transformation of primary rat cells. Oncogene 24:55-64. [DOI] [PubMed] [Google Scholar]
- 17.Endter, C., J. Kzhyshkowska, R. Stauber, and T. Dobner. 2001. SUMO-1 modification required for transformation by adenovirus type 5 early region 1B 55-kDa oncoprotein. Proc. Natl. Acad. Sci. USA 98:11312-11317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Flint, S. J., and R. A. Gonzalez. 2003. Regulation of mRNA production by the adenoviral E1B 55-kDa and E4 Orf6 proteins. Curr. Top. Microbiol. Immunol. 272:287-330. [DOI] [PubMed] [Google Scholar]
- 19.Gonzalez, R. A., and S. J. Flint. 2002. Effects of mutations in the adenoviral E1B 55-kilodalton protein coding sequence on viral late mRNA metabolism. J. Virol. 76:4507-4519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Graham, F. L., J. Smiley, W. C. Russel, and R. Nairn. 1977. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J. Gen. Virol. 36:59-72. [DOI] [PubMed] [Google Scholar]
- 21.Groitl, P., and T. Dobner. 2007. Construction of adenovirus type 5 early region 1 and 4 virus mutants, p. 29-39. In W. S. Wold and A. E. Tollefson (ed.), Adenovirus methods and protocols, 2nd edition ed., vol. 1. Humana Press Inc., Totowa, NJ. [DOI] [PubMed] [Google Scholar]
- 22.Härtl, B., T. Zeller, P. Blanchette, E. Kremmer, and T. Dobner. 2008. Adenovirus type 5 early region 1B 55-kDa oncoprotein can promote cell transformation by a mechanism independent from blocking p53-activated transcription. Oncogene 27:3673-3684. [DOI] [PubMed] [Google Scholar]
- 23.Kindsmuller, K., P. Groitl, B. Hartl, P. Blanchette, J. Hauber, and T. Dobner. 2007. Intranuclear targeting and nuclear export of the adenovirus E1B-55K protein are regulated by SUMO1 conjugation. Proc. Natl. Acad. Sci. USA 104:6684-6689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kozak, M. 1999. Initiation of translation in prokaryotes and eukaryotes. Gene 234:187-208. [DOI] [PubMed] [Google Scholar]
- 25.Kozak, M. 2002. Pushing the limits of the scanning mechanism for initiation of translation. Gene 299:1-34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kzhyshkowska, J., E. Kremmer, M. Hofmann, H. Wolf, and T. Dobner. 2004. Protein arginine methylation during lytic adenovirus infection. Biochem. J. 383:259-265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lethbridge, K. J., G. E. Scott, and K. N. Leppard. 2003. Nuclear matrix localization and SUMO-1 modification of adenovirus type 5 E1b 55K protein are controlled by E4 Orf6 protein. J. Gen. Virol. 84:259-268. [DOI] [PubMed] [Google Scholar]
- 28.Lewis, J. B., and C. W. Anderson. 1987. Identification of adenovirus type 2 early region 1B proteins that share the same amino terminus as do the 495R and 155R proteins. J. Virol. 61:3879-3888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu, Y., A. Shevchenko, A. Shevchenko, and A. J. Berk. 2005. Adenovirus exploits the cellular aggresome response to accelerate inactivation of the MRN complex. J. Virol. 79:14004-14016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Marton, M. J., S. B. Baim, D. A. Ornelles, and T. Shenk. 1990. The adenovirus E4 17-kilodalton protein complexes with the cellular transcription factor E2F, altering its DNA-binding properties and stimulating E1A-independent accumulation of E2 mRNA. J. Virol. 64:2345-2359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mitsudomi, T., S. M. Steinberg, M. M. Nau, D. Carbone, D. D'Amico, H. K. Bodner, H. K. Oie, R. I. Linnoila, J. L. Mulshine, J. D. Minna, and A. F. Gazdar. 1992. p53 gene mutations in non-small-lung cell cancer cell lines and their correlation with the presence of ras mutations and clinical features. Oncogene 7:171-180. [PubMed] [Google Scholar]
- 32.Montell, C., E. F. Fisher, M. H. Caruthers, and A. J. Berk. 1984. Control of adenovirus E1B mRNA synthesis by a shift in the activities of RNA splice sites. Mol. Cell. Biol. 4:966-972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nevels, M., S. Rubenwolf, T. Spruss, H. Wolf, and T. Dobner. 1997. The adenovirus E4orf6 protein can promote E1A/E1B-induced focus formation by interfering with p53 tumor suppressor function. Proc. Natl. Acad. Sci. USA 94:1206-1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ornelles, D. A., and T. Shenk. 1991. Localization of the adenovirus early region 1B 55-kilodalton protein during lytic infection: association with nuclear viral inclusions requires the early region 4 34-kilodalton protein. J. Virol. 65:424-429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pardo-Mateos, A., and C. S. Young. 2004. A 40 kDa isoform of the type 5 adenovirus IVa2 protein is sufficient for virus viability. Virology 324:151-164. [DOI] [PubMed] [Google Scholar]
- 36.Pilder, S., K. Leppard, J. Logan, and T. Shenk. 1986. Functional analysis of the adenovirus E1B 55K polypeptide. Cancer Cells 4:285-290. [Google Scholar]
- 37.Querido, E., P. Blanchette, Q. Yan, T. Kamura, M. Morrison, D. Boivin, W. G. Kaelin, R. C. Conaway, J. W. Conaway, and P. E. Branton. 2001. Degradation of p53 by adenovirus E4orf6 and E1B55K proteins occurs via a novel mechanism involving a Cullin-containing complex. Genes Dev. 15:3104-3117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Reich, N. C., P. Sarnow, E. Duprey, and A. J. Levine. 1983. Monoclonal antibodies which recognize native and denatured forms of the adenovirus DNA-binding protein. Virology 128:480-484. [DOI] [PubMed] [Google Scholar]
- 39.Rubenwolf, S., H. Schütt, M. Nevels, H. Wolf, and T. Dobner. 1997. Structural analysis of the adenovirus type 5 E1B 55-kilodalton-E4orf6 protein complex. J. Virol. 71:1115-1123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ryabova, L. A., M. M. Pooggin, and T. Hohn. 2002. Viral strategies of translation initiation: ribosomal shunt and reinitiation. Prog. Nucleic Acid Res. Mol. Biol. 72:1-39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sarnow, P., C. A. Sullivan, and A. J. Levine. 1982. A monoclonal antibody detecting the adenovirus type 5-E1b-58Kd tumor antigen: characterization of the E1b-58Kd tumor antigen in adenovirus-infected and -transformed cells. Virology 120:510-517. [DOI] [PubMed] [Google Scholar]
- 42.Schmid, S. I., and P. Hearing. 1999. Adenovirus DNA packaging. Construction and analysis of viral mutants, p. 47-59. In W. S. Wold (ed.), Adenovirus methods and protocols, vol. 21. Humana Press Inc., Totowa, NJ. [Google Scholar]
- 43.Sieber, T., and T. Dobner. 2007. Adenovirus type 5 early region 1B 156R protein promotes cell transformation independently of repression of p53-stimulated transcription. J. Virol. 81:95-105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Stracker, T. H., C. T. Carson, and M. D. Weitzman. 2002. Adenovirus oncoproteins inactivate the Mre11 Rad50 NBS1 DNA repair complex. Nature 418:348-352. [DOI] [PubMed] [Google Scholar]
- 45.Takayesu, D., J. G. Teodoro, S. G. Whalen, and P. E. Branton. 1994. Characterization of the 55K adenovirus type 5 E1B product and related proteins. J. Gen. Virol. 75:789-798. [DOI] [PubMed] [Google Scholar]
- 46.Tollefson, A. E., T. W. Hermiston, and W. S. Wold. 1999. Preparation and titration of CsCl-banded adenovirus stocks, p. 1-9. In W. S. Wold (ed.), Adenovirus methods and protocols, vol. 21. Humana Press Inc., Totowa, NJ. [DOI] [PubMed] [Google Scholar]
- 47.Touriol, C., S. Bornes, S. Bonnal, S. Audigier, H. Prats, A. C. Prats, and S. Vagner. 2003. Generation of protein isoform diversity by alternative initiation of translation at non-AUG codons. Biol. Cell 95:169-178. [DOI] [PubMed] [Google Scholar]
- 48.Virtanen, A., and U. Pettersson. 1985. Organization of early region 1B of human adenovirus type 2: identification of four differentially spliced mRNAs. J. Virol. 54:383-391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Weitzman, M. D., and D. A. Ornelles. 2005. Inactivating intracellular antiviral responses during adenovirus infection. Oncogene 24:7686-7696. [DOI] [PubMed] [Google Scholar]
- 50.White, E. 2001. Regulation of the cell cycle and apoptosis by the oncogenes of adenovirus. Oncogene 20:7836-7846. [DOI] [PubMed] [Google Scholar]
- 51.Woo, J. L., and A. J. Berk. 2007. Adenovirus ubiquitin-protein ligase stimulates viral late mRNA nuclear export. J. Virol. 81:575-587. [DOI] [PMC free article] [PubMed] [Google Scholar]