Abstract
The promyelocytic leukemia (PML) protein gathers other cellular proteins, such as Daxx and Sp100, to form subnuclear structures termed PML-nuclear bodies (PML-NBs) or ND10 domains. Many infecting viral genomes localize to PML-NBs, leading to speculation that these structures may represent the most efficient subnuclear location for viral replication. Conversely, many viral proteins modify or disrupt PML-NBs, suggesting that viral replication may be more efficient in the absence of these structures. Thus, a debate remains as to whether PML-NBs inhibit or enhance viral replication. Here we review and discuss recent data indicating that for herpesviruses, PML-NB proteins inhibit viral replication in cell types where productive, lytic replication occurs, while at the same time may enhance the establishment of lifelong latent infections in other cell types.
Keywords: HCMV, HSV-1, intrinsic immunity, PML-NB
There are proteins present in cells that directly inhibit the ability of infecting viral pathogens to replicate. Such intrinsic immune mechanisms are mediated by constitutively expressed cellular proteins, operate in both the cytoplasm and the nucleus, and represent one of the first lines of antiviral defenses in naive hosts [1]. To date, cell intrinsic defenses that inhibit the productive, lytic replication of retroviruses [1] and herpes-viruses [2-4] have been described, and members from other viral classes are likely subject to similar immune measures as well. Intrinsic immune proteins that restrict the replication of herpesviruses belong to a class of factors that localize within the nucleus to dot-like structures arranged by the promyelocytic leukemia (PML) protein [5] termed PML-nuclear bodies (PML-NBs; also called ND10s because there are often approximately ten of these nuclear domains per cell) [6]. This review focuses on the antiviral, and perhaps proviral, activities of PML-NB proteins toward herpesviruses.
The eight human herpesviruses cause diseases that range from cold sores to cancers. They are divided into alpha, beta and gamma subfamilies based upon sequence, cellular tropism and the length of the replication cycle [7]. In addition to the productive, lytic replication that generates infectious viral progeny, herpesviruses also establish latent infections where few, if any, viral proteins are expressed and no viral progeny are produced. In latently infected cells, viral genomes are efficiently maintained in a quiescent state allowing for the lifelong colonization of the host. Periodic reactivation of latent viruses to produce infectious viral particles permits the virus to spread within and among hosts. The ability of herpesviruses to establish latency contributes to the host’s failure to clear the infection by rendering infected cells undetectable to immune surveillance and resistant to antiviral drugs.
Herpesviruses have large (150–230 kb) linear DNA genomes protected in virions by both icosahedral capsids and lipid envelopes. Between the capsid and the envelope is a layer consisting mostly of viral proteins termed the tegument [7]. Upon viral entry, tegument proteins are delivered directly to cells and mediate the initial stages of infection, both in the cytoplasm and the nucleus. Tegument proteins are important for the initiation of viral gene expression after the viral genomes are delivered to the nucleus. These proteins stimulate expression of the immediate-early (IE) genes, the first viral genes to be expressed during lytic replication. The major viral IE proteins subsequently autoregulate their own expression, as well as activating the expression of the viral early genes involved in genome replication. The proteins produced from the IE transcripts commit the virus to a lytic replication cycle, so when latency is established, either the expression of the IE genes or the activities of their encoded proteins must be suppressed. This review will focus on how PML-NB and viral proteins regulate viral gene expression during lytic replication and the role they may play during latency for the prototypical members of the alphaherpesviruses (herpes simplex virus [HSV] type 1 [HSV-1]) and the betaherpesviruses (human cytomegalovirus [HCMV]).
Shortly after they enter the nucleus herpesviral genomes are found in association with PML-NBs
The intranuclear sites where herpesviral DNA replication occurs are called replication factories or compartments. The observation of symmetrical patterns of replication factories in each nucleus of binucleated cells led to the conclusion that the spatial arrangement of an unidentified, but pre-existing nuclear structure was conserved during nuclear division, with which herpesviral genomes specifically associated [8]. Subsequent experiments indicated that the structure in question was likely to be PML-NBs [9,10]. PML-NBs appear as numerous small spots within the nucleus when their resident proteins are observed by indirect immunofluorescence microscopy. Three major constituent proteins of PML-NBs are the PML protein itself, which is required to maintain the PML-NB structure [5], Sp100 [11] and Daxx [5]. All three of these proteins have the ability to activate and repress transcription [12]. Daxx also regulates apoptosis [13,14], and PML is implicated in many other processes, including the DNA damage and stress responses, apoptosis [15], tumor suppression [16] and cellular senescence. The functions of PML-NBs and constituent proteins have been recently reviewed [17-21].
Herpesviral genomes are found to be associated with PML-NBs in the absence of viral gene expression and at early times after infection [10,22,23]. Only HCMV genomes located near PML-NBs appeared to be transcriptionally active, leading to the hypothesis that PML-NBs represented positively acting, pre-existing, sub-nuclear sites that provided a structural environment conducive for viral gene expression [22], and perhaps viral DNA replication as well. However, contrary evidence supported a completely opposite view, one in which PML-NBs are detrimental to viral infections. While the mRNAs encoding PML, Sp100 and Daxx are constitutively expressed, they are upregulated in response to the innate immune system agonist interferon [24-26], implying that these proteins may play a role in cellular immunity to infecting pathogens. Furthermore, IE proteins from both HSV-1 and HCMV disrupt PML-NBs [27-29], also implying that these structures may be detrimental to replicating viruses. Finally, an elegant series of experiments clearly demonstrated that PML-NBs found associated with herpesviral genomes do not represent pre-existing subnuclear sites, but are actually newly assembled structures built around infecting viral genomes [30]. This work argued that viral genomes do not migrate through the nucleus in search of preformed PML-NBs, but that PML-NB proteins relocalize to sites of deposited infecting viral genomes. However, the determination as to whether PML-NBs and their resident proteins antagonize or facilitate herpesviral replication required more direct experimental examination.
Several proteins that localize to PML-NBs repress the transcription of herpesvirus lytic phase genes & inhibit their productive replication cycles
The observation that viruses encode proteins that disrupt PML-NB structures provided evidence that these domains may be inhibitory to viral infection. For example, the HSV-1 ICP0 protein is a ubiquitin ligase that induces the proteasomal degradation of PML and Sp100, leading to the disruption of PML-NBs [28,31-33]. A reduction in viral replication occurred when wildtype ICP0 was either deleted [34] or replaced with point mutants that fail to disrupt PML-NBs [35], indicating the importance of ICP0 and its ability to disrupt PML-NBs for HSV-1 infection. A similar growth defect occurred when the HCMV tegument protein pp71 was deleted [36] or its ability to interact with Daxx was disrupted [37]. Additional evidence demonstrated that pp71 induced the degradation of Daxx [2,4] through an unusual proteasome dependent, ubiquitin independent manner [38], but did not disrupt PML-NBs [2]. However, pp71-mediated Daxx degradation induces the expression of the viral IE1 protein [2,39-42], and IE1 subsequently disrupts PML-NBs by altering the SUMOylation state of PML [27,43,44] and perhaps other proteins as well. As stated above, the fact that these herpesviral proteins degraded and disrupted PML-NBs implied, but did not demonstrate, that PML-NB proteins antagonize herpesviral infections. However, experiments in which PML-NB proteins were overexpressed [39,45,46], or in which their levels were diminished by RNA interference [2-4,39,40,47,48], indicated that they indeed inhibit herpesviral lytic replication because they repress the transcription of lytic phase genes (Figure 1).
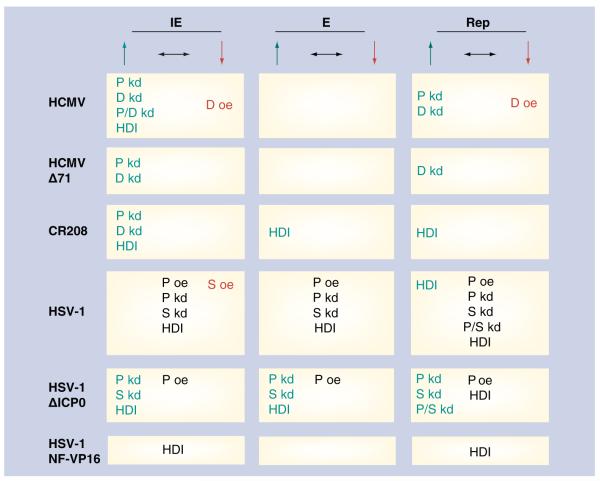
Figure 1. Effects of PML-NB proteins on wild-type and mutant HCMV and HSV-1 gene expression.
The effects of oe or kd of P, S, or D, or the inhibition of histone deacetylase on viral IE and E gene expression, or the completion of Rep are summarized. Conditions which increase gene expression are denoted in green (↑) and left-aligned; those which have no effect are black (↔) and centered; and those which decrease gene expression are right-aligned and in red (↓). Wild-type viruses and mutants are shown, which lack pp71 (HCMV Δ71), lack IE1 (CR208), lack ICP0 (HSV-1 ΔICP0), or contain nonfunctional VP16 (HSV-1 NF-VP16). D: Daxx; E: Early; HCMV: Human cytomegalovirus; HDI: Histone deacetylase inhibition; HSV: Herpes simplex virus; IE: Immediate-early; kd: knockdown; oe: Overexpression; P: Promyelocytic leukemia protein; Rep: Replication cycle; S: Sp100.
Knockdown of either PML or Sp100 individually or in combination increased viral lytic gene expression and replication of an ICP0-null mutant of HSV-1, but had no effect on wild- type virus [47,48]. In addition, overexpression of Sp100 inhibited IE gene expression from wild-type HSV-1 infections [46]. HSV-1 replicated to slightly higher titers in mouse cells devoid of PML as compared with wild-type mouse cells, and this differential replication was magnified when cells were treated with exogenously added interferon prior to infection [49]. A report in which no effect of PML overexpression on HSV-1 replication was observed is difficult to interpret because of the inclusion of the histone deacetylase (HDAC) inhibitor sodium butyrate in those experiments [50]. For HCMV, knockdown of PML and Daxx, either alone or in combination, increased viral IE gene expression and replication of the wild-type virus, and increased IE gene expression in pp71-null or IE1-null mutants [2-4,39,40]. Overexpression of Daxx inhibited wild-type HCMV IE gene expression and replication [39,45]. These experiments demonstrated that the constitutively expressed PML-NB proteins inhibit herpesviral gene expression and replication (Figure 1), and therefore they have been described as mediators of an intrinsic immune defense against these viruses. These experiments also seem to indicate that the inhibitory effects are resident in the proteins themselves, not in the overall structure of the PML-NB, because in the absence of PML, proteins that normally localize to PML-NBs, such as Daxx and Sp100, are found diffusely scattered throughout the nucleus, but assemble into PML-NB-like structures around infecting viral genomes [3,47,48].
While PML-NB proteins clearly repress the expression of HSV-1 and HCMV genes, the exact mechanism of how they do so is still unclear. For HCMV, the ability of PML-NBs to recruit HDACs to modify chromatin structure of the viral genome seems to be a likely mechanism. Treatment with chemical HDAC inhibitors mimics the functional consequences of pp71 and IE1 expression, and PML-NB protein knockdown [2,45,51]. The effects of HDAC inhibitors on HSV-1 gene expression from either wild-type, VP16 defective or ICP0-null genomes appear to show a cell type dependence, failing to stimulate HSV-1 replication in certain cell types [48,52] while enhancing replication or reactivation in others, particularly neuronal cells [53-56]. Thus, while ICP0 is thought to associate with and modify the function of cellular chromatin remodeling complexes that contain HDACs [57-60], it is evident that ICP0 may also possess additional mechanisms to alter viral gene expression.
It is interesting that the knockdown of PML-NB proteins can enhance wild-type HCMV gene expression, but not wild-type HSV-1 gene expression. This may mean that HSV-1 ICP0 is more efficient at neutralizing PML-NB proteins than HCMV pp71 and IE1, or that PML-NB proteins play a more prominent role in controlling HCMV gene expression in comparison with HSV-1 gene expression. While both explanations may be palatable, the later prediction seems particularly salient as the initial activation of viral IE gene expression for these two viruses happens through very different mechanisms. For HCMV, IE gene expression is derepressed by the pp71-mediated degradation of Daxx [2]. For HSV-1, IE gene expression is activated by VP16 through the recruitment of components of the RNA polymerase II transcriptional machinery to viral IE promoters [61-64], and perhaps additionally by preventing cellular histones from associating with them [65]. Further amplification of IE gene expression and the initiation of early and late gene expression are more similar for these two viruses. HSV-1 IE genes (ICP4 and ICP0) further activate the expression of viral genes [64]. ICP4 activates viral gene expression by promoting the formation of pre-initiation complexes at early and late promoters [66-69], but can also function to repress its own expression [64,70-72]. ICP0 is a ubiquitin ligase that induces the proteasomal degradation of PML and Sp100 leading to the disruption of PML-NBs [28,31-33], disrupts HDAC complexes [58,59] and changes the acetylation status, either directly or indirectly, of viral promoters [60]. The two major HCMV IE proteins, IE1 and IE2, also transactivate the expression of both cellular and viral genes through association with the basal transcription machinery [73,74]. IE1 further disrupts PML-NBs that accumulate at the viral genome [3,4,27,29,43,44], inactivating any additional repressive effects of these domains on downstream viral gene expression. Both IE1 and IE2 bind to HDACs [51,75], potentially negating their repressive effects, although IE2’s ability to bind HDAC1 and the major immediate-early promoter (MIEP) crs element allows for repression of its own expression [76].
In summary, HSV-1 and HCMV use similar strategies to initiate lytic gene expression (Figure 2). Tegument-delivered proteins stimulate the expression of the IE proteins, which in turn amplify their own expression and activate the expression of other classes of viral genes. While the overall strategy is similar there are subtle, but important, differences. For HCMV, the inactivation of intrinsic defenses mediated by PML-NB proteins is critical for both of these steps. pp71 degrades Daxx to initiate IE gene expression, and IE1 inactivates PML to amplify IE gene expression and activate the expression of the viral early genes. To date, the need to inactivate PML-NB proteins appears to apply only to HSV-1 IE gene amplification and early gene expression mediated by ICP0, and not to the initiation of IE gene expression stimulated by VP16. In addition to HSV-1 and HCMV, PML-NB proteins affect other classes of viruses, which encode proteins that can bind to and/or affect the integrity of PML-NBs. A compilation of different viruses and their encoded gene products that target PML-NBs are summarized in Table 1.
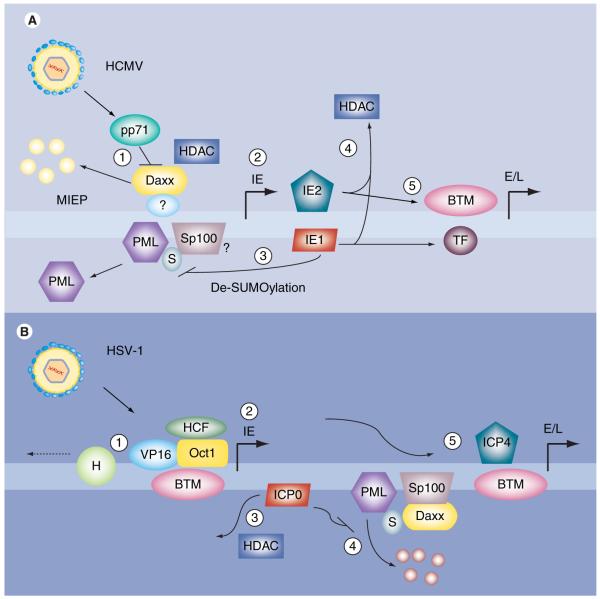
Figure 2. Activation of HCMV and HSV-1 viral gene expression.
(A) Upon HCMV fusion and content delivery to the infected cell, the tegument protein pp71 binds to and induces the degradation of Daxx (1). This de-represses the viral MIEP and promotes the expression of the IE genes (2). IE1 disrupts the remaining PML-NB proteins by preventing/disrupting the SUMOylation status of PML, and possibly Sp100, further increasing IE gene expression (3). Both IE1 and IE2 negate the effect of HDACs by binding to and sequestering them away from viral promoters (4). Finally, the IE proteins can recruit BTM and TFs to early and late viral promoters to activate their respective genes (5). (B) Upon HSV-1 fusion and content delivery to the infected cells, the tegument protein VP16 binds to the cellular Oct1 and HCF proteins and targets to the viral IE promoter, where it displaces cellular H (1), and activates viral gene expression by recruiting the BTM (2). ICP0 has multiple functions to activate subsequent viral gene expression, including dissociating HDAC complexes (3) and inducing the degradation of PML and Sp100 (4). ICP4 promotes the expression of early and late viral genes by recruiting BTM to targeted promoters (5). BTM: Basal transcriptional machinery; E/L: Early/late; H: Histones; HCF: Host-cell factor; HCMV: Human cytomegalovirus; HDAC: Histone deacetylase; HSV: Herpes simplex virus; IE: Immediate-early; MIEP: Major immediate-early promoter; PML: Promyelocytic leukemia; PML-NB: PML-nuclear body; TF: Transcription factor.
Table 1.
Viruses which associate with or affect PML-NB structures.
| Virus | Viral protein | PML-NB protein | Function | Ref. |
|---|---|---|---|---|
| HCMV | pp71 | Daxx | Degradation, ↑ IE | [2,39,40,45] |
| IE1 | PML | Disrupt PML-NB, ↑ IE | [3,4,27,29,44] | |
| HSV-1 | ICP0 | PML/Sp100 | Degradation, ↑ replication | [31,32,46–49] |
| EBV | BZLF-1 | PML | Disrupt PML-NB | [108] |
| HHV-6 | IE1 | PML-NB | ? | [109] |
| Adenovirus | E4 ORF3 | PML | Reorganizes PML-NB | [110,111] |
| HPV | E6 | PML | ↓ PML IV senescence | [112] |
| E7 | PML | ↓ PML IV senescence | [113] | |
| L2 | Sp100/Daxx | Reorganizes PML-NB | [114] | |
| HFV | Tas | PML | ↓ Tas transactivation | [115] |
| ASV | Integrase | Daxx | ↓ viral gene expression | [116,117] |
| VSV | ? | PML | PML overexpression, ↓ viral replication | [118] |
| Influenza A | ? | PML | PML overexpression, ↓ viral replication | [118] |
| HDV | L-HDAg | PML-NB | Reorganizes PML-NB | [119] |
| RSV | ? | PML/Sp100 | Cytoplasmic redistribution | [120] |
| PUUV | N protein | Daxx | ? | [121] |
| Dengue | C protein | Daxx | Sensitization to apoptosis | [122] |
| HCV | Core protein | PML | ↓ PML IV-induced apoptosis | [123] |
| Phage ΦC31 | Integrase | Daxx | ↓ recombination | [124] |
Viruses known to have proteins which localize to and/or disrupt PML-NB structures are listed with references provided.
ASV: Avian sarcoma virus; EBV: Epstein—Barr virus; HCMV: Human cytomegalovirus; HDV: Hepatitis delta virus; HFV: Human foamy virus; HHV-6: Human herpesvirus 6; HPV: Human papillomavirus; HSV: Herpes simplex virus; IE: Immediate-early; PML: Promyelocytic leukemia; PML-NB: PML-nuclear body; PUUV: Puumala virus; RSV: Respiratory syncytial virus; VSV: Vesicular stomatitis virus.
While they inhibit lytic replication, do PML-NB proteins facilitate herpesviral latency?
Approaches using chemical HDAC inhibitors, RNA interference methodologies and protein overexpression systems have now clearly demonstrated that PML-NB proteins inhibit herpesvirus lytic phase gene expression and productive viral replication. Such experiments have led to the description of PML-NB proteins as cellular mediators of intrinsic immunity against HSV-1 and HCMV. In order to productively replicate, these viruses encode proteins that efficiently inactivate PML-NB-mediated intrinsic defenses. We noted that, while the silencing of lytic phase gene expression would inhibit the productive replication cycle of these viruses, such silencing is actually required for their ability to establish latently-infected cells. Thus it is possible that the same PML-NB-based mechanisms that would silence herpesviral gene expression at the start of lytic infections (unless they are neutralized by viral countermeasures) may also be used to silence gene expression when latency is established.
HCMV is thought to establish latency in undifferentiated cells of the myeloid lineage [77-80]. In particular, CD34+ hematopoeitic progenitor cells are becoming the model system of choice for HCMV latency studies [81-85]. Latent viral genomes can be detected in and reactivated in vitro from CD34+ cells isolated from sero-positive patients [84] and from cultured CD34+ cells infected in vitro [81-83,85]. The HCMV genomes carried in CD34+ cells show a condensed chromatin structure that is conserved regardless of whether they are naturally infected or infected in vitro [83,84]. A similar chromatin structure is also observed on viral genomes when HCMV infects NT2 and THP-1 cells in vitro, two model systems that are used to study quiescent HCMV infections [86-88]. Infections in NT2 and THP-1 cells are referred to as quiescent (or latent like) because treatments that can reactivate expression from the silenced viral genomes in these cells (if they exist) have not yet been identified [89-92]. Additionally, in lytically-infected fibroblasts prior to pp71-mediated degradation of Daxx, HCMV genomes adopt a chromatin structure indistinguishable from that observed in CD34+ and NT2 cells [45].
The analysis of HCMV genomes during latent and quiescent infections, and at the very start of lytic infections indicated that their chromatin structure is indistinguishable by chromatin immunoprecipitation (ChIP) assays, perhaps indicating that similar mechanisms may be used to silence HCMV IE gene expression in all cell types. The mechanism by which the repressive chromatin structure is established on latent viral genomes in CD34+ cells has yet to be analyzed, but it is now evident that PML-NB proteins and HDACs contribute to this process at the start of lytic and quiescent HCMV infections. Chemical HDAC inhibitors permit IE gene expression in NT2 and THP-1 cells [87,90,92,93], indicating that HDACs play a role in transcriptional repression during HCMV quiescence. A study from our laboratory found that the knockdown of Daxx prior to HCMV infection also permits IE gene expression in NT2 and THP-1 cells [92]. Artificial knockdown of Daxx is not required for HCMV IE gene expression upon lytic infection of fibroblasts (as it is in quiescent infections of NT2 and THP-1 cells) because tegument-delivered pp71 migrates to the nucleus, binds to Daxx and induces its degradation in these cells [2,4]. Daxx is not degraded upon HCMV infection of NT2 and THP-1 cells because tegument-delivered pp71 is trapped in the cytoplasm [92]. Interestingly, pp71 proteins that are newly synthesized in NT2 and THP-1 cells localize to the nucleus, degrade Daxx and permit IE gene expression upon HCMV infection. In the absence of Daxx, these cells that would normally establish a quiescent infection initiate a productive, lytic replication program, but the replication cycle cannot be completed in these cells, and so an abortive infection is observed [92]. These experiments not only show that the PML-NB intrinsic defense mediated by Daxx operates in quiescent HCMV infections, but also that it must be active to prevent an abortive infection that may result in the immune detection and clearance of the virus.
A subsequent report [94] showed that Daxx knockdown did not rescue viral IE gene expression in a subclone of NT2 cells termed D1, and concluded that additional transcriptional repressors combined with the absence of transcriptional activators resulted in the silencing of IE gene expression during viral quiescence. Noteworthy differences between that study and ours [92] include our additional analysis of THP-1 cells, our controls for viral entry and our rescue of IE gene expression with an HDAC inhibitor. Different methods to deliver the siRNA (lipid-based transfection reagent [94] vs nonlipid based transfection or retroviral transduction [92]) were also used. Additionally, different viral strains were employed. Obviously, more work will be needed to define the roles of the PML-NB proteins during quiescent infection, and more importantly during true latent HCMV infections, as well as the potential contribution of positively-acting transcription factors perhaps only expressed in differentiated cells.
The contribution of PML-NBs to the establishment of HSV-1 latency has not been examined in neurons, the in vivo reservoir for latent infections. However, using quiescent viral genomes, which lack functional VP16, ICP0 and ICP4, the role of PML-NBs during quiescence in fibroblasts has been investigated [23]. These three viral proteins are the principal viral transcriptional transactivators necessary for the initiation of lytic viral replication. Infection of fibroblasts with the quiescent virus leads to the assembly of new PML-NBs around the infecting viral genome, which last for many days postinfection [23]. In the absence of PML, Sp100 and Daxx initially co-localize with the viral genome, but do not remain associated with it for extended periods of time postinfection [23]. A caveat of the HSV-1 quiescence experiments as a model for the establishment of latency is the absence of three functional viral transactivators, especially VP16, which stimulates IE gene expression through a mechanism that may not be related to PML-NB proteins. Interestingly, VP16 nuclear localization is dependent, in part, on the cellular protein host-cell factor (HCF), which is found in the cytoplasm of cells where latency is established [95,96], but can localize to the nucleus during reactivation. This suggests that tegument-delivered VP16 may also have cytoplasmic localization during the establishment of latency. A putative cytoplasmic protein localization due to sequestration by HCF may also explain why newly-expressed VP16 failed to prevent the establishment of latency in murine trigeminal ganglia [97]. Thus, it may be a common theme that tegument transactivators for both HSV-1 and HCMV are excluded from the nucleus in cells where latent infections are established; but further confirmation using cell systems that represent the true site of viral latency is required.
In addition to the establishment of latency, PML-NB proteins may play a role in maintaining the quiescent state of latent genomes. The exogenous expression of ICP0 or pp71 can activate gene expression from quiescent HSV-1 genomes [98-101]. Furthermore, latently-infected cells express antisense transcripts that may affect the expression of ICP0 [102-105] or pp71 [106], implying that repressing the expression of these proteins may be important for the maintenance of the latent state. For HSV-1, chromatin remodeling to achieve or maintain viral genome silencing also seems to occur through additional processes independent of PML-NB proteins [107].
Conclusion
All available evidence indicates that PML-NB proteins associate with HSV-1 and HCMV genomes at the very beginning of both lytic and quiescent infections, and implies that this association is also likely to occur at the start of latent infections. For HCMV, it is clear that the associated PML-NB proteins repress viral lytic gene expression at the start of a lytic infection through the action of HDACs, and that this silencing is eventually inactivated by the viral pp71 and IE1 proteins. Quiescent HCMV infections may also require the presence of Daxx to prevent the initiation of an abortive lytic replication program in these cells. Extending these results, one may predict that Daxx, and likely PML, may be responsible for silencing latent viral genomes as well. Thus, it seems possible that HCMV uses an intrinsic immune defense as one means to silence its genome and allow for the establishment of a latent infection.
The role of PML-NB proteins during HSV-1 infection is less clear. The initial activation of IE gene expression by VP16 may be independent of PML-NB proteins. However, the amplification of IE transcripts and the expression of early genes seem to rely, in part, on the ability of ICP0 negate the inhibitory effects of PML-NB proteins. While PML-NB proteins clearly associate with quiescent viral genomes, they do not appear to be absolutely required to maintain that quiescence. Thus, PML-NB proteins might be predicted to play a more limited role in HSV-1 latency than in HCMV latency.
While the reliance of these viruses on PML-NB proteins for transcriptional silencing during latency may be quite different, the general mechanisms by which their tegument transactivators are prevented from initiating lytic gene expression may be similar. During a quiescent infection, HCMV tegument-delivered pp71 remains in the cytoplasm. VP16 localizes to the nucleus by binding to HCF, and data suggests that HCF remains in the cytoplasm of cells where latency is established, suggesting that VP16 may also remain in the cytoplasm. Therefore, differential localization may be a similar theme, even though VP16 activates HSV-1 promoters while pp71 derepresses HCMV promoters. The different ways in which VP16 and pp71 facilitate the initiation of viral IE gene expression may be related to the cell types in which these two viruses establish latency, and to the promoters that drive expression of their IE genes. Perhaps neurons (HSV-1) are less transcriptionally active than CD34+ cells (HCMV), and, almost certainly, HSV-1 IE promoters have a lower basal expression level than the HCMV major IE promoter. Thus, when HSV-1 latency is established, the initial activation of relatively weak viral IE promoters (a process that PML-NB proteins may not be required for) must be inhibited (perhaps by sequestering tegument-delivered VP16 in the cytoplasm). Conversely, when HCMV latency is established, the repression of the strong IE promoters (a process that PML-NB proteins are known to be involved in) need only be maintained (perhaps by sequestering tegument-delivered pp71 in the cytoplasm). In conjunction with the PML-NB proteins, the differing milieu of transcriptional regulators present in the cell types where lytic or latent infections occur also likely contributes to the different viral gene expression patterns observed in these different types of infections.
Future perspective
Our understanding of PML-NB proteins and their relation to viral infection has evolved rapidly in recent years. A bevy of recent evidence demonstrates the negative influence PML-NB proteins have on herpesviral lytic gene expression and replication. These proteins seem to have more potent effects on HCMV than on HSV-1. A model is starting to emerge for a seemingly positive role of PML-NB proteins in the repression of viral gene expression during the establishment of latency. Clearly, PML-NBs and their associated proteins can affect the chromatin status of HCMV by recruiting chromatin modifiers in cells where productive infection occurs. Whether PML-NB proteins also influence the chromatin status of repressed latent genomes or whether they are necessary for the maintenance of a repressed state requires further examination. Determining the impact that PML-NB proteins have on herpesviral latency will advance our understanding of this process, which is critical for the lifelong infection of the host. Thus, this promises to be an area of intense focus for future research. In order to effectively address this topic, it will be necessary to use model systems that represent true sites of latency and to use clinical viral isolates in lieu of high-passage laboratory-adapted strains. Additionally, as more than one PML-NB protein affects lytic viral replication and potentially the establishment of latency, methods to simultaneously inactivate multiple PML-NB proteins will enhance our understanding of how these subnuclear sites affect both lytic and latent infections. Determining the sites where PML-NB proteins interact with the viral genome is also essential. This information could allow for the construction of mutant viruses immune to PML-NB proteins. Such mutants would provide yet another way to test the contribution of these proteins not only during latency, but also during lytic infection. Finally, the evolutionary implications of herpesviruses potential reliance on PML-NB proteins to repress their genomes during latency should be considered. In a cruel twist of fate, the pressures on cells to protect themselves from viral infections may have ultimately led to the ability of viruses to persist for the life of the host, avoiding immune clearance and causing little or no overt pathology, through the evolution of a latent stage of infection that can be subsequently reactivated to promote viral spread to new hosts.
Executive summary.
Introduction
Cells constitutively express proteins that have antiviral properties.
Such intrinsic immunity conferred by these proteins represents one of the first lines of defense against infecting viral pathogens.
Promyelocytic leukemia-nuclear body (PML-NB) proteins mediate a cellular intrinsic immune defense against herpesviruses.
Herpesviruses have dsDNA genomes and two different infectious cycles (lytic and latent).
Lytic (productive) replication allows for viral spread and latency (nonproductive) for lifelong persistence.
Viral immediate-early (IE) gene expression commits the virus to the lytic replication cycle, and therefore must be repressed when latency is established.
Shortly after they enter the nucleus, herpesviral genomes are found in association with PML-NBs
Three major PML-NB proteins are PML, Sp100 and Daxx.
PML, Sp100 and Daxx are constitutively expressed but also interferon inducible.
The genomes of herpes simplex virus (HSV)-1 and human cytomegalovirus (HCMV), as well as other DNA viruses, are found adjacent to PML-NBs.
One study showed that only HCMV genomes adjacent to PML-NBs produced transcripts.
HSV-1 and HCMV IE proteins disrupt PML-NBs and inactivate PML-NB proteins.
Reorganized PML-NBs assemble around infecting viral genomes.
Different sets of circumstantial evidence point to a positive or negative role for PML-NBs during herpesviral infections.
Several proteins that localize to PML-NBs repress the transcription of herpesvirus lytic phase genes & inhibit their productive replication cycles
Overexpression and knockdown studies reveal that PML-NB proteins inhibit HSV-1 and HCMV lytic replication by repressing viral gene expression.
The structure of the PML-NB is likely less important for the antiviral defense than the activities of the individual proteins that reside there.
HCMV escapes the PML-NB defense by pp71-mediated degradation of Daxx, leading to the expression of IE1, which further disrupts the remaining PML-NB structure.
HSV-1 escapes the PML-NB intrinsic defense through the degradation of PML and Sp100 by the viral ICP0 protein.
There are many similarities but also important differences in how HSV-1 and HCMV initiate lytic gene expression, the classes of viral genes silenced by PML-NB proteins and the mechanisms employed by the viruses to escape the repressive effects of the PML-NB proteins.
While they inhibit lytic replication, do PML-NB proteins facilitate herpesviral latency?
Unless inactivated, PML-NB proteins inhibit viral lytic gene expression, reminiscent of that which occurs at the start of a latent infection.
HCMV establishes a chromatin structure during natural latent infection of CD34+ cells that is indistinguishable from one formed in quiescent or latent-like infections in NT2 and THP-1 cells.
Daxx silences HCMV gene expression to establish quiescence and is not degraded because of the mislocalization of tegument-delivered pp71.
Although HCMV lytic gene expression initiates in the absence of Daxx in NT2 cells, the replication cycle is not completed and an abortive infection results.
Quiescent HSV-1 genomes become surrounded by PML-NB proteins and remain associated with them for multiple days postinfection.
Gene expression can be activated from quiescent HSV-1 genomes by ectopic-expression of ICP0 or pp71.
Conclusion
PML-NB proteins in conjunction with histone deacetylases repress HCMV gene expression. They are inactivated by pp71 and IE1 to initiate a lytic infection, but remain functional during quiescent, and possibly latent, infections.
PML-NB proteins appear to play an important, but less prominent, role during HSV-1 infection.
Sequestering virion-delivered transactivators in the cytoplasm may be a common theme for HCMV and HSV-1 to inhibit viral lytic gene expression when latency is established.
Future perspective
PML-NB proteins affect lytic replication, but their role, if any, during latency requires the study of relevant cells and clinical viral isolates.
Because multiple PML-NB proteins contribute to viral repression, the ability to downregulate multiple PML-NB proteins simultaneously will further clarify the true contribution of these structures.
Determining where and how PML-NB proteins exert their effect on viral DNA will allow for viral mutants to be constructed, providing another tool to examine the repressive effects of PML-NBs on viral gene expression.
Footnotes
Financial & competing interests disclosure
The authors have no relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript. This includes employment, consultancies, honoraria, stock ownership or options, expert testimony, grants or patents received or pending, or royalties.
No writing assistance was utilized in the production of this manuscript.
Contributor Information
Ryan T Saffert, University of Wisconsin-Madison, Institute for Molecular Virology & McArdle Laboratory for Cancer Research, Madison, WI, USA Tel.: +1 608 265 5546; Fax: +1 608 262 7414; rtsaffert@wisc.edu.
Robert F Kalejta, University of Wisconsin-Madison, Institute for Molecular Virology & McArdle Laboratory for Cancer Research, Madison, WI, USA Tel.: +1 608 265 5546; Fax: +1 608 262 7414; rfkalejta@wisc.edu.
Bibliography
Papers of special note have been highlighted as either of interest (•) or of considerable interest (••) to readers.
- 1•.Bieniasz PD. Intrinsic immunity: a front-line defense against viral attack. Nat. Immunol. 2004;5:1109–1115. doi: 10.1038/ni1125.Review defining intrinsic immunity.
- 2•.Saffert RT, Kalejta RF. Inactivating a cellular intrinsic immune defense mediated by Daxx is the mechanism through which the human cytomegalovirus pp71 protein stimulates viral immediate-early gene expression. J. Virol. 2006;80:3863–3871. doi: 10.1128/JVI.80.8.3863-3871.2006.First demonstration of the ability of a promyelocytic leukemia-nuclear body (PML-NB) protein to mediate intrinsic immune defense against a DNA virus.
- 3.Tavalai N, Papior P, Rechter S, Leis M, Stamminger T. Evidence for a role of the cellular ND10 protein PML in mediating intrinsic immunity against human cytomegalovirus infections. J. Virol. 2006;80:8006–8018. doi: 10.1128/JVI.00743-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4••.Tavalai N, Papior P, Rechter S, Stamminger T. Nuclear domain 10 components promyelocytic leukemia protein and hDaxx independently contribute to an intrinsic antiviral defense against human cytomegalovirus infection. J. Virol. 2008;82:126–137. doi: 10.1128/JVI.01685-07.Evidence that individual PML-NB proteins can have independent effects on herpesviral gene expression.
- 5.Ishov AM, Sotnikov AG, Negorev D, et al. PML is critical for ND10 formation and recruits the PML-interacting protein Daxx to this nuclear structure when modified by SUMO-1. J. Cell Biol. 1999;147:221–234. doi: 10.1083/jcb.147.2.221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ascoli CA, Maul GG. Identification of a novel nuclear domain. J. Cell Biol. 1991;112:785–795. doi: 10.1083/jcb.112.5.785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pellett P, Roizman B. The family Herpesviridae: a brief introduction. In: Knipe DM, Howley PM, editors. Fields Virology (2) Lippincott Williams and Wilkins; Philadelphia, PA, USA: 2007. pp. 2479–2500. [Google Scholar]
- 8.de Bruyn Kops A, Knipe DM. Preexisting nuclear architecture defines the intranuclear location of herpesvirus DNA replication structures. J. Virol. 1994;68:3512–3526. doi: 10.1128/jvi.68.6.3512-3526.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9•.Ishov AM, Maul GG. The periphery of nuclear domain 10 (ND10) as site of DNA virus deposition. J. Cell Biol. 1996;134:815–826. doi: 10.1083/jcb.134.4.815.Work demonstrating that herpesviral genomes associate with PML-NB at early times postinfection.
- 10•.Maul GG, Ishov AM, Everett RD. Nuclear domain 10 as pre-existing potential replication start sites of herpes simplex virus type-1. Virology. 1996;217:67–75. doi: 10.1006/viro.1996.0094.Work demonstrating that herpesviral genomes associate with PML-NB at early times postinfection.
- 11.Dyck JA, Maul GG, Miller WH, Jr, Chen JD, Kakizuka A, Evans RM. A novel macromolecular structure is a target of the promyelocyte-retinoic acid receptor oncoprotein. Cell. 1994;76:333–343. doi: 10.1016/0092-8674(94)90340-9. [DOI] [PubMed] [Google Scholar]
- 12.Zhong S, Salomoni P, Pandolfi PP. The transcriptional role of PML and the nuclear body. Nat. Cell Biol. 2000;2:E85–E90. doi: 10.1038/35010583. [DOI] [PubMed] [Google Scholar]
- 13.Michaelson JS. The Daxx enigma. Apoptosis. 2000;5:217–220. doi: 10.1023/a:1009696227420. [DOI] [PubMed] [Google Scholar]
- 14.Salomoni P, Khelifi AF. Daxx: death or survival protein? Trends Cell Biol. 2006;16:97–104. doi: 10.1016/j.tcb.2005.12.002. [DOI] [PubMed] [Google Scholar]
- 15.Wang ZG, Ruggero D, Ronchetti S, et al. PML is essential for multiple apoptotic pathways. Nat. Genet. 1998;20:266–272. doi: 10.1038/3073. [DOI] [PubMed] [Google Scholar]
- 16.Salomoni P, Pandolfi PP. The role of PML in tumor suppression. Cell. 2002;108:165–170. doi: 10.1016/s0092-8674(02)00626-8. [DOI] [PubMed] [Google Scholar]
- 17.Everett RD, Chelbi-Alix MK. PML and PML nuclear bodies: implications in antiviral defence. Biochimie. 2007;89:819–830. doi: 10.1016/j.biochi.2007.01.004. [DOI] [PubMed] [Google Scholar]
- 18•.Bernardi R, Pandolfi PP. Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies. Nat. Rev. Mol. Cell Biol. 2007;8:1006–1016. doi: 10.1038/nrm2277.Comprehensive review of PML-NB structures and protein functions.
- 19.Ching RW, Dellaire G, Eskiw CH, Bazett-Jones DP. PML bodies: a meeting place for genomic loci? J. Cell Sci. 2005;118:847–854. doi: 10.1242/jcs.01700. [DOI] [PubMed] [Google Scholar]
- 20.Negorev D, Maul GG. Cellular proteins localized at and interacting within ND10/PML nuclear bodies/PODs suggest functions of a nuclear depot. Oncogene. 2001;20:7234–7242. doi: 10.1038/sj.onc.1204764. [DOI] [PubMed] [Google Scholar]
- 21.Zimber A, Nguyen QD, Gespach C. Nuclear bodies and compartments: functional roles and cellular signalling in health and disease. Cell Signal. 2004;16:1085–1104. doi: 10.1016/j.cellsig.2004.03.020. [DOI] [PubMed] [Google Scholar]
- 22.Ishov AM, Stenberg RM, Maul GG. Human cytomegalovirus immediate early interaction with host nuclear structures: definition of an immediate transcript environment. J. Cell Biol. 1997;138:5–16. doi: 10.1083/jcb.138.1.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23•.Everett RD, Murray J, Orr A, Preston CM. Herpes simplex virus type 1 genomes are associated with ND10 nuclear substructures in quiescently infected human fibroblasts. J. Virol. 2007;81:10991–11004. doi: 10.1128/JVI.00705-07.Demonstrates the association of PML-NB proteins with herpes simplex virus (HSV)-1 DNA during a quiescent infection.
- 24.Lavau C, Marchio A, Fagioli M, et al. The acute promyelocytic leukaemia-associated PML gene is induced by interferon. Oncogene. 1995;11:871–876. [PubMed] [Google Scholar]
- 25.Guldner HH, Szostecki C, Grotzinger T, Will H. IFN enhance expression of Sp100, an autoantigen in primary biliary cirrhosis. J. Immunol. 1992;149:4067–4073. [PubMed] [Google Scholar]
- 26.Gongora R, Stephan RP, Zhang Z, Cooper MD. An essential role for Daxx in the inhibition of B lymphopoiesis by type I interferons. J. Immunol. 2001;14:727–737. doi: 10.1016/s1074-7613(01)00152-2. [DOI] [PubMed] [Google Scholar]
- 27.Ahn JH, Hayward GS. The major immediate-early proteins IE1 and IE2 of human cytomegalovirus colocalize with and disrupt PML-associated nuclear bodies at very early times in infected permissive cells. J. Virol. 1997;71:4599–4613. doi: 10.1128/jvi.71.6.4599-4613.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Everett RD, Maul GG. HSV-1 IE protein Vmw110 causes redistribution of PML. EMBO J. 1994;13:5062–5069. doi: 10.1002/j.1460-2075.1994.tb06835.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Korioth F, Maul GG, Plachter B, Stamminger T, Frey J. The nuclear domain 10 (ND10) is disrupted by the human cytomegalovirus gene product IE1. Exp. Cell Res. 1996;229:155–158. doi: 10.1006/excr.1996.0353. [DOI] [PubMed] [Google Scholar]
- 30••.Everett RD, Murray J. ND10 components relocate to sites associated with herpes simplex virus type 1 nucleoprotein complexes during virus infection. J. Virol. 2005;79:5078–5089. doi: 10.1128/JVI.79.8.5078-5089.2005.Video evidence for PML-NB protein migration to infecting HSV-1 genomes strengthened the argument that these proteins have antiviral activities.
- 31.Everett RD, Freemont P, Saitoh H, et al. The disruption of ND10 during herpes simplex virus infection correlates with the Vmw110- and proteasome-dependent loss of several PML isoforms. J. Virol. 1998;72:6581–6591. doi: 10.1128/jvi.72.8.6581-6591.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chelbi-Alix MK, de The H. Herpes virus induced proteasome-dependent degradation of the nuclear bodies-associated PML and Sp100 proteins. Oncogene. 1999;18:935–941. doi: 10.1038/sj.onc.1202366. [DOI] [PubMed] [Google Scholar]
- 33.Boutell C, Sadis S, Everett RD. Herpes simplex virus type 1 immediate-early protein ICP0 and its isolated RING finger domain act as ubiquitin E3 ligases in vitro. J. Virol. 2002;76:841–850. doi: 10.1128/JVI.76.2.841-850.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stow ND, Stow EC. Isolation and characterization of a herpes simplex virus type 1 mutant containing a deletion within the gene encoding the immediate early polypeptide Vmw110. J. Gen. Virol. 1986;67(Pt 12):2571–2585. doi: 10.1099/0022-1317-67-12-2571. [DOI] [PubMed] [Google Scholar]
- 35.Everett R, O’Hare P, O’Rourke D, Barlow P, Orr A. Point mutations in the herpes simplex virus type 1 Vmw110 RING finger helix affect activation of gene expression, viral growth, and interaction with PML-containing nuclear structures. J. Virol. 1995;69:7339–7344. doi: 10.1128/jvi.69.11.7339-7344.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bresnahan WA, Shenk TE. UL82 virion protein activates expression of immediate early viral genes in human cytomegalovirus-infected cells. Proc. Natl Acad. Sci. USA. 2000;97:14506–14511. doi: 10.1073/pnas.97.26.14506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Cantrell SR, Bresnahan WA. Interaction between the human cytomegalovirus UL82 gene product (pp71) and hDaxx regulates immediate-early gene expression and viral replication. J. Virology. 2005;79:7792–7802. doi: 10.1128/JVI.79.12.7792-7802.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hwang J, Kalejta RF. Proteasome-dependent, ubiquitin-independent degradation of Daxx by the viral pp71 protein in human cytomegalovirus-infected cells. Virology. 2007;367:334–338. doi: 10.1016/j.virol.2007.05.037. [DOI] [PubMed] [Google Scholar]
- 39.Cantrell SR, Bresnahan WA. Human cytomegalovirus (HCMV) UL82 gene product (pp71) relieves hDaxx-mediated repression of HCMV replication. J. Virol. 2006;80:6188–6191. doi: 10.1128/JVI.02676-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Preston CM, Nicholl MJ. Role of the cellular protein hDaxx in human cytomegalovirus immediate-early gene expression. J. Gen. Virol. 2006;87:1113–1121. doi: 10.1099/vir.0.81566-0. [DOI] [PubMed] [Google Scholar]
- 41.Liu B, Stinski MF. Human cytomegalovirus contains a tegument protein that enhances transcription from promoters with upstream ATF and AP-1 cis-acting elements. J. Virol. 1992;66:4434–4444. doi: 10.1128/jvi.66.7.4434-4444.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hofmann H, Sindre H, Stamminger T. Functional interaction between the pp71 protein of human cytomegalovirus and the PML-interacting protein human Daxx. J. Virol. 2002;76:5769–5783. doi: 10.1128/JVI.76.11.5769-5783.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lee HR, Kim DJ, Lee JM, et al. Ability of the human cytomegalovirus IE1 protein to modulate sumoylation of PML correlates with its functional activities in transcriptional regulation and infectivity in cultured fibroblast cells. J. Virol. 2004;78:6527–6542. doi: 10.1128/JVI.78.12.6527-6542.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Muller S, Dejean A. Viral immediate-early proteins abrogate the modification by SUMO-1 of PML and Sp100 proteins, correlating with nuclear body disruption. J. Virol. 1999;73:5137–5143. doi: 10.1128/jvi.73.6.5137-5143.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45•.Woodhall DL, Groves IJ, Reeves MB, Wilkinson G, Sinclair JH. Human Daxx-mediated repression of human cytomegalovirus gene expression correlates with a repressive chromatin structure around the major immediate early promoter. J. Biol. Chem. 2006;281:37652–37660. doi: 10.1074/jbc.M604273200.Combined with other studies from this laboratory, this paper demonstrated that the chromatin architecture assembled initially on infecting human cytomegalovirus (HCMV) genomes is indistinguishable in multiple cell types.
- 46•.Negorev DG, Vladimirova OV, Ivanov A, Rauscher F, 3rd, Maul GG. Differential role of Sp100 isoforms in interferon-mediated repression of herpes simplex virus type 1 immediate-early protein expression. J. Virol. 2006;80:8019–8029. doi: 10.1128/JVI.02164-05.Demonstrated that PML-NB proteins restrict HSV-1 infection.
- 47•.Everett RD, Rechter S, Papior P, Tavalai N, Stamminger T, Orr A. PML contributes to a cellular mechanism of repression of herpes simplex virus type 1 infection that is inactivated by ICP0. J. Virol. 2006;80:7995–8005. doi: 10.1128/JVI.00734-06.Demonstrated that PML-NB proteins restrict HSV-1 infection.
- 48.Everett RD, Parada C, Gripon P, Sirma H, Orr A. Replication of ICP0 null mutant herpes simplex virus type 1 is restricted by both PML and Sp100. J. Virol. 2008;82(6):2661–2672. doi: 10.1128/JVI.02308-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Chee AV, Lopez P, Pandolfi PP, Roizman B. Promyelocytic leukemia protein mediates interferon-based anti-herpes simplex virus 1 effects. J. Virol. 2003;77:7101–7105. doi: 10.1128/JVI.77.12.7101-7105.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lopez P, Jacob RJ, Roizman B. Overexpression of promyelocytic leukemia protein precludes the dispersal of ND10 structures and has no effect on accumulation of infectious herpes simplex virus 1 or its proteins. J. Virol. 2002;76:9355–9367. doi: 10.1128/JVI.76.18.9355-9367.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Nevels M, Paulus C, Shenk T. Human cytomegalovirus immediate-early 1 protein facilitates viral replication by antagonizing histone deacetylation. Proc. Natl Acad. Sci. USA. 2004;101:17234–17239. doi: 10.1073/pnas.0407933101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Preston CM, McFarlane M. Cytodifferentiating agents affect the replication of herpes simplex virus type 1 in the absence of functional VP16. Virology. 1998;249:418–426. doi: 10.1006/viro.1998.9314. [DOI] [PubMed] [Google Scholar]
- 53.Poon AP, Liang Y, Roizman B. Herpes simplex virus 1 gene expression is accelerated by inhibitors of histone deacetylases in rabbit skin cells infected with a mutant carrying a cDNA copy of the infected-cell protein no. 0. J. Virol. 2003;77:12671–12678. doi: 10.1128/JVI.77.23.12671-12678.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Terry-Allison T, Smith CA, DeLuca NA. Relaxed repression of herpes simplex virus type 1 genomes in murine trigeminal neurons. J. Virol. 2007;81:12394–12405. doi: 10.1128/JVI.01068-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Arthur JL, Scarpini CG, Connor V, Lachmann RH, Tolkovsky AM, Efstathiou S. Herpes simplex virus type 1 promoter activity during latency establishment, maintenance, and reactivation in primary dorsal root neurons in vitro. J. Virol. 2001;75:3885–3895. doi: 10.1128/JVI.75.8.3885-3895.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Danaher RJ, Jacob RJ, Steiner MR, Allen WR, Hill JM, Miller CS. Histone deacetylase inhibitors induce reactivation of herpes simplex virus type 1 in a latency-associated transcript-independent manner in neuronal cells. J. Neurovirol. 2005;11:306–317. doi: 10.1080/13550280590952817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lomonte P, Thomas J, Texier P, Caron C, Khochbin S, Epstein AL. Functional interaction between class II histone deacetylases and ICP0 of herpes simplex virus type 1. J. Virol. 2004;78:6744–6757. doi: 10.1128/JVI.78.13.6744-6757.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Gu H, Roizman B. Herpes simplex virus-infected cell protein 0 blocks the silencing of viral DNA by dissociating histone deacetylases from the CoREST-REST complex. Proc. Natl Acad. Sci. USA. 2007;104:17134–17139. doi: 10.1073/pnas.0707266104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Gu H, Liang Y, Mandel G, Roizman B. Components of the REST/CoREST/histone deacetylase repressor complex are disrupted, modified, and translocated in HSV-1-infected cells. Proc. Natl Acad. Sci. USA. 2005;102:7571–7576. doi: 10.1073/pnas.0502658102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Coleman HM, Connor V, Cheng ZS, Grey F, Preston CM, Efstathiou S. Histone modifications associated with herpes simplex virus type 1 genomes during quiescence and following ICP0-mediated de-repression. J. Gen. Virol. 2008;89:68–77. doi: 10.1099/vir.0.83272-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Triezenberg SJ. Structure and function of transcriptional activation domains. Curr. Opin. Genet. Dev. 1995;5:190–196. doi: 10.1016/0959-437x(95)80007-7. [DOI] [PubMed] [Google Scholar]
- 62.Wysocka J, Herr W. The herpes simplex virus VP16-induced complex: the makings of a regulatory switch. Trends Biochem. Sci. 2003;28:294–304. doi: 10.1016/S0968-0004(03)00088-4. [DOI] [PubMed] [Google Scholar]
- 63.Flint J, Shenk T. Viral transactivating proteins. Annu. Rev. Genet. 1997;31:177–212. doi: 10.1146/annurev.genet.31.1.177. [DOI] [PubMed] [Google Scholar]
- 64.Roizman B, Knipe DM, Whitley RJ. Herpes simplex viruses. In: Knipe DM, Howley PM, editors. Fields Virology (2) Lippincott Williams and Wilkins; Philadelphia, PA, USA: 2007. pp. 2501–2602. [Google Scholar]
- 65.Herrera FJ, Triezenberg SJ. VP16-dependent association of chromatin-modifying coactivators and underrepresentation of histones at immediate-early gene promoters during herpes simplex virus infection. J. Virol. 2004;78:9689–9696. doi: 10.1128/JVI.78.18.9689-9696.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Gu B, DeLuca N. Requirements for activation of the herpes simplex virus glycoprotein C promoter in vitro by the viral regulatory protein ICP4. J. Virol. 1994;68:7953–7965. doi: 10.1128/jvi.68.12.7953-7965.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Grondin B, DeLuca N. Herpes simplex virus type 1 ICP4 promotes transcription preinitiation complex formation by enhancing the binding of TFIID to DNA. J. Virol. 2000;74:11504–11510. doi: 10.1128/jvi.74.24.11504-11510.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Carrozza MJ, DeLuca NA. Interaction of the viral activator protein ICP4 with TFIID through TAF250. Mol. Cell Biol. 1996;16:3085–3093. doi: 10.1128/mcb.16.6.3085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Sampath P, Deluca NA. Binding of ICP4, TATA-binding protein, and RNA polymerase II to herpes simplex virus, type 1 immediate early, early and late promoters in virus-infected cells. J. Virol. 2008;82(5):2339–2349. doi: 10.1128/JVI.02459-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Roberts MS, Boundy A, O’Hare P, Pizzorno MC, Ciufo DM, Hayward GS. Direct correlation between a negative autoregulatory response element at the cap site of the herpes simplex virus type 1 IE175 (alpha 4) promoter and a specific binding site for the IE175 (ICP4) protein. J. Virol. 1988;62:4307–4320. doi: 10.1128/jvi.62.11.4307-4320.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Michael N, Roizman B. Repression of the herpes simplex virus 1 alpha 4 gene by its gene product occurs within the context of the viral genome and is associated with all three identified cognate sites. Proc. Natl Acad. Sci. USA. 1993;90:2286–2290. doi: 10.1073/pnas.90.6.2286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Leopardi R, Michael N, Roizman B. Repression of the herpes simplex virus 1 alpha 4 gene by its gene product (ICP4) within the context of the viral genome is conditioned by the distance and stereoaxial alignment of the ICP4 DNA binding site relative to the TATA box. J. Virol. 1995;69:3042–3048. doi: 10.1128/jvi.69.5.3042-3048.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Castillo JP, Kowalik TF. Human cytomegalovirus immediate early proteins and cell growth control. Gene. 2002;290:19–34. doi: 10.1016/s0378-1119(02)00566-8. [DOI] [PubMed] [Google Scholar]
- 74.Mocarski ES, Shenk T, Pass RF. Cytomegaloviruses. In: Knipe DM, Howley PM, editors. Fields Virology. Lippincott Williams and Wilkins; Philadelphia, PA, USA: 2007. pp. 2701–2772. [Google Scholar]
- 75.Park JJ, Kim YE, Pham HT, Kim ET, Chung YH, Ahn JH. Functional interaction of the human cytomegalovirus IE2 protein with histone deacetylase 2 in infected human fibroblasts. J. Gen. Virol. 2007;88:3214–3223. doi: 10.1099/vir.0.83171-0. [DOI] [PubMed] [Google Scholar]
- 76.Reeves M, Murphy J, Greaves R, Fairley J, Brehm A, Sinclair J. Autorepression of the human cytomegalovirus major immediate-early promoter/enhancer at late times of infection is mediated by the recruitment of chromatin remodeling enzymes by IE86. J. Virol. 2006;80:9998–10009. doi: 10.1128/JVI.01297-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Taylor-Wiedeman J, Sissons JG, Borysiewicz LK, Sinclair JH. Monocytes are a major site of persistence of human cytomegalovirus in peripheral blood mononuclear cells. J. Gen. Virol. 1991;72(Pt 9):2059–2064. doi: 10.1099/0022-1317-72-9-2059. [DOI] [PubMed] [Google Scholar]
- 78.Taylor-Wiedeman J, Sissons P, Sinclair J. Induction of endogenous human cytomegalovirus gene expression after differentiation of monocytes from healthy carriers. J. Virol. 1994;68:1597–1604. doi: 10.1128/jvi.68.3.1597-1604.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Hahn G, Jores R, Mocarski ES. Cytomegalovirus remains latent in a common precursor of dendritic and myeloid cells. Proc. Natl Acad. Sci. USA. 1998;95:3937–3942. doi: 10.1073/pnas.95.7.3937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Khaiboullina SF, Maciejewski JP, Crapnell K, et al. Human cytomegalovirus persists in myeloid progenitors and is passed to the myeloid progeny in a latent form. Br. J. Haematol. 2004;126:410–417. doi: 10.1111/j.1365-2141.2004.05056.x. [DOI] [PubMed] [Google Scholar]
- 81.Goodrum F, Reeves M, Sinclair J, High K, Shenk T. Human cytomegalovirus sequences expressed in latently infected individuals promote a latent infection in vitro. Blood. 2007;110:937–945. doi: 10.1182/blood-2007-01-070078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Goodrum FD, Jordan CT, High K, Shenk T. Human cytomegalovirus gene expression during infection of primary hematopoietic progenitor cells: a model for latency. Proc. Natl Acad. Sci. USA. 2002;99:16255–16260. doi: 10.1073/pnas.252630899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83••.Reeves MB, Lehner PJ, Sissons JG, Sinclair JH. An in vitro model for the regulation of human cytomegalovirus latency and reactivation in dendritic cells by chromatin remodelling. J. Gen. Virol. 2005;86:2949–2954. doi: 10.1099/vir.0.81161-0.Indication that the chromatin structure that appears to modulate HCMV gene expression during latency can be recapitulated in vitro.
- 84.Reeves MB, MacAry PA, Lehner PJ, Sissons JG, Sinclair JH. Latency, chromatin remodeling, and reactivation of human cytomegalovirus in the dendritic cells of healthy carriers. Proc. Natl Acad. Sci. USA. 2005;102:4140–4145. doi: 10.1073/pnas.0408994102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Cheung AK, Abendroth A, Cunningham AL, Slobedman B. Viral gene expression during the establishment of human cytomegalovirus latent infection in myeloid progenitor cells. Blood. 2006;108:3691–3699. doi: 10.1182/blood-2005-12-026682. [DOI] [PubMed] [Google Scholar]
- 86.Sinclair J, Sissons P. Latency and reactivation of human cytomegalovirus. J. Gen. Virol. 2006;87:1763–1779. doi: 10.1099/vir.0.81891-0. [DOI] [PubMed] [Google Scholar]
- 87.Murphy JC, Fischle W, Verdin E, Sinclair JH. Control of cytomegalovirus lytic gene expression by histone acetylation. EMBO J. 2002;21:1112–1120. doi: 10.1093/emboj/21.5.1112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Ioudinkova E, Arcangeletti MC, Rynditch A, et al. Control of human cytomegalovirus gene expression by differential histone modifications during lytic and latent infection of a monocytic cell line. Gene. 2006;384:120–128. doi: 10.1016/j.gene.2006.07.021. [DOI] [PubMed] [Google Scholar]
- 89.Keller MJ, Wu AW, Andrews JI, McGonagill PW, Tibesar EE, Meier JL. Reversal of human cytomegalovirus major immediate-early enhancer/promoter silencing in quiescently infected cells via the cyclic-AMP signaling pathway. J. Virol. 2007;81:6669–6681. doi: 10.1128/JVI.01524-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Meier JL. Reactivation of the human cytomegalovirus major immediate-early regulatory region and viral replication in embryonal NTera2 cells: role of trichostatin A, retinoic acid, and deletion of the 21-base-pair repeats and modulator. J. Virol. 2001;75:1581–1593. doi: 10.1128/JVI.75.4.1581-1593.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Yee LF, Lin PL, Stinski MF. Ectopic expression of HCMV IE72 and IE86 proteins is sufficient to induce early gene expression but not production of infectious virus in undifferentiated promonocytic THP-1 cells. Virology. 2007;363:174–188. doi: 10.1016/j.virol.2007.01.036. [DOI] [PubMed] [Google Scholar]
- 92••.Saffert RT, Kalejta RF. Human cytomegalovirus gene expression is silenced by Daxx-mediated intrinsic immune defense in model latent infections established in vitro. J. Virol. 2007;81:9109–9120. doi: 10.1128/JVI.00827-07.Evidence that a PML-NB protein regulates gene expression during a quiescent HCMV infection.
- 93.Meier JL, Stinski MF. Regulation of human cytomegalovirus immediate-early gene expression. Intervirology. 1996;39:331–342. doi: 10.1159/000150504. [DOI] [PubMed] [Google Scholar]
- 94.Groves IJ, Sinclair JH. Knockdown of hDaxx in normally non-permissive undifferentiated cells does not permit human cytomegalovirus immediate-early gene expression. J. Gen. Virol. 2007;88:2935–2940. doi: 10.1099/vir.0.83019-0. [DOI] [PubMed] [Google Scholar]
- 95.La Boissiere S, Hughes T, O’Hare P. HCF-dependent nuclear import of VP16. EMBO J. 1999;18:480–489. doi: 10.1093/emboj/18.2.480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Kristie TM, Vogel JL, Sears AE. Nuclear localization of the C1 factor (host cell factor) in sensory neurons correlates with reactivation of herpes simplex virus from latency. Proc. Natl Acad. Sci. USA. 1999;96:1229–1233. doi: 10.1073/pnas.96.4.1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Sears AE, Hukkanen V, Labow MA, Levine AJ, Roizman B. Expression of the herpes simplex virus 1 alpha transinducing factor (VP16) does not induce reactivation of latent virus or prevent the establishment of latency in mice. J. Virol. 1991;65:2929–2935. doi: 10.1128/jvi.65.6.2929-2935.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Samaniego LA, Neiderhiser L, DeLuca NA. Persistence and expression of the herpes simplex virus genome in the absence of immediate-early proteins. J. Virol. 1998;72:3307–3320. doi: 10.1128/jvi.72.4.3307-3320.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Preston CM, Nicholl MJ. Human cytomegalovirus tegument protein pp71 directs long-term gene expression from quiescent herpes simplex virus genomes. J. Virol. 2005;79:525–535. doi: 10.1128/JVI.79.1.525-535.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Minaker RL, Mossman KL, Smiley JR. Functional inaccessibility of quiescent herpes simplex virus genomes. Virol. J. 2005;2:85. doi: 10.1186/1743-422X-2-85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Hobbs WE, Brough DE, Kovesdi I, DeLuca NA. Efficient activation of viral genomes by levels of herpes simplex virus ICP0 insufficient to affect cellular gene expression or cell survival. J. Virol. 2001;75:3391–3403. doi: 10.1128/JVI.75.7.3391-3403.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Chen SH, Lee LY, Garber DA, Schaffer PA, Knipe DM, Coen DM. Neither LAT nor open reading frame P mutations increase expression of spliced or intron-containing ICP0 transcripts in mouse ganglia latently infected with herpes simplex virus. J. Virol. 2002;76:4764–4772. doi: 10.1128/JVI.76.10.4764-4772.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Maillet S, Naas T, Crepin S, et al. Herpes simplex virus type 1 latently infected neurons differentially express latency-associated and ICP0 transcripts. J. Virol. 2006;80:9310–9321. doi: 10.1128/JVI.02615-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Mador N, Goldenberg D, Cohen O, Panet A, Steiner I. Herpes simplex virus type 1 latency-associated transcripts suppress viral replication and reduce immediate-early gene mRNA levels in a neuronal cell line. J. Virol. 1998;72:5067–5075. doi: 10.1128/jvi.72.6.5067-5075.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Thompson RL, Shieh MT, Sawtell NM. Analysis of herpes simplex virus ICP0 promoter function in sensory neurons during acute infection, establishment of latency, and reactivation in vivo. J. Virol. 2003;77:12319–12330. doi: 10.1128/JVI.77.22.12319-12330.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Bego M, Maciejewski J, Khaiboullina S, Pari G, Jeor S. Characterization of an antisense transcript spanning the UL81–82 locus of human cytomegalovirus. J. Virol. 2005;79:11022–11034. doi: 10.1128/JVI.79.17.11022-11034.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107•.Knipe DM, Cliffe A. Chromatin control of herpes simplex virus lytic and latent infection. Nat. Rev. Microbiol. 2008;6:211–221. doi: 10.1038/nrmicro1794.Thorough review of how the chromatin structure of the viral genome affects lytic and latent HSV-1 infections.
- 108.Adamson AL, Kenney S. Epstein—barr virus immediate-early protein BZLF1 is SUMO-1 modified and disrupts promyelocytic leukemia bodies. J. Virol. 2001;75:2388–2399. doi: 10.1128/JVI.75.5.2388-2399.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Stanton R, Fox JD, Caswell R, Sherratt E, Wilkinson GW. Analysis of the human herpesvirus-6 immediate-early 1 protein. J. Gen. Virol. 2002;83:2811–2820. doi: 10.1099/0022-1317-83-11-2811. [DOI] [PubMed] [Google Scholar]
- 110.Hoppe A, Beech SJ, Dimmock J, Leppard KN. Interaction of the adenovirus type 5 E4 Orf3 protein with promyelocytic leukemia protein isoform II is required for ND10 disruption. J. Virol. 2006;80:3042–3049. doi: 10.1128/JVI.80.6.3042-3049.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Ullman AJ, Reich NC, Hearing P. Adenovirus E4 ORF3 protein inhibits the interferon-mediated antiviral response. J. Virol. 2007;81:4744–4752. doi: 10.1128/JVI.02385-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Guccione E, Lethbridge KJ, Killick N, Leppard KN, Banks L. HPV E6 proteins interact with specific PML isoforms and allow distinctions to be made between different POD structures. Oncogene. 2004;23:4662–4672. doi: 10.1038/sj.onc.1207631. [DOI] [PubMed] [Google Scholar]
- 113.Bischof O, Nacerddine K, Dejean A. Human papillomavirus oncoprotein E7 targets the promyelocytic leukemia protein and circumvents cellular senescence via the Rb and p53 tumor suppressor pathways. Mol. Cell Biol. 2005;25:1013–1024. doi: 10.1128/MCB.25.3.1013-1024.2005. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 114.Florin L, Schafer F, Sotlar K, Streeck RE, Sapp M. Reorganization of nuclear domain 10 induced by papillomavirus capsid protein l2. Virology. 2002;295:97–107. doi: 10.1006/viro.2002.1360. [DOI] [PubMed] [Google Scholar]
- 115.Regad T, Saib A, Lallemand-Breitenbach V, Pandolfi PP, de The H, Chelbi-Alix MK. PML mediates the interferon-induced antiviral state against a complex retrovirus via its association with the viral transactivator. EMBO J. 2001;20:3495–3505. doi: 10.1093/emboj/20.13.3495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Greger JG, Katz RA, Ishov AM, Maul GG, Skalka AM. The cellular protein Daxx interacts with avian sarcoma virus integrase and viral DNA to repress viral transcription. J. Virol. 2005;79:4610–4618. doi: 10.1128/JVI.79.8.4610-4618.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Poleshko A, Palagin I, Zhang R, et al. Identification of cellular proteins that maintain retroviral epigenetic silencing: evidence for an antiviral response. J. Virol. 2007;82(5):2313–2323. doi: 10.1128/JVI.01882-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Chelbi-Alix MK, Quignon F, Pelicano L, Koken MH, de The H. Resistance to virus infection conferred by the interferon-induced promyelocytic leukemia protein. J. Virol. 1998;72:1043–1051. doi: 10.1128/jvi.72.2.1043-1051.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Bell P, Brazas R, Ganem D, Maul GG. Hepatitis delta virus replication generates complexes of large hepatitis delta antigen and antigenomic RNA that affiliate with and alter nuclear domain 10. J. Virol. 2000;74:5329–5336. doi: 10.1128/jvi.74.11.5329-5336.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Brasier AR, Spratt H, Wu Z, et al. Nuclear heat shock response and novel nuclear domain 10 reorganization in respiratory syncytial virus-infected a549 cells identified by high-resolution two-dimensional gel electrophoresis. J. Virol. 2004;78:11461–11476. doi: 10.1128/JVI.78.21.11461-11476.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Li XD, Makela TP, Guo D, et al. Hantavirus nucleocapsid protein interacts with the Fas-mediated apoptosis enhancer Daxx. J. Gen. Virol. 2002;83:759–766. doi: 10.1099/0022-1317-83-4-759. [DOI] [PubMed] [Google Scholar]
- 122.Limjindaporn T, Netsawang J, Noisakran S, et al. Sensitization to Fas-mediated apoptosis by dengue virus capsid protein. Biochem. Biophys. Res. Commun. 2007;362:334–339. doi: 10.1016/j.bbrc.2007.07.194. [DOI] [PubMed] [Google Scholar]
- 123.Herzer K, Weyer S, Krammer PH, Galle PR, Hofmann TG. Hepatitis C virus core protein inhibits tumor suppressor protein promyelocytic leukemia function in human hepatoma cells. Cancer Res. 2005;65:10830–10837. doi: 10.1158/0008-5472.CAN-05-0880. [DOI] [PubMed] [Google Scholar]
- 124.Chen JZ, Ji CN, Xu GL, et al. Daxx interacts with phage PhiC31 integrase and inhibits recombination. Nucleic Acids Res. 2006;34:6298–6304. doi: 10.1093/nar/gkl890. [DOI] [PMC free article] [PubMed] [Google Scholar]