Abstract
Listeria monocytogenes is a Gram-positive bacterium which grows in the cytoplasm of eukaryotic cells and can cause severe disease in immunocompromised individuals1,2. In murine systems CD8+ T lymphocytes have been shown to be important effectors of acquired protective immunity against L. monocytogenes3–5. Class I MHC-restricted CD8+ cytotoxic T lymphocytes (CTL), which lyse J774 macrophage-like targets infected with L. monocytogenes, are induced following in vivo injection of live organisms. Natural peptide epitopes derived from L. monocytogenes can be acid-extracted from heavily infected BALB/c spleens and detected by CTL. A CTL clone, B9, derived from a (BALB/c × C57BL/6)F1, (H–2d×b) mouse, recognizes one of these natural epitopes in an H–2Kd-restricted fashion. B9 also recognizes P815 (H–2d) mastocytoma cells transfected with the listeriolysin gene. To identify the region of the listeriolysin recognized by CTL we used the H–2Kd peptide-binding motif described by Rammensee and colleagues6 to synthesize 11 nonamer peptides. One of these peptides, listeriolysin 91–99, was recognized very efficiently by B9. This represents the first identified class I MHC-restricted epitope of bacteria and demonstrates the utility of the allele-specific motif for predicting CTL epitopes.
Strains of L. monocytogenes that express listeriolysin (LLO; relative molecular mass 59,000 (Mr 59K)) are virulent in mice and, when administered at a sublethal dose, induce protective immunity7. L. monocytogenes lacking LLO are avirulent and cannot induce immunity8–10. (BALB/c × C57BL/6)F1 mice were infected with a sublethal dose of virulent L. monocytogenes (ATCC 43251) and eight days later splenocytes were taken and stimulated in vitro with J774 macrophage-like cells (H–2d) infected with L. monocytogenes. After three cycles of stimulation the resulting T cells were >94% CD8+ with no detectable CD4+ cells (results not shown). One of these T cell lines, named LmT, lysed J774 cells only if they were infected with live, virulent L. monocytogenes (Fig. 1). J774 cells infected with avirulent L. monocytogenes that do not express LLO (ATCC 43250)11 or treated with heat-killed virulent L. monocytogenes were not lysed significantly above background (Fig. 1). These findings agree with recent studies showing that the stimulation of γ-interferon production by L. monocytogenes-specific CD8+ T cells requires infection of presenting cells with live, virulent organisms12.
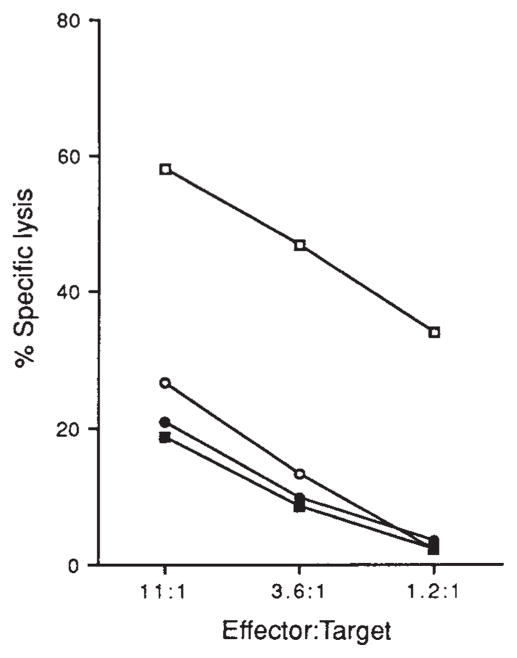
FIG. 1.
CTL lysis of J774 cells infected with L. monocytogenes. J774 macrophages were treated with virulent L. monocytogenes (□), avirulent L. monocytogenes (○), heat-killed virulent L. monocytogenes (●) or left untreated (■). Effectors were added to 51Cr-labelled targets at the ratios indicated and percentage 51Cr release was determined as described23.
METHODS. CTL line LmT was obtained from a (BALB/c × C57BL/6) F1 mouse 8 days after infection with 5 × 103 virulent L. monocytogenes (ATCC 43251). Flasks for in vitro stimulation were prepared by adding 5 × 106 J774 cells in 10 ml RP10 lacking antibiotics23. After 24 h the flasks were infected with 1 ml virulent L. monocytogenes in mid-log phase (A600 of 0.1) for 25 min, at which time the medium was replaced with RP10 containing 5μg ml−1 gentamicin. Three hours later medium was replaced with RP10 containing 100 U ml−1 penicillin, 100 μg ml−1 streptomycin and 50 μg ml−1 gentamicin and incubated overnight. Immune splenocytes (3×107) were added to the flasks (in 20 ml volume) and incubated upright. CTL were restimulated every 7–12 days and after the first two stimulations the media was suplemented with interleukin-2. CTL were assayed by labelling 106 J774 cells with 100 μCi 51Cr for 45 min in antibiotic-free media. Cells (104) were placed into wells and allowed to adhere for 30 min. Cells were then exposed to either 2 × 106 live or heat-killed (60 °C for 30 min) virulent or avirulent (ATCC 43250) L. monocytogenes for 25 min. Medium was replaced with RP10 containing 5 μg ml−1 gentamicin, CTL were added and lysis was allowed to proceed for 3 h. Spontaneous lysis of all target cells was <30%.
To define the target antigen specificity of the LmT CTL line, spleens from BALB/c (H–2d) mice infected with L. monocytogenes were extracted with 0.1% trifluoroacetic acid and material of less than Mr 5K was separated by reversed-phase high-pressure liquid chromatography (HPLC). Individual fractions were then tested for the presence of peptide antigens derived from L. monocytogenes by their ability to target CTL killing. LmT detected two main peaks (fractions 28 and 30) and one minor peak (fraction 35) of targeting activity on P815 (H–2d) cells (Fig. 2a). This targeting activity is MHC-restricted as the BALB/c trifluoroacetic acid extracts did not sensitize EL4 (H–2b) cells and is L. monocytogenes dependent as uninfected BALB/c spleen extracts did not sensitize P815 targets. Additionally, trifluoroacetic acid extracts of infected C57BL/6 (H–2b) spleens did not sensitize H–2d targets. L. monocytogenes- specific CD8+ T lymphocytes that are not H–2 restricted have been described13,14. Although CTL line LmT is H–2 restricted, we have generated other L. monocytogenes-specific CTL lines that recognize peptides extracted from infected allogeneic spleens in an H–2 unrestricted manner. The specificities of these CTL lines are being investigated.
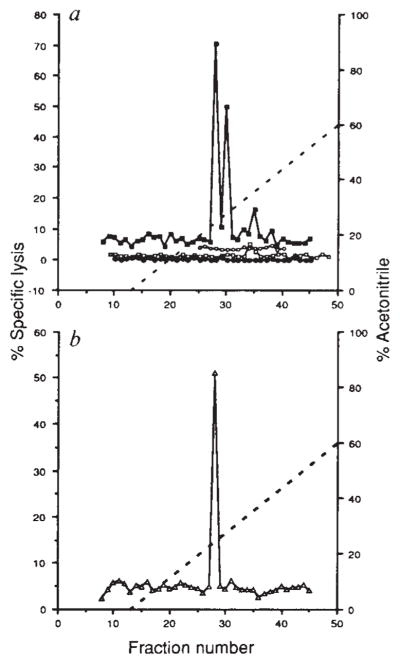
FIG. 2.
Recognition by CTL line LmT and clone B9 of natural peptides derived from L. monocytogenes by acid extraction of infected spleens, a, CTL line LmT was used to lyse 51Cr-labelled P815 cells coated with reversed-phase HPLC fractions of acid extracts from infected BALB/c (■) and C57BL/6 (○) spleens and uninfected BALB/c (□) spleens. EL4 cells were also used with HPLC fractions from infected BALB/c spleens (●). Effector to target ratio was 20:1. b, CTL clone B9 was assayed on P815 cells coated with HPLC fractions of acid extracts from infected BALB/c spleens (△). Effector to target ratio was 20:1.
METHODS. BALB/c mice were infected with 5×105 L. monocytogenes and C57BL/6 mice were infected with 106 L. monocytogenes. After 48 h the spleens were taken and homogenized sequentially with a tissue grinder and dounce homogenizer in 0.1% trifluoroacetic acid (TFA) and sonicated as described24. This material was centrifuged at 100,000g for 30 min and the supernatant was passed over a Sephadex G-25 column. Material of less than 5K was collected and lyophilized. HPLC of the acid extract from one spleen in 0.1% TFA was performed with a C18 300A reversed-phase column using a 0–60% gradient of acetonitrile with 0.1% TFA at a rate of 1 ml min−1 and 1 ml fractions were collected. Fractions were lyophilized and suspended in 100 μl PBS. CTL assays were performed for 4 h with LmT and B9 effectors using either P815 or EL4 target cells in 200-μ1 wells, of which 50 μl represented the HPLC fraction.
CTL clone B9, which was derived from the LmT line by limiting dilution, is specific only for fraction 28 of the infected BALB/c spleen extract (Fig. 2b). H–2Kd-specific antibodies blocked lysis of P815 cells coated with fraction 28 whereas the H–2Dd-specific antibodies did not (Fig. 3a), indicating that the L. monocytogenes-derived peptide in fraction 28 is presented by H–2Kd. We were able to determine which bacterial protein provided this epitope as a P815 cell line transfected with the LLO gene (PHem3) was lysed by LmT and the B9 CTL clone. Lysis of PHem3 was also blocked by H-2Kd-specific antibodies (Fig. 3b).
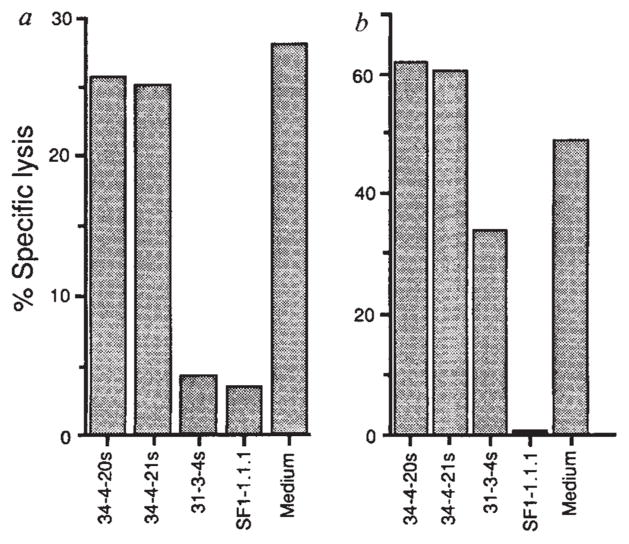
FIG. 3.
Presentation of HPLC Fraction 28 by H-2Kd and lysis of P815 cells transfected with LLO. a, P815 cells were coated with HPLC fraction 28 peptides and exposed to LmT CTL in the presence of monoclonal antibodies 34-4-20s and 34-4-21s, which are specific for H-2Da, and 31-3-4s25 and SF1-1.1.1, which are specific for H-2Kd. b, P815 cells transfected with LLO (PHem3) were used as targets for LmT CTL in the presence and absence of anti-H-2d antibodies as in a.
METHODS. P815 and PHem3 cells were labelled with 51Cr and suspended in culture supernatants from hybridomas 34-4-20s, 34-4-21s, 31-3-4s, SF1-1.1.1 and control medium. Fraction 28 peptides were added to the P815 targets. LmT effectors were added to targets at a ratio of 20:1 and the specific lysis was determined after 3 h. PHem3 was obtained by transfecting P815 cells with linearized pHβAPr-1-neo26 containing the entire LLO gene17. Bases 1-1,587 of the hlyA gene encoding LLO were cloned into the BamHI site of the pHβAPr-1-neo expression vector after polymerase chain reaction (PCR) amplification from L. monocytogenes DNA. The oligonucleotides used for priming were 5′-CCCGGGATCCACCATGAAAAAAATAATGCTAG-3′ and 5′-GGATCCGGATCCTTATTCGATTGGATTATC-3′ which introduced flanking BamHI sites. Electroporation of PvuI linearized plasmid containing the insert into P815 cells and selection of G418 resistant clones was performed as described23. P815 cells transfected with the vector without the insert were not lysed by the LmT CTL line (not shown).
The exact sequences of certain peptides that are derived from endogenously synthesized viral proteins and associate with class I MHC molecules have recently been determined15,16. Furthermore, sequencing the mixture of peptides associated with purified class I molecules has led to the prediction of MHC allele-specific peptide-binding motifs6. Thus, it has been proposed that H-2Kd-associated peptides are nine residues long and contain tyrosine at position 2 and either leucine or isoleucine at the C terminus6. The predicted LLO sequence17 contains five such motifs (Fig. 4a). In addition, some identified foreign epitopes have valine, alanine or threonine at the C terminus. According to these criteria we synthesized 11 nonamers, representing only 18% of the LLO molecule, and tested them for their ability to sensitize P815 target cells to lysis by H-2Kd-restricted, LLO-specific CTL. Amino-acid residues 91–99 represent the H-2Kd LLO epitope recognized by B9 and this peptide can target cells at concentrations less than 10−11 M (Fig. 4b). The precise molecular mass and partial amino-acid sequence of synthetic LLO 91–99 were confirmed by tandem ion spray mass spectroscopy. The other 10 peptides were not detected by LmT, suggesting that LLO contains only one H-2Kd-restricted peptide. The exact comigration of synthetic LLO 91–99 with the CTL targeting activity in trifluoroacetic acid extracts from infected BALB/c spleens on a very shallow acetonitrile gradient (Fig. 4c, d) indicates that we have identified the natural epitope. Other regions of the LLO molecule contain epitopes recognized by L. monocytogenes-specific CD4+ T lymphocytes18. LLO 91–99 was not, however, contained within any of the previously predicted T lymphocyte epitopes18.
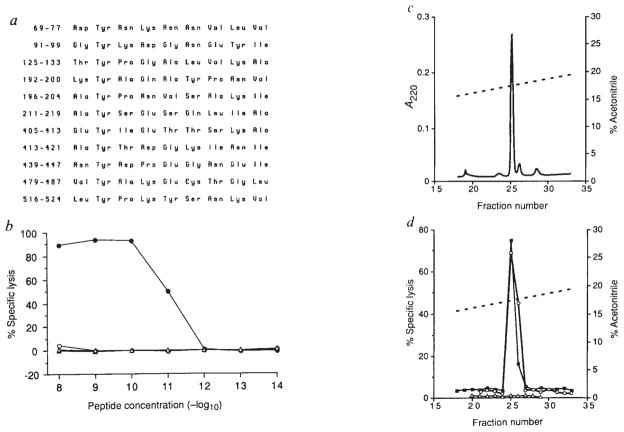
FIG. 4.
Exact sequence determination of the H-2Kd-restricted LLO peptide. a, Eleven nonamer peptides that conform to the H-2Kd-binding motif were found in the LLO amino-acid sequence and were synthesized on resin with the F-moc multiple peptide synthesis method (Cambridge Research Biochemicals). The amino acids are numbered according to the sequence of LLO. b, All 11 peptides were used with P815 target cells and CTL assays were performed for 3 h with B9 CTL at an effector to target ratio of 20:1. LLO 91–99 sensitized targets (●) whereas LLO 196–204 (○), LLO 211–219 (△) and all other eight peptides (not shown) did not. c, HPLC A220 profile of 2 μg synthetic LLO 91–99 (—) on a shallow acetonitrile gradient (broken line), d, CTL assays of shallow gradient HPLC fractions of 2 μg LLO 91–99 (each fraction diluted 10−6 for assay) (○), mock HPLC fractionation to assure that peptide contamination of the HPLC system had not occurred (△), and a TFA extract of a spleen from an L. monocytogenes-infected BALB/c mouse (■). B9 effectors were used to detect targeting activity on 51Cr-labelled P815 cells at a ratio of 10:1.
Phagocytosed, virulent L. monocytogenes secrete LLO which disrupts the phagolysosomal membrane and allows the bacterium to enter the host cell cytoplasm1. It has been proposed that only upon entry into the cytoplasm can L. monocytogenes antigens become processed and presented by class I MHC molecules12. This would explain why L. monocytogenes strains or mutants lacking LLO do not induce protective immunity. Although our findings are consistent with this hypothesis, they could also be interpreted to suggest that the absence of LLO in a virulent L. monocytogenes is responsible for their lack of immunogenicity. Although the immunogenicity of LLO has been demonstrated 18,19, further studies will be required to settle this controversy.
Identification of the CTL epitopes of infectious agents is important for vaccine development20–22. The diversity and number of proteins expressed by bacterial and protozoal pathogens has made the task of identifying CTL epitopes formidable. Determination of LLO 91–99 as a major H-2d-restricted CTL epitope was possible because of the recent identification of the binding motif of H-2Kd for endogenous peptides6. Our findings demonstrate the predictive power of this motif for the determination of a pathogen-derived CTL epitope and argue the case for the systematic determination of binding motifs of other class I molecules.
Acknowledgments
We thank S. Jameson for discussion. Supported by the National Institute of Allergy and Infectious Diseases of the NIH, and the Howard Hughes Medical Institute.
References
- 1.Tilney LG, Portnoy DA. J Cell Biol. 1989;109:1597–1608. doi: 10.1083/jcb.109.4.1597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Marget W, Seeliger HPR. Infection. 1988;16:175–177. [Google Scholar]
- 3.Kaufmann SHE, Hug E, DeLibero G. J exp Med. 1986;164:363–368. doi: 10.1084/jem.164.1.363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bishop DK, Hinrichs DJ. J Immun. 1987;139:2005–2009. [PubMed] [Google Scholar]
- 5.Mielke MEA, Ehlers S, Hahn H. Infect Immun. 1988;56:1920–1925. doi: 10.1128/iai.56.8.1920-1925.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Falk K, Roetzschke O, Stevanovic S, Jung G, Rammensee H-G. Nature. 1991;351:290–296. doi: 10.1038/351290a0. [DOI] [PubMed] [Google Scholar]
- 7.Mackaness GB. J exp Med. 1962;116:381–406. [Google Scholar]
- 8.Gaillard JL, Berche P, Sansonetti P. Infect Immun. 1986;52:50–55. doi: 10.1128/iai.52.1.50-55.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kathariou S, Metz P, Hof H, Goebel W. J Bact. 1987;169:1291–1297. doi: 10.1128/jb.169.3.1291-1297.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Portnoy DA, Jacks PS, Hinrichs DJ. J exp Med. 1988;167:1459–1471. doi: 10.1084/jem.167.4.1459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pine L, et al. J clin Microbiol. 1987;25:2247–2251. doi: 10.1128/jcm.25.11.2247-2251.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Brunt LM, Portnoy DA, Unanue ER. J Immun. 1990;145:3540–3546. [PubMed] [Google Scholar]
- 13.DeLibero G, Kaufmann SHE. J Immun. 1986;137:2688–2694. [Google Scholar]
- 14.Lukacs K, Kurlander RJ. J Immun. 1989;143:3731–3736. [PubMed] [Google Scholar]
- 15.Van Bleek GM, Nathenson SG. Nature. 1990;348:213–216. doi: 10.1038/348213a0. [DOI] [PubMed] [Google Scholar]
- 16.Roetzschke O, et al. Nature. 1990;348:252–254. doi: 10.1038/348252a0. [DOI] [PubMed] [Google Scholar]
- 17.Mengaud J, et al. Infect Immun. 1988;56:766–772. doi: 10.1128/iai.56.4.766-772.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Safley SA, Cluff CW, Marshall NE, Ziegler HK. J Immun. 1991;146:3604–3616. [PubMed] [Google Scholar]
- 19.Berche P, Gaillard J, Geoffrey C, Alouf JE. J Immun. 1987;139:3813–3821. [PubMed] [Google Scholar]
- 20.Deres K, Schild H, Wiesmueller K, Jung G, Rammensee HG. Nature. 1989;342:561–564. doi: 10.1038/342561a0. [DOI] [PubMed] [Google Scholar]
- 21.Aichele P, Hengartner H, Zinkernagel RF, Schulz M. J exp Med. 1990;171:1815–1820. doi: 10.1084/jem.171.5.1815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kast M, et al. Proc natn Acad Sci USA. 1991;88:2283–2287. doi: 10.1073/pnas.88.6.2283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Moore MW, Carbone FR, Bevan MJ. Cell. 1988;54:777–785. doi: 10.1016/s0092-8674(88)91043-4. [DOI] [PubMed] [Google Scholar]
- 24.Roetzschke O, Falk K, Wallny HJ, Faath S, Rammensee HG. Science. 1990;249:283–287. doi: 10.1126/science.1695760. [DOI] [PubMed] [Google Scholar]
- 25.Ozato K, Mayer NM, Sachs DH. Transplantation. 1982;34:113–120. doi: 10.1097/00007890-198209000-00001. [DOI] [PubMed] [Google Scholar]
- 26.Gunning P, Leavitt J, Muscat G, Sun-Yu N, Kedes L. Proc natn Acad Sci USA. 1987;84:4831–4835. doi: 10.1073/pnas.84.14.4831. [DOI] [PMC free article] [PubMed] [Google Scholar]