Abstract
The nuclear actin-related proteins (ARPs) share overall structure and low-level sequence homology with conventional actin. They are indispensable subunits of macromolecular machines that control chromatin remodeling and modification leading to dynamic changes in DNA structure, transcription, and DNA repair. Cellular, genetic, and biochemical studies suggest that the nuclear ARPs are essential to the epigenetic control of the cell cycle and cell proliferation in all eukaryotes, while in plants and animals they also exert epigenetic controls over most stages of multicellular development including organ initiation, the switch to reproductive development, and senescence and programmed cell death. A theme emerging from plants and animals is that in addition to their role in controlling the general compaction of DNA and gene silencing, isoforms of nuclear ARP-containing chromatin complexes have evolved to exert dynamic epigenetic control over gene expression and different phases of multicellular development. Herein, we explore this theme by examining nuclear ARP phylogeny, activities of ARP-containing chromatin remodeling complexes that lead to epigenetic control, expanding developmental roles assigned to several animal and plant ARP-containing complexes, the evidence that thousands of ARP complex isoforms may have evolved in concert with multicellular development, and ARPs in human disease.
Keywords: ARPs, Nucleosome, Epigenetic control, Chromatin remodeling, Histone variant, Multicellular development, SWI/SNF, SWR1, INO80
1. Introduction
The nuclear actin-related proteins (ARPs) were first discovered in the early 1990s, but their central role as components of chromatin remodeling and modifying machines has emerged only recently. Various studies in yeast, animals, and plants reveal that nuclear ARPs are essential to the epigenetic control of chromatin structure. Nuclear ARP-dependent epigenetic controls participate in DNA repair, chromosome segregation, and gene expression, which in turn regulate the cell cycle and cell proliferation, phase transitions in development, and senescence and programmed cell death (PCD). In plants and animals, these processes are at the heart of multicellular tissue and organ development. Due to the combinatorial complexity of the numerous subunits of these chromatin complexes, the nuclear ARPs may participate in hundreds if not thousands of different complex isoforms, with the potential to evolve novel activities essential to the macroevolution of new tissues and organs. Nuclear ARP-containing machines are involved in an organism’s responses to environment, diet, and age, which are known to be key factors in the onset of numerous human diseases. In this broad context, we have attempted to review the evolution and functions of the nuclear ARPs in regulating cell and organismal development.
2. Nuclear ARPs as Epigenetic Factors
2.1. Structural and sequence identity
The ARPs share limited sequence identity with conventional actins, but they appear to maintain the actin-fold, a potential nucleotide-binding pocket and hinged structure that enables a conformational change in actin family members. Eight to ten ancient classes of ARPs are found in most eukaryotes that have been examined, and all appear to participate in protein complexes (Chen and Shen, 2007; Muller et al., 2005). A subset of the more divergent ARPs can be identified in most eukaryotes as homologues of yeast Arp4, Arp5, Arp6, and Arp8, which are found primarily in the nucleus (Harata et al., 2000; Kandasamy et al., 2003; Shen et al., 2003b). The predominantly nuclear localization distinguishes them from actin or Arp2 and Arp3, which are detected in the nucleus, but are primarily concentrated in the cytoplasm and participate in cytoskeletal functions (Harries et al., 2005). None of the nuclear ARPs are known to form polymers such as F-actin microfilaments or the short filaments formed from ARP1 and ARP10 in the centractin complex (Eckley and Schroer, 2003). Instead, nuclear ARPs act as essential subunits of macromolecular machines controlling chromatin dynamics with subsequent effects on transcription and DNA repair.
2.2. Exclusive function in chromatin remodeling and modifying machines
The only functions demonstrated for nuclear ARPs are as subunits of chromatin remodeling and modifying complexes (Chen and Shen, 2007). Specifically, ARP-containing remodeling complexes are responsible for nucleosome phasing and movement and exchange of histone subunit isovariants within nucleosomes (nucleosome remodeling (NR) complexes). ARP-containing chromatin modification complexes direct posttranscriptional modifications to histone amino acid residues such as the acetylation or methylation of lysine residues. The dynamic activities of chromatin remodeling and modifying complexes serve a basal regulatory function by reinforcing or alleviating the nucleosomal suppression of transcription that affects most genes due to the extreme compaction of DNA in nuclei or nucleoids (Yuan et al., 2005). Conversely, because there is only room for a small fraction of any genome to be in an open chromatin conformation and transcriptionally active at any one time, chromatin remodeling is a dynamic process necessary to unfold and refold various loci throughout the cell and developmental cycles of the organism. Although most of these chromatin activities appear global in nature, the nuclear ARPs may also exert more precise epigenetic control over the activities of particular regulatory genes that have an overriding influence on cell and organ ontogeny or on the response to environmental stress (Jonsson et al., 2004; Meagher et al., 2007; Oma et al., 2003; Wu et al., 2007).
2.3. Distinguishing genetic from epigenetic controls
For most of the last century, it was presumed that only genetic mutation provided the variation acted on by natural selection and neutral drift, thus leading to the evolution of new organisms and new structures within organisms. After the 1960s, with the knowledge that regulatory genes encode transcription factors that act on specific DNA-binding motifs, a more defined molecular genetic model emerged. Such interactions between transcription factors and cis-DNA elements produce linear or branched information cascades that directly control a few to dozens of genes and subsequently development. Mutations in these sequences are then acted on during evolution. However, as the gene compositions of multicellular plants and animals were determined, it became increasingly difficult to explain the significant physical differences among organisms and their rapid rates of morphological change from common ancestors as resulting only from gene duplications and mutant alleles affecting linear pathways of developmental control.
There were problems with this simple gene and cis-element mutation model for morphological evolution. If chimpanzees have the same genes and nearly identical protein sequences as humans (Kehrer-Sawatzki and Cooper, 2007; Wilson and Sarich, 1969) and if the Tetraodontoid fish, Fugu rubripes, has a similar gene composition as mammals (Aparicio et al., 2002), how do we account for the remarkable diversity of organisms and organ structures that evolved within chordates and vertebrates over the last 650 million years? Similar questions are asked about plant genomes and the rates of plant morphological evolution for the last 600 million years, since vascular plants evolved from common ancestors with moss. Again, the rapid rates of morphological evolution and macroevolution in higher plants and animals far out strips the observed rates of gene duplication, mutation, and evolution of new protein sequences. Regulatory genes and DNA motifs may accumulate mutations at slightly faster rates than structural genes (Elango et al., 2008), but probably not fast enough to fully explain these rapid changes in morphology. This dilemma is at least partially addressed by the view that natural selection may be acting on mutations in epigenetic control systems that network chromatin structure and gene expression. Mutations in a single epigenetic factor can alter the expression of thousands of genes in multiple gene networks and can have widely varied and pleiotropic consequences on phenotype in yeast, plants, or animals (Meagher et al., 2005; Morgan et al., 1999; Roux-Rouquie, 2000; True et al., 2004). Epigenetic control adds a level of complexity to the inter regulation of gene activities that is now widely recognized as essential to multicellular development and macroevolution.
2.4. The nuclear ARPs are epigenetic factors
The majority of epigenetic controls are exerted by changes to chromatin structure leading to alterations in gene expression. As mentioned earlier, the nuclear ARP-dependent modifications of chromatin include histone modifications and NR. These activities contribute to “epigenetic control” of cell development, a phrase that David Nanney used to define “inherited differences between two cells that are not due to changes in DNA sequence” (Haig, 2004; Nanney, 1958). Nanney’s definition of epigenetic control and the roles we infer for nuclear ARPs applys to two differentiating yeast or protozoan cells that have divided, just as well as it does to two differentiating cells within a mammalian limb bud, insect imaginal disk, or plant organ primordia. Nuclear ARPs also function in the epigenetic control of chromosome segregation and DNA repair (Minoda et al., 2005). Therefore, most of the direct functions of nuclear ARPs fit within this definition making them fundamental if not essential factors in epigenetic control.
However, the indirect roles of nuclear ARPs in the regulation of pathways of multicellular development also fit a slightly older definition of epigenetics as “the interactions among cells and cell products that lead to morphogenesis and differentiation” (Haig, 2004; Waddington, 1957). Waddington focused on defining the inherited differences and interactions among cells of a differentiating multicellular organ. This definition of epigenetics particularly takes into account the interactions and networking of information that results from diffusible signaling molecules moving between subsets of cells in an organ primordia. While nuclear ARPs do not diffuse between cells or act as receptors themselves, their chromatin activities are associated with the differential expression of many genes in receptor-mediated intracellular signaling pathways directing plant and animal organ development (Mehler, 2008b; Walley et al., 2008; Williams and Fletcher, 2005). Furthermore, most scientists today do not include this concept of signals diffusing between cells as part of modern epigenetics. Waddington’s definition of epigenetics also does not encompass the role of nuclear ARPs in chromosome segregation or DNA repair, in generational epigenetic inheritance, or in inheritance of epigenetic differences between dividing single cell fungi or protists.
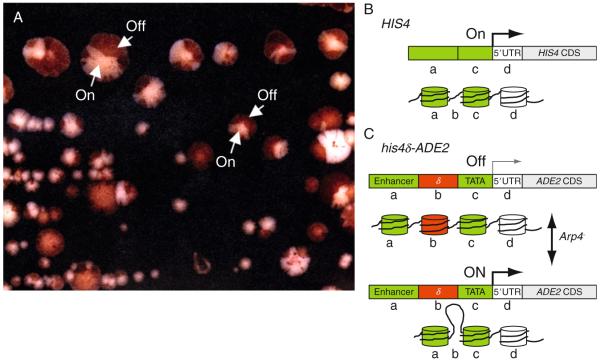
While Nanny and Waddington had intimate familiarity with particular protist and mammalian models of development, yeast emerged early as model genetic system to dissect mechanisms of epigenetic control (Pillus and Rine, 1989). Shortly after the discovery of the nuclear ARPs, mutants in yeast ARP4 (originally Act3b) were isolated as suppressors of an epigenetic reporter his4δ-ADE2 (Jiang and Stillman, 1996). ARP4-defective yeast containing this reporter in an ade2− background show reversible and stochastic changes in the morphology of colored sectors of cells within single yeast colonies, red to white and back to red as shown in Fig. 5.1. The his4δ promoter contains an insertion δ (Fig. 5.1C) separating the HIS4 gene’s transcriptional enhancers from its TATA box (Fig. 5.1B) rendering the his4δ-ADE2 gene inactive “Off” in a wild-type yeast background. The cells produce red pigment when the reporter is inactive because they are still ade2−. In ARP4-defective yeast cells, the variegated colony color results from stochastic reversible modifications to the chromatin structure of the his4δ-ADE2 promoter turning the gene “On” (white cells) and “Off” again. By the model shown in Fig. 5.1C, in the arp4− background, changes occur to chromatin structure that allow the HIS4-enhancer to interact with the HIS4-TATA sequence turning “On” ADE2. The results of this study demonstrate that ARP4’s activities are involved in epigenetic control and not classical genetic control. In another more widely known example of loss of normal epigenetic control, stochastic changes to Drosophila eye color result from position effect variegation of white gene expression (Csink and Henikoff, 1996; Henikoff, 1979). The yeast colony color and eye color phenotypes highlight the stochastic changes in gene expression often associated with epigenetic defects. All the activities discovered so far for nuclear ARPs and their constituent remodeling complexes involve making transient changes to chromatin structure and are consistent with ARPs being defined as factors in epigenetic control.
Figure 5.1.
Sectored colony color morphology of ARP4-defective yeast results from an epigenetic defect. (A) Sectored colony color phenotype resulting from the loss of ARP4 function in an ade2− yeast strain containing the epigenetic reporter his4δ-ADE2 (Jiang and Stillman, 1996). Portions of two colonies where the his4δ-ADE2 reporter gene is “Off” (red) or “On” (white) are indicated with arrows. This reporter with a dysfunctional promoter is normally “Off” producing red colonies in ade2− yeast cells. The reporter is turned “On” stochastically in arp4− yeast cells. Photo, courtesy of David Stillman. (B) Normal HIS4 promoter and gene structure. (C) A model showing the structure of the his4δ-ADE2 epigenetic reporter gene in relation to possible changes in chromatin structure that render the gene transcriptionally “On” or “Off”.
3. Evolutionary Origin and Phylogeny of Nuclear ARPs
3.1. A class of nuclear proteins
3.1.1. ARP localization to the nucleus
Considering the decades of research on conventional actin as a cytoplasmic cytoskeletal protein, it was at first surprising to find a class of ARPs had evolved activity in the nucleus. Yeast ARP4 (ACT3b) was the first ARP to be found localized to the nucleus (Weber et al., 1995), followed closely by Drosophila ARP4 (Frankel et al., 1997), two isovariants of human ARP4 (Baf53A and Baf53B) (Harata et al., 1999a), and Drosophila and vertebrate ARP6 (Kato et al., 2001; Ohfuchi et al., 2006). Harata et al. (2000) delineate and define the entire class of nuclear ARPs by showing that yeast Arp5, Arp6, Arp7, Arp8, and Arp9 are also localized to the nucleus, while fungal Arp1, Arp2, Arp3, and Arp10 are predominantly in the cytoplasm (Eckley et al., 1999; Zhang et al., 2008). We have shown that Arabidopsis ARP4, ARP5, ARP6, ARP7, and ARP8 are also localized to the nucleus as illustrated in Fig. 5.2 (Deal et al., 2005; Kandasamy et al., 2003, 2008, 2009). Arabidopsis ARP8 is the first ARP found to be concentrated in the nucleolus and not in the nucleoplasm (Fig. 5.2).
Figure 5.2.
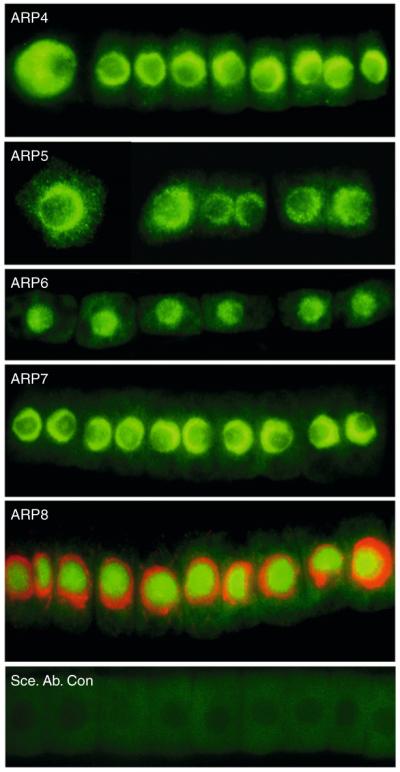
Nuclear localization of five plant ARPs. The nuclear localization of Arabidopsis ARP4, ARP5, ARP6, ARP7, and ARP8 were demonstrated using ARP-class-specific monoclonal antibodies prepared against the various recombinant plant proteins or synthetic peptides. ARP4, ARP5, ARP6, and ARP7 are concentrated in the nucleoplasm, while ARP8 is concentrated in the nucleolus. For the ARP8 image, DNA staining with DAPI (red) is merged with the ARP8-specific monoclonal antibody immunostaining (green). A control strip of root cells with secondary antibody labeling is shown in the bottom panel.
Most published studies used immunofluorescence microscopy or fluorescently tagged proteins to show the nuclear ARPs are concentrated in the nucleus during interphase, though they are mostly dissociated from chromatin during cell division (e.g., metaphase). For example, during telophase, anaphase, and metaphase we found weak immunofluorescent staining throughout most of the cell for Arabidopsis ARP4, ARP5, ARP6, ARP7, and ARP8 (Deal et al., 2005; Kandasamy et al., 2003, 2004, 2005a,b, 2008, 2009). Most of the nuclear ARPs appear to diffuse into the cytoplasm after the nuclear membrane breaks down during cell division and then are transported back and associated with chromatin before the nuclear membrane is reformed. This ebb and flow of ARP proteins is in contrast, for example, to the four core histones that stay concentrated on chromatin. The dissociation of the bulk of nuclear ARPs may be contrary to expectation, because chromatin is rapidly remodeled during all stages of the cell cycle.
Human ARP8 is the first among the nuclear ARPs to be identified as associating almost exclusively with mitotic chromatin in a dividing cell, and human ARP8 is essential to mitotic alignment of chromosomes (Aoyama et al., 2008). In addition, ARP4 was found weakly localized to the surface of mitotic chromatin in cultured mouse and human cells in more detailed immunofluorescence analyses of metaphase (Lee et al., 2007b). Genetic and biochemical studies also demonstrate that yeast ARP4 is associated with centromeric DNA during metaphase (Ogiwara et al., 2007b; Steinboeck et al., 2006). Its location here in previous studies may have been missed using standard microscopic techniques.
3.1.2. Nuclear transport
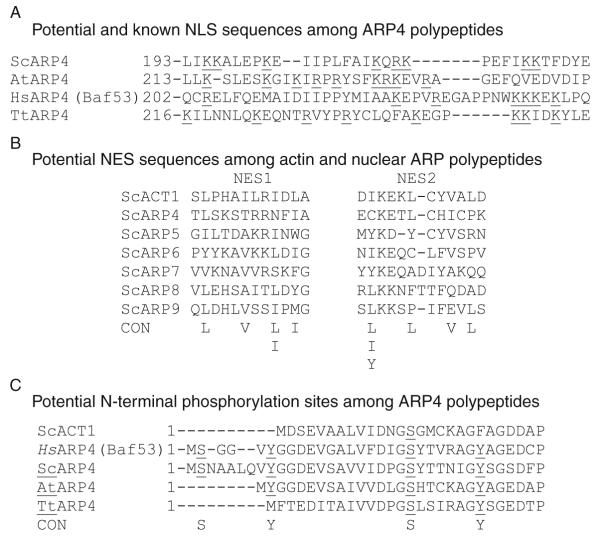
Nuclear ARPs are synthesized in the cytoplasm and transported into the nucleus, but the ratio of nuclear to cytoplasmic ARP4 concentration varies dramatically among mammalian cell types in culture (Lee et al., 2003). A bipartite nuclear localization signal (NLS) sequence (195KKALE199, 210KQRK213) was first identified in yeast ARP4 within the conserved insertion shared among all ARP4s (Fig. 5.3A) (Stefanov, 2000). The NLS is essential to yeast ARP4 nuclear localization and function. However, alignment of this region with homologous domains from Arabidopsis, human, and Tetrahymena ARP4 sequences reveals poor conservation of these particular NLS sequences across kingdoms. This same general region in each of the plant, animal, and protist ARP4 proteins does contain alternative Lys- and Arg-rich motifs with potential NLS activities. Thus, different NLSs may have evolved independently among distant members of the ARP4 class. Similarly, all ARP5 sequences contain potential NLSs that are not well conserved across kingdoms in the very large conserved insertion located after actin amino acid 246. Lee et al. (2003) have shown that the shuttling of human ARP4 (Baf53) between the nucleus and the cytoplasm is energy dependent. Again the location of independently evolved NLS motifs in ARP4- and ARP5-specific insertions argues that ARP4 and ARP5 evolved independently from an ancestral actin sequence, and that ARP5 did not evolve from ARP4.
Figure 5.3.
Potential NLS, NES, and phosphorylation sites. (A) The proposed nuclear localization sequences (NLS) in various ARP4s are underlined and appear to be only moderately well conserved across the four eukaryotic kingdoms. The human (Hs), Arabidopsis (At), yeast (Sc), and protist (Tetrahymena thermophila, Tt) sequences are compared. (B) Proposed nuclear export sequences are compared among the yeast nuclear ARPs and actin and a consensus (CON) sequence is given. (C) Potential N-terminal phosphorylation sequences in ARP4 class members from the various kingdoms are compared to actin sequence. The conserved serine (S) and tyrosine (Y) residues that may be phosphorylated are underlined.
Conventional actins contain two leucine-rich nuclear export signal sequences (NESs), located between residues 85 and 137 (Fig. 5.3B). The first of these NESs is easily identifiable in the N-terminal region of all nuclear ARPs, while the second is less frequently conserved among the ARPs (Harata et al., 2000, 2001). NES sequences might mediate ATP-dependent ARP export. However, initial studies suggest that, at least for human ARP4, nuclear export is not inhibited by leptomycin B an inhibitor of the export receptor CRM1 that generally controls the export of leucine-rich NES proteins (Lee et al., 2003). At least two interesting questions remain: (1) To what purpose are the nuclear ARPs shuttled between the nucleus and the cytoplasm? (2) What regulates their concentrations in the two compartments? Perhaps their transport is essential for cytoplasmic– nuclear communication controlling the cell cycle and multicellular development.
3.2. Overall relationship to actin
The nuclear ARP sequences evolved from a common ancestral actin or actin-related gene prior to the divergence of the four eukaryotic kingdoms. They share only 17–35% amino acid sequence identity with conventional actin and are significantly more divergent from conventional actin than the cytoplasmic ARPs (Kandasamy et al., 2004; McKinney et al., 2002; Muller et al., 2005). For comparison, the cytoplasmic ARP1, ARP2, and ARP3 sequences average 38–52% identity with actin. Because the initial characterization of all the classes of ARPs was performed in yeast, most nuclear ARPs in other organisms are named relative to the yeast nuclear ARPs: Arp4, Arp5, Arp6, Arp7, Arp8, and Arp9 (Harata et al., 2000; Poch and Winsor, 1997). The larger numbers in this ARP nomenclature represent increasing divergence from conventional actin, with ARP4 being most closely related (approximately 35% amino acid identity to actin) and ARP8 and ARP9 the most divergent (an average of 18% identity to actin) (Goodson and Hawse, 2002; Kandasamy et al., 2004; McKinney et al., 2002; Muller et al., 2005). Protein sequence divergence within each nuclear ARP class is also relatively high. For example, ARP8 sequences from different kingdoms usually share less than 30% identity (Muller et al., 2005). These data demonstrate that nuclear ARP sequences evolve much more rapidly than conventional actin sequences.
3.3. Relationships among the nuclear ARPs
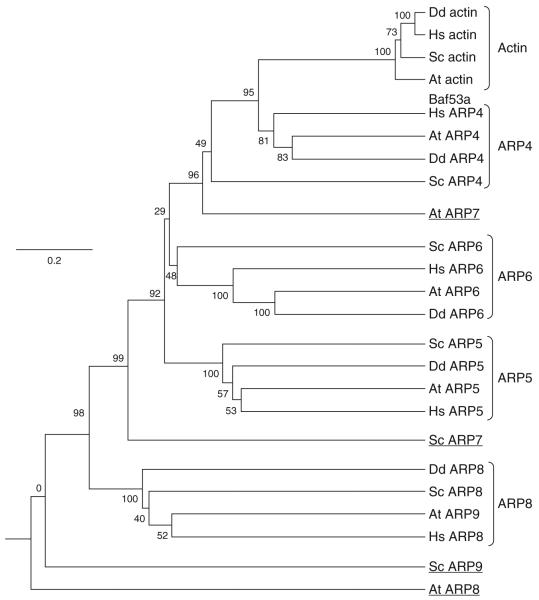
There are four classes of ARPs whose origins clearly predate the divergence of the four eukaryotic kingdoms. The yeast representatives of these four classes are ARP4, ARP5, ARP6, and ARP8. The phylogenetic relationships among the nuclear ARPs from humans, Saccharomyces cerevisiae (Sc), Arabidopsis thaliana (At), and Dictyostelium discoideum representing animal, plant, fungal, and protist kingdoms are illustrated in Fig. 5.4. Humans, Arabidopsis, and Dicytostelium contain clear homologs of yeast ARP4, ARP5, ARP6, and ARP8.
Figure 5.4.
Phylogenetic relationships among nuclear ARPs from the four eukaryotic kingdoms. The phylogenetic relationships among the nuclear ARPs encoded by the animal (human, Hs), yeast (Saccharomyces cerevisiae, Sc), plant (Arabidopsis thaliana, At), and protist (Dictyostelium discoideum, Dd) genomes are illustrated. The four classes of nuclear ARPs—ARP4, ARP5, ARP6, and ARP8—that are generally conserved among animals, plants, fungi, and some protists are indicated. A few examples of orphaned ARPs are indicated by underlining. Clustal was used to align the sequences. The phylogram presented used the unweighted pair-group method with arithmetic means (UPGMA) to create the tree’s topography based on sequence similarity (Tamura et al., 2007). The neighbor joining tree building method also yields a tree with very similar, but not identical, branching patterns. Human β-actin and Arabidopsis ACTIN2 were used as conventional actin gene representatives from within the divergent families of animal and plant actins.
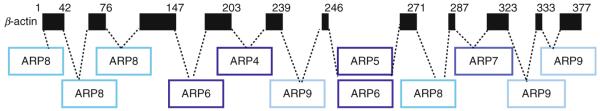
Relative to conventional actin, the nuclear ARPs contain numerous insertions, some very large, and a few small deletions. Figure 5.5 maps the location of 12 insertions that help define the four conserved nuclear ARP classes relative to the sequence of conventional mammalian β-actin (cytoplasmic actin) as first summarized by Muller et al. (2005). The insertion after the equivalent of mammalian β-actin amino acid 246 is found in all ARP5 and ARP6 sequences and the insertion results in a large increased in the size of all ARP5 proteins, however, ARP7 contains an insertion in this position in only a few species. All ARP4 sequences contain a small insertion after actin amino acid 203, but ARP4s in a few species contain insertions in three other locations (not shown). Numerous other kingdom and/or species-specific insertions are also reported and the orphaned ARPs may share the insertions shown and other novel insertions (Muller et al., 2005). For example, the novel insertions found in yeast ARP7 and ARP9 are shown in Fig. 5.5. The resulting variation in amino acid length of the four conserved classes of nuclear ARPs is summarized in Table 5.1. Overall, the low level of sequence identity among the nuclear ARPs in a class and between classes and the variation in size and location of INDELs create difficulties in aligning, identifying, and naming them. In many genome databases, the nuclear ARPs are annotated simply as “actin-like” or “actin-related” sequences.
Figure 5.5.
Coding sequence insertions in ARPs with respect to amino acid positions in conventional human β-actin. The locations of polypeptide insertions in the ARP4, ARP5, ARP6, and ARP8 classes that are conserved across all four eukaryotic kingdoms are shown. Numbered positions refer to amino acids in human β-actin. Insertions in yeast orphaned ARPs—ARP7 and ARP9—that are conserved in fungi are also shown for comparison.
Table 5.1. Across kingdom comparisons of the amino acid lengths of four ARP classes of known phylogenic origin.
| ARP | Species | length a.a. |
|---|---|---|
| ARP4 (isovariant) | ||
| “ | Sc | 498 |
| “ | Dd | 440 |
| “ (Baf53a) | Hs | 429 |
| “ (Baf53b) | Hs | 426 |
| “ | At | 441 |
| ARP5 | ||
| “ | Sc | 755 |
| “ | Dd | 684 |
| “ | Hs | 607–620 |
| “ | At | 726 |
| ARP6 | ||
| “ | Sc | 438 |
| “ | Dd | 490 |
| “ | Hs | 396 |
| “ | At | 420 |
| ARP8 | ||
| Sc | 881 | |
| Dd | 873 | |
| Hs | 624 | |
| (ARP9) | At | 596 |
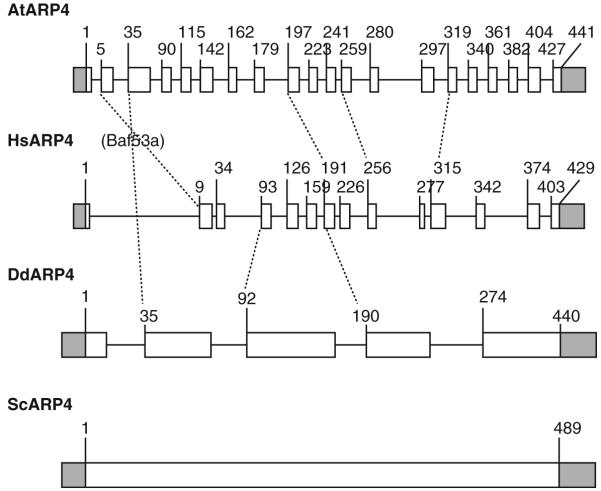
Considering the ancient common origin of four classes of nuclear ARPs, it seemed possible that intron–exon positions might reveal something about their evolutionary history. We compared the 19 intron positions in the Arabidopsis ARP4 gene, to the 13 in the human ARP4 (Baf53a) gene, and the 4 in the Dictyostelium ARP4 gene as shown in Fig. 5.6. Like most yeast genes, yeast ARP4 lacks any introns. Overall, it is clear that intron–exon positions are poorly conserved even between plant and human ARP4s that each contains large number of introns. We observed seven intron locations (dotted lines, Fig. 5.6) that were conserved between pairs of ARP4 genes in different kingdoms.
Figure 5.6.
Comparison of intron–exon structures for ARP4 genes across the four eukaryotic kingdoms. The intron/exon structure of ARP4 genes from Arabidopsis (At), human (Hs, Baf53a gene), protist (Dictyostelium discoideum, Dd), and yeast (Sc) genomes are compared. Coding exons are shown as white boxes, introns as lines, and transcribed but untranslated flanking regions (UTRs) in light gray boxes. The accession numbers of the particular transcript sequence compared are AT1G18450.1, BAG51043, XP_640964, and NP_012454, respectively, distinguishing these data from other possible transcript variants that exist for the plant and animal sequences. Conserved intron–exon junction positions are indicated by dashed lines. To generate these data, the intron positions identified from transcript and gene sequence alignments were compared to the amino acid sequence alignment of the four ARP4 protein sequences.
3.4. Inconsistent composition of the nuclear ARP classes in various protists
Considering that protist development was an early model of epigenetic control, we and others have looked for homologs of nuclear ARPs encoded in several representative and relatively complete protist genomes (Hedges et al., 2004; Muller et al., 2005). However, unlike the other eukaryotic kingdoms, the protists comprise an ancient, complex, and polyphyletic group of organisms (Adl et al., 2007). Hence, no single species or group can be considered as truly representative of the protist kingdom. We found clear homologs of ARP4 and ARP6 in representative species from diverse protist groups including Tetrahymena thermophila and Toxoplasma gondii representing the chromalveolates, Entamoeba histolytica representing the amoeba, Trypanosoma brucei representing the kinetoplastids (Gordon and Sibley, 2005), and Chlamydomonas reinhardtii representing the euglenids. The T. thermophila and D. discoideum genomes contained ARP5 and ARP8 homologs, while most protist species lacked them. Dictyostelium sequences were used in the nuclear ARP phylogeny (Fig. 5.4), because this protist had one of the most complete complements. Without inferring that the sequences were lost repeatedly, it is hard to reconcile the lack of ARP5 and ARP8 homologs in most protist genomes with the likely basal ancestry of protists to the other eukaryotic kingdoms. As might be expected, the extremely simple and possibly basal genomes of the parabasalids, Giardia lamblia and Trichomonas vaginalis, contained only an ARP4 homolog and no nuclear ARP homologs, respectively (Adam, 2000). We cannot yet say if this is a derived state representing loss of ARP sequences from an ancestral protist ancestor, or if this represents the early evolutionary state of chromatin remodeling systems, or some combination of both.
3.5. Orphaned ARPs
The orphaned nuclear ARPs are so named, because their phylogenetic relationships across kingdoms are less clear. Arabidopsis ARP7 and ARP8, yeast ARP7 and ARP9, and human ARP7 and ARP11 are examples of orphaned ARPs. In particular, Arabidopsis ARP7 shares 39% amino acid identity with actin and the basal structure of a conventional actin, but is only 23–27% identical to yeast ARP4, ARP5, ARP6, ARP7, or ARP8. As a result of numerous small insertions and deletions and small sequence similarities, ARP7 is phylogenetically positioned among the nuclear ARPs (Blessing et al., 2004; Kandasamy et al., 2004), and yet not clearly allied with any one yeast or animal nuclear ARP. Arabidopsis ARP8 and other plant ARP8 homologs are distinct from any ARP found outside the plant kingdom in that they combine a 40 amino acid long hydrophopic leader followed by a 50 amino acid long complete F-box domain and a complete 380 amino acid long C-terminal actin-related sequence (Kandasamy et al., 2008; McKinney et al., 2002). F-box domains target protein ubiquitination in other proteins, but how the F-box and the hydrophobic domain function together as part of any nuclear ARP is unknown. Moreover, this novel ARP is localized to the nucleolus. It seems possible that the N-terminal hydrophobic domain targets ARP8 to the nucleolus. Thus, plant ARP7 and ARP8 belong to plant-specific (Kandasamy et al., 2004; McKinney et al., 2002) or orphaned (Blessing et al., 2004) ARP classes. This does not mean that ARP7 and ARP8 do not share a common evolutionary origin with other specific nuclear ARPs (Fig. 5.4), but only that phylogenetic analysis has not unambiguously resolved these evolutionary relationships, if they exist. Similarly, yeast nuclear ARP7 and ARP9 also share no immediate homologs in other kingdoms and have an orphaned status (Blessing et al., 2004). Yeast and Arabidopsis ARP7 both appear more closely related in sequence to ARP4. Considering that yeast ARP7 is known to be part of SWI/SNF complexes, while ARP4 is a component in animal counterparts, it is reasonable to speculate that ARP7 evolved directly from ARP4. Many tree building programs place yeast and Arabidopsis ARP7 most closely allied to ARP4 and Arabidopsis ARP8 as most closely related to other ARP9s. Some protist genomes including Tetrahymena and Disctyostelium contained several other orphaned ARPs with questionable phylogenies and unknown functions.
3.6. Ancient origins of nuclear ARP sequences
The three-dimensional structures of the actins and ARPs reveal that they belong to a diverse and older family of proteins found in archaea, prokaryotes, and eukaryotes (Muller et al., 2005). This family includes glycerol kinases, hexokinases, and some chaperones like the 70 kDa heat shock protein, all of which share the actin-fold and the theoretical capacity for conformational change (Galkin et al., 2002). While the origin of conventional actin has been traced to a common ancestor of the bacterial structural protein MreB (van den Ent et al., 2001), a bacterial origin for an ARP with chromatin remodeling activity has not yet been identified. Some bacteria contain nucleosomal DNA and may remodel their chromatin, thus the bacterial origin of nuclear ARPs as part of chromatin remodeling machines remains a possibility (Bendich and Drlica, 2000; Champion and Higgins, 2007).
The topographies of the tree in Fig. 5.4 and other phylogenies comparing actins and nuclear ARP sequences (Kandasamy et al., 2004; Muller et al., 2005) generally position ARP4 closest to conventional actin and basal to the other nuclear ARPs. This topography not only reflects the rooting of the tree with conventional actin, and ARP4’s relatively high level of conservation and sequence identity with actin, but might also reflect the possibility that ARP4 was the first nuclear ARP to evolve. Supporting the view that ARP4 evolved first, ARP4 is found in more kinds of chromatin remodeling complexes (e.g., SWI/SNF, INO80, and SWR1) than any other nuclear ARP (Chen and Shen, 2007). Because ARP4 is involved in various kinds of complexes and perhaps is the most ancient, it may be the most indispensable nuclear ARP. Further, ARP4 homologs are the only nuclear ARPs present in a few simple protist genomes suggesting again that ARP4 may be the most ancestral (Gordon et al., 2008). It is reasonable to speculate that ARP5, ARP6, and ARP8 evolved from ARP4, but have become more subfunctionalized. ARP5 and ARP8 are part of the yeast and human INO80 NR complexes, while ARP6 is only found in SWR1-related histone variant exchange (HVE) complexes.
However, ARP4s contain an insertion after actin amino acid 203 that is conserved across all eukaryotes and not found in any other nuclear ARPs, except for a subset of ARP7 sequences in a few species. For ARP4 to be ancestral to the other nuclear ARPs, the founding ARP4 genes would need to have acquired this same insertion in multiple ancient phyla independently, after the evolution of ARP5, ARP6, and/or ARP8. The distinct insertions shared among each of these three ARP classes can be used to make similar arguments about the independent evolution of each classes from actin. Thus, it seems unlikely that the other nuclear ARPs are descended from ARP4 and, more likely, they evolved directly from conventional actin. The binding of conventional actins to Swi2- and Vid21-related ATPases and their role in chromatin remodeling and modification may have provided the target for natural selection of new nuclear ARPs (Szerlong et al., 2008). By this model, duplication of conventional actin genes followed by selective modification and subfunctionalization for nuclear chromatin activity would have independently produced ARP4, ARP5, ARP6, ARP8, and ARP9.
4. Function of the Nuclear ARPs in Chromatin Remodeling and Modifying Complexes
4.1. Nuclear ARPs bind Swi2-related DNA-dependent ATPases in chromatin remodeling machines
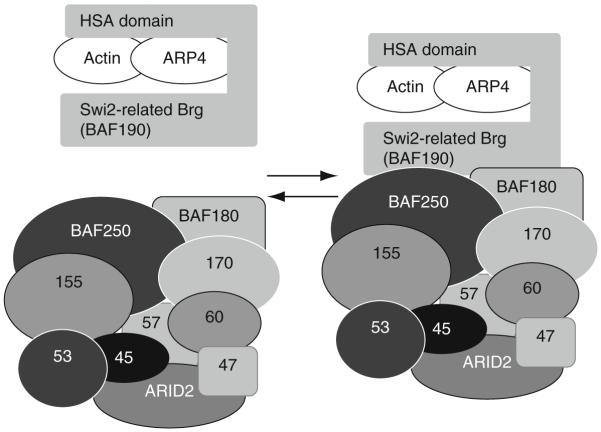
The nuclear ARPs are multifunctional proteins and some appear to have their own distinct properties, but only recently has a single common activity emerged that may be conserved among all members of the family. The nuclear ARPs are constituents of ATP-dependent chromatin remodeling complexes (e.g., SWI/SNF, SWR1, RSC, INO80, p400) and/or chromatin modifying complexes (NuA4 HAT) (Chen and Shen, 2007; Krogan et al., 2003; Mizuguchi et al., 2004; Olave et al., 2002b). Indeed, all chromatin remodeling complexes that contain a Swi2-related DNA-dependent ATPase subunit also include one or more nuclear ARPs, and if they contain only a single nuclear ARP subunit, this is in association with an actin subunit (Szerlong et al., 2008). Figure 5.7 models the assembly of one possible isoform of the mammalian Swi/Snf BRG chromatin remodeling complex containing approximately 13 subunits. It will be used as an example in the following discussion. Nucleosomal movement is powered by ATP hydrolysis mediated by the Swi2-related ATPase (Dang and Bartholomew, 2007). Swi2-related proteins share a rapidly evolving HSA (helicase SANT-associated) domain and an adjacent post-HSA domain of approximately 100 amino acid in length and located within a few hundred amino acid of the N-terminus of these very large proteins (Szerlong et al., 2008).
Figure 5.7.
Nuclear ARP4 (Baf53) and actin bind the HSA domain of the Swi2-related Brg DNA-dependent ATPase in the mammalian Swi/Snf BRG chromatin remodeling complex. This model illustrates that β-actin and ARP4 bind Brg, and that the β-actin/ARP4/Brg subcomplex binds to a second subcomplex containing several other Brg proteins to form an active chromatin-remodeling machine (Lessard et al., 2007; Szerlong et al., 2008). A related model may be proposed for a large number of chromatin-active complexes. Two nuclear ARPs or a nuclear ARP and actin bind as heterodimers to the helicase-SANT (HSA) domain of the large Swi2-related DNA-dependent ATPase subunit in chromatin remodeling complexes or the Vid21-related helicase subunit in chromatin modifying complexes. Again, the ARP-containing subcomplex then binds a second subcomplex with a larger number of subunits.
A number of early studies lent strong support to the particular binding of nuclear ARPs and Swi2-related ATPases. For example, the BAF complex of mutant human cells lacking the Swi2-related Brg1 subunit also were missing the ARP4 (Baf53) and actin subunits, while a BAF subcomplex lacking all three of these components still formed (Zhao et al., 1998). Furthermore, the binding of ARP8 in the INO80 complex and the binding of actin and ARP4 in the SWR1 complex both require N-terminal regions of the Swi2-related Ino80 and Swr1 subunits, respectively (Shen et al., 2003b; Wu et al., 2005). INO80 complexes purified from yeast arp5Δ or arp8Δ strains still retain most other subunits, but are deficient in the Ino80 ATPase, ARP4, and actin, indicating that ARP5 and ARP8 may be needed to recruit these other subunits into the complex (Shen et al., 2003b). In Arabidopsis, the N-terminal half of Swi2-related SPLAYED retains activities necessary for normal meristem development and flowering (Su et al., 2006). Finally, yeast ARP7 and ARP9 were shown to both interact with Swi2-related Sth1 subunit to form a catalytic SWI/SNF subcomplex (Yang et al., 2007).
Isolation of yeast suppressor mutations of ARP deletions Δarp7 and Δarp9 provided genetic proof of the interaction between nuclear ARP and HSA domain of Swi2-related proteins. All 10 suppressors of these null mutants mapped as amino acid substitutions within the 83 amino acid HSA domain or an immediately adjacent 20 amino acid post-HSA domain of the Swi2-related Sth1 subunit of a SWI/SNF complex (Szerlong et al., 2008). Biochemical protein interaction assays suggest that ARP7 and ARP9 bind the HSA/post-HSA domain as a heterodimer similar to the interaction of actin and ARP4 in the BRG complex (Fig. 5.7). Yeast ARP7 and ARP9 also bind the HSA domain of yeast Swi2-related subunit Snf2 of the RSC complex. Similarly, human and yeast actin and ARP4 subunits of SWI/SNF complexes bind the HSA domains of Swi2-related human Brg1 and yeast Swr1, respectively. Furthermore, yeast actin, ARP4 and ARP8 subunits of the INO80 complex bind the HSA domain of the Ino80 subunit. Of particular note is the significant specificity within the various divergent HSA domains that directs binding of the correct nuclear ARP and actin partners to the corresponding Swi2-related sequence, and not to irrelevant HSA domain proteins. Szerlong et al. (2008) propose that ARPs bind as dimers to an HSA domain protein in all complexes in which they participate.
4.2. Nuclear ARPs bind Vid21-related helicase subunits in chromatin modifying machines
These data beg the question: why are nuclear ARP4 and actin found in large histone acetylation complexes that lack a Swi2-homolog like the various fungal NuA4 HAT and mammalian TIP60 complexes? Szerlong et al. (2008) show that the Eaf1 subunit of NuA4 and other Vid21-related subunits of fungal HAT complexes each contain a HSA domain and the HSA domain of Eaf1 binds ARP4 and actin as this domain binds Brg in the BRG complex (Fig. 5.7). Hence, an alternate HSA domain protein interacts with two actin-related sequences in these fungal HAT complexes. Sequence conservation among the Vid21 sequences and their HSA domains is weak even among distant fungi and thus structural homologs in other kingdoms may be difficult to identify. Eaf1 has some weak and previously undetected homology of its HSA and SANT domains with a few Swi2-related proteins such as human p400/Domino (Auger et al., 2008). We observed that the human genome encodes a few other proteins similar in size to the 982 amino acid long yeast Eaf1 protein with low-complexity sequence matches to Eaf1’s HSA domain (e.g., BAC04759) that are not yet identified as HSA-containing homologs. These and other HSA domain proteins have the potential to interact with nuclear ARPs and actins in novel chromatin modifying complexes. Finally, mammalian TIP60 complexes contain Ruvbl1 (RuvB-like 1, Tap54, Tip49) with both DNA helicase and ATPase domains (Ikura et al., 2000). Although human Ruvbl1 (456 amino acid) shows no statistically significant amino acid sequence homology with the larger Swi2-related proteins or Vid21-related sequences, it is reasonable to speculate that actin and ARP4 might interact via a poorly conserved and ancient HSA domain.
4.3. Other activities and interactions with chromatin
Most models for ARP and actin function in chromatin remodeling and modifying complexes center on the actin-fold and hinge region that impart the ability to shift between distinct conformational states upon nucleotide binding, ATP hydrolysis, and ATP/ADP exchange. Nuclear ARPs belong to the actin superfamily of proteins that include cytoskeletal actins, heat shock proteins, ATPases, and sugar kinases that all appear to retain the structural potential to bind a nucleotide and to change conformation (Boyer and Peterson, 2000; Sunada et al., 2005). Conventional actin, for example, is converted from an inactive ADP bound conformational state to an ATP bound active form via its interaction with profilin. ATP-actin is added to actin filaments, but subsequent ATP hydrolysis within monomers lowers filament stability, alters filament turnover rates, and changes the affinity for different actin-binding proteins. The ARP and actin subunits of chromatin remodeling machines may undergo similar nucleotide-dependent changes in the nucleus to act as molecular switches controlling the assembly, stability, and/or activity of these complexes. To date, only ARP4 has been shown to bind nucleotides, but it is possible that more evidence for nucleotide binding will follow for other nuclear ARP family members. Biochemical and genetic studies in yeast demonstrate that mutant ARP4 proteins defective in nucleotide binding are more concentrated in high-molecular-weight protein fractions containing remodeling complexes than wild-type ARP4 protein (Sunada et al., 2005). By contrast, mutant forms of ARP4 enhanced for ATP binding remain mostly as unbound monomers. Perhaps the nucleotide bound state of ARP4 and other ARPs may affect their assembly and disassembly as ARP-ARP-HSA or ARP-actin-HSA trimer subcomplexes.
ARPs are required to associate some complexes with the nuclear matrix (Zhao et al., 1998). Yeast ARP4 and ARP8 bind histones directly, so ARPs may help recruit complexes directly to nucleosomes (Harata et al., 1999b; Shen et al., 2003b). In addition, ARPs and actins may connect one chromatin complex to another to create higher order chromatin complexes, acting similarly to conventional actin, only in the “polymerization” of higher order chromatin structures (Olave et al., 2002b). Silencing of mammalian ARP4 (Baf53) or conditional knockdown mutations of yeast ARP4 causes a loss of chromatin compaction, as revealed by increases in the nuclease sensitivity of nucleosomes (Georgieva et al., 2008; Lee et al., 2007a). Furthermore, higher order interactions of distinct chromatin remodeling complexes INO80, SRCAP, and TIP60 are supported by initial proteomic data. Analyses of the proteins bound to large numbers of individually epitope tagged subunits of these complexes revealed interactions between and among complexes (Sardiu et al., 2008), further supporting the idea that nuclear ARP complexes contribute to the formation and integrity of higher ordered chromatin structures.
4.4. Activities of particular ARPs
4.4.1. ARP4
ARP4 is an essential gene in yeast and appears to be essential in Arabidopsis, because severe knockdown plants with less than 15–20% of wild-type levels of ARP4 protein are extremely dwarfed and highly sterile (Kandasamy et al., 2005a). ARP4 is found in more diverse chromatin remodeling complexes than any other ARP in plants, animals, and fungi. Yeast ARP4 is found in the NUA4 HAT (histone acetyltransferase) nucleosome modifying complex, the INO80 NR complex and the SWR1 HVE and NR complex (Galarneau et al., 2000; Harata et al., 1994; Huang et al., 1996; Krogan et al., 2003; Minoda et al., 2005; Mizuguchi et al., 2004; Shen et al., 2000). The 1.3 MDa NUA4 complex contains 11 subunits, including ARP4 and conventional actin, and it is known to primarily acetylate histone H4. In temperature-sensitive yeast arp4 mutants, the full NUA4 complex is absent at the restrictive temperature (Galarneau et al., 2000). These same mutants display increases or decreases in the transcription of a number of target genes at the restrictive temperature, coinciding with changes in the normal chromatin structure around those genes (Harata et al., 2002; Jiang and Stillman, 1996). ARP4 orthologs are also found in several different mammalian ATP-dependent NR and HVE complexes (Fuchs et al., 2001; Lee et al., 2005; Nie et al., 2000; Oma et al., 2003; Rando et al., 2002; Ruhl et al., 2006), in mammalian INO80 NR DNA repair complexes (Cai et al., 2006), in mammalian Tip60 NR HAT complexes (Feng et al., 2003; Ikura et al., 2000), and in Drosophila NR complexes (Mohrmann and Verrijzer, 2005; Papoulas et al., 1998).
ARP4 binds all four core histones in vitro (Harata et al., 1999b) although yeast ARP4 shows some preference for H2A. The particular histone-binding preferences reported for ARP4 vary among the different species in which binding was measured. How histone modification may affect ARP4’s preferential binding is only now being addressed. It has been well established that a phosphoserine residue in the C-terminal domain of H2A variant H2AX tags nucleosomes at sites of severely damaged DNA (e.g., double-strand breaks (DSBs) requiring recombination repair) (Kuo and Yang, 2008; Morrison et al., 2004; van Attikum et al., 2004). ARP4 binds this residue within nucleosomes and directs the binding of yeast ARP4-containing complexes, NUA1, INO80, and SWR1, to remodel chromatin at these damaged sites (Bird et al., 2002; Downs et al., 2004). Thus, another reason that ARP4 is such a widely conserved subunit of diverse complexes may be its ability to bind different core histones and histone modifications, and hence, target complexes to chromatin.
Molecular genetic evidence suggests that the N-terminal Ser2 and Tyr6 residues of human ARP4 (Baf53) may be phosphorylated for at least some ARP4 activities (Lee et al., 2005). A small N-terminal extension encodes Ser and/or Tyr residues in the human, S. cerevisiae and A. thaliana sequences, but not in T. thermophila ARP4 homologs as shown in Fig. 5.3C. Mammalian ARP4 sequences also contain Ser20 and Tyr27 residues in the first actin homology domain that are conserved among ARP4 homologs from all four eukaryotic kingdoms. The possible conservation of the phosphorylation of N-terminal Ser and Tyr residues and the significance of these modified residues on ARP4 activities outside of human cell culture have yet to be determined. Phosphoinositol (PIP2) addition to cultured mouse lymphocytes rapidly targets ARP4-containing BAF complexes to chromatin, but it is not known if PIP2 signaling affects phosphorylation of ARP4 or if this phosphorylation independently affects localization of the complex (Zhao et al., 1998). Surprisingly, expression of a truncated ARP4 (Baf53b) mutant protein lacking this short N-terminal domain has dominant negative effects and is lethal to human cells (Choi et al., 2001).
4.4.2. ARP5 and ARP8
ARP5 and ARP8 proteins are not essential to the viability of budding yeast (Shen et al., 2003b; van Attikum et al., 2004). Yeast, mammalian, plant, and protist ARP5 and ARP8 (Arabidopsis ARP9) are highly modified by insertions of protein sequences relative to actin (Fig. 5.5) making them the largest nuclear ARPs, typically 600–800 amino acids (Muller et al., 2005). In addition to ARP5 and ARP8, the yeast and mammalian INO80 NR complexes also contain ARP4, monomeric actin, a Swi2-related DNA-dependent ATPase Ino80, and several other subunits (Cai et al., 2006; Jin et al., 2005). The Arabidopsis genome encodes homologous of all the subunits of the yeast INO80 complex (Fritsch et al., 2004; Meagher et al., 2005).
Because ARP4, ARP5, ARP8, and Ino80 genes and most other subunits are conserved across the fungal, animal, and plant kingdoms, it appears that INO80 complexes will be relatively universal. Roles for INO80 complexes are reported in dynamic control of transcription, DNA repair, and DNA replication (Conaway and Conaway, 2009). Yeast loss-of-function mutations for the Ino80 Swi2-related subunit are defective in transcribing the genes needed for inositol biosynthesis (e.g., the INO1 gene) and require inositol, signifying that the INO80 complex is involved in transcriptional control as expected for a NR complex. Ino80 defective cells are hypersensitive to several DNA damaging agents demonstrating INO80 is also involved in DNA damage repair (Ebbert et al., 1999; Shen et al., 2000). Deletion mutants lacking the ARP5 and ARP8 genes are defective in the DNA-dependent ATPase activity of the Ino80 subunit and require inositol. This undoubtedly occurs because these ARPs are critical to binding the HSA domain on Ino80 (Szerlong et al., 2008) and assembly of a subcomplex needed for the full activity of INO80 (Fig. 5.7). ARP8 is also required for the INO80 complex to bind the promoter of the INO1 gene affording some mechanistic explanation for the phenotype of Arp8 and Ino80 defective mutants (Ford et al., 2008). The ATP-binding Rvb helicase proteins bind ARP5 and are essential for its assembly into INO80 complexes and for formation of a functional complex (Jonsson et al., 2004). ARP5- and ARP8-defective mutants in yeast are dramatically altered in DNA repair and the cell cycle (van Attikum et al., 2004, 2007).
In mammalian and yeast systems, extensive effort has gone into studying the role of INO80 in repairing radiation-induced double-strand breaks in DNA (Cairns, 2004; Morrison et al., 2004). Less is known about the particular roles of ARP5 and ARP8 in these processes. Cells with mutations in Arp8 are defective in end-processing of gamma radiation-induced DSBs (van Attikum et al., 2007) and in recombination repair of sister chromatids (Kawashima et al., 2007). The finding that ARP8 binds histones H3 and H4 in vitro suggests that along with ARP4, ARP8 may also serve as a point of contact between the complex and chromatin (Shen et al., 2003b). Human ARP8 was one of the first nuclear ARPs to be shown to accumulate on mitotic chromatin. The transient silencing of ARP8 or expression of truncated ARP8 variants causes misalignment of metaphase chromosomes, while the silencing of ARP5 or Ino80 homologs did not (Aoyama et al., 2008). These data suggest novel functions for ARP8 outside those known for INO80 complexes.
Despite the low level of homology, human ARP5 partially complements the loss of UV resistance of yeast arp5Δ strains (Kitayama et al., 2009). HeLa cells partially silenced for ARP5 expression are significantly more sensitive to DSBs caused by bleomycin than control cells. Kitayama et al. (2009) also show that ARP5 appears to bind directly to both H2AX and phosphorylated γH2AX after treatment with bleomycin. Finally, overexpression of ARP5 causes an increase in γH2AX levels. Arabidopsis plants defective in ARP5 are hypersensitive to DNA damaging agents HU, MMS, and bleomycin, and show dramatic defects in multicellular development (Kandasamy et al., 2009). Together these data suggest that ARP5 may interact directly with chromatin in regulating the activities of the INO80 complex in chromatin remodeling and DSB repair.
4.4.3. ARP6
ARP6 is not essential for the viability of budding or fission yeast or Arabidopsis, but it is essential for their normal growth and development (Deal et al., 2005; Kawashima et al., 2007; Ueno et al., 2004). ARP6 is a universally conserved subunit of HVE complexes found in fungi and plants (SWR1), Drosophila (dISWI), and mammals (SRCAP) (Cai et al., 2005; Ruhl et al., 2006). The S. cerevisiae SWR1 and human SRCAP complexes share a large part of their subunit composition with other ATP-dependent NR complexes including the ARP4 and actin subunit (Kobor et al., 2004). The biochemically characterized ARP6-HVE complexes from animals and yeast also contain a large and conserved Swi2-related Swr1 subunit that is distinct among the families of fungal and animal DNA-dependent ATPases (Mizuguchi et al., 2004). Genetic and biochemical evidence support the existence of the SWR1 complex in plants. The Arabidopsis genome encodes homologs of all the subunits of yeast SWR1 (Meagher et al., 2005). Null mutants in ARP6 and the Swr1-related PIE1 (Snf2/Swr1), and another conserved SWR1 subunit, Swc6, share most of the same suite of developmental phenotypes (Choi et al., 2005, 2007; Deal et al., 2005, 2007; Lazaro et al., 2008; March-Diaz et al., 2007; Noh and Amasino, 2003). We have shown that ARP6- and PIE1-defective mutants lack measurable levels of histone variant H2AZ deposition at a number of target loci, lending strong support for the presence of a plant SWR1-like complex (Deal et al., 2007).
Despite sharing many subunits with other NR complexes, SWR1 homologs have the unique ability to catalyze the replacement of histone H2A with the variant H2AZ at specific chromosomal locations (Krogan et al., 2003). In yeast and Arabidopsis, the H2AZ variant plays important roles in promoting transcription and probably in antagonizing the spread of heterochromatin into euchromatic regions (Adam et al., 2001; Deal et al., 2007; Larochelle and Gaudreau, 2003; Meneghini et al., 2003; Santisteban et al., 2000). In both yeast and Arabidopsis, ARP6 is essential for the activity of the complex depositing H2AZ into chromatin (Deal et al., 2007; Wu et al., 2005). Within the yeast SWR1 complex, ARP6 recruits other critical subunits, one of which, Swc2, interacts directly with the C-terminal end of H2AZ, and hence, ARP6 is indirectly required for binding of the complex to nucleosomes. There are numerous other histone variants (e.g., H2AX), but it is unknown if SWR1 controls their deposition into nucleosomes.
Work in animals shows an association of ARP6 activity and HETEROCHROMATIN PROTEIN1 in hetrochromatic regions (Kato et al., 2001; Ohfuchi et al., 2006). The histone variant H2AZ exchange activity of ARP6-containing SWR1 complexes is interpreted as antagonizing the spread of inactive heterochromatin into active euchromatic regions of the genome and as maintaining gene silencing (Kato et al., 2001; Ohfuchi et al., 2006; Ueno et al., 2004). Contrary to this view, detailed mapping at individual loci in yeast and Arabidopsis shows H2AZ distributed at the 5′ ends of most genes, both active and inactive (Deal et al., 2007; Raisner and Madhani, 2006; Raisner et al., 2005), and is not particularly concentrated in heterochromatic regions (Dryhurst et al., 2004; Fan et al., 2004; Li et al., 2005; Meneghini et al., 2003). Thus, ARP6 appears to have multiple and contrasting functions that must be viewed as part of an integrated system of chromatin remodeling activities, with activities in heterochromatic regions that are not yet fully defined. Supporting the view that ARP6 can have opposing functions on different genes, we found that loss of ARP6 activity in Arabidopsis caused loss of H2AZ deposition at the 5′ and 3′ ends of three active MADS box loci that are generally active in nongametic tissues. Loss of ARP6 results in significant repression of their transcript levels (Deal et al., 2007). We have found other classes of transcription factors where the loss of 5′ H2AZ deposition in ARP6-deficient plants caused activation (unpublished data), fitting the original view of ARP6 and H2AZ as silencing gene activities. Perhaps ARP6-dependent H2AZ deposition can potentiate transcriptional activation serving an epigenetic memory function by marking the ends of active genes and preparing silenced genes for reactivation (Meagher et al., 2007).
4.4.4. Orphan ARPs
As discussed previously, a number of nuclear ARPs are classified as orphaned sequences, because they cannot be reliably grouped in phylogenies across the various eukaryotic kingdoms (Fig. 5.4, Section 3.5). Yeast orphaned ARPs, ARP7 and ARP9, are found among the 11–15 subunits of the SWI/SNF and RSC ATP-dependent NR complexes (Cairns et al., 1998; Peterson et al., 1998; Szerlong et al., 2003). It is unclear how human or Arabidopsis ARP7 and ARP8 sequences relate to these yeast sequences (Kandasamy et al., 2004, 2008). Unlike the complexes described here, neither yeast SWI/SNF nor RSC contain monomeric actin. The SWI/SNF complex was identified independently in genetic screens for genes involved in mating-type switching and sucrose fermentation, and the RSC complex was later isolated based on homology to SWI/SNF complex components. Strains lacking ARP7 or ARP9 show phenotypes typical of swi/snf− mutants, indicating that these proteins play an essential role in the function of SWI/SNF.
In addition to the swi/snf− phenotype, mutations in yeast ARP7 or ARP9 also lead to other transcriptional defects, indicating that RSC plays a role in general transcriptional regulation (Cairns et al., 1998). Surprisingly, RSC complexes isolated from arp7Δ/arp9Δ cells are otherwise fully intact and retain the ability to remodel nucleosomes in vitro. However, a screen for suppressors of arp7 and arp9 mutations identified the architectural transcription factor Nhp6, which interacts physically with RSC and enhances the activity of the complex in vitro (Szerlong et al., 2003). These data suggest that ARP7 and ARP9 serve to connect the RSC complex to interacting proteins or other complexes, allowing functionality in vivo. The characterized animal SWI/SNF (NR) complexes contain homologs of yeast ARP4 instead of homologs of the orphaned yeast ARP7 and ARP9 found in the yeast complex. This suggests at least a functional relationship exists between the orphaned ARPs and the reasonably well-conserved ARP4, if not an undetected closer phylogenetic relationship.
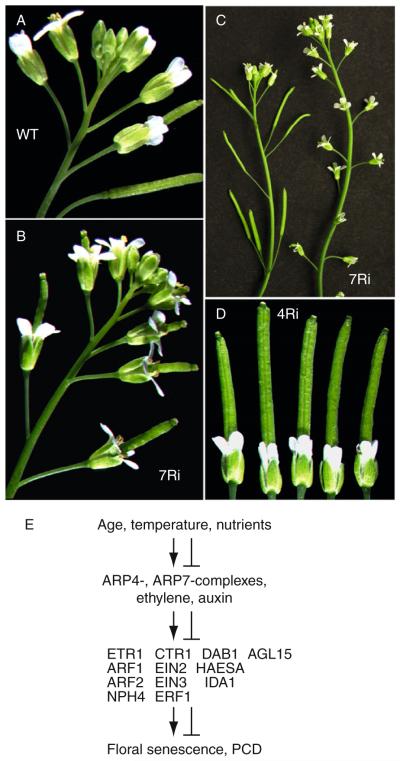
Arabidopsis ARP7 is an essential gene. Nearly normal heterozygous plants carrying a null allele produce homozygous arp7-1/arp7-1 embryos that abort at the torpedo stage, and mutant seeds containing defective embryos arrest their development. It is possible that a small amount of residual ARP7 from the parental egg cell cytoplasm may support early embryonic development. Plants partially silenced for ARP7 expression display a wide variety of dramatic phenotypes (Table 5.2) some of which are discussed in the following sections. Even though the phylogenic origin of plant ARP7 is not clear, this ARP has important role(s) in multicellular development.
Table 5.2. Defects in Arabidopsis actin-related protein expression lead to numerous alterations in multicellular development.
| Nuclear ARP defect |
||||
|---|---|---|---|---|
| Phenotype | ARP4 | ARP5 | ARP6 | ARP7 |
| Small leaves | Yes | Yes | Yes | Yes |
| Small leaf cells | Yes | Yes | No | Yes |
| Upward curled leaves | No | Yes | No | Yes |
| Excess stomata relative to other epidermal cells |
No | Yes | No | No |
| Delayed stomatal development | No | Yes | No | No |
| Flower early with long days | Yes | No | Yes | No |
| Flower early with short days | No | No | Yes | No |
| Small flowers and floral organs | Yes | Yes | Yes | Yes |
| Delayed floral senescence | Yes | No | No | Yes |
| Excessive root hairs | ND | Yes | Yes | No |
| Shortened root apical zone | ND | No | Yes | Yes |
| HU hypersensitivity | Yes | Yes | ND | ND |
| MMS hypersensitivity | Yes | Yes | ND | ND |
| Bleomycin hypersensitivity | Yes | Yes | ND | ND |
ND, no data.
5. Isoforms of ARP Complexes
5.1. Defining isoforms of chromatin complexes
A theme emerging in plants and animals is that diverse isoforms of nuclear ARP-containing chromatin complexes exert epigenetic control over multicellular development (Brown et al., 2007; Dirscherl and Krebs, 2004; Meagher et al., 2005; Nie et al., 2000; Olave et al., 2002b). Furthermore, there are increasing combinatorial possibilities in the numbers of isoforms of nuclear ARP complexes, paralleling the macroevolution of organismal complexity from single-celled organisms like yeast and Tetrahymena to more complex eukaryotes like humans and Arabidopsis (Dirscherl and Krebs, 2004; Iyer et al., 2008). To illustrate what is meant by isoforms of chromatin complexes, consider the ARP-containing SWI/SNF (NR), INO80 (NR), and SWR1 (HVE) complexes. Based on their composition in yeast or mouse, these three complexes contain 12 or more different protein subunits. Small gene families frequently encode these individual subunits. Thus, isoforms of a complex may be defined as those complexes containing altered isovariant subunit compositions, but composed of the same general subunit makeup. In our usage, protein “isovariants” or “variants” are closely related polypeptides with altered sequences encoded by different gene family members. Besides being derived from different members of gene families, protein isovariants also can be generated from single genes by alternate RNA splicing and poly(A) site selection, by alternate initiation and termination of translation, and by posttranslational protein modifications such as phosphorylation. In plants and animals, divergent gene families encode multiple isovariants of several subunits in each of these three types of complexes and perhaps even all major classes of complexes. Substituting one single subunit with a second different isovariant would generate a new isoform of a complex. A new isoform has the potential to recognize a new target gene based on local histone modification or nucleosome composition or bound protein factors. Alternatively, a new isoform may carry out a slightly different chromatin modifying reaction such as different phasing of nucleosomes, exchange of a different histone isovariant, or different histone modifications. For example, substituting two different isovariants of the large Swi2-related DNA-dependent ATPase subunit into a SWI/SNF complex will undoubtedly generate two isoforms of a complex with different activities on nucleosomes.
5.2. The numbers of isoforms increase with organismal complexity
Chromatin remodeling proteins in single-celled organisms like yeast (~6000 genes) are encoded by singlet genes and a few small gene families. Most of these genes were expanded into much larger gene families in complex multicellular organisms like humans encoding ~37,000 genes and A. thaliana encoding ~36,000 genes. Again, using the Swi2-related subunit as an example, there are 10 Swi2-related sequences in yeast, but more than 30 in the mouse and human genomes and 44 in Arabidopsis. Thus, by increasing the number of Swi2-related isovariants the number of isoforms of remodeling complexes was increased during the evolution of multicellular plants and animals. By extension, holding one of these Snf2-related subunits constant and substituting two isovariants of another subunit generates two other novel isoforms. Supporting the reality of this view are numerous papers on the importance of Swi2-related isovariants. For example, in embryonal cells, Swi2-related β-BRG1, but not α-BRG1, plays an essential role in basal processes involved in cell proliferation (Sumi-Ichinose et al., 1997). Finally, it should be mentioned that the tissue- or organ-specific expression of even one isovariant within a complex has the potential to refine epigenetic control by a complex.
In fact, several human and mouse chromatin remodeling complexes were isolated as mixtures of isoforms. For example, the purified mammalian BAF or SWI/SNF complex was shown to have a basic composition of about 9–12 proteins, but it existed in several isoforms. Isoforms purified from various organs varied in their Baf60 subunit and Swi2-related subunit compositions (Debril et al., 2004; Wang et al., 1996a,b). BAF60a is ubiquitously expressed, whereas the BAF60b and BAF60c isovariants are expressed in a subset of tissues and organs. The Baf60 isovariants appear to contribute to the target gene specificity of the complex. Furthermore, phosphorylation of BAF60 subunits creates additional isovariants that appear to target a subset of BAF complex isoforms to particular myogenic target genes (Simone et al., 2004).
Surprisingly, the nuclear ARP genes themselves are seldom found duplicated into gene families, with a notable exception. Mammalian isovariants of ARP4, Baf53a, and Baf53b, participate in SWI/SNF and TRAAP chromatin remodeling complexes and are differentially expressed. Baf53a expression is restricted primarily to brain and other neuronal tissues, while the Baf53b isovariant, which is the product of a separate gene, is more broadly expressed (Kuroda et al., 2002; Lessard et al., 2007; Olave et al., 2002a; Wu et al., 2007). Baf53a interacts specifically with a transcriptional corepressor CtPB, but Baf53b does not. The activity of SWI/SNF complex and its target genes are repressed in the presence of Baf53a, but not Baf53b (Oma et al., 2003). Baf53a isovariant cannot be replaced by Baf53b, because particular sequences in subdomain 2 of Baf53a add necessary and specific protein–protein interactions to chromatin remodeling in the nervous system. Baf53b is assembled into multiple PBAF complexes in postmitotic neurons (Choi et al., 2001; Lee et al., 2003; Ohfuchi et al., 2002; Olave et al., 2002a). In summary, remodeling complexes with the Baf53a isovariant are functionally distinct from those containing the Baf53b isovariant.
Analyses of mutants in the four Arabidopsis Swi3 isovariants suggest different developmental roles for four likely SWI/SNF complex isoforms, each with a different Swi3 isovariant (Sarnowski et al., 2005; Zhou et al., 2003). Each mutant with defects in swi3a, swi3b, swi3c,or swi3d displays a distinct set of phenotypes, including embryo lethality and vegetative and reproductive defects. The sum of the phenotypes described for these four mutants include most of the phenotypes we described for plants deficient in ARP7, which is likely to be a member of this SWI/SNF complex (Kandasamy et al., 2005b). For example, the swi3a and swi3b mutants display an embryo lethal phenotype similar to the homozygous null arp7-1 Arabidopsis mutant. In addition, the swi3d mutations cause severe dwarfism and complete male and female infertility like strong ARP7 knockdown epialleles. Plants carrying swi3c mutations share retarded root and plant growth, curly leaf, and reduced fertility phenotypes with the moderate ARP7 epialleles.
We have shown previously that homologs of most subunit proteins within the yeast SWI/SNF, SWR1 and INO80 complexes are encoded by gene families in Arabidopsis (Meagher et al., 2005). There are 12 likely subunit isovariants in five gene families that may produce 64 SWI/SNF isoforms. There are 19 subunit isovariants in six gene families with the potential to produce 600 SWR1 isoforms. There are 13 isovariants in four gene families with the potential to produce 56 INO80 isoforms. These estimates exclude variations in the possible actin and Swi2-related subunit isovariant composition, each variation having the potential to add significantly to the complexity of isoforms. Each individual ARP complex isoform with a different isovariant composition may only control a small subset of all epigenetically controlled target genes with a specific impact on development. Using the same yeast query sequences (Meagher et al., 2005) to explore the mouse and human genomes reveals that a similar high level of combinatorial diversity exists among the subunits of these remodeling complexes in mammals (not shown). Hence, it is reasonable to propose that in plants and animals there are hundreds of functional isoforms of each type of basic remodeling or modifying complex and that each isoform of a complex may differ in some way in its activity.
5.3. Contingency, macroevolution, and the origin of new isoforms
One evolutionary view connecting ARP complexes to the macroevolution of multicellular development is that as plants and animals diversified from single-celled protist ancestors, there was a combinatorial expansion in the number of nuclear ARP complex isoforms following an expansion and diversification of the gene families encoding their various subunit isovariants. Besides normal gene endoduplication, a number of ancient genome wide duplications are identified in the ancient ancestry of higher plants and animals. Thus, gene duplication is not likely to have limited increases in the numbers of protein isovariants (Meagher et al., 1999b). An increase in chromatin complex isoforms allowed the natural selection of more specialized control over chromatin dynamics and target-gene transcription, which generated more specialized epigenetic control over multicellular development. Greater target gene specificity and more finely tuned epigenetic control should be a selective advantage to multicellular organisms. Contingency theory can be used to explain the seemingly improbable pathways of gene duplication, gene divergence, neutral drift, natural selection, and stepwise increases in the complexity of chromatin complex isoforms that lead to increasing complexity of tissue and organ development in animals and plants (Bak and Paczuski, 1995). Previous discussion linking the long pathways of genetic events to the evolution of protein–protein interactions and macroevolution of developmental pathways rely heavily upon arguments of historical contingency (Meagher, 1995; Meagher et al., 1999a, 2008; Muller and Newman, 2005).
6. Role of Nuclear ARPs in the Epigenetic Control of Morphological Development
6.1. ARP complexes control development
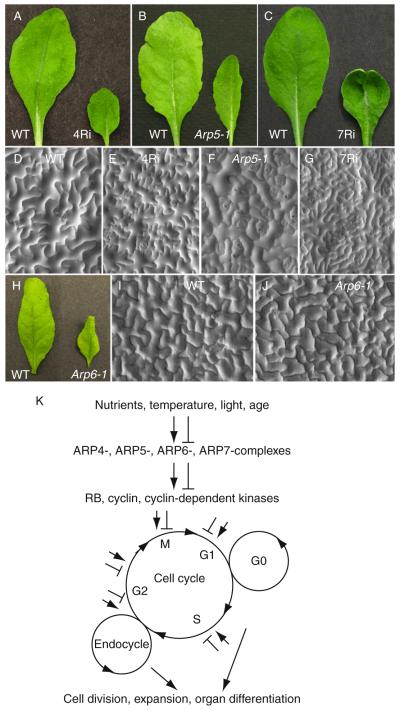
The following section summarizes several functional studies demonstrating an essential role for nuclear ARPs in different pathways of yeast, animal, and plant development. In plants, for example, chromatin remodeling and epigenetic control are directly linked to numerous pathways of multicellular development (Bezhani et al., 2007; Chen, 2007; Cuzin et al., 2008; Henderson and Jacobsen, 2007; Kwon and Wagner, 2007; Saze et al., 2008). Analysis of mutants in Swi2-related family members likely to interact directly with ARPs via their HSA domains (Fig. 5.7) led the way in these studies, including defects in SYD (Wagner and Meyerowitz, 2002), PIE1 (Swr1 homolog) (Noh and Amasino, 2003), DDM1 (Bartee and Bender, 2001; Brzeski and Jerzmanowski, 2003; Jeddeloh et al., 1999; Johnson et al., 2002; Mittelsten Scheid and Paszkowski, 2000; Miura et al., 2001), CR12 (Iswi) (Huanca-Mamani et al., 2005), DRD1 (Kanno et al., 2004), Ino80 (Fritsch et al., 2004), and Rad54 (Shaked et al., 2006). Considering that Arabidopsis has 44 predicted Swi2-like genes, but only six nuclear ARPs, it was not surprising to find that knocking down or knocking out individual Arabidopsis nuclear ARPs in whole plants resulted in even more severe and pleiotropic defects in more pathways of development than Swi2-related gene mutations (Choi et al., 2005; Deal et al., 2005, 2007; Kandasamy et al., 2005a,b; March-Diaz et al., 2007; Martin-Trillo et al., 2006; Meagher et al., 2005, 2007). A sampling of the cell proliferation and developmental abnormalities associated with Arabidopsis ARP4, ARP5, ARP6, and ARP7 deficiencies is summarized in Table 5.2. In animals, the majority of studies on nuclear ARPs are focused on conditional and knockdown mutants in cultured cells, where organismal viability is not compromised. It is possible that the plasticity of Arabidopsis development may allow individuals partially compromised for nuclear ARP-dependent epigenetic control to survive. New conditional knockdown technologies in the mouse should allow organismal- or organ-level gene silencing of nuclear ARPs to proceed.
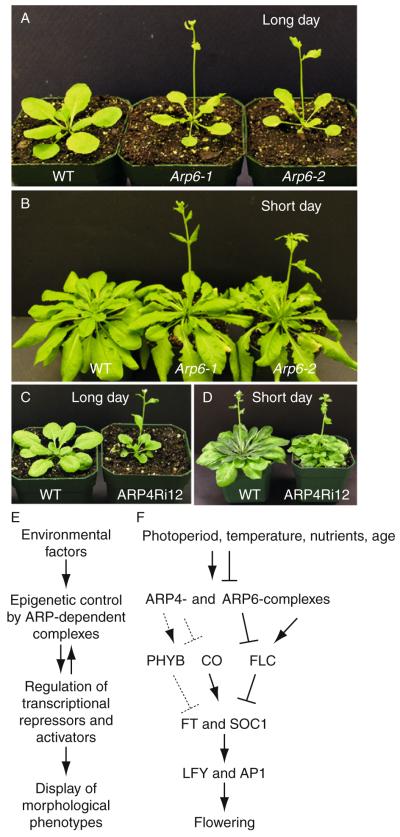
A theme presented at the beginning of this chapter states that in addition to their role in controlling the compaction of DNA and associated gene silencing, isoforms of nuclear ARP-containing chromatin complexes have evolved to exert dynamic epigenetic control over gene expression and different phases of multicellular development. A general model for the position of the nuclear ARPs in various pathways of developmental control is outlined in Fig. 5.8E. Chromatin remodeling factors like the ARPs affect epigenetic controls over the regulation of transcriptional repressors and activators leading to the display of various morphological phenotypes. However, the epigenetic controls themselves appear to be influenced by environmental factors such as age and diet in animals and age, nutrients, temperature, and light in plants.
Figure 5.8.
ARP6- and ARP4-defective Arabidopsis plants misregulate the phase transition from vegetative to reproductive growth. ARP6-defective plants flower early when grown under long- or short-day growth conditions, while ARP4-defective plants flower early only when grown under long-day conditions. (A) Twenty-day-old WT and arp6-1 (null) and arp6-2 (strong knockdown allele) plants grown under long-day conditions. (B) Fifty-day-old WT and arp6-1 and arp6-2 plants grown under short-day conditions. (C) Twenty-day-old WT and an ARP4 RNA interference silenced line (ARP4-Ri) grown under long-day conditions. (D) Sixty-eight-day-old WT and ARP4-Ri plant grown under short-day conditions. Long day = 16 h light/24 h. Short day = 9h light/24 h. (E) A general outline depicting the flow of information for developmental pathways under ARP-dependent epigenetic control. (F) Specific model for the ARP4- and ARP6-dependent control of flowering time. ARP6-containing SWR1 complexes potentiate the expression of the central repressor of flowering FLC. In ARP6 mutants when FLC levels are down, the levels of the transcriptional activators of flowering FT, SOC1, LFY, and AP1 are up and the plants flower early. ARP4-mediated epigenetic control of other high-level regulators of flowering including PhyB and CO is proposed in this model.
6.2. Developmental transitions
Epigenetic controls are essential to most major developmental phase transitions in animals and plants including the transitions from undifferentiated stem cells and primordia to organ development, from embryonic to vegetative growth, and from vegetative to reproductive development (Bezhani et al., 2007; Ooi and Henikoff, 2007; Vignon et al., 2002). Homeiotic genes that act early to determine organ identity appear to be under particularly strong epigenetic control (O’Dor et al., 2006; Papoulas et al., 1998; Pien et al., 2008; Vasanthi and Mishra, 2008). While ARP-containing complexes are clearly required for these processes, the role of nuclear ARPs themselves has been demonstrated in only a limited number of cases.
The transition from vegetative growth to flowering is one of the most important and most highly regulated developmental phase change in the life of a plant and appears exquisitely sensitive to changes in chromatin remodeling and modification. The decision to make this transition is regulated by a variety of environmental and endogenous plant cues including day length (photoperiod), temperature, nutrient status, the age of the plant and for many plants even the number of hours of exposure to cold weather (Fig. 5.8F). Each of these stimuli is sensed by separate genetic pathways, which feed into a central regulatory module that controls the switch from vegetative growth to flowering. This central module consists of the flowering repressor gene FLOWERING LOCUS C (FLC), which acts to repress the floral promoting genes FLOWERING LOCUS T (FT) and SUPRESSOR OF OVEREXPRESSION OF CONSTANS 1 (SOC1). When FLC levels are high, the FT and SOC1 genes are repressed, and vegetative growth continues. However, when FLC is epigenetically silenced, the FT and SOC1 genes are expressed and act to promote the expression of the floral meristem identity genes, resulting in flowering. Thus, these sensory pathways can promote flowering by repressing the expression of FLC or activating the expression of FT and SOC1, or they can repress flowering by activating FLC expression (Putterill et al., 2004; Simpson and Dean, 2002). In a parallel pathway, day length is perceived by the photoperiod pathway, and when the light period reaches a critical length in the spring, the transcription factor CONSTANS (CO) is produced, and serves to activate FT and SOC1, leading to the induction of flowering (Putterill et al., 1995; Samach et al., 2000). Another important sensory pathway, known as the autonomous pathway, is comprised of a collection of factors that repress FLC expression in response to diverse day length-independent stimuli, resulting in flowering (Simpson, 2004).
Interestingly, ARP4 and ARP6 have important, but separate roles in repressing the transition to flowering in Arabidopsis. Plants in which ARP4 expression has been silenced by RNAi, show early flowering in long days, but not in short days, indicating that the photoperiod-dependent pathway is aberrantly activated to promote flowering (Fig. 5.8C and D) (Kandasamy et al., 2005a). While the molecular basis of this phenotype is currently unknown, it is likely that ARP4-containing chromatin remodeling complexes may act to promote the expression of genes that repress CO, CO-like genes, or other regulatory genes, such as PHYTOCHROME B (Putterill et al., 1995). Alternatively, ARP4 may normally act to repress CO-related gene expression directly. In either case, CO could be activated inappropriately in ARP4-defective lines, leading to early flowering only under long-day conditions.
In contrast to the photoperiod-dependent ARP4 phenotype, a null mutant (arp6-1) and a severe knockdown mutant (arp6-2) defective in ARP6 expression result in early flowering under both long- and short-day photoperiods (Fig. 5.8A and B) (Choi et al., 2005; Deal et al., 2005; Martin-Trillo et al., 2006). This photoperiod-independent early flowering phenotype of arp6 plants was shown to result from a drastic 5- to 10-fold reduction in the expression of FLC, indicating that ARP6 is normally required to promote the expression of FLC. As mentioned previously, yeast and mammalian ARP6 proteins function with the SWR1 and SRCAP complexes, respectively, which act to deposit the histone variant H2AZ into chromatin. In support for the activity of Arabidopsis ARP6 in a homologous complex, mutations in the Arabidopsis homologs of two other components of this complex, PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1 (PIE1) and SERRATED LEAVES AND EARLY FLOWERING (SEF), also cause photoperiod-independent early flowering due to reduced FLC expression (March-Diaz et al., 2007; Noh and Amasino, 2003). Furthermore, we and others have recently shown that ARP6 and SEF interact with each other and with PIE1 (March-Diaz et al., 2007), and that ARP6 and PIE1 are required for deposition of H2AZ at the FLC locus (Deal et al., 2007). In addition, the loss of H2AZ from FLC chromatin in the arp6 and pie1 plants correlates with reduced FLC expression, indicating that plant H2AZ deposition in nucleosomes can potentiate transcriptional activation, similar to its role in yeast (Deal et al., 2007). These observations suggest that a SWR1-related complex is conserved in plants, and support a model in which dynamic ARP6/PIE1-mediated deposition of H2AZ is required for repression of flowering and maintenance of the vegetative growth phase.
A few subunits of ARP-containing chromatin remodeling complexes have roles in the phase transitions leading to animal organ and tissue development such as the development of bone, heart, striated muscle, T cells, and brain (Barak et al., 2004; Brown et al., 2007; Chi et al., 2002; Gebuhr et al., 2003; Lickert et al., 2004; Tromans, 2004; Yeh et al., 2002; Young et al., 2005). Solid evidence for the developmental roles for nuclear ARPs, however, is limited. Olave et al. (2002a) show that Arp4 isovariant Baf53b is first expressed in postmitotic neurons, and as part of a BAF (SWI/SNF) complex Baf53b interacts with the transcriptional corepressor CtBP to repress glucocorticoid receptor expression and alter development.
As in plants, the phase transition in yeast from vegetative to reproductive development and in particular the yeast mating type switch are under strong and complex epigenetic control (Klar, 2007). The mating type switch requires both DNA rearrangement and transcription of the MAT locus, with both INO80 and SWR1 complexes being recruited to the region of DNA with DSBs. ARP8 mutants are defective in the end processing of DNA strands during DSB repair at MAT (van Attikum et al., 2007). In summary, nuclear ARPs play essential roles in developmental phase transitions, however, the severity of some ARP-defective phenotypes may hinder a more complete understanding of their importance in these subtle processes.
6.3. Senescence and PCD
In plants and animals, chromatin remodeling and histone modification are known to be key to the epigenetic control of senescence and PCD (Adams, 2007a; Burzynski, 2005; Lewis et al., 2006; Sarnowski et al., 2005; Wu et al., 2008). As illustrated in Fig. 5.9E, floral senescence is controlled by a complex pathway that is influenced by age, nutrient status, temperature, and dozens of transcription factors. Clear floral senescence phenotypes are demonstrated for two classes of ARP-defective plants. In most flowering plants, the shedding of sepals and petals generally follows fertilization of the flower and development of fruit and seeds. As a consequence, when wild-type Arabidopsis are grown under long-day conditions the inflorescence displays only four to five opened flowers that retain intact sepals and petals (Fig. 5.9A). This small number of flowers with sepals and petals remains essentially constant from the time the first fruits develop until as many as several dozen fruits develop on an older inflorescence. Knocking down ARP4 or ARP7 expression with RNA interference (Ri) results in plants with defects in this senescence phase of flower development and often produces inflorescences that are fully covered with flowers bearing sepals and petals (Fig. 5.9B) (Kandasamy et al., 2005a,b). These ARP4-Ri and ARP7-Ri plant lines maintain fully turgid sepals and petals on 12–20 or more flowers during long-day growth and even fully mature fruit may retain sepals and petals (Fig. 5.9B–D). During short-day growth, six to eight wild-type flowers retain these organs at any one time, whereas, in the ARP4-Ri and ARP7-Ri lines 20 or more flowers retain sepals and petals.
Figure 5.9.
ARP4- and ARP7-defficient Arabidopsis plants display delayed floral organ senescence and abscission. (A) WT retains sepals and petals on only four to five flowers when grown under long-day conditions. (B) A moderate ARP7 RNA interference knockdown line (7Ri) retains petals and sepals after fertilization and even after the fruits are fully developed. (C) A strong ARP7 RNAi line (right) is relatively sterile and retains floral organs on all flowers as compared to wild type (left). (D) Close-up examination of developing fruits in a moderate ARP4 RNA interference line (4Ri) reveals retention of sepals and petals for a longer period than wild type (WT, A). (E) A general pathway proposed for the ARP4- and ARP7-dependent epigenetic control of floral organ senescence. A large number of transcription factors are known to control floral senescence and it is likely that the influence of age, nutrients, and temperature are processed through ARP-dependent epigenetic mechanisms. ARP7-dependent chromatin remodeling activities function downstream of ethylene perception, whereas the sites of ARP4s activities are unclear (Meagher et al., 2007).
These senescence phenotypes are the result of a failure of the cluster of cells at the base of sepals and petals, known as the abscission zone, to die and produce the required organ release, but not from a failure to develop the abscission zone of tissue itself (Kandasamy et al., 2005b). PCD at the abscission zones of petals and sepals is thought to proceed by autophagous mechanisms in which proteases are released from endoplasmic reticulum-derived vesicles to kill cells (Rogers, 2006). ARP4 and ARP7 may act within one or more chromatin modifying complexes to regulate the pathways of PCD leading to floral organ abscission. The hormone ethylene is known to play a central role in the regulation of flower senescence and abscission of floral organs in many plants (Bleecker and Patterson, 1997; Chang et al., 1993; Klee, 2002; Roberts, 2000). However, exogenous application of ethylene does not suppress the delayed floral organ abscission in ARP7-deficient lines (Kandasamy et al., 2005b) suggesting that ARP7 participates in controlling floral organ abscission in an ethylene-independent pathway. Epigenetic models for the normal action of ARP7-containing chromatin remodeling complexes may include the maintenance of ETHYLENE RECEPTOR 1 (ETR1) in an activated state poised to respond to ethylene or regulation of any number of transcription factors activating or repressing floral senescence (Fig. 5.9E). We feel it is most likely that many of the other transcription factors known to control floral senescence will also fall under nuclear ARP control (Meagher et al., 2007).
Deficiencies, overexpression, and misexpression of various subunits and activators of mammalian ARP-containing chromatin remodeling and modifying complexes induce or suppress apoptosis and various pathways of senescence (Samuelson et al., 2005). Ectopic expression of a histone acetyltransferase-defective Tip60 complex in HeLa cells causes a defect in DNA DSB repair, leading to an inability to carry out apoptosis (Ikura et al., 2000; Jehn and Osborne, 1997). The human snf5-homolog Ini1 is a subunit of the SWI/SNF NR complex. Overexpression of Ini1 in human ini1-deficeint cell lines increased the rates of apoptosis and arrested many cells in G1 (Ae et al., 2002; Klochendler-Yeivin et al., 2006). In heterozygous Ini1/ini1-null mice, p53 overexpression causes increased rates of tumor formation over the rates of tumor formation in wild type. These results lead the authors to suggest that Ini1 normally acts as a tumor suppressor. Similarly, overexpression of the Swi2 homolog Brg1 (Smarca4) increased p53 and retinoblastoma (Rb) expression and induced cell cycle arrest and apoptosis (Okazaki et al., 2008). Conversely, disruption of the Swi2-subunit homolog ATRX causes p53-mediated apoptosis (Seah et al., 2008). Normal SWI/SNF chromatin remodeling activity is essential to prevent apoptosis of cells undergoing DNA damage repair (Gong et al., 2008; Park et al., 2009).
In yeast cells, DSB repair is defective in ARP5- or ARP8-deficient cells or cells deficient in other subunits of the INO80 complex (van Attikum et al., 2004). RNAi silencing of the Swi2-related p400 in untransformed human fibroblasts causes G1 arrest, induction of p21, and increased levels of senescence-associated heterochromatic foci (Chan et al., 2005). Overexpression of Swi2-related Brg1 expression in mesenchymal stem cells induced a significant cell cycle arrest of MSCs in culture (Napolitano et al., 2007). The activities of both Rb and p53, commonly associated with senescence, are essential to this Brg1-mediated increase in stem cell senescence. Thus, there is strong direct and indirect evidence to suggest that mammalian nuclear ARP-containing chromatin remodeling complexes play roles in the control of apoptosis and senescence (Burzynski, 2005; Lin et al., 2006).
6.4. Cell proliferation and the cell cycle
Epigenetic controls over cell proliferation and the cell cycle are well established in animals, plants, and fungi (Chen et al., 2008; Francis, 2007). Roles for nuclear ARPs in cell cycle control are recently emerging as illustrated by a few studies in Arabidopsis and yeast, and for animal cells in culture. When grown under long-day conditions A. thaliana (Columbia) normally develops a rosette of about 12 leaves just prior to the initiation of flowering. The largest wild-type leaves average 4–5 cm in length at this developmental transition. Arabidopsis plants defective in ARP4, ARP5, ARP6, or ARP7 expression produce rosettes with leaves that are approximately half the size of wild type (Table 5.2, Fig. 5.10). Microscopic observations reveal that smaller leaves result from various abnormalities in leaf cell proliferation and/or expansion. Ri was used to generate stable epiallelic lines (ARP4-Ri), where ARP4 transcript levels are reduced to less than 20% of wild-type levels (Kandasamy et al., 2005a). The leaf epidermis of these strongly silenced epiallelic lines are composed of small cells that are less than half the size of wild type yet fairly normal in shape (Fig. 5.10A, D, and E). A null mutant lacking ARP5 (arp5-1) produces stochastic patches of very small epidermal cells interspersed among moderately sized cells to produce small leaves (Fig. 5.10B and F) (Kandasamy et al., 2009). Strongly silenced ARP7 RNAi lines (ARP7-Ri) also produce epidermal leaf cells that are essentially normally shaped, but average half the size of wild-type cells, with some clusters of very small cells (Fig. 5.10C and G) (Kandasamy et al., 2005b). Because of nonuniform epidermal cell expansion and clustering of extremely small and moderately sized cells and cell size differences on the abaxial and adaxial surfaces, ARP5- and ARP7-deficient leaves are often curled.
Figure 5.10.
Nuclear ARP-defects alter the cell cycle and/or cell proliferation with effects on leaf morphology. ARP4-, ARP5-, ARP6-, and ARP7-deficient Arabidopsis plants all produce dwarf leaves, but the small organs result from different epidermal leaf cell morphologies. (A, D, E) ARP4-defective RNAi lines (4Ri) produce very small leaves composed of mixtures of small and tiny cells (E) compared to wild type (WT, D). (B, F) The ARP5-defective null mutant arp5-1 displays small, elongated leaves with mixtures of large and small poorly lobed cells and a significant excess of underdeveloped stomatal complexes relative to WT. (C, G) ARP7-defective RNAi lines (7Ri) develop small curled leaves composed of a few normal sized cells interspersed with large numbers of very small cells, as compared with the WT. (H–J) The ARP6 null mutant (arp6–1) produces small curled leaves composed of fewer cells, but of relatively normal cell sizes and with only slightly reduced number of lobes relative to wild type. (D–G, I, J) Scanning electron micrographs compare the epidermis of WT to ARP-deficient lines. All light and scanning microscope images were prepared using the largest rosette leaves from 3-week-old plants. (K) Possible pathways for epigenetic control of the cell cycle and endocycle are outlined. The phenotypes of ARP4-, ARP5-, ARP6-, and ARP7-defective plants suggest that cell division, cell expansion, and cell differentiation are under the influence of complex epigenetic control. By this model, ARP-containing chromatin remodeling machines respond to environmental influences and regulate the expression of transcription factors and signaling proteins controlling the cell cycle, cell proliferation, and cell and organ development.
Stable knockout (arp6-1) and severe knockdown (arp6-2) lines of Arabidopsis were isolated that were defective in ARP6 expression (Deal et al., 2005). The ARP6-defective leaves were less than half the size of wild type (Fig. 5.10H). However, in contrast to defects in ARP4, ARP5, and ARP7, scanning electron microscopic examination of ARP6 mutant leaves reveals epidermal cell sizes and shapes that are indistinguishable from wild type (Fig. 5.10I and J). Thus, rather than producing smaller cells, ARP6-defective plants produce leaves composed of fewer total cells. These data suggest that during wild-type rosette leaf development, ARP6 promotes or extends the duration of cell proliferation to produce full-sized leaves. ARP4, ARP5, and ARP7 govern and balance rates of cell expansion, cell division, and the endocycle as suggested in the model pathway shown in Fig. 5.10K. We speculate that ARP deficiencies may produce these plant phenotypes, because of defects in epigenetic control including loss of normal chromatin remodeling essential for chromosomal segregation and changes in chromatin structure causing inappropriate expression of specific regulators of the cell cycle or endocycle including RBs, cyclins, and cyclin-dependent kinases (Francis, 2007; Meagher et al., 2007). Other than leaves, plant organs such as roots, stems, sepals, petals, stamens, and carpels are also smaller in these plants, but their cellular phenotypes have not yet been studied in detail.
Evidence from budding and fission yeast suggests that there is an essential role for nuclear ARPs in mitotic segregation, a process that impacts cell size and proliferation, which in multicellular organisms, helps determine organ size. In fission yeast, ARP4 and the DNA-dependent ATPase subunits of INO80 and SWI/SNF and SWR1 are all associated with centromeric heterochromatin. ARP4-defective cells are more sensitive to drugs known to inhibit progression through the cell cycle and arrest at the G2/M transition. The abnormally large budded ARP4-defective yeast cells display other mitotic phenotypes such as 2N DNA content and numerous broken spindles (Ogiwara et al., 2007b; Steinboeck et al., 2006). In addition, ARP4 associates with other known proteins in the kinetochore and is essential for its assembly. Fission yeast cells with defects in ARP4 expression also display mitotic phenotypes, including altered chromosome condensation and segregation and desilencing of centromeric transcription (Minoda et al., 2005). While a defect in ARP8 impairs the remodeling activity of INO80 and alters sister chromatid cohesion, it does not affect kinetochore assembly (Ogiwara et al., 2007a). Finally, ARP4-defective cells misregulated a number of genes in the TOR and NCR pathways that regulate cell growth (Steinboeck et al., 2006). Thus, it appears that ARP-containing complexes may be involved in the epigenetic control of chromatin remodeling and associated gene expression necessary for kinetochore assembly, chromosomal segregation, and progression through the cell cycle.
Strong indirect support also exists for the animal nuclear ARPs participating in the cell cycle and in cell proliferation. The ARP4 (BAF53)-containing mammalian BAF complexes were often classified as tumor suppressors (Sansam and Roberts, 2006) and by one view they may control the cell cycle by alternatively activating and repressing epigenetic states of the cell (Metivier et al., 2003). In flies and mammals, subunits of related SWI/SNF complexes act in the same pathway as E2F, a major regulator of cell cycle genes like the cyclins that impact, among other things, the G2/M transition (Staehling-Hampton et al., 1999). In human cell lines, subunits of the SWI/SNF complex interact with cyclin complexes and the tumor suppressor p53 to control cell cycle progression (Shanahan et al., 1999; Zhang et al., 2000). Human cultured cells expressing mutant forms of ARP4 show significant loss of viability and altered control of p53 (Lee et al., 2005). In addition, human ARP4 acts to repress p53-dependent p21-gene transcription by interacting directly with p53 (Wang et al., 2007) suggesting a specific role for this human ARP in the epigenetic control of the cell cycle and cell proliferation.
Using a combination of conditional mutations and drug inhibition of microtubule assembly, yeast ARP4 has now been shown to be associated with centromeric heterochromatin during the entire cell cycle and to be essential to cell division (Ogiwara et al., 2007b; Steinboeck et al., 2006). Moreover, defects in ARP4 cause arrest in the G2/M boundary of the cell cycle. These data suggest that perhaps the amount of ARP4 associated with the relatively small amount centromeric DNA was missed in previous microscopic localization studies showing that ARP4 was found concentrated in nucleoplasm of interphase cells.
7. Nuclear ARPs and Epigenetics in Human Disease
Altered chromatin structure has been allied with human disease for the last 30 years (Crapper et al., 1979; Fujii et al., 1980; Smith et al., 1979; Voitenko, 1980). One may predict with confidence that loss in the epigenetic control of gene expression for high-level transcription factors and the resulting alteration in tissue and organ development could account for many of the stochastic and complex properties associated with the onset of disease (Bottardi et al., 2003; Grewal and Moazed, 2003; Groudine et al., 1983; Nanney, 1958; Waddington, 1957). By considering epigenetic controls, we are better able to model how genes interact with the environment and how gene expression changes with the age of the individual. Otherwise, the influence of environment and age are difficult to interpret with purely genetic and biochemical models (Bjornsson et al., 2004; Ferluga, 1989; Graff and Mansuy, 2008; Mehler, 2008a; Weinhold, 2006). In the last decade, a strong connection has been made between epigenetic defects and diverse cancers and a variety of other human disorders including diabetes, cognitive dysfunction, autoimmune diseases, and neurobehavioral illnesses, all of which have age-related components associated with their frequency of onset (Adams, 2007b; Fraga et al., 2007; Garcia et al., 2005; Jirtle et al., 2000; Mehler and Purpura, 2008; Minucci and Pelicci, 1999; Remus et al., 2005; Robertson and Wolffe, 2000; Schulz et al., 2006; Shames et al., 2007; Soragni et al., 2008; Wong et al., 2007; Zeisig et al., 2008). ARP-containing chromatin remodeling complexes control directly or indirectly the majority of those changes to chromatin structure known to be associated with these diseases.
In particular, deficiencies in mutant forms of specific subunits of ARP-containing remodeling chromatin complex are linked to neoplastic cell growth and many subunits were initially identified as tumor suppressors (Olave et al., 2002b). DeCristofaro et al. (1999, 2001) and others have shown that a number of human tumors and immortal tumor-derived cells lines are deficient in subunits of the SWI/SNF family of complexes including defects in snf5, baf57, brg-1, and brm-1. For example, extracts from human adrenal gland carcinoma cells lack expression of the normally constitutive Swi2-related brg-1 and brm-1 proteins and form incomplete BAF and PBAF subcomplexes lacking ARP4 and actin. Transformation of the brg-1 gene into these cells disrupts their neoplastic state of growth and these genetically complemented cells begin to differentiate (Liu et al., 2001). Among its numerous activities on target genes, the BAF complex binds to the promoter of COLONY-STIMULATING FACTOR 1 (CSF1) promoter with a resulting alteration in DNA structure that appears essential for normal CSF1 expression. In another example, Ruvbl1 (RUVB-like 1, TIP49), an ATPase subunit in an ARP4-containing remodeling complex, has been implicated as a cofactor for oncogenic transformation via its binding to β-catenin. β-Catenin mediates the neoplastic growth of cancer cells through its role in Wnt signaling that, among other things, affects T-cell factor (TCF)-mediated transcription. Expression of an ATPase-deficient mutant form of Ruvbl1 (TIP49D302N) substantially inhibited β-catenin cell transformation (Feng et al., 2003). As another example of tumor suppressor activity, silencing the catalytic core subunits of SWI/SNF greatly reduces the ability of cells to carry out radiation-induced DSB repair and causes reduced survival of cells (Park et al., 2006).
DNA repair is essential to genome integrity and genetic inheritance, and inefficient or inaccurate repair results in mutations that may lead to carcinogenesis or inherited genetic diseases. Depending on the kind and severity of DNA damage, DNA repair proceeds through at least two major pathways: nucleotide excision repair (NER) and DSB repair. Evidence from diverse model systems suggests that reorganization of chromatin structure during NER and DSB repair may require the SWI/SNF, BAF, NEF, SWR1, RSC, INO80, SWR, and TIP60 complexes (Fritsch et al., 2004; Groth et al., 2007; Linger and Tyler, 2007; Morrison and Shen, 2006; Osley et al., 2007), whose compositions taken together include yeast and mammalian ARP4, ARP5, ARP6, ARP8, and yeast ARP7 and ARP9.
ARP4 is a common essential subunit of most of these complexes. The combined processes of chromatin remodeling and DNA damage repair have been modeled as proceeding through three major steps: access, repair, and restore (Smerdon and Lieberman, 1978). Access to the damaged site, arresting transcription, and arresting the cell cycle during DSB repair are accomplished by placing epigenetic marks on chromatin surrounding the damaged DNA. These marks include methylation, acetylation, ubiquitination, and phosphorylation of H2, H3, and/or H4 histone isovariants (Dinant et al., 2008). ARP4 is part of the TIP60 and NuA4 histone modifying complexes and is essential to localizing the yeast Nu4A complex to the sites of damaged DNA (Downs et al., 2004; Minoda et al., 2005). The early steps of access may also involve exchange of H2A, H2AX, and H2AZ isovariants within nucleosomes.
As mentioned earlier (Section 4.4), ARP4, ARP5, and ARP8 are essential components of the INO80 complex. INO80 is particularly associated with repair and phosphorylation of H2AX, a universal and well-characterized mark identifying DNA surrounding DSBs. Kitayama et al. (2009) have shown recently that human ARP5 partially complements the UV sensitivity of yeast ARP5 deletions. Silencing of human ARP5 in HeLa cells dramatically increases sensitivity to DNA DSBs. Furthermore, ARP5 depletion dramatically reduces phosphorylation of H2AX in these cells and the affinity of the Swi2-related Ino80 subunit of INO80 for DNA (Kitayama et al., 2009).
During the second step, repair, ARP8’s activity in the INO80 complex is implicated in converting DSBs to ssDNA and to recombination repair (van Attikum et al., 2004). Phosphoryated H2AX is essential to the recruitment of the ARP6-containing SWR1 complex to DSB sites (van Attikum et al., 2007). Studies on the third step, restore, in which normal chromatin dynamics and DNA activities are resumed is just beginning. Following the repair process, ARP5- and ARP8-containing INO80 complexes aid in restarting stalled replication forks (Shimada et al., 2008; Trujillo and Osley, 2008). These various steps of addressing DNA damage, access, repair, and restore require integrated activities of several nuclear ARPs and their chromatin remodeling complexes.
The nuclear ARPs themselves are described as tumor suppressors, because constituent ARP remodeling complexes activate genes with tumor suppressor activity. However, considering the complexity and dynamic nature of epigenetic control systems, it seems just as likely that ARP complexes may silence tumor suppressor genes as silence genes with other beneficial activities. For example, during ARP4 deficiency in budding yeast, a large number of stress response genes are increased or decreased in expression, relative to wild type (Gorzer et al., 2003; Shen et al., 2003a; Steinboeck et al., 2006). Similarly, ARP4-deficient fission yeast cells are defective in diverse histone H4 acetylation activities, resulting in the desilencing of genes linked to the centromere (Minoda et al., 2005).
ARP4 (Baf53) appears to be a necessary cofactor for c-Myc and p53 to participate in both tumor suppression and oncogenic transformation (Lee et al., 2005; Park et al., 2002; Wang et al., 2007). The cellular oncogenic proteins c-Myc and p53 play essential roles in the cell cycle, in DNA repair, and in apoptosis by regulating two relatively distinct and essential sets of genes and gene networks controlling these processes. Gene networks regulated by c-Myc and p53 and the normal expression of these two genes themselves are frequently disrupted in human and other animal cell cancers. ARP4 (Baf53) interacts with an N-terminal subdomain of c-Myc, the Myc-homology domain II (MBII) to form subcomplexes on chromatin and activate gene expression perhaps by recruiting ARP4-containing chromatin remodeling complexes to the promoters of these genes.
Mammalian ARP4 also interacts directly with p53 (Lee et al., 2005; Wang et al., 2007). Expression of a functional ARP4 protein with an intact N-terminal domain is essential for the normal expression of human p53. Alanine substitution at either the conserved Ser2 or Tyr6 residue in N-terminus of ARP4 inhibits human cell growth in culture, measured as reduced colony formation and growth rate (Lee et al., 2005). Expression of the Ser2Ala or Tyr6Ala forms show dramatically reduced transcription of p53. By contrast, the phosphomemetic form with Ser2Asp and Tyr6Glu substitutions complemented loss of ARP4 function restoring both normal cell growth and p53 expression levels to those of the parental cells. These data suggest that a phosphorylated form of ARP4 is essential for the control of the widely important activity of p53. Thus, ARP4 appears to be essential to regulating a secondary modification of p53 protein that affects p53’s activity as a transcriptional activator (Wang et al., 2007).
The nuclear ARPs are ubiquitously expressed at low levels and have no known functions that do not require their interaction with other subunits of chromatin remodeling complexes. Hence, the few examples of ARP expression levels being regulated in diseased cells are intriguing. Aging mice with the lethal yellow (LY) mutation in the agouti gene locus (Ay) show progressive obesity and reproductive dysfunction, similar to women with polycystic ovary syndrome (POS). When POS women are treated with insulin sensitizing agents like pioglitazone, they often have improved ovulatory function. Transcriptional array data on AyLY mice treated with pioglitazone revealed that a number of potentially important regulatory genes were misregulated, including a 50% drop in ARP6 transcript levels (Brannian et al., 2008). It is unknown if altered ARP6 levels affect insulin responsiveness or ovarian function. However, any reduction in nuclear ARP levels is likely to have a significant effect on the chromatin remodeling in these cells. We found, for example, that the Arabidopsis arp4-1 knockdown mutation, with only a 40% reduction in ARP4 levels throughout the plant, dramatically altered anther development and caused male sterility (Kandasamy et al., 2005a). Fission yeast expressing a temperature-sensitive arp4 allele (alp5-1134) display serious mitotic phenotypes at the nonpermissive temperature, including missegregation of chromosomes due to altered structures in kinetochores (Minoda et al., 2005). These plant and fungal data suggest that downregulation of ARP4 homologs in mammalian cells could result in cancerous cell growth.
ARP-containing chromatin remodeling and modifying complexes are essential to normal epigenetic control of gene expression, chromosomal segregation, and DNA repair—basal functions necessary to human health. Significant roles for nuclear ARPs and the numerous other subunits of ARP complexes are emerging in cancer and many other human diseases. It is likely that the impact of aging and environment on human disease, in particular, act via nuclear ARP-mediated epigenetic controls.
8. Conclusions
Fungal, plant, animal, and many protist genomes encode several ancient classes of nuclear ARPs that participate in large macromolecular machines directing chromatin dynamics. The nuclear ARPs are variously required for the assembly and the activities of these complexes via their binding to the HSA domains of Swi2- and Vid21-related proteins with helicase and ATPase activities. The nuclear ARPs help these complexes bind to chromatin, and in the cases of ARP4, ARP5, and ARP8, the ARPs appear to bind nucleosomal histones directly. Nuclear ARP complexes carry out NR including nucleosomal phasing and HVE reactions and the chemical modifications of histone termini. These activities exert an epigenetic control over basal levels of transcription affecting cell division and proliferation in all eukaryotes, but in particular, in animals and plants neoplastic cell growth and multicellular development. Genome integrity is maintained by the activities of nuclear ARP complexes in DNA replication, chromosome segregation, and DNA repair. In Arabidopsis, developmental pathways affecting nearly every plant tissue and organ and important organismal phase transitions require normal levels of ARP4, ARP5, ARP6, and/or ARP7 expression, providing strong evidence for their epigenetic role in multicellular development. In animals, among many roles, specific ARP-containing complexes are essential to heart, T cell, and brain development, but most studies on the nuclear ARPs themselves have been restricted primarily to cultured cells. Diverse isoforms of ARP complexes in animals and plants may provide greater target gene specificity and a wider variety of chromatin modifying activities than found in single-celled organisms like yeast. We propose that the greater epigenetic control provided by increases in novel ARP-containing complex isoforms was a major factor in the macroevolution of new tissues and organs in higher plants and animals.
ACKNOWLEDGMENTS
We thank Lori King for her critical reading of the manuscript and Jessie Kissinger (Genetics Department, University of Georgia) for her insights into the analysis of protist genomes. The National Institutes of Health (GM 36397) and the Department of Energy supported this work.
REFERENCES
- Adam RD. The Giardia lamblia genome. Int. J. Parasitol. 2000;30:475–484. doi: 10.1016/s0020-7519(99)00191-5. [DOI] [PubMed] [Google Scholar]
- Adam M, Robert F, Larochelle M, Gaudreau L. H2A.Z is required for global chromatin integrity and for recruitment of RNA polymerase II under specific conditions. Mol. Cell. Biol. 2001;21:6270–6279. doi: 10.1128/MCB.21.18.6270-6279.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams PD. Remodeling chromatin for senescence. Aging Cell. 2007a;6:425–427. doi: 10.1111/j.1474-9726.2007.00313.x. [DOI] [PubMed] [Google Scholar]
- Adams PD. Remodeling of chromatin structure in senescent cells and its potential impact on tumor suppression and aging. Gene. 2007b;397:84–93. doi: 10.1016/j.gene.2007.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adl SM, Leander BS, Simpson AG, Archibald JM, Anderson OR, Bass D, et al. Diversity, nomenclature, and taxonomy of protists. Syst. Biol. 2007;56:684–689. doi: 10.1080/10635150701494127. [DOI] [PubMed] [Google Scholar]
- Ae K, Kobayashi N, Sakuma R, Ogata T, Kuroda H, Kawaguchi N, et al. Chromatin remodeling factor encoded by ini1 induces G1 arrest and apoptosis in ini1-deficient cells. Oncogene. 2002;21:3112–3120. doi: 10.1038/sj.onc.1205414. [DOI] [PubMed] [Google Scholar]
- Aoyama N, Oka A, Kitayama K, Kurumizaka H, Harata M. The actin-related protein hArp8 accumulates on the mitotic chromosomes and functions in chromosome alignment. Exp. Cell Res. 2008;314:859–868. doi: 10.1016/j.yexcr.2007.11.020. [DOI] [PubMed] [Google Scholar]
- Aparicio S, Chapman J, Stupka E, Putnam N, Chia JM, Dehal P, et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science. 2002;297:1301–1310. doi: 10.1126/science.1072104. [DOI] [PubMed] [Google Scholar]
- Auger A, Galarneau L, Altaf M, Nourani A, Doyon Y, Utley RT, et al. Eaf1 is the platform for NuA4 molecular assembly that evolutionarily links chromatin acetylation to ATP-dependent exchange of histone H2A variants. Mol. Cell. Biol. 2008;28:2257–2270. doi: 10.1128/MCB.01755-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bak P, Paczuski M. Complexity, contingency, and criticality. Proc. Natl. Acad. Sci. USA. 1995;92:6689–6696. doi: 10.1073/pnas.92.15.6689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barak O, Lazzaro MA, Cooch NS, Picketts DJ, Shiekhattar R. A tissue-specific, naturally occurring human SNF2L variant inactivates chromatin remodeling. J. Biol. Chem. 2004;279:45130–45138. doi: 10.1074/jbc.M406212200. [DOI] [PubMed] [Google Scholar]
- Bartee L, Bender J. Two Arabidopsis methylation-deficiency mutations confer only partial effects on a methylated endogenous gene family. Nucleic Acids Res. 2001;29:2127–2134. doi: 10.1093/nar/29.10.2127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bendich AJ, Drlica K. Prokaryotic and eukaryotic chromosomes: what’s the difference? Bioessays. 2000;22:481–486. doi: 10.1002/(SICI)1521-1878(200005)22:5<481::AID-BIES10>3.0.CO;2-T. [DOI] [PubMed] [Google Scholar]
- Bezhani S, Winter C, Hershman S, Wagner JD, Kennedy JF, Kwon CS, et al. Unique, shared, and redundant roles for the Arabidopsis SWI/SNF chromatin remodeling ATPases BRAHMA and SPLAYED. Plant Cell. 2007;19:403–416. doi: 10.1105/tpc.106.048272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bird AW, Yu DY, Pray-Grant MG, Qiu Q, Harmon KE, Megee PC, et al. Acetylation of histone H4 by Esa1 is required for DNA double-strand break repair. Nature. 2002;419:411–415. doi: 10.1038/nature01035. [DOI] [PubMed] [Google Scholar]
- Bjornsson HT, Fallin MD, Feinberg AP. An integrated epigenetic and genetic approach to common human disease. Trends Genet. 2004;20:350–358. doi: 10.1016/j.tig.2004.06.009. [DOI] [PubMed] [Google Scholar]
- Bleecker AB, Patterson SE. Last exit: senescence, abscission, and meristem arrest in Arabidopsis. Plant Cell. 1997;9:1169–1179. doi: 10.1105/tpc.9.7.1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blessing CA, Ugrinova GT, Goodson HV. Actin and ARPs: action in the nucleus. Trends Cell Biol. 2004;14:435–442. doi: 10.1016/j.tcb.2004.07.009. [DOI] [PubMed] [Google Scholar]
- Bottardi S, Aumont A, Grosveld F, Milot E. Developmental stage-specific epigenetic control of human beta-globin gene expression is potentiated in hematopoietic progenitor cells prior to their transcriptional activation. Blood. 2003;102:3989–3997. doi: 10.1182/blood-2003-05-1540. [DOI] [PubMed] [Google Scholar]
- Boyer LA, Peterson CL. Actin-related proteins (Arps): conformational switches for chromatin-remodeling machines? Bioessays. 2000;22:666–672. doi: 10.1002/1521-1878(200007)22:7<666::AID-BIES9>3.0.CO;2-Y. [DOI] [PubMed] [Google Scholar]
- Brannian JD, Eyster KM, Weber M, Diggins M. Pioglitazone administration alters ovarian gene expression in aging obese lethal yellow mice. Reprod. Biol. Endo-crinol. 2008;6:10. doi: 10.1186/1477-7827-6-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown E, Malakar S, Krebs JE. How many remodelers does it take to make a brain? Diverse and cooperative roles of ATP-dependent chromatin-remodeling complexes in development. Biochem. Cell Biol. 2007;85:444–462. doi: 10.1139/O07-059. [DOI] [PubMed] [Google Scholar]
- Brzeski J, Jerzmanowski A. Deficient in DNA methylation 1 (DDM1) defines a novel family of chromatin-remodeling factors. J. Biol. Chem. 2003;278:823–828. doi: 10.1074/jbc.M209260200. [DOI] [PubMed] [Google Scholar]
- Burzynski SR. Aging: gene silencing or gene activation? Med. Hypotheses. 2005;64:201–208. doi: 10.1016/j.mehy.2004.06.010. [DOI] [PubMed] [Google Scholar]
- Cai Y, Jin J, Florens L, Swanson SK, Kusch T, Li B, et al. The mammalian YL1 protein is a shared subunit of the TRRAP/TIP60 histone acetyltransferase and SRCAP complexes. J. Biol. Chem. 2005;280:13665–13670. doi: 10.1074/jbc.M500001200. [DOI] [PubMed] [Google Scholar]
- Cai Y, Jin J, Gottschalk AJ, Yao T, Conaway JW, Conaway RC. Purification and assay of the human INO80 and SRCAP chromatin remodeling complexes. Methods. 2006;40:312–317. doi: 10.1016/j.ymeth.2006.06.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairns BR. Around the world of DNA damage INO80 days. Cell. 2004;119:733–735. doi: 10.1016/j.cell.2004.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairns BR, Erdjument-Bromage H, Tempst P, Winston F, Kornberg RD. Two actin-related proteins are shared functional components of the chromatin-remodeling complexes RSC and SWI/SNF. Mol. Cell. 1998;2:639–651. doi: 10.1016/s1097-2765(00)80162-8. [DOI] [PubMed] [Google Scholar]
- Champion K, Higgins NP. Growth rate toxicity phenotypes and homeostatic supercoil control differentiate Escherichia coli from Salmonella enterica serovar Typhimurium. J. Bacteriol. 2007;189:5839–5849. doi: 10.1128/JB.00083-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan HM, Narita M, Lowe SW, Livingston DM. The p400 E1A-associated protein is a novel component of the p53–> p21 senescence pathway. Genes Dev. 2005;19:196–201. doi: 10.1101/gad.1280205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang C, Kwok SF, Bleecker AB, Meyerowitz EM. Arabidopsis ethylene-response gene ETR1: similarity of product to two-component regulators. Science. 1993;262:539–544. doi: 10.1126/science.8211181. [DOI] [PubMed] [Google Scholar]
- Chen ZJ. Genetic and epigenetic mechanisms for gene expression and phenotypic variation in plant polyploids. Annu. Rev. Plant Biol. 2007;58:377–406. doi: 10.1146/annurev.arplant.58.032806.103835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M, Shen X. Nuclear actin and actin-related proteins in chromatin dynamics. Curr. Opin. Cell Biol. 2007;19:326–330. doi: 10.1016/j.ceb.2007.04.009. [DOI] [PubMed] [Google Scholar]
- Chen Z, Tan JL, Ingouff M, Sundaresan V, Berger F. Chromatin assembly factor 1 regulates the cell cycle but not cell fate during male gametogenesis in Arabidopsis thaliana. Development. 2008;135:65–73. doi: 10.1242/dev.010108. [DOI] [PubMed] [Google Scholar]
- Chi TH, Wan M, Zhao K, Taniuchi I, Chen L, Littman DR, et al. Reciprocal regulation of CD4/CD8 expression by SWI/SNF-like BAF complexes. Nature. 2002;418:195–199. doi: 10.1038/nature00876. [DOI] [PubMed] [Google Scholar]
- Choi EY, Park JA, Sung YH, Kwon H. Generation of the dominant-negative mutant of hArpNbeta: a component of human SWI/SNF chromatin remodeling complex. Exp. Cell Res. 2001;271:180–188. doi: 10.1006/excr.2001.5355. [DOI] [PubMed] [Google Scholar]
- Choi K, Kim S, Kim SY, Kim M, Hyun Y, Lee H, et al. SUPPRESSOR OF FRIGIDA3 encodes a nuclear ACTIN-RELATED PROTEIN6 required for floral repression in Arabidopsis. Plant Cell. 2005;17:2647–2660. doi: 10.1105/tpc.105.035485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi K, Park C, Lee J, Oh M, Noh B, Lee I. Arabidopsis homologs of components of the SWR1 complex regulate flowering and plant development. Development. 2007;134:1931–1941. doi: 10.1242/dev.001891. [DOI] [PubMed] [Google Scholar]
- Conaway RC, Conaway JW. The INO80 chromatin remodeling complex in transcription, replication and repair. Trends Biochem. Sci. 2009;34:71–77. doi: 10.1016/j.tibs.2008.10.010. [DOI] [PubMed] [Google Scholar]
- Crapper DR, Quittkat S, de Boni U. Altered chromatin conformation in Alzheimer’s disease. Brain. 1979;102:483–495. doi: 10.1093/brain/102.3.483. [DOI] [PubMed] [Google Scholar]
- Csink AK, Henikoff S. Genetic modification of heterochromatic association and nuclear organization in Drosophila. Nature. 1996;381:529–531. doi: 10.1038/381529a0. [DOI] [PubMed] [Google Scholar]
- Cuzin F, Grandjean V, Rassoulzadegan M. Inherited variation at the epigenetic level: paramutation from the plant to the mouse. Curr. Opin. Genet Dev. 2008;18:193–196. doi: 10.1016/j.gde.2007.12.004. [DOI] [PubMed] [Google Scholar]
- Dang W, Bartholomew B. Domain architecture of the catalytic subunit in the ISW2-nucleosome complex. Mol. Cell. Biol. 2007;27:8306–8317. doi: 10.1128/MCB.01351-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deal RB, Kandasamy MK, McKinney EC, Meagher RB. The nuclear actin-related protein ARP6 is a pleiotropic developmental regulator required for the maintenance of FLOWERING LOCUS C expression and repression of flowering in Arabidopsis. Plant Cell. 2005;17:2633–2646. doi: 10.1105/tpc.105.035196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deal RB, Topp CN, McKinney EC, Meagher RB. Repression of flowering in Arabidopsis requires activation of FLOWERING LOCUS C expression by the histone variant H2A.Z. Plant Cell. 2007;19:74–83. doi: 10.1105/tpc.106.048447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debril MB, Gelman L, Fayard E, Annicotte JS, Rocchi S, Auwerx J. Transcription factors and nuclear receptors interact with the SWI/SNF complex through the BAF60c subunit. J. Biol. Chem. 2004;279:16677–16686. doi: 10.1074/jbc.M312288200. [DOI] [PubMed] [Google Scholar]
- DeCristofaro MF, Betz BL, Wang W, Weissman BE. Alteration of hSNF5/INI1/BAF47 detected in rhabdoid cell lines and primary rhabdomyosarcomas but not Wilms’ tumors. Oncogene. 1999;18:7559–7565. doi: 10.1038/sj.onc.1203168. [DOI] [PubMed] [Google Scholar]
- DeCristofaro MF, Betz BL, Rorie CJ, Reisman DN, Wang W, Weissman BE. Characterization of SWI/SNF protein expression in human breast cancer cell lines and other malignancies. J. Cell Physiol. 2001;186:136–145. doi: 10.1002/1097-4652(200101)186:1<136::AID-JCP1010>3.0.CO;2-4. [DOI] [PubMed] [Google Scholar]
- Dinant C, Houtsmuller AB, Vermeulen W. Chromatin structure and DNA damage repair. Epigen. Chrom. 2008;1:9. doi: 10.1186/1756-8935-1-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dirscherl SS, Krebs JE. Functional diversity of ISWI complexes. Biochem. Cell Biol. 2004;82:482–489. doi: 10.1139/o04-044. [DOI] [PubMed] [Google Scholar]
- Downs JA, Allard S, Jobin-Robitaille O, Javaheri A, Auger A, Bouchard N, et al. Binding of chromatin-modifying activities to phosphorylated histone H2A at DNA damage sites. Mol. Cell. 2004;16:979–990. doi: 10.1016/j.molcel.2004.12.003. [DOI] [PubMed] [Google Scholar]
- Dryhurst D, Thambirajah AA, Ausio J. New twists on H2A.Z: a histone variant with a controversial structural and functional past. Biochem. Cell Biol. 2004;82:490–497. doi: 10.1139/o04-043. [DOI] [PubMed] [Google Scholar]
- Ebbert R, Birkmann A, Schuller HJ. The product of the SNF2/SWI2 paralogue INO80 of Saccharomyces cerevisiae required for efficient expression of various yeast structural genes is part of a high-molecular-weight protein complex. Mol. Microbiol. 1999;32:741–751. doi: 10.1046/j.1365-2958.1999.01390.x. [DOI] [PubMed] [Google Scholar]
- Eckley DM, Schroer TA. Interactions between the evolutionarily conserved, actin-related protein, Arp11, actin, and Arp1. Mol. Biol. Cell. 2003;14:2645–2654. doi: 10.1091/mbc.E03-01-0049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eckley DM, Gill SR, Melkonian KA, Bingham JB, Goodson HV, Heuser JE, et al. Analysis of dynactin subcomplexes reveals a novel actin-related protein associated with the arp1 minifilament pointed end. J. Cell Biol. 1999;147:307–320. doi: 10.1083/jcb.147.2.307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elango N, Kim SH, Vigoda E, Yi SV. Mutations of different molecular origins exhibit contrasting patterns of regional substitution rate variation. PLoS Comput. Biol. 2008;4:e1000015. doi: 10.1371/journal.pcbi.1000015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan JY, Rangasamy D, Luger K, Tremethick DJ. H2A.Z alters the nucleosome surface to promote HP1alpha-mediated chromatin fiber folding. Mol. Cell. 2004;16:655–661. doi: 10.1016/j.molcel.2004.10.023. [DOI] [PubMed] [Google Scholar]
- Feng Y, Lee N, Fearon ER. TIP49 regulates beta-catenin-mediated neoplastic transformation and T-cell factor target gene induction via effects on chromatin remodeling. Cancer Res. 2003;63:8726–8734. [PubMed] [Google Scholar]
- Ferluga J. Possible organ and age-related epigenetic factors in Huntington’s disease and colorectal carcinoma. Med. Hypotheses. 1989;29:51–54. doi: 10.1016/0306-9877(89)90168-0. [DOI] [PubMed] [Google Scholar]
- Ford J, Odeyale O, Shen CH. Activator-dependent recruitment of SWI/SNF and INO80 during INO1 activation. Biochem. Biophys. Res. Commun. 2008;373:602–606. doi: 10.1016/j.bbrc.2008.06.079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fraga MF, Agrelo R, Esteller M. Cross-talk between aging and cancer: the epigenetic language. Ann. NY Acad. Sci. 2007;1100:60–74. doi: 10.1196/annals.1395.005. [DOI] [PubMed] [Google Scholar]
- Francis D. The plant cell cycle—15 years on. New Phytol. 2007;174:261–278. doi: 10.1111/j.1469-8137.2007.02038.x. [DOI] [PubMed] [Google Scholar]
- Frankel S, Sigel EA, Craig C, Elgin SC, Mooseker MS, Artavanis-Tsakonas S. An actin-related protein in Drosophila colocalizes with heterochromatin protein 1 in pericentric heterochromatin. J. Cell Sci. 1997;110(Pt 17):1999–2012. doi: 10.1242/jcs.110.17.1999. [DOI] [PubMed] [Google Scholar]
- Fritsch O, Benvenuto G, Bowler C, Molinier J, Hohn B. The INO80 protein controls homologous recombination in Arabidopsis thaliana. Mol. Cell. 2004;16:479–485. doi: 10.1016/j.molcel.2004.09.034. [DOI] [PubMed] [Google Scholar]
- Fuchs M, Gerber J, Drapkin R, Sif S, Ikura T, Ogryzko V, et al. The p400 complex is an essential E1A transformation target. Cell. 2001;106:297–307. doi: 10.1016/s0092-8674(01)00450-0. [DOI] [PubMed] [Google Scholar]
- Fujii K, Yonemasu Y, Kitamura K, Tsubota Y. A genetic aspect of chromatin proteins of human brain tumors. J. Neurol. 1980;223:23–34. doi: 10.1007/BF00313137. [DOI] [PubMed] [Google Scholar]
- Galarneau L, Nourani A, Boudreault AA, Zhang Y, Heliot L, Allard S, et al. Multiple links between the NuA4 histone acetyltransferase complex and epigenetic control of transcription. Mol. Cell. 2000;5:927–937. doi: 10.1016/s1097-2765(00)80258-0. [DOI] [PubMed] [Google Scholar]
- Galkin VE, VanLoock MS, Orlova A, Egelman EH. A new internal mode in F-actin helps explain the remarkable evolutionary conservation of actin’s sequence and structure. Curr. Biol. 2002;12:570–575. doi: 10.1016/s0960-9822(02)00742-x. [DOI] [PubMed] [Google Scholar]
- Garcia BA, Busby SA, Shabanowitz J, Hunt DF, Mishra N. Resetting the epigenetic histone code in the MRL-lpr/lpr mouse model of lupus by histone deacetylase inhibition. J. Proteome Res. 2005;4:2032–2042. doi: 10.1021/pr050188r. [DOI] [PubMed] [Google Scholar]
- Gebuhr TC, Kovalev GI, Bultman S, Godfrey V, Su L, Magnuson T. The role of Brg1, a catalytic subunit of mammalian chromatin-remodeling complexes, in T cell development. J. Exp. Med. 2003;198:1937–1949. doi: 10.1084/jem.20030714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georgieva M, Harata M, Miloshev G. The nuclear actin-related protein Act3p/Arp4 influences yeast cell shape and bulk chromatin organization. J. Cell Biochem. 2008;104:59–67. doi: 10.1002/jcb.21600. [DOI] [PubMed] [Google Scholar]
- Gong F, Fahy D, Liu H, Wang W, Smerdon MJ. Role of the mammalian SWI/SNF chromatin remodeling complex in the cellular response to UV damage. Cell Cycle. 2008;7:1067–1074. doi: 10.4161/cc.7.8.5647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodson HV, Hawse WF. Molecular evolution of the actin family. J. Cell Sci. 2002;115:2619–2622. doi: 10.1242/jcs.115.13.2619. [DOI] [PubMed] [Google Scholar]
- Gordon JL, Sibley LD. Comparative genome analysis reveals a conserved family of actin-like proteins in apicomplexan parasites. BMC Genom. 2005;6:179. doi: 10.1186/1471-2164-6-179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon JL, Beatty WL, Sibley LD. A novel actin-related protein is associated with daughter cell formation in Toxoplasma gondii. Eukaryot. Cell. 2008;7:1500–1512. doi: 10.1128/EC.00064-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorzer I, Schuller C, Heidenreich E, Krupanska L, Kuchler K, Wintersberger U. The nuclear actin-related protein Act3p/Arp4p of Saccharomyces cerevisiae is involved in transcription regulation of stress genes. Mol. Microbiol. 2003;50:1155–1171. doi: 10.1046/j.1365-2958.2003.03759.x. [DOI] [PubMed] [Google Scholar]
- Graff J, Mansuy IM. Epigenetic codes in cognition and behaviour. Behav. Brain Res. 2008;192:70–87. doi: 10.1016/j.bbr.2008.01.021. [DOI] [PubMed] [Google Scholar]
- Grewal SI, Moazed D. Heterochromatin and epigenetic control of gene expression. Science. 2003;301:798–802. doi: 10.1126/science.1086887. [DOI] [PubMed] [Google Scholar]
- Groth A, Rocha W, Verreault A, Almouzni G. Chromatin challenges during DNA replication and repair. Cell. 2007;128:721–733. doi: 10.1016/j.cell.2007.01.030. [DOI] [PubMed] [Google Scholar]
- Groudine M, Kohwi-Shigematsu T, Gelinas R, Stamatoyannopoulos G, Papayannopoulou T. Human fetal to adult hemoglobin switching: changes in chromatin structure of the beta-globin gene locus. Proc. Natl. Acad. Sci. USA. 1983;80:7551–7555. doi: 10.1073/pnas.80.24.7551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haig D. The (dual) origin of epigenetics. Cold Spring Harb. Symp. Quant. Biol. 2004;69:67–70. doi: 10.1101/sqb.2004.69.67. [DOI] [PubMed] [Google Scholar]
- Harata M, Karwan A, Wintersberger U. An essential gene of Saccharomyces cerevisiae coding for an actin-related protein. Proc. Natl. Acad. Sci. USA. 1994;91:8258–8262. doi: 10.1073/pnas.91.17.8258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harata M, Mochizuki R, Mizuno S. Two isoforms of a human actin-related protein show nuclear localization and mutually selective expression between brain and other tissues. Biosci. Biotechnol. Biochem. 1999a;63:917–923. doi: 10.1271/bbb.63.917. [DOI] [PubMed] [Google Scholar]
- Harata M, Oma Y, Mizuno S, Jiang YW, Stillman DJ, Wintersberger U. The nuclear actin-related protein of Saccharomyces cerevisiae, Act3p/Arp4, interacts with core histones. Mol. Biol. Cell. 1999b;10:2595–2605. doi: 10.1091/mbc.10.8.2595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harata M, Oma Y, Tabuchi T, Zhang Y, Stillman DJ, Mizuno S. Multiple actin-related proteins of Saccharomyces cerevisiae are present in the nucleus. J. Biochem. (Tokyo) 2000;128:665–671. doi: 10.1093/oxfordjournals.jbchem.a022799. [DOI] [PubMed] [Google Scholar]
- Harata M, Nishimori K, Hatta S. Identification of two cDNAs for human actin-related proteins (Arps) that have remarkable similarity to conventional actin. Biochim. Biophys. Acta. 2001;1522:130–133. doi: 10.1016/s0167-4781(01)00315-3. [DOI] [PubMed] [Google Scholar]
- Harata M, Zhang Y, Stillman DJ, Matsui D, Oma Y, Nishimori K, et al. Correlation between chromatin association and transcriptional regulation for the Act3p/Arp4 nuclear actin-related protein of Saccharomyces cerevisiae. Nucleic Acids Res. 2002;30:1743–1750. doi: 10.1093/nar/30.8.1743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harries PA, Pan A, Quatrano RS. Actin-related protein2/3 complex component ARPC1 is required for proper cell morphogenesis and polarized cell growth in Physcomitrella patens. Plant Cell. 2005;17:2327–2339. doi: 10.1105/tpc.105.033266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedges SB, Blair JE, Venturi ML, Shoe JL. A molecular timescale of eukaryote evolution and the rise of complex multicellular life. BMC Evol. Biol. 2004;4:2. doi: 10.1186/1471-2148-4-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson IR, Jacobsen SE. Epigenetic inheritance in plants. Nature. 2007;447:418–424. doi: 10.1038/nature05917. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Position effects and variegation enhancers in an autosomal region of Drosophila melanogaster. Genetics. 1979;93:105–115. doi: 10.1093/genetics/93.1.105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huanca-Mamani W, Garcia-Aguilar M, Leon-Martinez G, Grossniklaus U, Vielle-Calzada JP. CHR11, a chromatin-remodeling factor essential for nuclear proliferation during female gametogenesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA. 2005;102:17231–17236. doi: 10.1073/pnas.0508186102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang ME, Souciet JL, Chuat JC, Galibert F. Identification of ACT4, a novel essential actin-related gene in the yeast Saccharomyces cerevisiae. Yeast. 1996;12:839–848. doi: 10.1002/(SICI)1097-0061(199607)12:9%3C839::AID-YEA982%3E3.0.CO;2-8. [DOI] [PubMed] [Google Scholar]
- Ikura T, Ogryzko VV, Grigoriev M, Groisman R, Wang J, Horikoshi M, et al. Involvement of the TIP60 histone acetylase complex in DNA repair and apoptosis. Cell. 2000;102:463–473. doi: 10.1016/s0092-8674(00)00051-9. [DOI] [PubMed] [Google Scholar]
- Iyer LM, Anantharaman V, Wolf MY, Aravind L. Comparative genomics of transcription factors and chromatin proteins in parasitic protists and other eukaryotes. Int. J. Parasitol. 2008;38:1–31. doi: 10.1016/j.ijpara.2007.07.018. [DOI] [PubMed] [Google Scholar]
- Jeddeloh JA, Stokes TL, Richards EJ. Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nat. Genet. 1999;22:94–97. doi: 10.1038/8803. [DOI] [PubMed] [Google Scholar]
- Jehn BM, Osborne BA. Gene regulation associated with apoptosis. Crit. Rev. Eukaryot. Gene Exp. 1997;7:179–193. doi: 10.1615/critreveukargeneexpr.v7.i1-2.100. [DOI] [PubMed] [Google Scholar]
- Jiang YW, Stillman DJ. Epigenetic effects on yeast transcription caused by mutations in an actin-related protein present in the nucleus. Genes Dev. 1996;10:604–619. doi: 10.1101/gad.10.5.604. [DOI] [PubMed] [Google Scholar]
- Jin J, Cai Y, Yao T, Gottschalk AJ, Florens L, Swanson SK, et al. A mammalian chromatin remodeling complex with similarities to the yeast INO80 complex. J. Biol. Chem. 2005;280:41207–41212. doi: 10.1074/jbc.M509128200. [DOI] [PubMed] [Google Scholar]
- Jirtle RL, Sander M, Barrett JC. Genomic imprinting and environmental disease susceptibility. Environ. Health Perspect. 2000;108:271–278. doi: 10.1289/ehp.00108271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson L, Cao X, Jacobsen S. Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr. Biol. 2002;12:1360–1367. doi: 10.1016/s0960-9822(02)00976-4. [DOI] [PubMed] [Google Scholar]
- Jonsson ZO, Jha S, Wohlschlegel JA, Dutta A. Rvb1p/Rvb2p recruit Arp5p and assemble a functional Ino80 chromatin remodeling complex. Mol. Cell. 2004;16:465–477. doi: 10.1016/j.molcel.2004.09.033. [DOI] [PubMed] [Google Scholar]
- Kandasamy MK, McKinney EC, Meagher RB. Cell cycle-dependent association of Arabidopsis actin-related proteins AtARP4 and AtARP7 with the nucleus. Plant J. 2003;33:939–948. doi: 10.1046/j.1365-313x.2003.01691.x. [DOI] [PubMed] [Google Scholar]
- Kandasamy MK, Deal RB, McKinney EC, Meagher RB. Plant actin-related proteins. Trends Plant Sci. 2004;9:196–202. doi: 10.1016/j.tplants.2004.02.004. [DOI] [PubMed] [Google Scholar]
- Kandasamy MK, Deal RB, McKinney EC, Meagher RB. Silencing the nuclear actin-related protein AtARP4 in Arabidopsis has multiple effects on plant development, including early flowering and delayed floral senescence. Plant J. 2005a;41:845–858. doi: 10.1111/j.1365-313X.2005.02345.x. [DOI] [PubMed] [Google Scholar]
- Kandasamy MK, McKinney EC, Deal RB, Meagher RB. Arabidopsis ARP7 is an essential actin-related protein required for normal embryogenesis, plant architecture, and floral organ abscission. Plant Physiol. 2005b;138:2019–2032. doi: 10.1104/pp.105.065326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandasamy MK, McKinney EC, Meagher RB. ACTIN-RELATED PROTEIN8 encodes an F-box protein localized to the nucleolus in Arabidopsis. Plant Cell Physiol. 2008;49:858–863. doi: 10.1093/pcp/pcn053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandasamy MK, McKinney LC, Deal RB, Smith AP, Meagher RB. Arabidopsis actin-related protein ARP5 in multicellular development and DNA repair. Developmental Biology. 2009 doi: 10.1016/j.ydbio.2009.08.006. In Press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanno T, Mette MF, Kreil DP, Aufsatz W, Matzke M, Matzke AJ. Involvement of putative SNF2 chromatin remodeling protein DRD1 in RNA-directed DNA methylation. Curr. Biol. 2004;14:801–805. doi: 10.1016/j.cub.2004.04.037. [DOI] [PubMed] [Google Scholar]
- Kato M, Sasaki M, Mizuno S, Harata M. Novel actin-related proteins in vertebrates: similarities of structure and expression pattern to Arp6 localized on Drosophila heterochromatin. Gene. 2001;268:133–140. doi: 10.1016/s0378-1119(01)00420-6. [DOI] [PubMed] [Google Scholar]
- Kawashima S, Ogiwara H, Tada S, Harata M, Wintersberger U, Enomoto T, et al. The INO80 complex is required for damage-induced recombination. Biochem. Biophys. Res. Commun. 2007;355:835–841. doi: 10.1016/j.bbrc.2007.02.036. [DOI] [PubMed] [Google Scholar]
- Kehrer-Sawatzki H, Cooper DN. Understanding the recent evolution of the human genome: insights from human-chimpanzee genome comparisons. Hum. Mutat. 2007;28:99–130. doi: 10.1002/humu.20420. [DOI] [PubMed] [Google Scholar]
- Kitayama K, Kamo M, Oma Y, Matsuda R, Uchida T, Ikura T, et al. The human actin-related protein hArp5: nucleo-cytoplasmic shuttling and involvement in DNA repair. Exp. Cell Res. 2009;315:206–217. doi: 10.1016/j.yexcr.2008.10.028. [DOI] [PubMed] [Google Scholar]
- Klar AJ. Lessons learned from studies of fission yeast mating-type switching and silencing. Annu. Rev. Genet. 2007;41:213–236. doi: 10.1146/annurev.genet.39.073103.094316. [DOI] [PubMed] [Google Scholar]
- Klee HJ. Control of ethylene-mediated processes in tomato at the level of receptors. J. Exp. Bot. 2002;53:2057–2063. doi: 10.1093/jxb/erf062. [DOI] [PubMed] [Google Scholar]
- Klochendler-Yeivin A, Picarsky E, Yaniv M. Increased DNA damage sensitivity and apoptosis in cells lacking the Snf5/Ini1 subunit of the SWI/SNF chromatin remodeling complex. Mol. Cell. Biol. 2006;26:2661–2674. doi: 10.1128/MCB.26.7.2661-2674.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kobor MS, Venkatasubrahmanyam S, Meneghini MD, Gin JW, Jennings JL, Link AJ, et al. A protein complex containing the conserved Swi2/Snf2-related ATPase Swr1p deposits histone variant H2A.Z into euchromatin. PLoS Biol. 2004;2:E131. doi: 10.1371/journal.pbio.0020131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogan NJ, Keogh MC, Datta N, Sawa C, Ryan OW, Ding H, et al. A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol. Cell. 2003;12:1565–1576. doi: 10.1016/s1097-2765(03)00497-0. [DOI] [PubMed] [Google Scholar]
- Kuo LJ, Yang LX. Gamma-H2AX—a novel biomarker for DNA double-strand breaks. In Vivo. 2008;22:305–309. [PubMed] [Google Scholar]
- Kuroda Y, Oma Y, Nishimori K, Ohta T, Harata M. Brain-specific expression of the nuclear actin-related protein ArpNalpha and its involvement in mammalian SWI/SNF chromatin remodeling complex. Biochem. Biophys. Res. Commun. 2002;299:328–334. doi: 10.1016/s0006-291x(02)02637-2. [DOI] [PubMed] [Google Scholar]
- Kwon CS, Wagner D. Unwinding chromatin for development and growth: a few genes at a time. Trends Genet. 2007;23:403–412. doi: 10.1016/j.tig.2007.05.010. [DOI] [PubMed] [Google Scholar]
- Larochelle M, Gaudreau L. H2A.Z has a function reminiscent of an activator required for preferential binding to intergenic DNA. Embo. J. 2003;22:4512–4522. doi: 10.1093/emboj/cdg427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazaro A, Gomez-Zambrano A, Lopez-Gonzalez L, Pineiro M, Jarillo JA. Mutations in the Arabidopsis SWC6 gene, encoding a component of the SWR1 chromatin remodelling complex, accelerate flowering time and alter leaf and flower development. J. Exp. Bot. 2008;59:653–666. doi: 10.1093/jxb/erm332. [DOI] [PubMed] [Google Scholar]
- Lee JH, Chang SH, Shim JH, Lee JY, Yoshida M, Kwon H. Cytoplasmic localization and nucleo-cytoplasmic shuttling of BAF53, a component of chromatin-modifying complexes. Mol. Cells. 2003;16:78–83. [PubMed] [Google Scholar]
- Lee JH, Lee JY, Chang SH, Kang MJ, Kwon H. Effects of Ser2 and Tyr6 mutants of BAF53 on cell growth and p53-dependent transcription. Mol. Cells. 2005;19:289–293. [PubMed] [Google Scholar]
- Lee K, Kang MJ, Kwon SJ, Kwon YK, Kim KW, Lim JH, et al. Expansion of chromosome territories with chromatin decompaction in BAF53-depleted interphase cells. Mol. Biol. Cell. 2007a;18:4013–4023. doi: 10.1091/mbc.E07-05-0437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee K, Shim JH, Kang MJ, Kim JH, Ahn JS, Yoo SJ, et al. Association of BAF53 with mitotic chromosomes. Mol. Cells. 2007b;24:288–293. [PubMed] [Google Scholar]
- Lessard J, Wu JI, Ranish JA, Wan M, Winslow MM, Staahl BT, et al. An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron. 2007;55:201–215. doi: 10.1016/j.neuron.2007.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis MW, Leslie ME, Liljegren SJ. Plant separation: 50 ways to leave your mother. Curr. Opin. Plant Biol. 2006;9:59–65. doi: 10.1016/j.pbi.2005.11.009. [DOI] [PubMed] [Google Scholar]
- Li B, Pattenden SG, Lee D, Gutierrez J, Chen J, Seidel C, et al. Preferential occupancy of histone variant H2AZ at inactive promoters influences local histone modifications and chromatin remodeling. Proc. Natl. Acad. Sci. USA. 2005;102:18385–18390. doi: 10.1073/pnas.0507975102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lickert H, Takeuchi JK, Von Both I, Walls JR, McAuliffe F, Adamson SL, et al. Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature. 2004;432:107–112. doi: 10.1038/nature03071. [DOI] [PubMed] [Google Scholar]
- Lin HY, Chen CS, Lin SP, Weng JR. Targeting histone deacetylase in cancer therapy. Med. Res. Rev. 2006;26:397–413. doi: 10.1002/med.20056. [DOI] [PubMed] [Google Scholar]
- Linger JG, Tyler JK. Chromatin disassembly and reassembly during DNA repair. Mutat. Res. 2007;618:52–64. doi: 10.1016/j.mrfmmm.2006.05.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu R, Liu H, Chen X, Kirby M, Brown PO, Zhao K. Regulation of CSF1 promoter by the SWI/SNF-like BAF complex. Cell. 2001;106:309–318. doi: 10.1016/s0092-8674(01)00446-9. [DOI] [PubMed] [Google Scholar]
- March-Diaz R, Garcia-Dominguez M, Florencio FJ, Reyes JC. SEF, a new protein required for flowering repression in Arabidopsis, interacts with PIE1 and ARP6. Plant Physiol. 2007;143:893–901. doi: 10.1104/pp.106.092270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin-Trillo M, Lazaro A, Poethig RS, Gomez-Mena C, Pineiro MA, Martinez-Zapater JM, et al. EARLY IN SHORT DAYS 1 (ESD1) encodes ACTIN-RELATED PROTEIN 6 (AtARP6), a putative component of chromatin remodelling complexes that positively regulates FLC accumulation in Arabidopsis. Development. 2006;133:1241–1252. doi: 10.1242/dev.02301. [DOI] [PubMed] [Google Scholar]
- McKinney EC, Kandasamy MK, Meagher RB. Arabidopsis contains ancient classes of differentially expressed actin-related protein genes. Plant Physiol. 2002;128:997–1007. doi: 10.1104/pp.010906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meagher RB. The impact of historical contingency on gene phylogeny: plant actin diversity. In: Hecht M, MacIntyre R, Clegg M, editors. Evolutionary Biology. Vol. 28. Plenum Press; NY: 1995. pp. 195–215. [Google Scholar]
- Meagher RB, McKinney EC, Kandasamy MK. Isovariant dynamics expand and buffer the responses of complex systems: the diverse plant actin gene family. Plant Cell. 1999a;11:995–1006. doi: 10.1105/tpc.11.6.995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meagher RB, McKinney EC, Vitale AV. The evolution of new structures: clues from plant cytoskeletal genes. Trends Genet. 1999b;15:278–284. doi: 10.1016/s0168-9525(99)01759-x. [DOI] [PubMed] [Google Scholar]
- Meagher RB, Deal RB, Kandasamy MK, McKinney EC. Nuclear actin-related proteins as epigenetic regulators of development. Plant Physiol. 2005;139:1576–1585. doi: 10.1104/pp.105.072447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meagher RB, Kandasamy MK, Deal RB, McKinney EC. Actin-related proteins in chromatin-level control of the cell cycle and developmental transitions. Trends Cell Biol. 2007;17:325–332. doi: 10.1016/j.tcb.2007.06.001. [DOI] [PubMed] [Google Scholar]
- Meagher RB, Kandasamy MK, McKinney EC. Multicellular development and protein–protein interactions. Plant Signal Behav. 2008;3:333–336. doi: 10.4161/psb.3.5.5343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehler MF. Epigenetic principles and mechanisms underlying nervous system functions in health and disease. Prog. Neurobiol. 2008a;86:305–341. doi: 10.1016/j.pneurobio.2008.10.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehler MF. Epigenetics and the nervous system. Ann. Neurol. 2008b;64:602–617. doi: 10.1002/ana.21595. [DOI] [PubMed] [Google Scholar]
- Mehler MF, Purpura DP. Autism, fever, epigenetics and the locus coeruleus. Brain Res. Rev. 2008;59:388–392. doi: 10.1016/j.brainresrev.2008.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meneghini MD, Wu M, Madhani HD. Conserved histone variant H2A.Z protects euchromatin from the ectopic spread of silent heterochromatin. Cell. 2003;112:725–736. doi: 10.1016/s0092-8674(03)00123-5. [DOI] [PubMed] [Google Scholar]
- Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M, et al. Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell. 2003;115:751–763. doi: 10.1016/s0092-8674(03)00934-6. [DOI] [PubMed] [Google Scholar]
- Minoda A, Saitoh S, Takahashi K, Toda T. BAF53/Arp4 homolog Alp5 in fission yeast is required for histone H4 acetylation, kinetochore-spindle attachment, and gene silencing at centromere. Mol. Biol. Cell. 2005;16:316–327. doi: 10.1091/mbc.E04-06-0519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minucci S, Pelicci PG. Retinoid receptors in health and disease: co-regulators and the chromatin connection. Semin. Cell Dev. Biol. 1999;10:215–225. doi: 10.1006/scdb.1999.0303. [DOI] [PubMed] [Google Scholar]
- Mittelsten Scheid O, Paszkowski J. Transcriptional gene silencing mutants. Plant Mol. Biol. 2000;43:235–241. doi: 10.1023/a:1006487529698. [DOI] [PubMed] [Google Scholar]
- Miura A, Yonebayashi S, Watanabe K, Toyama T, Shimada H, Kakutani T. Mobilization of transposons by a mutation abolishing full DNA methylation in Arabidopsis. Nature. 2001;411:212–214. doi: 10.1038/35075612. [DOI] [PubMed] [Google Scholar]
- Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science. 2004;303:343–348. doi: 10.1126/science.1090701. [DOI] [PubMed] [Google Scholar]
- Mohrmann L, Verrijzer CP. Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochim. Biophys. Acta. 2005;1681:59–73. doi: 10.1016/j.bbaexp.2004.10.005. [DOI] [PubMed] [Google Scholar]
- Morgan HD, Sutherland HG, Martin DI, Whitelaw E. Epigenetic inheritance at the agouti locus in the mouse. Nat. Genet. 1999;23:314–318. doi: 10.1038/15490. [DOI] [PubMed] [Google Scholar]
- Morrison AJ, Shen X. Chromatin modifications in DNA repair. Results Probl. Cell Differ. 2006;41:109–125. doi: 10.1007/400_008. [DOI] [PubMed] [Google Scholar]
- Morrison AJ, Highland J, Krogan NJ, Arbel-Eden A, Greenblatt JF, Haber JE, et al. INO80 and gamma-H2AX interaction links ATP-dependent chromatin remodeling to DNA damage repair. Cell. 2004;119:767–775. doi: 10.1016/j.cell.2004.11.037. [DOI] [PubMed] [Google Scholar]
- Muller GB, Newman SA. The innovation triad: an EvoDevo agenda. J. Exp. Zool. B Mol. Dev. Evol. 2005;304:487–503. doi: 10.1002/jez.b.21081. [DOI] [PubMed] [Google Scholar]
- Muller J, Oma Y, Vallar L, Friederich E, Poch O, Winsor B. Sequence and comparative genomic analysis of actin-related proteins. Mol. Biol. Cell. 2005;16:5736–5748. doi: 10.1091/mbc.E05-06-0508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nanney DL. Epigenetic control systems. Proc. Natl. Acad. Sci. USA. 1958;44:712–717. doi: 10.1073/pnas.44.7.712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Napolitano MA, Cipollaro M, Cascino A, Melone MA, Giordano A, Galderisi U. Brg1 chromatin remodeling factor is involved in cell growth arrest, apoptosis and senescence of rat mesenchymal stem cells. J. Cell Sci. 2007;120:2904–2911. doi: 10.1242/jcs.004002. [DOI] [PubMed] [Google Scholar]
- Nie Z, Xue Y, Yang D, Zhou S, Deroo BJ, Archer TK, et al. A specificity and targeting subunit of a human SWI/SNF family-related chromatin-remodeling complex. Mol. Cell. Biol. 2000;20:8879–8888. doi: 10.1128/mcb.20.23.8879-8888.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noh YS, Amasino RM. PIE1,an ISWI family gene, is required for FLC activation and floral repression in Arabidopsis. Plant Cell. 2003;15:1671–1682. doi: 10.1105/tpc.012161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Dor E, Beck SA, Brock HW. Polycomb group mutants exhibit mitotic defects in syncytial cell cycles of Drosophila embryos. Dev. Biol. 2006;290:312–322. doi: 10.1016/j.ydbio.2005.11.015. [DOI] [PubMed] [Google Scholar]
- Ogiwara H, Enomoto T, Seki M. The INO80 chromatin remodeling complex functions in sister chromatid cohesion. Cell Cycle. 2007a;6:1090–1095. doi: 10.4161/cc.6.9.4130. [DOI] [PubMed] [Google Scholar]
- Ogiwara H, Ui A, Kawashima S, Kugou K, Onoda F, Iwahashi H, et al. Actin-related protein Arp4 functions in kinetochore assembly. Nucleic Acids Res. 2007b;35:3109–3117. doi: 10.1093/nar/gkm161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohfuchi E, Nishimori K, Harata M. Alternative splicing products of the gene for a human nuclear actin-related protein, hArpNbeta/Baf53, that encode a protein isoform, hArpNbetaS, in the cytoplasm. Biosci. Biotechnol. Biochem. 2002;66:1740–1743. doi: 10.1271/bbb.66.1740. [DOI] [PubMed] [Google Scholar]
- Ohfuchi E, Kato M, Sasaki M, Sugimoto K, Oma Y, Harata M. Vertebrate Arp6, a novel nuclear actin-related protein, interacts with heterochromatin protein 1. Eur. J. Cell Biol. 2006;85:411–421. doi: 10.1016/j.ejcb.2005.12.006. [DOI] [PubMed] [Google Scholar]
- Okazaki N, Ikeda S, Ohara R, Shimada K, Yanagawa T, Nagase T, et al. The novel protein complex with SMARCAD1/KIAA1122 binds to the vicinity of TSS. J. Mol. Biol. 2008;382:257–265. doi: 10.1016/j.jmb.2008.07.031. [DOI] [PubMed] [Google Scholar]
- Olave I, Wang W, Xue Y, Kuo A, Crabtree GR. Identification of a polymorphic, neuron-specific chromatin remodeling complex. Genes Dev. 2002a;16:2509–2517. doi: 10.1101/gad.992102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olave IA, Reck-Peterson SL, Crabtree GR. Nuclear actin and actin-related proteins in chromatin remodeling. Annu. Rev. Biochem. 2002b;71:755–781. doi: 10.1146/annurev.biochem.71.110601.135507. [DOI] [PubMed] [Google Scholar]
- Oma Y, Nishimori K, Harata M. The brain-specific actin-related protein ArpN alpha interacts with the transcriptional co-repressor CtBP. Biochem. Biophys. Res. Commun. 2003;301:521–528. doi: 10.1016/s0006-291x(02)03073-5. [DOI] [PubMed] [Google Scholar]
- Ooi SL, Henikoff S. Germline histone dynamics and epigenetics. Curr. Opin. Cell Biol. 2007;19:257–265. doi: 10.1016/j.ceb.2007.04.015. [DOI] [PubMed] [Google Scholar]
- Osley MA, Tsukuda T, Nickoloff JA. ATP-dependent chromatin remodeling factors and DNA damage repair. Mutat. Res. 2007;618:65–80. doi: 10.1016/j.mrfmmm.2006.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papoulas O, Beek SJ, Moseley SL, McCallum CM, Sarte M, Shearn A, et al. The Drosophila trithorax group proteins BRM, ASH1 and ASH2 are subunits of distinct protein complexes. Development. 1998;125:3955–3966. doi: 10.1242/dev.125.20.3955. [DOI] [PubMed] [Google Scholar]
- Park J, Wood MA, Cole MD. BAF53 forms distinct nuclear complexes and functions as a critical c-Myc-interacting nuclear cofactor for oncogenic transformation. Mol. Cell. Biol. 2002;22:1307–1316. doi: 10.1128/mcb.22.5.1307-1316.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park JH, Park EJ, Lee HS, Kim SJ, Hur SK, Imbalzano AN, et al. Mammalian SWI/SNF complexes facilitate DNA double-strand break repair by promoting gamma-H2AX induction. Embo J. 2006;25:3986–3997. doi: 10.1038/sj.emboj.7601291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park JH, Park EJ, Hur SK, Kim S, Kwon J. Mammalian SWI/SNF chromatin remodeling complexes are required to prevent apoptosis after DNA damage. DNA Repair (Amst) 2009;8:29–39. doi: 10.1016/j.dnarep.2008.08.011. [DOI] [PubMed] [Google Scholar]
- Peterson CL, Zhao Y, Chait BT. Subunits of the yeast SWI/SNF complex are members of the actin-related protein (ARP) family. J. Biol. Chem. 1998;273:23641–23644. doi: 10.1074/jbc.273.37.23641. [DOI] [PubMed] [Google Scholar]
- Pien S, Fleury D, Mylne JS, Crevillen P, Inze D, Avramova Z, et al. ARABIDOPSIS TRITHORAX1 dynamically regulates FLOWERING LOCUS C activation via histone 3 lysine 4 trimethylation. Plant Cell. 2008;20:580–588. doi: 10.1105/tpc.108.058172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pillus L, Rine J. Epigenetic inheritance of transcriptional states in S. cerevisiae. Cell. 1989;59:637–647. doi: 10.1016/0092-8674(89)90009-3. [DOI] [PubMed] [Google Scholar]
- Poch O, Winsor B. Who’s who among the Saccharomyces cerevisiae actin-related proteins? A classification and nomenclature proposal for a large family. Yeast. 1997;13:1053–1058. doi: 10.1002/(SICI)1097-0061(19970915)13:11<1053::AID-YEA164>3.0.CO;2-4. [DOI] [PubMed] [Google Scholar]
- Putterill J, Robson F, Lee K, Simon R, Coupland G. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell. 1995;80:847–857. doi: 10.1016/0092-8674(95)90288-0. [DOI] [PubMed] [Google Scholar]
- Putterill J, Laurie R, Macknight R. It’s time to flower: the genetic control of flowering time. Bioessays. 2004;26:363–373. doi: 10.1002/bies.20021. [DOI] [PubMed] [Google Scholar]
- Raisner RM, Madhani HD. Patterning chromatin: form and function for H2A.Z variant nucleosomes. Curr. Opin. Genet. Dev. 2006;16:119–124. doi: 10.1016/j.gde.2006.02.005. [DOI] [PubMed] [Google Scholar]
- Raisner RM, Hartley PD, Meneghini MD, Bao MZ, Liu CL, Schreiber SL, et al. Histone variant H2A.Z marks the 5′ ends of both active and inactive genes in euchromatin. Cell. 2005;123:233–248. doi: 10.1016/j.cell.2005.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rando OJ, Zhao K, Janmey P, Crabtree GR. Phosphatidylinositol-dependent actin filament binding by the SWI/SNF-like BAF chromatin remodeling complex. Proc. Natl. Acad. Sci. USA. 2002;99:2824–2829. doi: 10.1073/pnas.032662899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Remus R, Kanzaki A, Yawata A, Nakanishi H, Wada H, Sugihara T, et al. DNA methylation in promoter regions of red cell membrane protein genes in healthy individuals and patients with hereditary membrane disorders. Int. J. Hematol. 2005;81:385–395. doi: 10.1532/ijh97.04171. [DOI] [PubMed] [Google Scholar]
- Roberts JA. Abscission and dehiscence. Symp. Soc. Exp. Biol. 2000;52:203–211. [PubMed] [Google Scholar]
- Robertson KD, Wolffe AP. DNA methylation in health and disease. Nat. Rev. Genet. 2000;1:11–19. doi: 10.1038/35049533. [DOI] [PubMed] [Google Scholar]
- Rogers HJ. Programmed cell death in floral organs: how and why do flowers die? Ann. Bot. (Lond) 2006;97:309–315. doi: 10.1093/aob/mcj051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roux-Rouquie M. Genetic and epigenetic regulation schemes: need for an alternative paradigm. Mol. Genet. Metab. 2000;71:1–9. doi: 10.1006/mgme.2000.3048. [DOI] [PubMed] [Google Scholar]
- Ruhl DD, Jin J, Cai Y, Swanson S, Florens L, Washburn MP, et al. Purification of a human SRCAP complex that remodels chromatin by incorporating the histone variant H2A.Z into nucleosomes. Biochemistry. 2006;45:5671–5677. doi: 10.1021/bi060043d. [DOI] [PubMed] [Google Scholar]
- Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz-Sommer Z, Yanofsky MF, et al. Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science. 2000;288:1613–1616. doi: 10.1126/science.288.5471.1613. [DOI] [PubMed] [Google Scholar]
- Samuelson AV, Narita M, Chan HM, Jin J, de Stanchina E, McCurrach ME, et al. p400 is required for E1A to promote apoptosis. J. Biol. Chem. 2005;280:21915–21923. doi: 10.1074/jbc.M414564200. [DOI] [PubMed] [Google Scholar]
- Sansam CG, Roberts CW. Epigenetics and cancer: altered chromatin remodeling via Snf5 loss leads to aberrant cell cycle regulation. Cell Cycle. 2006;5:621–624. doi: 10.4161/cc.5.6.2579. [DOI] [PubMed] [Google Scholar]
- Santisteban MS, Kalashnikova T, Smith MM. Histone H2A.Z regulates transcription and is partially redundant with nucleosome remodeling complexes. Cell. 2000;103:411–422. doi: 10.1016/s0092-8674(00)00133-1. [DOI] [PubMed] [Google Scholar]
- Sardiu ME, Cai Y, Jin J, Swanson SK, Conaway RC, Conaway JW, et al. Probabilistic assembly of human protein interaction networks from label-free quantitative proteomics. Proc. Natl. Acad. Sci. USA. 2008;105:1454–1459. doi: 10.1073/pnas.0706983105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarnowski TJ, Rios G, Jasik J, Swiezewski S, Kaczanowski S, Li Y, et al. SWI3 subunits of putative SWI/SNF chromatin-remodeling complexes play distinct roles during Arabidopsis development. Plant Cell. 2005;17:2454–2472. doi: 10.1105/tpc.105.031203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saze H, Sasaki T, Kakutani T. Negative regulation of DNA methylation in plants. Epigenetics. 2008;3:122–124. doi: 10.4161/epi.3.3.6355. [DOI] [PubMed] [Google Scholar]
- Schulz WA, Steinhoff C, Florl AR. Methylation of endogenous human retro-elements in health and disease. Curr. Top. Microbiol. Immunol. 2006;310:211–250. doi: 10.1007/3-540-31181-5_11. [DOI] [PubMed] [Google Scholar]
- Seah C, Levy MA, Jiang Y, Mokhtarzada S, Higgs DR, Gibbons RJ, et al. Neuronal death resulting from targeted disruption of the Snf2 protein ATRX is mediated by p53. J. Neurosci. 2008;28:12570–12580. doi: 10.1523/JNEUROSCI.4048-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaked H, Avivi-Ragolsky N, Levy AA. Involvement of the Arabidopsis SWI2/SNF2 chromatin remodeling gene family in DNA damage response and recombination. Genetics. 2006;173:985–994. doi: 10.1534/genetics.105.051664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shames DS, Minna JD, Gazdar AF. DNA methylation in health, disease, and cancer. Curr. Mol. Med. 2007;7:85–102. doi: 10.2174/156652407779940413. [DOI] [PubMed] [Google Scholar]
- Shanahan F, Seghezzi W, Parry D, Mahony D, Lees E. Cyclin E associates with BAF155 and BRG1, components of the mammalian SWI-SNF complex, and alters the ability of BRG1 to induce growth arrest. Mol. Cell. Biol. 1999;19:1460–1469. doi: 10.1128/mcb.19.2.1460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen X, Mizuguchi G, Hamiche A, Wu C. A chromatin remodelling complex involved in transcription and DNA processing. Nature. 2000;406:541–544. doi: 10.1038/35020123. [DOI] [PubMed] [Google Scholar]
- Shen J, Hsu CM, Kang BK, Rosen BP, Bhattacharjee H. The Saccharomyces cerevisiae Arr4p is involved in metal and heat tolerance. Biometals. 2003a;16:369–378. doi: 10.1023/a:1022504311669. [DOI] [PubMed] [Google Scholar]
- Shen X, Ranallo R, Choi E, Wu C. Involvement of actin-related proteins in ATP-dependent chromatin remodeling. Mol. Cell. 2003b;12:147–155. doi: 10.1016/s1097-2765(03)00264-8. [DOI] [PubMed] [Google Scholar]
- Shimada K, Oma Y, Schleker T, Kugou K, Ohta K, Harata M, et al. Ino80 chromatin remodeling complex promotes recovery of stalled replication forks. Curr. Biol. 2008;18:566–575. doi: 10.1016/j.cub.2008.03.049. [DOI] [PubMed] [Google Scholar]
- Simone C, Forcales SV, Hill DA, Imbalzano AN, Latella L, Puri PL. p38 pathway targets SWI-SNF chromatin-remodeling complex to muscle-specific loci. Nat. Genet. 2004;36:738–743. doi: 10.1038/ng1378. [DOI] [PubMed] [Google Scholar]
- Simpson GG. The autonomous pathway: epigenetic and post-transcriptional gene regulation in the control of Arabidopsis flowering time. Curr. Opin. Plant Biol. 2004;7:570–574. doi: 10.1016/j.pbi.2004.07.002. [DOI] [PubMed] [Google Scholar]
- Simpson GG, Dean C. Arabidopsis, the Rosetta stone of flowering time? Science. 2002;296:285–289. doi: 10.1126/science.296.5566.285. [DOI] [PubMed] [Google Scholar]
- Smerdon MJ, Lieberman MW. Nucleosome rearrangement in human chromatin during UV-induced DNA- reapir synthesis. Proc. Natl. Acad. Sci. USA. 1978;75:4238–4241. doi: 10.1073/pnas.75.9.4238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith HS, Springer EL, Hackett AJ. Nuclear ultrastructure of epithelial cell lines derived from human carcinomas and nonmalignant tissues. Cancer Res. 1979;39:332–344. [PubMed] [Google Scholar]
- Soragni E, Herman D, Dent SY, Gottesfeld JM, Wells RD, Napierala M. Long intronic GAA*TTC repeats induce epigenetic changes and reporter gene silencing in a molecular model of Friedreich ataxia. Nucleic Acids Res. 2008;36:6056–6065. doi: 10.1093/nar/gkn604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staehling-Hampton K, Ciampa PJ, Brook A, Dyson N. A genetic screen for modifiers of E2F in Drosophila melanogaster. Genetics. 1999;153:275–287. doi: 10.1093/genetics/153.1.275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stefanov SA. Further characterization of the actin-related protein Act3p/Arp4 of S. cerevisiae through mutational analysis. Mol. Biol. Rep. 2000;27:35–43. doi: 10.1023/a:1007173530124. [DOI] [PubMed] [Google Scholar]
- Steinboeck F, Krupanska L, Bogusch A, Kaufmann A, Heidenreich E. Novel regulatory properties of Saccharomyces cerevisiae Arp4. J. Biochem. (Tokyo) 2006;139:741–751. doi: 10.1093/jb/mvj080. [DOI] [PubMed] [Google Scholar]
- Su Y, Kwon CS, Bezhani S, Huvermann B, Chen C, Peragine A, et al. The N-terminal ATPase AT-hook-containing region of the Arabidopsis chromatin-remodeling protein SPLAYED is sufficient for biological activity. Plant J. 2006;46:685–699. doi: 10.1111/j.1365-313X.2006.02734.x. [DOI] [PubMed] [Google Scholar]
- Sumi-Ichinose C, Ichinose H, Metzger D, Chambon P. SNF2beta-BRG1 is essential for the viability of F9 murine embryonal carcinoma cells. Mol. Cell. Biol. 1997;17:5976–5986. doi: 10.1128/mcb.17.10.5976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sunada R, Gorzer I, Oma Y, Yoshida T, Suka N, Wintersberger U, et al. The nuclear actin-related protein Act3p/Arp4p is involved in the dynamics of chromatin-modulating complexes. Yeast. 2005;22:753–768. doi: 10.1002/yea.1239. [DOI] [PubMed] [Google Scholar]
- Szerlong H, Saha A, Cairns BR. The nuclear actin-related proteins Arp7 and Arp9: a dimeric module that cooperates with architectural proteins for chromatin remodeling. EMBO J. 2003;22:3175–3187. doi: 10.1093/emboj/cdg296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szerlong H, Hinata K, Viswanathan R, Erdjument-Bromage H, Tempst P, Cairns BR. The HSA domain binds nuclear actin-related proteins to regulate chromatin-remodeling ATPases. Nat. Struct. Mol. Biol. 2008;15:469–476. doi: 10.1038/nsmb.1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- Tromans A. Cardiovascular biology: how genes know their place. Nature. 2004;432:29. doi: 10.1038/432029a. [DOI] [PubMed] [Google Scholar]
- True HL, Berlin I, Lindquist SL. Epigenetic regulation of translation reveals hidden genetic variation to produce complex traits. Nature. 2004;431:184–187. doi: 10.1038/nature02885. [DOI] [PubMed] [Google Scholar]
- Trujillo KM, Osley MA. INO80 meets a fork in the road. Nat. Struct. Mol. Biol. 2008;15:332–334. doi: 10.1038/nsmb0408-332. [DOI] [PubMed] [Google Scholar]
- Ueno M, Murase T, Kibe T, Ohashi N, Tomita K, Murakami Y, et al. Fission yeast Arp6 is required for telomere silencing, but functions independently of Swi6. Nucleic Acids Res. 2004;32:736–741. doi: 10.1093/nar/gkh234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Attikum H, Fritsch O, Hohn B, Gasser SM. Recruitment of the INO80 complex by H2A phosphorylation links ATP-dependent chromatin remodeling with DNA double-strand break repair. Cell. 2004;119:777–788. doi: 10.1016/j.cell.2004.11.033. [DOI] [PubMed] [Google Scholar]
- van Attikum H, Fritsch O, Gasser SM. Distinct roles for SWR1 and INO80 chromatin remodeling complexes at chromosomal double-strand breaks. EMBO J. 2007;26:4113–4125. doi: 10.1038/sj.emboj.7601835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Ent F, Amos L, Lowe J. Prokaryotic origin of the actin cytoskeleton. Nature. 2001;413:39–44. doi: 10.1038/35092500. [DOI] [PubMed] [Google Scholar]
- Vasanthi D, Mishra RK. Epigenetic regulation of genes during development: a conserved theme from flies to mammals. J. Genet. Genomics. 2008;35:413–429. doi: 10.1016/S1673-8527(08)60059-4. [DOI] [PubMed] [Google Scholar]
- Vignon X, Zhou Q, Renard JP. Chromatin as a regulative architecture of the early developmental functions of mammalian embryos after fertilization or nuclear transfer. Cloning Stem Cells. 2002;4:363–377. doi: 10.1089/153623002321025041. [DOI] [PubMed] [Google Scholar]
- Voitenko VP. Aging, diseases and X-chromatin. Z. Gerontol. 1980;13:18–23. [PubMed] [Google Scholar]
- Waddington CH. The Strategy of the Genes. George Allen & Unwin Ltd; London, Ruskin House: 1995. The strategy of the genes; p. 257. [Google Scholar]
- Wagner D, Meyerowitz EM. SPLAYED, a novel SWI/SNF ATPase homolog, controls reproductive development in Arabidopsis. Curr. Biol. 2002;12:85–94. doi: 10.1016/s0960-9822(01)00651-0. [DOI] [PubMed] [Google Scholar]
- Walley JW, Rowe HC, Xiao Y, Chehab EW, Kliebenstein DJ, Wagner D, et al. The chromatin remodeler SPLAYED regulates specific stress signaling pathways. PLoS Pathog. 2008;4:e1000237. doi: 10.1371/journal.ppat.1000237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W, Cote J, Xue Y, Zhou S, Khavari PA, Biggar SR, et al. Purification and biochemical heterogeneity of the mammalian SWI-SNF complex. EMBO J. 1996a;15:5370–5382. [PMC free article] [PubMed] [Google Scholar]
- Wang W, Xue Y, Zhou S, Kuo A, Cairns BR, Crabtree GR. Diversity and specialization of mammalian SWI/SNF complexes. Genes Dev. 1996b;10:2117–2130. doi: 10.1101/gad.10.17.2117. [DOI] [PubMed] [Google Scholar]
- Wang M, Gu C, Qi T, Tang W, Wang L, Wang S, et al. BAF53 interacts with p53 and functions in p53-mediated p21-gene transcription. J. Biochem. 2007;142:613–620. doi: 10.1093/jb/mvm176. [DOI] [PubMed] [Google Scholar]
- Weber V, Harata M, Hauser H, Wintersberger U. The actin-related protein Act3p of Saccharomyces cerevisiae is located in the nucleus. Mol. Biol. Cell. 1995;6:1263–1270. doi: 10.1091/mbc.6.10.1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinhold B. Epigenetics: the science of change. Environ. Health Perspect. 2006;114:A160–A167. doi: 10.1289/ehp.114-a160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams L, Fletcher JC. Stem cell regulation in the Arabidopsis shoot apical meristem. Curr. Opin. Plant Biol. 2005;8:582–586. doi: 10.1016/j.pbi.2005.09.010. [DOI] [PubMed] [Google Scholar]
- Wilson AC, Sarich VM. A molecular time scale for human evolution. Proc. Natl. Acad. Sci. USA. 1969;63:1088–1093. doi: 10.1073/pnas.63.4.1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong JJ, Hawkins NJ, Ward RL. Colorectal cancer: a model for epigenetic tumorigenesis. Gut. 2007;56:140–148. doi: 10.1136/gut.2005.088799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu WH, Alami S, Luk E, Wu CH, Sen S, Mizuguchi G, et al. Swc2 is a widely conserved H2AZ-binding module essential for ATP-dependent histone exchange. Nat. Struct. Mol. Biol. 2005;12:1064–1071. doi: 10.1038/nsmb1023. [DOI] [PubMed] [Google Scholar]
- Wu JI, Lessard J, Olave IA, Qiu Z, Ghosh A, Graef IA, et al. Regulation of dendritic development by neuron-specific chromatin remodeling complexes. Neuron. 2007;56:94–108. doi: 10.1016/j.neuron.2007.08.021. [DOI] [PubMed] [Google Scholar]
- Wu K, Zhang L, Zhou C, Yu CW, Chaikam V. HDA6 is required for jasmonate response, senescence and flowering in Arabidopsis. J. Exp. Bot. 2008;59:225–234. doi: 10.1093/jxb/erm300. [DOI] [PubMed] [Google Scholar]
- Yang X, Zaurin R, Beato M, Peterson CL. Swi3p controls SWI/SNF assembly and ATP-dependent H2A–H2B displacement. Nat. Struct. Mol. Biol. 2007;14:540–547. doi: 10.1038/nsmb1238. [DOI] [PubMed] [Google Scholar]
- Yeh JH, Spicuglia S, Kumar S, Sanchez-Sevilla A, Ferrier P, Imbert J. Control of IL-2Ralpha gene expression: structural changes within the proximal enhancer/core promoter during T-cell development. Nucleic Acids Res. 2002;30:1944–1951. doi: 10.1093/nar/30.9.1944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young DW, Pratap J, Javed A, Weiner B, Ohkawa Y, van Wijnen A, et al. SWI/SNF chromatin remodeling complex is obligatory for BMP2-induced, Runx2-dependent skeletal gene expression that controls osteoblast differentiation. J. Cell Biochem. 2005;94:720–730. doi: 10.1002/jcb.20332. [DOI] [PubMed] [Google Scholar]
- Yuan GC, Liu YJ, Dion MF, Slack MD, Wu LF, Altschuler SJ, et al. Genome-scale identification of nucleosome positions in S. cerevisiae. Science. 2005;309:626–630. doi: 10.1126/science.1112178. [DOI] [PubMed] [Google Scholar]
- Zeisig BB, Cheung N, Yeung J, So CW. Reconstructing the disease model and epigenetic networks for MLL-AF4 leukemia. Cancer Cell. 2008;14:345–347. doi: 10.1016/j.ccr.2008.10.008. [DOI] [PubMed] [Google Scholar]
- Zhang HS, Gavin M, Dahiya A, Postigo AA, Ma D, Luo RX, et al. Exit from G1 and S phase of the cell cycle is regulated by repressor complexes containing HDAC-Rb-hSWI/SNF and Rb-hSWI/SNF. Cell. 2000;101:79–89. doi: 10.1016/S0092-8674(00)80625-X. [DOI] [PubMed] [Google Scholar]
- Zhang J, Wang L, Zhuang L, Huo L, Musa S, Li S, et al. Arp11 affects dynein-dynactin interaction and is essential for dynein function in Aspergillus nidulans. Traffic. 2008;9:1073–1087. doi: 10.1111/j.1600-0854.2008.00748.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao K, Wang W, Rando OJ, Xue Y, Swiderek K, Kuo A, et al. Rapid and phosphoinositol-dependent binding of the SWI/SNF-like BAF complex to chromatin after T lymphocyte receptor signaling. Cell. 1998;95:625–636. doi: 10.1016/s0092-8674(00)81633-5. [DOI] [PubMed] [Google Scholar]
- Zhou C, Miki B, Wu K. CHB2, a member of the SWI3 gene family, is a global regulator in Arabidopsis. Plant Mol. Biol. 2003;52:1125–1134. doi: 10.1023/b:plan.0000004305.60407.8b. [DOI] [PubMed] [Google Scholar]