Abstract
Ghrelin (GHRL) and its receptor (GHSR) are involved in various bioactivities. In this study, the complete cDNA and 5′ flanking region of the duck GHRL (dGHRL) gene and a 3717 bp fragment of the duck GHSR (dGHSR) gene were obtained. A total of 19, 8, 43, and 48 SNPs identified in 2751, 1358, 3671, and 3567 bp of the chicken GHRL (cGHRL), chicken GHSR (cGHSR), dGHRL, and dGHSR genes, respectively. Both cGHRL and dGHRL were expressed predominantly in the proventriculus, whereas the highest mRNA levels of cGHSR and dGHSR were detected in the breast muscle and pituitary. Association analysis showed that C-2047G, A-2355C, and A-2220C of the cGHRL gene were significantly associated with abdominal fat weight (AFW; P = .01), crude protein content of leg muscle (CPCLM; P = .02), and CPCLM (P = .0009), respectively. C-1459T of the cGHSR gene was also significantly associated with CPCLM (P = .0004). C-729T of dGHRL and A3427T of dGHSR were both significantly associated with subcutaneous fat thickness (SFT; P = .04). It was indicated by this study that the GHRL and GHSR genes were related to fat deposition in both chicken and duck.
1. Introduction
Small synthetic molecules termed Growth Hormone Secretagogues (GHSs) act on the pituitary gland and the hypothalamus to regulate growth hormone (GH) release. The Growth Hormone Secretagogue Receptor (GHSR or GHS-R), a G protein-coupled receptor, has been cloned from human and pig and is mainly expressed in the pituitary, hypothalamus, and hippocampus [1]. In 1999, a novel 28-amino acid gut-brain peptide named Ghrelin (GHRL) was first isolated from the mammalian stomach [2]. Similar to the synthetic molecules of GHSs, GHRL proved to be an endogenous ligand for GHSR and involved in GH secretion, food intake, and energy homeostasis [2, 3]. Even though the mature GHRL varies in structure among mammals, birds, and fish, the n-octanoylation of the Ser-3 residue is highly conserved and essential for biological activity [2, 4]. It is now well known that GHRL interacts with GHSR to mediate a variety of bioactivities through the neuroendocrine pathway.
The GHRL and GHSR genes are reportedly related to obesity in human and mouse. Body mass index and body fat were shown to be negatively correlated with plasma GHRL levels [5–8]. Two polymorphisms, Arg51Gln and Leu72Met, in the human GHRL gene were identified in obese subjects, and Leu72Met was significantly associated with body mass index and fat mass in some human populations [9, 10]. In addition to these two nonsynonymous SNPs, two other variations of Gln90Leu and a frameshift mutation (2 bp deletion at codon 34) were also reported in the human GHRL gene, and the 90Leu allele frequency was significantly higher in extremely obese children and adolescents (0.063) than in normal-weight students [11]. A novel SNP, T3056C in intron 2 of the GHRL gene, was significantly associated with the acylation of GHRL and some fatness traits, including high body mass index, body weight, fat mass, and skinfold thickness [12]. Among 11 SNPs identified in the 5′ flanking region of the GHRL gene, SNP-501A > C was significantly associated with body mass index in human [13]. In mice, the peripheral administration of GHRL was found to regulate body weight, adiposity, and UCP mRNA expression [14]. It was also interesting that central administration of GHRL enhanced rat fat ingestion [15].
Until now, the research progress on the GHRL and GHSR genes was achieved in chicken. The chicken GHRL (cGHRL) gene cDNA was first cloned by Kaiya et al. [16], and it encoded a 116-amino acid GHRL precursor and a 26-amino acid mature GHRL. The complete cGHRL gene comprised five exons and four introns. The first exon did not encode any amino acids, a characteristic that was conserved in mammals [17, 18]. The cGHSR gene was composed of two exons separated by an intron and encoded a normal GHSR with seven transmembrane domains (TM1 to TM7); however, a variant GHSR without TM6 due to a 48-bp deletion was found [19]. A total of 19 and 37 SNPs were identified in the cGHRL and cGHSR genes, respectively [20], and an 8-bp indel in the 5′ untranslated region (UTR) and some SNPs of the cGHRL gene were related to chicken growth and muscle fiber traits [21–23]. A 6-bp indel of the cGHSR gene was significantly linked with fat traits [21]. The duck GHRL (dGHRL) gene cDNA was also cloned, and the encoded dGHRL was highly similar to cGHRL [24]. Nevertheless, no further studies on the dGHRL and duck GHSR (dGHSR) genes have been reported.
Fat deposition is significant in both chicken and duck. Most fat accumulates as abdominal fat in chicken but as subcutaneous fat in duck; these characteristics make the chicken and duck ideal for use in this study. RACE, genome walking, real time PCR, and association analysis were performed to characterize the GHRL and GHSR genes and reveal their effects on fat deposition in chicken and duck.
2. Materials and Methods
2.1. Animals and Sample Preparation
A total of 411 birds (219 males and 193 females) from 8 chicken populations (POP1 to POP8) and 139 birds from 9 duck populations (POP9 to POP17) were used in this study (Table 1). These samples consisted of six chicken breeds, including Xinghua (XH), White Recessive Rock (WRR), Taihe Silkies (TS), Leghorn (LH), Lingnan Yellow (LY), Huiyang Beard (HB), and an F2 full-sib hybrid population (XH&WRR) described by Fang et al. [23], as well as four duck breeds, Sanshui White (SW), Peking (PK), Partridge (PT), and Lake (LK). Either blood or live tissues were collected from these birds at 9 weeks of age for the purpose of sequence cloning, variation, mRNA quantitative expression, cell culture, and marker trait association analyses (Table 1).
Table 1.
Chicken and duck samples used in this study.
| No. | Species | Population(1) | Samples | Blood/live tissues | Purpose |
|---|---|---|---|---|---|
| POP1 | Chicken | XH | 6 (3 ♂; 3 ♀) | Blood for DNA | GHRL and GHSR gene variation |
| POP2 | Chicken | WRR | 6 (3 ♂; 3 ♀) | ||
| POP3 | Chicken | TS | 6 (3 ♂; 3 ♀) | ||
| POP4 | Chicken | LH | 6 (3 ♂; 3 ♀) | ||
| POP5 | Chicken | XH&WRR | 373 (200 ♂; 173 ♀) | Blood for DNA | Association analysis |
| POP6 | Chicken | LY | 6 (3 ♂; 3 ♀) | 18 tissues (2) | Real time PCR |
| POP7 | Chicken | HY | 6 (3 ♂; 3 ♀) | ||
| POP8 | Chicken | LY | 2 ♀ | Subcutaneous and abdominal adipose tissues | Cell culture |
| POP9 | Duck | SW | 1 ♂ | Blood for DNA and proventriculus tissue | GHRL gene sequence cloning |
| POP10 | Duck | SW | 6 (3 ♂; 3 ♀) | Blood for DNA | GHRL and GHSR gene variation |
| POP11 | Duck | PK | 6 (3 ♂; 3 ♀) | ||
| POP12 | Duck | PT | 6 (3 ♂; 3 ♀) | ||
| POP13 | Duck | LK | 6 (3 ♂; 3 ♀) | ||
| POP14 | Duck | SW | 100 (52 ♂; 48 ♀) | Blood for DNA | Association analysis |
| POP15 | Duck | SW | 6 (3 ♂; 3 ♀) | 18 tissues | Real time PCR |
| POP16 | Duck | PT | 6 (3 ♂; 3 ♀) | ||
| POP17 | Duck | SW | 2 ♀ | Subcutaneous and abdominal adipose tissues | Cell culture |
(1)XH = Xinghua chicken, WRR = White Recessive Rock, TS = Taihe Silkies, LH = Leghorn, XH&WRR = an F2 population crossed by XH and WRR, LY = Lingnan Yellow, HY = Huiyang Beard, SW = Sanshui White duck, PK = Peking duck, PT = Partridge duck, and LK = Lake duck.
(2) The 18 tissues used were pituitary, cerebrum, lung, abdominal fat, liver, testis, proventriculus, ovary, subcutaneous fat, spleen, kidney, oviduct, leg muscle, uropygial gland, hypothalamus, glandularis, small intestine, cerebellum, heart and breast muscle.
The SW commercial population (POP14), including 100 ducks (52 males and 48 females), were slaughtered at 90 days old, and body weights at hatch (g) and 90 days old (g), carcass weight (g), fat thickness under skin (mm), fat width (mm), eviscerated weight (g), abdominal fat pad weight (g), and abdominal fat pad ratio (%) were recorded. All birds were raised in separate pens and fed with commercial corn-soybean-based diets that met NRC requirements. All chickens were collected from Guangdong Wens Foodstuff Corporation Ltd. (Guangdong, China), and all ducks were provided by Guangdong Sanshui Lianke Duck Farm (Guangdong, China).
Genomic DNA was extracted from EDTA-anticoagulated blood. Total RNA was extracted from the isolated tissues using an improved Trizol isolation method (Invitrogen, California, USA) following the manufacturer's instructions. One microgram of total RNA was used to obtain cDNA by reverse transcription with use of the Rever Tra Ace Transcriptase Kit (Toyobo, Japan) and random hexamers.
2.2. Primers
Thirty-three primer pairs (PM1 to PM33) were designed and consisted of PM1 to PM6 (6) for the cGHRL gene, PM7 to PM10 (4) for the cGHSR gene, PM11 for the chicken β-actin gene, PM12 to PM24 (13) for the dGHRL gene, PM25 to PM32 (8) for the dGHSR gene, and PM33 for the duck β-actin gene (Table 2). PM12 and PM26 were both genomic primers, each of which consisting of three reverse primers and a random forward primer supplied by the Genomic Walking Kit. PM14 was a primer pair for 3′ RACE, and the remaining 30 primer pairs were gene-specific primers. The GENETOOL software (http://www.biologysoft.com/) was used to design the primers based on reported sequences of the chicken GHRL (GenBank AB075215), GHSR (GenBank AB095994), and β-actin genes (GenBank NM_205518), as well as duck GHRL (GenBank AY338466), GHSR (GenBank EU005225), β-actin (GenBank EF667345), and goose GHRL (GenBank AY338465). All primers were commercially synthesized by Shanghai Biosune Biotechnology Co. Ltd. (Shanghai, China).
Table 2.
Descriptions of the 33 primers used in this study.
| Gene | Primer | Forward/reverse | Sequence (5′ to 3′) | Annealing temp (°C) | Purpose |
|---|---|---|---|---|---|
| cGHRL | PM1 | PM1F | tgctgaaggaccgaaaacaaa | 56 | SNP identification |
| PM1R | gccttgacagatgccttagtg | ||||
| PM2 | PM2F | aagaaagctggtaactgcactag | 55 | SNP identification | |
| PM2R | ggtgggctggtggagtta | ||||
| PM3 | PM3F | tgcgttctgctactctttttcat | 63 | SNP identification | |
| PM3R | gggccaaggaggagtgtct | ||||
| PM4 | PM4F | cggcagacactcctcctt | 57 | SNP identification | |
| PM4R | gccatccttccaactgtgtatat | ||||
| PM5 | PM5F | tatgcgttctgctactcttt | 58 | Genotyping by sequencing | |
| PM5R | tggaagcgatcactatacc | ||||
| PM6 | PM6F | catacagcaacaaaaggatac | 63 | Real time PCR | |
| PM6R | tgtggttgtccttcagct | ||||
| cGHSR | PM7 | PM7F | gtgggtcagggcatcaaactc | 58 | SNP identification and PCR-RFLP |
| PM7R | tcgagggctgctgaattttatg | ||||
| PM8 | PM8F | ggctcttcctttttggtttgtct | 60 | SNP identification and PCR-RFLP | |
| PM8R | tcgccctctctctgattcacct | ||||
| PM9 | PM9F | tgatgccactggtctgagaat | 58 | Genotyping by PCR-RFLP | |
| PM9R | tatccagctgcccatgtaaat | ||||
| PM10 | PM10F | tgggcgtcgagcatgagaat | 63 | Real time PCR | |
| PM10R | ccacgactagcatcttcacag | ||||
| Chicken β-actin | PM11 | PM11F | ccccaaagccaacagagaga | 63 | Real time PCR |
| PM11R | ggtggtgaagctgtagcctctc | ||||
| dGHRL | PM12 | PM12F | Genomic Walking Kit | According to the kit | Genomic Walking |
| PM12R1 | ttccatcagttctagacgagt | ||||
| PM12R2 | tgtttgtccatactcttgatact | ||||
| PM12R3 | gcccctgctttatctgtatc | ||||
| PM13 | PM13F | ccgcctggtgaagaaaac | 56 | RT-PCR | |
| PM13R | aagcctacacatccacctgcaat | ||||
| PM14 | PM14F | gaggcaagctgaaggacaaccaca | 54 | 3′ RACE | |
| PM14R | 3′ RACE Kit | ||||
| PM15 | PM15F | gctggttttcccgtgtaattc | 56 | RT-PCR | |
| PM15R | tgtttgtccatactcttgatact | ||||
| PM16 | PM16F | gctggttttcccgtgtaattc | 54 | Intron 1 amplification | |
| PM16R | tgtttgtccatactcttgatact | ||||
| PM17 | PM17F | tggtttggctggctctagt | 56 | Intron 2 amplification | |
| PM7R | PM12R3 | ||||
| PM18 | PM18F | aaagcaggggcagaagat | 57 | Intron 3 amplification | |
| PM18R | PM12R2 | ||||
| PM19 | PM19F | agggtcctggtccaaaaat | 54 | Intron 4 amplification | |
| PM19R | PM12R1 | ||||
| PM20 | PM20F | cgcatggtagccttcacacac | 56 | SNP identification and PCR-RFLP | |
| PM20R | agccgatgggttagcagagag | ||||
| PM21 | PM21F | tcggctccatcagttgcagttat | 56 | SNP identification | |
| PM21R | cggcgatgtatttttgctgttg | ||||
| PM22 | PM22F | tccccacgacagagtagtttgag | 56 | SNP identification and PCR-RFLP | |
| PM22R | gccttcccctgcttcctaa | ||||
| PM23 | PM23F | atagcagcattttagaagtga | 54 | SNP identification | |
| PM23R | ttgcctcagcttttcact | ||||
| PM24 | PM24F | cgtgtaattcctctctgctaa | 63 | Real time PCR | |
| PM24R | cgatgtatttttgctgttgtt | ||||
| dGHSR | PM25 | PM25F | cccctgagcaccaacgagt | 56 | Intron 1 amplification |
| PM25R | ggcaaccagcagagtatga | ||||
| PM26 | PM26F | Genomic Walking Kit | According to the kit | Genomic Walking | |
| PM26R1 | cggaccgatgttctttctcttt | ||||
| PM26R2 | gacgggcaggaaaaagaagac | ||||
| PM26R3 | ggcccagaggatgaggatga | ||||
| PM27 | PM27F | tggcgttctccgacctgctcat | 57 | SNP identification | |
| PM27R | acacagaccctcagaaacac | ||||
| PM28 | PM28F | cgggtggcttaaataacaacag | 56 | SNP identification | |
| PM28R | tgccctttttcctcagtttctc | ||||
| PM29 | PM29F | cggaccataattaaacacctgaa | 56 | SNP identification | |
| PM29R | caggtagaagaggacaaaggaca | ||||
| PM30 | PM30F | tggcgttctccgacctgctcat | 56 | Genotyping by PCR-RFLP | |
| PM30R | acacagaccctcagaaacac | ||||
| PM31 | PM31F | gcaggttgtttagatatggct | 56 | Genotyping by PCR-RFLP | |
| PM31R | cgtgaaaaggcaaccagcagag | ||||
| PM32 | PM32F | cccctgagcaccaacgagt | 63 | Real time PCR | |
| PM32R | ccaccacaactaacattttcaca | ||||
| Duck β-actin | PM33 | PM33F | acgccaacacggtgctg | 63 | Real time PCR |
| PM33R | gggtccggattcatcatactc | ||||
2.3. PCR, PCR-RFLP, and Sequencing
PCR was performed in 25 μL reactions containing 50 ng chicken genomic DNA, 1 × PCR buffer, 12.5 pmol primers, 100 μM of each dNTP, 1.5 mM MgCl2, and 1.0 U Taq DNA polymerase (Sangon Biological Engineering Technology Company, Shanghai, China). PCR was run in a Mastercycler gradient (M. J. Research Co. Ltd., USA) with the following procedure: 3 minutes at 94°C, followed by 35 cycles of 30 seconds at 94°C, 45 seconds at the annealing temperature (58–62°C) and 1 minute at 72°C, and a final extension of 5 minutes at 72°C. PCR products were checked for size and quality on 1% agarose gels.
The PCR product of PM5 was sequenced for genotyping of the A-2220C, A-2246G, A-2355C, C-2399del, C-2030T, and C-2047G polymorphisms in the cGHRL gene. The PCR product of the PM7 reaction was digested with Tat I with the PCR-RFLP method for genotyping C-1896A in the cGHSR gene. The PCR product of the PM8 reaction was digested by Hin1 II with the PCR-RFLP method for genotyping the C-1459T SNP in the cGHSR gene. The PCR product of the PM9 reaction was digested with Tail I with the PCR-RFLP method for genotyping the A-1022C SNP in the cGHSR gene. The PCR product of the PM20 reaction was digested with Csp6 I with the PCR-RFLP method for genotyping the C-729T SNP in the dGHRL gene. The PCR product of the PM22 reaction was digested with Tail I with the PCR-RFLP method for genotyping the T985C SNP in the dGHRL gene. The PCR product of the PM30 reaction was digested with Csp6 I with the PCR-RFLP method for genotyping the T404C SNP in the dGHSR gene. The PCR product of the PM31 reaction was digested with Csp6 I with the PCR-RFLP method for genotyping the A3427G SNP in the dGHSR gene. Digestion products were detected by agarose gel (1.5%) electrophoresis, and genotypes were determined by specific profiles (Supplementary Figure 1 in Supplementary Material available on-line at doi: 10.1155/2009/567120).
2.4. RT-PCR and 3′ RACE
Total RNA was extracted from the proventriculus from an SW duck by the TRIzol reagent (Invitrogen, Carlsbad, CA) following the manufacturer's instructions. A proventriculus cDNA library was prepared using the SMART RACE cDNA amplification kit (Clontech Laboratories, Mountain View, CA). Primer-directed RT-PCR and 3′ rapid amplification of cDNA ends (RACE) were used to generate duck GHRL cDNAs, amplified using primers P1 to P3 (Table 2). A cDNA library was prepared using the SMART RACE cDNA amplification kit (Clontech Laboratories, Mountain View, CA). The 3′ ends of the cDNAs were amplified with a specific forward primer and reverse random primers from the SMART RACE cDNA amplification kit following the manufacturer's instructions.
2.5. Genome Walking
To obtain the 5′ flanking region of the duck GHRL and GHSR genes, genome walking was performed using the Genome Walking Kit (TakaRa, Japan). The duck genomic DNA was purified using the Genomic DNA Purification Kit (U-gene, China) and was then used for genome walking. The genome walking procedure included three reactions: the first PCR reaction used the outer adaptor primer AP1 from the kit and an outer gene-specific primer PM12R1 (or PM26R1); the second reaction used primers AP2 and PM26R2; the third reaction used primers AP3 and PM26R3. Each reaction was performed step-by-step following the kit instructions. PCR products of the third reaction were purified using the Gel Extraction Kit (U-gene, China), cloned into the pGEM-T easy vector (Promega, USA), and sequenced by Shanghai Biosune Biotechnology Co. Ltd. (Shanghai, China).
2.6. Quantitative Real Time PCR
The mRNA levels of GHRL and GHSR in 18 tissues of chickens and ducks were determined by quantitative real time PCR with SYBR green dye using β-actin as an internal positive control (IPC). The 15 μL PCR mixture contained 1 μL of cDNA, 7.5 μM Super Mix (Toyobo, Japan), 10 μM of each primer (PM6, PM10, PM11, PM24, PM32, and PM33), and 6.0 μL DEPC H2O. Real time PCR was run on an ABI 7500 (Applied Biosystems, Foster City, CA, USA) using the following program: initial denaturation at 95°C for 4 minutes, then 40 cycles of 94°C for 30 seconds, 63°C for 30 seconds, and 72°C for 45 seconds. Each sample was repeated three times and the average mRNA level was obtained for further analysis. Two random PCR products of PM6, PM10, PM11, PM24, PM32, and PM33 were sequenced by Shanghai Biosune Biotechnology Co. Ltd. (Shanghai, China) to confirm that specific fragments of the cGHRL, cGHSR, chicken β-actin, dGHRL, dGHSR, and duck β-actin genes were obtained. Quantitative values were obtained from the Ct values, which were the inverse ratio relative to the starting PCR product. The relative quantification was obtained by 2−ΔCt in which ΔCt = Cttarget gene − CtIPC. The data are expressed as the mean ± S.E. Statistics were analyzed by SAS Students t-test that measures analysis of variance with a significance level of 0.05.
2.7. Statistics Methods
2.7.1. Sequence BLAST, Prediction of Transcription Factors, and Haplotypes
The homology of GHRL among human, mouse, chicken, turkey, emu, goose, and duck was studied. Identity percentages were obtained from amino acid BLAST with the DNASTAR software (http://www.dnastar.com/). The obtained sequences of GHRL and GHSR were aligned with the Mega 3.1 software (http://www.megasoftware.net/). The 798-bp 5′ flanking region of the dGHRL gene obtained in this study was used to predict the potential binding sites for transcription factors by an online service (http://www.fruitfly.org/cgi-bin/seq_tools/promoter.pl). In the chicken F2 population (XH&WRR) and the duck commercial population (SW), haplotypes were inferred based on the genotype data with the PHASE 2.1 software (http://en.wikipedia.org/wiki/Phase) [25].
2.7.2. Nucleotide Diversity (θ)
To evaluate the nucleotide diversity, the normalized numbers of SNPs (θ) were calculated by dividing K, the number of observed nucleotide changes, by the total sequence length in base pairs (L) and correcting for the sample size (n) as described in [26], using the following formula:
| (1) |
2.7.3. Marker-Trait Association Analysis
The effects of 13 SNPs identified in this study on fatty traits were estimated by marker-trait association analysis. These SNPs included six in the cGHRL gene (C-2030T, C-2047G, A-2220C, A-2246G, A-2355C, and C-2399del), three in the cGHSR gene (C-1896A, C-1459T, and A-1022C), two in the dGHRL gene (C-729T and T985C), and two in the dGHSR gene (T404C and A3427G). Marker-trait association analysis was performed using the SAS GLM procedure (SAS Institute, 1996) and the genetic effects were analyzed by a mixed procedure according to the following model:
| (2) |
where Y is an observation on the trait, μ is the overall population mean, G is the fixed effect of the genotype, D is the random effect of dam, H is the fixed effect of hatch, S is the fixed effect of sex (male or female), and e is the residual random error.
3. Results
3.1. Cloning the Full-Length cDNA of the dGHRL Gene
The full-length cDNA of the dGHRL gene (835 bp) was amplified using primers PM13 to PM15. It comprised 115 bp of the 5′ untranslated region (5′ UTR), the 351 bp open reading frame (ORF) encoding a 116-amino acid (aa) peptide, and 369 bp of the 3′ UTR (Figure 1) (GenBank EF613551). The SW duck GHRL precursor (116 aa) included a signal peptide (23 aa), the mature GHRL (26 aa), and a long C-terminal peptide (67 aa). Compared with the other reported mallard dGHRL precursor sequence (NCBI accession number: AY338466), there were some different amino acids (W16R, G21A, A53T, T59A, A64T, and G71S). The putative SW dGHRL precursor showed identities of 88.8%, 72.4%, 69.8%, 71.6%, 37.9%, and 37.1% with its counterparts of goose, chicken, turkey, emu, mouse, and human, respectively (Figure 2). As far as the mature peptides are concerned, however, dGHRL had much higher homology with the other six species (92.3%, 65.4%, 65.4%, 69.2%, 53.8%, and 50.0% with goose, chicken, turkey, emu, mouse, and human, resp.) (Figure 2).
Figure 1.
cDNA and deduced amino acid sequence of the dGHRL gene. The arrowheads indicate the locations of introns. Polyadenylation signal is boxed. The asterisk (*) represents the putative stop codon (TGA). The nucleotide sequence is deposited in GenBank (EF613551).
Figure 2.
Alignment of GHRL sequences among 7 species. The mature peptides were boxed.
3.2. Genomic Organization of the dGHRL and dGHSR Genes
The sequence ensemble generated with the PM12 to PM19 products gave rise to a 3670 bp genomic fragment of the dGHRL gene, which included 798 bp of the 5′ flanking region, 883 bp of five exons (134, 137, 114, 109, and 389 bp for exon 1 to exon 5) and 1989 bp of four introns (168, 469, 458, and 894 bp for intron 1 to intron 4). Like other species, the four introns of the dGHRL gene all followed the “GT-AG” rule, and moreover, exon 1 did not encode any amino acids (Supplementary Figure 2). The obtained dGHRL genomic sequences were submitted to the NCBI database (GenBank EF613552).
The 3717 bp sequence of the dGHSR gene was deduced by genomic walking (PM26), PCR amplification (PM25) and sequence ensemble with reported sequences (Genbank EU005225). It comprised two incomplete exons (591 and 148 bp for exon 1 and exon 2) and one intron (2978 bp) that followed the “GT-AG” intron rule (Supplementary Figure 2). The obtained dGHSR gene sequences were also submitted to the NCBI database (GenBank FJ194548).
3.3. Online Prediction of Transcription Factor Binding Sites in the dGHRL Gene
Online prediction of transcription factor binding sites in the 5′ flanking region of the dGHRL gene showed that a typical TATA box was present in 23 bp upstream of the transcription start site, and potential binding sites for some transcription factors were found, including one AML-1a site (runt-factor AML-1), two cap 1 sites (cap signal for transcription initiation), two CdxA 1 sites, three GATA-1 sites (GATA-binding factor 1), as well as one Oct-1 site (octamer factor 1) (Figure 3).
Figure 3.
Online prediction of transcription factor binding sites and the TATA box in the 5′ flanking region of the dGHRL gene.
3.4. SNPs and Nucleotide Diversity
A total of 19 SNPs (10 transitions, 8 transversions, and 1 indel) and 1 polyA polymorphism were identified in 2751 bp of the 5′ UTR region of the cGHRL gene, while 8 SNPs (5 transitions and 3 transversions) were identified in 1358 bp of the 5′ UTR of the cGHSR gene (Table 3). Forty-three SNPs (27 transitions, 15 transversions, and 1 triallelic SNP), 1 SSR, and 1 9-bp indel were identified in 3671 bp of the dGHRL gene, while 48 SNPs (35 transitions and 13 transversions) and 5 indels were found in 3567 bp of the dGHSR gene (Table 3). Eleven SNPs were located in the coding regions of the dGHRL gene, but only four SNPs were nonsynonymous and led to amino acid changes in the C-terminal extension peptide. For the dGHSR gene, however, no nonsynonymous SNPs were found within five SNPs in the coding regions (Table 3). A triallelic SNP (GA/C at nt 1774) was observed in the coding region of the dGHRL gene, which was related to three different amino acids (A/P/T at aa 59) (Table 3).
Table 3.
SNPs identified in the cGHRL, cGHSR, dGHRL, and dGHSR genes.
| Gene | No. | SNP1 | Region | Note (2, 3,4) | No. | SNP | Region | Note (2, 3,4) |
|---|---|---|---|---|---|---|---|---|
| cGHRL | 1 | C-495T | 5′ flanking | 19369777(2)Dde I (3) | 11 | A-2220C | 5′ flanking | 19371502 |
| 2 | C-517T | 5′ flanking | 19369799 | 12 | A-2246G | 5′ flanking | 19371528 Tas I | |
| 3 | C-873G | 5′ flanking | 19370155 | 13 | A-2355C | 5′ flanking | 19371637 | |
| 4 | T-1038C | 5′ flanking | 19370320 | 14 | C-2399del | 5′ flanking | 19371681 | |
| 5 | C-1253T | 5′ flanking | 19370535 Hin1 II | 15 | G-2861T | 5′ flanking | 19372143 | |
| 6 | T-1562G | 5′ flanking | 19370844 | 16 | G-3264C | 5′ flanking | 19372546 | |
| 7 | A-1589G | 5′ flanking | 19370871 | 17 | C-3290T | 5′ flanking | 19372572 Tai I | |
| 8 | G-1950A | 5′ flanking | 19371232 | 18 | G-3321T | 5′ flanking | 19372603 | |
| 9 | C-2030T | 5′ flanking | 19371312 | 19 | T-3324C | 5′ flanking | 19372606 | |
| 10 | C-2046G | 5′ flanking | 19371329 | |||||
| cGHSR | 1 | A-1907G | 5′ flanking | 18787994 | 5 | T-1288C | 5′ flanking | 18789055 |
| 2 | C-1896A | 5′ flanking | 18788005 Tat I | 6 | A-1022C | 5′ flanking | 18789321 Tail I | |
| 3 | C-1608T | 5′ flanking | 18788293 | 7 | T-892C | 5′ flanking | 18789762 Bsa JI | |
| 4 | C-1459T | 5′ flanking | 18788442 Hin1 II | 8 | G-834C | 5′ flanking | 18789820 Hin6 I | |
| dGHRL | 1 | G221A | 5′ flanking | 23 | T1128C | Exon 2 | ||
| 2 | T256C | 5′ flanking | 24 | T1145C | Exon 2 | G5 Syn(4), Csp6 I | ||
| 3 | T263C | 5′ flanking | 25 | C1179G | Exon 2 | R17G, Msp I | ||
| 4 | G273T | 5′ flanking | 26 | A1244T | Intron 2 | |||
| 5 | A276C | 5′ flanking | 27 | T1278C | Intron 2 | |||
| 6 | A303G | 5′ flanking | 28 | G1390A | Intron 2 | |||
| 7 | A342C | 5′ flanking | 29 | A1756G | Exon 3 | T53A, Hin 6 I | ||
| 8 | T401C | 5′ flanking | Csp6 I | 30 | G1761A | Exon 3 | E54 Syn | |
| 9 | C405T | 5′ flanking | 31 | G1774A/C | Exon 3 | A59P/T | ||
| 10 | A532C | 5′ flanking | 32 | A1789G | Exon 3 | T64A | ||
| 11 | A628G | 5′ flanking | 33 | C1803T | Exon 3 | N68 Syn | ||
| 12 | C638T | 5′ flanking | 34 | A1807G | Exon 3 | G70S | ||
| 13 | A645G | 5′ flanking | 35 | A1943C | Intron 3 | HpyCH4IV | ||
| 14 | C686G | 5′ flanking | 36 | C2070T | Intron 3 | |||
| 15 | C691A | 5′ flanking | 37 | T2115C | Intron 3 | TailI | ||
| 16 | A736T | 5′ flanking | 38 | C2179T | Intron 3 | |||
| 17 | C778A | 5′ flanking | 39 | A2324G | Exon 4 | E89 Syn | ||
| 18 | C1028A | Intron 1 | 40 | G2345A | Exon 4 | T96 Syn | ||
| 19 | A1070G | Intron 1 | 41 | A2351G | Exon 4 | E98 Syn | ||
| 20 | A1108G | Exon 2 | 42 | G2409C | Intron 4 | |||
| 21 | A1118G | Exon 2 | 43 | T2509C | Intron 4 | |||
| 22 | A1117G | Exon 2 | ||||||
| dGHSR | 1 | C320T | Exon 1 | A Syn, Msp I(4) | 25 | C2815A | Intron | |
| 2 | T359C | Exon 1 | N Syn | 26 | C2843T | Intron | ||
| 3 | G398C | Exon 1 | T Syn, Bsa JI | 27 | C2861T | Intron | ||
| 4 | T404C | Exon 1 | Y Syn, Csp6 I | 28 | G2863C | Intron | ||
| 5 | G548A | Exon 1 | T Syn | 29 | C2883T | Intron | ||
| 6 | C779T | Intron | 30 | T2906C | Intron | |||
| 7 | T1364A | Intron | 31 | C2939A | Intron | |||
| 8 | C1437T | Intron | 32 | T2958A | Intron | |||
| 9 | T1456C | Intron | 33 | G2993A | Intron | |||
| 10 | T1469C | Intron | 34 | G3001A | Intron | |||
| dGHSR | 11 | C1478T | Intron | 35 | A3304G | Intron | ||
| 12 | A2393T | Intron | 36 | A3037G | Intron | |||
| 13 | C2430T | Intron | 37 | T3050G | Intron | |||
| 14 | A2431G | Intron | 38 | G3080A | Intron | |||
| 15 | T2578C | Intron | 39 | T3208A | Intron | Taa I | ||
| 16 | T2591A | Intron | Tail I | 40 | C3329T | Intron | Taa I | |
| 17 | T2619A | Intron | 41 | T3340A | Intron | |||
| 18 | C2657T | Intron | 42 | G3362A | Intron | |||
| 19 | C2681T | Intron | 43 | G3363A | Intron | |||
| 20 | A2682G | Intron | 44 | T3380C | Intron | |||
| 21 | G2746A | Intron | 45 | C3424T | Intron | Csp6 I | ||
| 22 | T2792C | Intron | 46 | A3427G | Intron | |||
| 23 | G2794T | Intron | 47 | A3451G | Intron | |||
| 24 | G2803A | Intron | 48 | C3543T | Intron | |||
(1)The first nucleotide of start codon was marked as +1, and the next upstream nucleotide was −1. SNP position was determined based on reported sequences of GenBank EF613552 (dGHRL gene) and FJ194548 (dGHSR gene), respectively. (2)SNP position was determined according to the released chicken genomic sequence (http://genome.ucsc.edu/). (3)Restriction enzyme was used for PCR-RFLP. (4)SNP caused a codon and amino acid change.
The average bps per SNP were 145, 170, 86, and 75 for the cGHRL, cGHSR, dGHRL, and dGHSR genes, respectively. The nucleotide diversities (θ) corrected for sample size were 1.83 × 10−3, 1.56 × 10−3, 3.10 × 10−3, and 3.56 × 10−3 for the cGHRL, cGHSR, dGHRL, and dGHSR genes, respectively.
3.5. Tissue-Specific Expression of the cGHRL, cGHSR, dGHRL, and dGHSR Genes
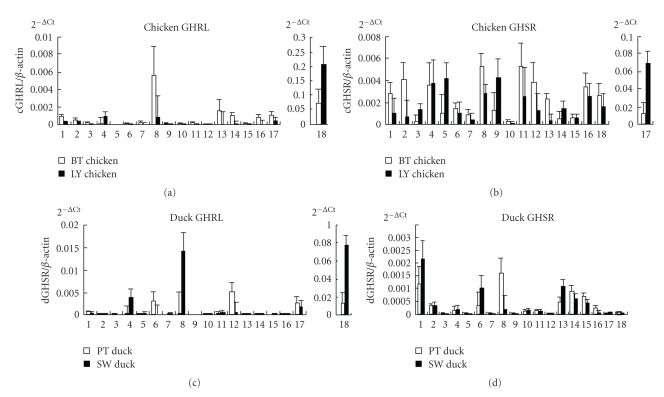
The highest mRNA level of the cGHRL gene was found in the proventriculus and then the pituitary, cerebrum, abdominal fat, subcutaneous fat, hypothalamus, small intestine, heart, and breast muscle. The highest mRNA level of the dGHRL gene was also found in the proventriculus and then the abdominal fat, testicles, subcutaneous fat, uropygial gland, and breast muscle (Figure 4; Supplementary Table 1).
Figure 4.
Relative mRNA expression levels of the GHRL and GHSR genes in adult chicken and duck tissues. The vertical axis indicates the 2−Ct value (mean ± S.E.M., n = 6). Pituitary, Cerebrum, Lung, Abdominal fat, Liver, Testis, Gizzard, Subcutaneous fat, Spleen, Kidney, Leg muscle, Uropygial gland, Hypothalamus, Small intestine, Cerebellum, Heart, Breast muscle, and Proventriculus are considered.
The highest mRNA level of the cGHSR gene was detected in the breast muscle and then the subcutaneous fat, leg muscle, abdominal fat, heart, spleen, liver, uropygial gland, cerebrum, proventriculus, pituitary, and testicles. The highest mRNA level of the dGHSR gene was detected in the pituitary and then the subcutaneous fat, hypothalamus, small intestine, testis, cerebellum, and cerebrum, whereas little mRNA was found in the other tissues (Figure 4; Supplementary Table 1).
When the HB and LY chicken breeds were compared, significant differences in the GHRL mRNA levels were found in the proventriculus, subcutaneous fat, lung, and kidney (P < .05), whereas significant differences in the GHSR mRNA levels were found in the hypothalamus and breast muscle (P < .05). When the PT duck and SW duck breeds were compared, significant differences in the GHRL mRNA levels were found in the proventriculus and subcutaneous fat (P < .05), whereas no significant differences in the GHSR mRNA levels were found in any tissues (P > .05) (Figure 4; Supplementary Table 1).
3.6. Associations of SNPs with Fatty Traits in Chicken and Duck
For the cGHRL gene, C-2047G and A-2355C were found to be significantly associated with abdominal fat weight (AFW; P = .01 < .05) and crude protein content of leg muscle (CPCLM; P = .02 < .05), and A-2220C was significantly associated with CPCLM (P = .0009 < .01). In addition, haplotypes across six SNPs (C-2030T, C-2047G, A-2220C, A-2246G, A-2355C, and C-2399del) were significantly associated with AFW (P = .04 < .05), abdominal fat ratio (AFR; P = .04 < .05), and CPCLM (P < .0001). As far as the three genotypes of each locus were concerned, CC of C-2047G, CC of A-2220C, and AA of A-2355C had the highest values of association with AFW and CPCLM, indicating that the advantageous alleles for fat deposition were C-2047, A-2220, and A-2355 (Table 4). For the cGHSR gene, C-1459T was shown to be significantly associated with CPCLM (P = .0004 < .01). Individuals with the CC genotype had the highest CPCLM association, indicating that C rather than T was the dominant allele for CPCLM (Table 4).
Table 4.
Effects of SNPs on fatty traits in chicken and duck.
| Gene | SNP | Traits | P-value | Genotype and traits | ||
|---|---|---|---|---|---|---|
| cGHRL | C-2047G | AFW | .01 | 29.01 ± 1.089a (C/C) | 25.57 ± 3.46b (C/G) | 23.30 ± 5.676b (G/G) |
| A-2220C | CPCLM | .0009 | 20.71 ± 0.1892a (A/A) | 20.03 ± 0.319a (A/C) | 22.47 ± 0.7554B (C/C) | |
| A-2355C | CPCLM | .0203 | 21.83 ± 0.4864a (A/A) | 20.75 ± 0.228b (A/C) | 20.58 ± 0.1889b (C/C) | |
| cGHSR | C-1456T | CPCLM | .0004 | 67.77 ± 4.598a (C/C) | 66.88 ± 1.097a (C/T) | 63.40 ± 0.7162B (T/T) |
| dGHRL | C-729T | SFT | .0408 | 1.52 ± 0.22a (T/T) | 2.44 ± 0.32b (C/T) | 2.02 ± 0.48ab (C/C) |
| dGHSR | C-3427T | SFT | .0407 | 0.80 ± 0.77a (C/C) | 1.79 ± 0.30b (C/T) | 1.93 ± 0.23b (T/T) |
Note that only significant associations are shown in this table. AFW = abdominal fat weight; CPCLM = crude protein content of leg muscle; SFT = subcutaneous fat thickness. Values within a row without a common superscript letter differ and values with superscript letters differ significantly (P < .05 and P < .01 each). Genotypes of a and b mean they differed significantly (P < .05) and genotypes of a and B mean they differed great significantly (P < .01).
In duck, C-729T and haplotypes of C-729T and T985C of the dGHRL gene were both significantly associated with subcutaneous fat thickness (SFT; P = .04 and .05, resp.). Individuals with the CT genotype of C-729T had the highest SFT. For the dGHSR gene, A3427G and haplotypes of T404C and A3427G were both significantly associated with SFT (P = .04 and .05, resp.). Individuals with the TT genotype of C3427T had the highest SFT, indicating that T rather than C was the dominant allele for fat deposition (Table 4).
4. Discussion
4.1. Molecular Characterization of the GHRL and GHSR Genes
The genomic organization of the GHRL and GHSR genes was conserved in chicken and duck (Supplementary Figure 2). The cDNAs of the cGHRL and cGHSR genes were reported by Kaiya et al. [16] and Tanaka et al. [19], respectively. The genome sequences of the cGHRL and cGHSR genes were also previously known. The reported cGHRL and cGHSR genes were 2706 and 4121 bp long, respectively, and comprised five and two exons each [17, 19, 27]. From the cloning and sequencing performed in this study, the dGHRL cDNA (GenBank EF613551) had a 351-bp ORF that encodes a 116-aa precursor. The predicted mature dGHRL was a 26-aa peptide, and only two amino acids were different from cGHRL. The first seven amino acids, including a serine residue at position 3 (site of n-octanoylation), of the mature dGHRL are identical to those of chicken, turkey, goose, and emu (Figure 2) [17, 24].
The obtained dGHRL gene sequence (GenBank EF613552) also comprised five exons and four introns, and the first exon did not encode any amino acids. This characteristic was conserved in chicken, turkey, human, and rodent [16–18, 28, 29]. The obtained 3717 bp sequence of the dGHSR gene (GenBank FJ194548) consisted of two exons and one intron and encoded a 347-aa mature receptor protein. All introns of the cGHRL, cGHSR, dGHRL, and dGHSR genes followed the GT-AG rule, which is conserved in eukaryotic genes.
Online prediction indicated that a typical TATA box and a number of transcriptional factor binding sites (AML-1a, cap 1, CdxA 1, GATA-1, and Oct-1) were present in the 5′ flanking region of the dGHRL gene. These transcriptional factor binding sites were also found in the GHRL gene promoter regions of human and chicken [29, 30], and a “TATATAA” sequence was reported to be located at the cGHRL gene promoter [18]. GATA regulatory motifs were first found in the promoters of globulin and other erythroid-specific genes [31]. AML-1 is also a key regulator of hematopoiesis and plays an important role in development of all hematopoietic lineages [32]. It is possible that dGHRL gene expression in hematopoietic cells would be regulated by GATA-1 and AML-1a. The transcription factor cap-1 is a key factor of the cap-dependent translation initiation mechanism. Oct-1 is one of the transcription factors with a POU homeodomain and is expressed in most tissues including the brain [33]. Moreover, Oct-1 plays very important roles in the transcriptional regulation of gonadotropin-releasing hormone and aldolase C gene expression in the brain [34, 35]. CdxA is one of the vertebrate caudal proteins that play important roles in establishment of the body plan during early development [36]. These transcription factor binding sites in the putative promoter region might affect dGHRL gene expression.
The GHRL and GHSR genes were found to be more variable in poultry than in mammals. As indicated by this study, the SNP densities were 145, 170, 86, and 75 bp per SNP for the cGHRL (2751 bp/19 SNP), cGHSR (1358/8), dGHRL (3671/43), and dGHSR (3567/48) genes, respectively (Table 3). In humans, a total of 11 SNPs were found in 1657 bp of GHRL gene with an SNP density of 1 SNP/151 bp [13]. The reported density for the human GHSR gene (5 SNP of 1101 bp) was 220 bp per SNP [37]. Based on the dbSNP data (http://www.ncbi.nlm.nih.gov), a total of 8 and 32 SNPs were reported in 2922 and 6494 bp of the mouse GHRL and GHSR genes, with estimated densities of 365 and 203 bp per SNP, respectively. It was indicated that the higher variations of the GHRL and GHSR genes were found in chicken and duck compared with those of human and mouse. Moreover, as far as these two species were concerned, higher SNP densities were found in duck compared to chicken, as the nucleotide diversity of the dGHRL (3.10 × 10−3) and dGHSR (3.56 × 10−3) genes were much higher than those of cGHRL (1.83 × 10−3) and cGHSR (1.56 × 10−3). Our recent study on the chicken and duck THRSP α genes also showed higher variations in the duck genome (data not shown).
4.2. Effects of the GHRL and GHSR Genes on Fat Deposition
As indicated in this study, the GHRL and GHSR genes were associated with fatness traits in chicken and duck, which is consistent with some previous studies in humans. Many studies showed associations of the GHRL gene with adiposity [10–12, 38] and Type 2 diabetes [39] in humans. Additionally, some polymorphisms in the GHSR gene were found to be associated with obesity, bulimia nervosa, and pharmacological abnormalities in humans [40–43]. In chickens, the association of cGHRL polymorphisms with growth traits has been reported [22, 23]. Moreover, significant association of the cGHSR gene with several fatness traits was reported by a recent study [21]. In this study, the GHRL and GHSR genes were found to be associated with several fatness traits like AFW, CPCLM, and SFT in both chicken and duck.
Analysis of mRNA levels also showed the effects of the GHRL and GHSR genes on fat deposition in poultry. Even though the GHRL gene was predominantly expressed in the proventriculus, high mRNA levels of the GHRL and GHSR genes were detected in subcutaneous fat and abdominal fat in both chicken and duck (Figure 4). This suggests that both GHRL and GHSR genes may be involved in fat mobilization in poultry. Moreover, much higher GHRL mRNA levels were found in LY (2.07 ± 0.06) rather than HB (0.07 ± 0.05) chickens; higher levels were found in SW (0.78 ± 0.01) compared to those in PT (0.13 ± 0.01) ducks (Supplementary Table 1). SW ducks and LY chickens are two fast-growing commercial breeds with higher fat contents compared with those of the other two native breeds of PT ducks and HB chickens. The higher GHRL mRNA in higher-fat breeds suggested that GHRL was probably advantageous for fat deposition in poultry.
5. Conclusion
From these investigations, the variations and expression patterns of the cGHRL, cGHSR, dGHRL, and dGHSR genes were characterized, and some SNPs of these genes were related to fatty traits.
Supplementary Material
Supplementary Figure 1 indicated that genotyps of SNPs were determined based on different profiles by either directly sequencing or PCR‐RFLP technology. Supplementary Figure 2 indicated the genomic structure of the cGHRL, cGHSR, dGHRL, and dGHSR genes. Supplementary Table 1 showed the relative mRNA expression levels of the cGHRL, cGHSR, dGHRL and dGHSR genes in 18 tissues of adult birds.
Acknowledgments
This work was funded by projects under the Major State Basic Research Development (973) Program of China (2006CB102107) and the National Natural Science Foundation of China (30600429). Q. Nie and M. Fang contributed this work equally.
References
- 1.Howard AD, Feighner SD, Cully DF, et al. A receptor in pituitary and hypothalamus that functions in growth hormone release. Science. 1996;273(5277):974–977. doi: 10.1126/science.273.5277.974. [DOI] [PubMed] [Google Scholar]
- 2.Kojima M, Hosoda H, Date Y, Nakazato M, Matsuo H, Kangawa K. Ghrelin is a growth-hormone-releasing acylated peptide from stomach. Nature. 1999;402(6762):656–660. doi: 10.1038/45230. [DOI] [PubMed] [Google Scholar]
- 3.Date Y, Kojima M, Hosoda H, et al. Ghrelin, a novel growth hormone-releasing acylated peptide, is synthesized in a distinct endocrine cell type in the gastrointestinal tracts of rats and humans. Endocrinology. 2000;141(11):4255–4261. doi: 10.1210/endo.141.11.7757. [DOI] [PubMed] [Google Scholar]
- 4.Matsumoto M, Hosoda H, Kitajima Y, et al. Structure-activity relationship of ghrelin: pharmacological study of ghrelin peptides. Biochemical and Biophysical Research Communications. 2001;287(1):142–146. doi: 10.1006/bbrc.2001.5553. [DOI] [PubMed] [Google Scholar]
- 5.Monti V, Carlson JJ, Hunt SC, Adams TD. Relationship of ghrelin and leptin hormones with body mass index and waist circumference in a random sample of adults. Journal of the American Dietetic Association. 2006;106(6):822–828. doi: 10.1016/j.jada.2006.03.015. [DOI] [PubMed] [Google Scholar]
- 6.Stylianou C, Galli-Tsinopoulou A, Farmakiotis D, et al. Ghrelin and leptin levels in obese adolescents. Relationship with body fat and insulin resistance. Hormones. 2007;6(4):295–303. doi: 10.14310/horm.2002.1111025. [DOI] [PubMed] [Google Scholar]
- 7.Lindeman JHN, Pijl H, van Dielen FMH, Lentjes EGWM, van Leuven C, Kooistra T. Ghrelin and the hyposomatotropism of obesity. Obesity Research. 2002;10(11):1161–1166. doi: 10.1038/oby.2002.157. [DOI] [PubMed] [Google Scholar]
- 8.Foster CM, Barkan A, Kasa-Vubu JZ, Jaffe C. Ghrelin concentrations reflect body mass index rather than feeding status in obese girls. Pediatric Research. 2007;62(6):731–734. doi: 10.1203/PDR.0b013e3181598cad. [DOI] [PubMed] [Google Scholar]
- 9.Ukkola O, Ravussin E, Jacobson P, et al. Role of ghrelin polymorphisms in obesity based on three different studies. Obesity Research. 2002;10(8):782–791. doi: 10.1038/oby.2002.106. [DOI] [PubMed] [Google Scholar]
- 10.Ukkola O, Ravussin E, Jacobson P, et al. Mutations in the preproghrelin/ghrelin gene associated with obesity in humans. The Journal of Clinical Endocrinology & Metabolism. 2001;86(8):3996–3999. doi: 10.1210/jcem.86.8.7914. [DOI] [PubMed] [Google Scholar]
- 11.Hinney A, Hoch A, Geller F, et al. Ghrelin gene: identification of missense variants and a frameshift mutation in extremely obese children and adolescents and healthy normal weight students. The Journal of Clinical Endocrinology & Metabolism. 2002;87(6):2716–2719. doi: 10.1210/jcem.87.6.8672. [DOI] [PubMed] [Google Scholar]
- 12.Ando R, Kawakami S-I, Bungo T, et al. Feeding responses to several neuropeptide Y receptor agonists in the neonatal chick. European Journal of Pharmacology. 2001;427(1):53–59. doi: 10.1016/s0014-2999(01)01201-8. [DOI] [PubMed] [Google Scholar]
- 13.Vartiainen J, Kesäniemi YA, Ukkola O. Sequencing analysis of ghrelin gene 5′ flanking region: relations between the sequence variants, fasting plasma total ghrelin concentrations, and body mass index. Metabolism. 2006;55(10):1420–1425. doi: 10.1016/j.metabol.2006.06.014. [DOI] [PubMed] [Google Scholar]
- 14.Tsubone T, Masaki T, Katsuragi I, Tanaka K, Kakuma T, Yoshimatsu H. Leptin downregulates ghrelin levels in streptozotocin-induced diabetic mice. American Journal of Physiology. 2005;289(6):R1703–R1706. doi: 10.1152/ajpregu.00773.2004. [DOI] [PubMed] [Google Scholar]
- 15.Shimbara T, Mondal MS, Kawagoe T, et al. Central administration of ghrelin preferentially enhances fat ingestion. Neuroscience Letters. 2004;369(1):75–79. doi: 10.1016/j.neulet.2004.07.060. [DOI] [PubMed] [Google Scholar]
- 16.Kaiya H, van der Geyten S, Kojima M, et al. Chicken ghrelin: purification, cDNA cloning, and biological activity. Endocrinology. 2002;143(9):3454–3463. doi: 10.1210/en.2002-220255. [DOI] [PubMed] [Google Scholar]
- 17.Nie Q, Zeng H, Lei M, et al. Genomic organisation of the chicken ghrelin gene and its single nucleotide polymorphisms detected by denaturing high-performance liquid chromatography. British Poultry Science. 2004;45(5):611–618. doi: 10.1080/00071660400006263. [DOI] [PubMed] [Google Scholar]
- 18.Richards MP, Poch SM, McMurtry JP. Characterization of turkey and chicken ghrelin genes, and regulation of ghrelin and ghrelin receptor mRNA levels in broiler chickens. General and Comparative Endocrinology. 2006;145(3):298–310. doi: 10.1016/j.ygcen.2005.09.013. [DOI] [PubMed] [Google Scholar]
- 19.Tanaka M, Miyazaki T, Yamamoto I, et al. Molecular characterization of chicken growth hormone secretagogue receptor gene. General and Comparative Endocrinology. 2003;134(2):198–202. doi: 10.1016/s0016-6480(03)00247-8. [DOI] [PubMed] [Google Scholar]
- 20.Nie Q, Lei M, Ouyang J, Zeng H, Yang G, Zhang X. Identification and characterization of single nucleotide polymorphisms in 12 chicken growth-correlated genes by denaturing high performance liquid chromatography. Genetics Selection Evolution. 2005;37(3):339–360. doi: 10.1186/1297-9686-37-4-339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lei M, Luo C, Peng X, et al. Polymorphism of growth-correlated genes associated with fatness and muscle fiber traits in chickens. Poultry Science. 2007;86(5):835–842. doi: 10.1093/ps/86.5.835. [DOI] [PubMed] [Google Scholar]
- 22.Li CC, Li K, Li J, et al. Polymorphism of ghrelin gene in twelve Chinese indigenous chicken breeds and its relationship with chicken growth traits. Asian-Australasian Journal of Animal Sciences. 2006;19(2):153–159. [Google Scholar]
- 23.Fang M, Nie Q, Luo C, Zhang D, Zhang X. An 8 bp indel in exon 1 of ghrelin gene associated with chicken growth. Domestic Animal Endocrinology. 2007;32(3):216–225. doi: 10.1016/j.domaniend.2006.02.006. [DOI] [PubMed] [Google Scholar]
- 24.Yuan J, Zhou J, Hu X, Li N. Molecular cloning and comparison of avian preproghrelin genes. Biochemical Genetics. 2007;45(3-4):185–194. doi: 10.1007/s10528-006-9060-z. [DOI] [PubMed] [Google Scholar]
- 25.SAS Institute. SAS® User’s Guide. Statistics. Version 6.12. Cary, NC, USA: SAS Institute; 1996. [Google Scholar]
- 26.Cargill M, Altshuler D, Ireland J, et al. Characterization of single-nucleotide polymorphisms in coding regions of human genes. Nature Genetics. 1999;22(3):231–238. doi: 10.1038/10290. [DOI] [PubMed] [Google Scholar]
- 27.Geelissen SME, Beck IME, Darras VM, Kühn ER, van der Geyten S. Distribution and regulation of chicken growth hormone secretagogue receptor isoforms. General and Comparative Endocrinology. 2003;134(2):167–174. doi: 10.1016/s0016-6480(03)00250-8. [DOI] [PubMed] [Google Scholar]
- 28.Nakai N, Kaneko M, Nakao N, et al. Identification of promoter region of ghrelin gene in human medullary thyroid carcinoma cell line. Life Sciences. 2004;75(18):2193–2201. doi: 10.1016/j.lfs.2004.04.028. [DOI] [PubMed] [Google Scholar]
- 29.Tanaka M, Hayashida Y, Iguchi T, Nakao N, Nakai N, Nakashima K. Organization of the mouse ghrelin gene and promoter: occurrence of a short noncoding first exon. Endocrinology. 2001;142(8):3697–3700. doi: 10.1210/endo.142.8.8433. [DOI] [PubMed] [Google Scholar]
- 30.Kanamoto N, Akamizu T, Tagami T, et al. Genomic structure and characterization of the 5′-flanking region of the human ghrelin gene. Endocrinology. 2004;145(9):4144–4153. doi: 10.1210/en.2003-1718. [DOI] [PubMed] [Google Scholar]
- 31.Evans T, Reitman M, Felsenfeld G. An erythrocyte-specific DNA-binding factor recognizes a regulatory sequence common to all chicken globin genes. Proceedings of the National Academy of Sciences of the United States of America. 1988;85(16):5976–5980. doi: 10.1073/pnas.85.16.5976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ogawa E, Maruyama M, Kagoshima H, et al. PEBP2/PEA2 represents a family of transcription factors homologous to the products of the Drosophila runt gene and the human AML1 gene. Proceedings of the National Academy of Sciences of the United States of America. 1993;90(14):6859–6863. doi: 10.1073/pnas.90.14.6859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Groenen MAM, Dijkhof RJM, van der Poel JJ, van Diggelen R, Verstege E. Multiple octamer binding sites in the promoter region of the bovine αs2-casein gene. Nucleic Acids Research. 1992;20(16):4311–4318. doi: 10.1093/nar/20.16.4311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Eraly SA, Nelson SB, Huang KM, Mellon PL. Oct-1 binds promoter elements required for transcription of the GnRH gene. Molecular Endocrinology. 1998;12(4):469–481. doi: 10.1210/mend.12.4.0092. [DOI] [PubMed] [Google Scholar]
- 35.Skala H, Porteu A, Thomas M, et al. Upstream elements involved in vivo in activation of the brain-specific rat aldolase C gene: role of binding sites for POU and winged helix proteins. The Journal of Biological Chemistry. 1999;273(48):31806–31814. doi: 10.1074/jbc.273.48.31806. [DOI] [PubMed] [Google Scholar]
- 36.Margalit Y, Yarus S, Shapira E, Gruenbaum Y, Fainsod A. Isolation and characterization of target sequences of the chicken CdxA homeobox gene. Nucleic Acids Research. 1993;21(21):4915–4922. doi: 10.1093/nar/21.21.4915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Vartiainen J, Pöykkö SM, Räisänen T, Kesäniemi YA, Ukkola O. Sequencing analysis of the ghrelin receptor (growth hormone secretagogue receptor type 1a) gene. European Journal of Endocrinology. 2004;150(4):457–463. doi: 10.1530/eje.0.1500457. [DOI] [PubMed] [Google Scholar]
- 38.Miraglia del Giudice E, Santoro N, Cirillo G, et al. Molecular screening of the ghrelin gene in Italian obese children: the Leu72Met variant is associated with an earlier onset of obesity. International Journal of Obesity. 2004;28(3):447–450. doi: 10.1038/sj.ijo.0802572. [DOI] [PubMed] [Google Scholar]
- 39.Pöykkö SM, Ukkola O, Kauma H, Savolainen MJ, Kesäniemi YA. Ghrelin Arg51Gln mutation is a risk factor for Type 2 diabetes and hypertension in a random sample of middle-aged subjects. Diabetologia. 2003;46(4):455–458. doi: 10.1007/s00125-003-1058-z. [DOI] [PubMed] [Google Scholar]
- 40.Wang H-J, Geller F, Dempfle A, et al. Ghrelin receptor gene: identification of several sequence variants in extremely obese children and adolescents, healthy normal-weight and underweight students, and children with short normal stature. The Journal of Clinical Endocrinology & Metabolism. 2004;89(1):157–162. doi: 10.1210/jc.2003-031395. [DOI] [PubMed] [Google Scholar]
- 41.Baessler A, Hasinoff MJ, Fischer M, et al. Genetic linkage and association of the growth hormone secretagogue receptor (ghrelin receptor) gene in human obesity. Diabetes. 2005;54(1):259–267. doi: 10.2337/diabetes.54.1.259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Miyasaka K, Hosoya H, Sekime A, et al. Association of ghrelin receptor gene polymorphism with bulimia nervosa in a Japanese population. Journal of Neural Transmission. 2006;113(9):1279–1285. doi: 10.1007/s00702-005-0393-2. [DOI] [PubMed] [Google Scholar]
- 43.Liu G, Fortin J-P, Beinborn M, Kopin AS. Four missense mutations in the ghrelin receptor result in distinct pharmacological abnormalities. The Journal of Pharmacology and Experimental Therapeutics. 2007;322(3):1036–1043. doi: 10.1124/jpet.107.123141. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Figure 1 indicated that genotyps of SNPs were determined based on different profiles by either directly sequencing or PCR‐RFLP technology. Supplementary Figure 2 indicated the genomic structure of the cGHRL, cGHSR, dGHRL, and dGHSR genes. Supplementary Table 1 showed the relative mRNA expression levels of the cGHRL, cGHSR, dGHRL and dGHSR genes in 18 tissues of adult birds.