Abstract
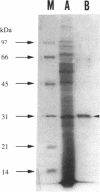
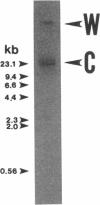
A clinical isolate of Serratia marcescens (TN9106) produced a metallo beta-lactamase (IMP-1) which conferred resistance to imipenem and broad-spectrum beta-lactams. The blaIMP gene providing imipenem resistance was cloned and expressed in Escherichia coli HB101. The IMP-1 was purified from E. coli HB101 that harbors pSMBNU24 carrying blaIMP, and its apparent molecular mass was calculated to be about 30 kDa by sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Kinetic studies of IMP-1 against various beta-lactams revealed that this enzyme hydrolyzes not only various broad-spectrum beta-lactams but also carbapenems. However, aztreonam was relatively stable against IMP-1. Although clavulanate or cloxacillin failed to inhibit IMP-1, Hg2+, Fe2+, or Cu2+ blocked the enzyme's activity. Moreover, the presence of EDTA in the reaction buffer resulted in a decrease in the enzyme's activity. Carbapenem resistance was not transferred from S. marcescens TN9106 to E. coli CSH2 by conjugation. A hybridization study confirmed that blaIMP was encoded on the chromosome of S. marcescens TN9106. By nucleotide sequencing analysis, blaIMP was found to encode a protein of 246 amino acid residues and was shown to have considerable homology to the metallo beta-lactamase genes of Bacillus cereus, Bacteroides fragilis, and Aeromonas hydrophila. The G+C content of blaIMP was 39.4%. Four consensus amino acid residues, His-95, His-97, Cys-176, and His-215, which form putative zinc ligands, were conserved in the deduced amino acid sequence of IMP-1. By determination of the amino acid sequence at the N terminus of purified mature IMP-1, 18 amino acid residues were found to be processed from the N terminus of the premature enzyme as a signal peptide. These results clearly show that IMP-1 is an enterobacterial metallo beta-lactamase, of which the primary structure has been completely determined, that confers resistance to carbapenems and other broad-spectrum beta-lactams.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arakawa Y., Ohta M., Kido N., Fujii Y., Komatsu T., Kato N. Close evolutionary relationship between the chromosomally encoded beta-lactamase gene of Klebsiella pneumoniae and the TEM beta-lactamase gene mediated by R plasmids. FEBS Lett. 1986 Oct 20;207(1):69–74. doi: 10.1016/0014-5793(86)80014-x. [DOI] [PubMed] [Google Scholar]
- Arakawa Y., Ohta M., Kido N., Mori M., Ito H., Komatsu T., Fujii Y., Kato N. Chromosomal beta-lactamase of Klebsiella oxytoca, a new class A enzyme that hydrolyzes broad-spectrum beta-lactam antibiotics. Antimicrob Agents Chemother. 1989 Jan;33(1):63–70. doi: 10.1128/aac.33.1.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bandoh K., Watanabe K., Muto Y., Tanaka Y., Kato N., Ueno K. Conjugal transfer of imipenem resistance in Bacteroides fragilis. J Antibiot (Tokyo) 1992 Apr;45(4):542–547. doi: 10.7164/antibiotics.45.542. [DOI] [PubMed] [Google Scholar]
- Bush K., Macalintal C., Rasmussen B. A., Lee V. J., Yang Y. Kinetic interactions of tazobactam with beta-lactamases from all major structural classes. Antimicrob Agents Chemother. 1993 Apr;37(4):851–858. doi: 10.1128/aac.37.4.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bush K., Sykes R. B. Methodology for the study of beta-lactamases. Antimicrob Agents Chemother. 1986 Jul;30(1):6–10. doi: 10.1128/aac.30.1.6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cuchural G. J., Jr, Malamy M. H., Tally F. P. Beta-lactamase-mediated imipenem resistance in Bacteroides fragilis. Antimicrob Agents Chemother. 1986 Nov;30(5):645–648. doi: 10.1128/aac.30.5.645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galleni M., Lindberg F., Normark S., Cole S., Honore N., Joris B., Frere J. M. Sequence and comparative analysis of three Enterobacter cloacae ampC beta-lactamase genes and their products. Biochem J. 1988 Mar 15;250(3):753–760. doi: 10.1042/bj2500753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horii T., Arakawa Y., Ohta M., Ichiyama S., Wacharotayankun R., Kato N. Plasmid-mediated AmpC-type beta-lactamase isolated from Klebsiella pneumoniae confers resistance to broad-spectrum beta-lactams, including moxalactam. Antimicrob Agents Chemother. 1993 May;37(5):984–990. doi: 10.1128/aac.37.5.984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacoby G. A., Medeiros A. A. More extended-spectrum beta-lactamases. Antimicrob Agents Chemother. 1991 Sep;35(9):1697–1704. doi: 10.1128/aac.35.9.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarlier V., Nicolas M. H., Fournier G., Philippon A. Extended broad-spectrum beta-lactamases conferring transferable resistance to newer beta-lactam agents in Enterobacteriaceae: hospital prevalence and susceptibility patterns. Rev Infect Dis. 1988 Jul-Aug;10(4):867–878. doi: 10.1093/clinids/10.4.867. [DOI] [PubMed] [Google Scholar]
- Kitzis M. D., Billot-Klein D., Goldstein F. W., Williamson R., Tran Van Nhieu G., Carlet J., Acar J. F., Gutmann L. Dissemination of the novel plasmid-mediated beta-lactamase CTX-1, which confers resistance to broad-spectrum cephalosporins, and its inhibition by beta-lactamase inhibitors. Antimicrob Agents Chemother. 1988 Jan;32(1):9–14. doi: 10.1128/aac.32.1.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim H. M., Pène J. J., Shaw R. W. Cloning, nucleotide sequence, and expression of the Bacillus cereus 5/B/6 beta-lactamase II structural gene. J Bacteriol. 1988 Jun;170(6):2873–2878. doi: 10.1128/jb.170.6.2873-2878.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindberg F., Normark S. Sequence of the Citrobacter freundii OS60 chromosomal ampC beta-lactamase gene. Eur J Biochem. 1986 May 2;156(3):441–445. doi: 10.1111/j.1432-1033.1986.tb09601.x. [DOI] [PubMed] [Google Scholar]
- Livermore D. M. Interplay of impermeability and chromosomal beta-lactamase activity in imipenem-resistant Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1992 Sep;36(9):2046–2048. doi: 10.1128/aac.36.9.2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lodge J. M., Minchin S. D., Piddock L. J., Busby S. J. Cloning, sequencing and analysis of the structural gene and regulatory region of the Pseudomonas aeruginosa chromosomal ampC beta-lactamase. Biochem J. 1990 Dec 15;272(3):627–631. doi: 10.1042/bj2720627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Massidda O., Rossolini G. M., Satta G. The Aeromonas hydrophila cphA gene: molecular heterogeneity among class B metallo-beta-lactamases. J Bacteriol. 1991 Aug;173(15):4611–4617. doi: 10.1128/jb.173.15.4611-4617.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto Y., Ikeda F., Kamimura T., Yokota Y., Mine Y. Novel plasmid-mediated beta-lactamase from Escherichia coli that inactivates oxyimino-cephalosporins. Antimicrob Agents Chemother. 1988 Aug;32(8):1243–1246. doi: 10.1128/aac.32.8.1243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nomura K., Yoshida T. Nucleotide sequence of the Serratia marcescens SR50 chromosomal ampC beta-lactamase gene. FEMS Microbiol Lett. 1990 Aug;58(3):295–299. doi: 10.1111/j.1574-6968.1990.tb13992.x. [DOI] [PubMed] [Google Scholar]
- Papanicolaou G. A., Medeiros A. A., Jacoby G. A. Novel plasmid-mediated beta-lactamase (MIR-1) conferring resistance to oxyimino- and alpha-methoxy beta-lactams in clinical isolates of Klebsiella pneumoniae. Antimicrob Agents Chemother. 1990 Nov;34(11):2200–2209. doi: 10.1128/aac.34.11.2200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petit A., Sirot D. L., Chanal C. M., Sirot J. L., Labia R., Gerbaud G., Cluzel R. A. Novel plasmid-mediated beta-lactamase in clinical isolates of Klebsiella pneumoniae more resistant to ceftazidime than to other broad-spectrum cephalosporins. Antimicrob Agents Chemother. 1988 May;32(5):626–630. doi: 10.1128/aac.32.5.626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinn J. P., Dudek E. J., DiVincenzo C. A., Lucks D. A., Lerner S. A. Emergence of resistance to imipenem during therapy for Pseudomonas aeruginosa infections. J Infect Dis. 1986 Aug;154(2):289–294. doi: 10.1093/infdis/154.2.289. [DOI] [PubMed] [Google Scholar]
- Rasmussen B. A., Gluzman Y., Tally F. P. Cloning and sequencing of the class B beta-lactamase gene (ccrA) from Bacteroides fragilis TAL3636. Antimicrob Agents Chemother. 1990 Aug;34(8):1590–1592. doi: 10.1128/aac.34.8.1590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanders C. C. Chromosomal cephalosporinases responsible for multiple resistance to newer beta-lactam antibiotics. Annu Rev Microbiol. 1987;41:573–593. doi: 10.1146/annurev.mi.41.100187.003041. [DOI] [PubMed] [Google Scholar]
- Seeberg A. H., Tolxdorff-Neutzling R. M., Wiedemann B. Chromosomal beta-lactamases of Enterobacter cloacae are responsible for resistance to third-generation cephalosporins. Antimicrob Agents Chemother. 1983 Jun;23(6):918–925. doi: 10.1128/aac.23.6.918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Spencer R. C., Wheat P. F., Winstanley T. G., Cox D. M., Plested S. J. Novel beta-lactamase in a clinical isolate of Klebsiella pneumoniae conferring unusual resistance to beta-lactam antibiotics. J Antimicrob Chemother. 1987 Dec;20(6):919–921. doi: 10.1093/jac/20.6.919. [DOI] [PubMed] [Google Scholar]
- Sutton B. J., Artymiuk P. J., Cordero-Borboa A. E., Little C., Phillips D. C., Waley S. G. An X-ray-crystallographic study of beta-lactamase II from Bacillus cereus at 0.35 nm resolution. Biochem J. 1987 Nov 15;248(1):181–188. doi: 10.1042/bj2480181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takeshita S., Sato M., Toba M., Masahashi W., Hashimoto-Gotoh T. High-copy-number and low-copy-number plasmid vectors for lacZ alpha-complementation and chloramphenicol- or kanamycin-resistance selection. Gene. 1987;61(1):63–74. doi: 10.1016/0378-1119(87)90365-9. [DOI] [PubMed] [Google Scholar]
- Watanabe M., Iyobe S., Inoue M., Mitsuhashi S. Transferable imipenem resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1991 Jan;35(1):147–151. doi: 10.1128/aac.35.1.147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto T., Murayama S. Y., Sawai T. Cloning and expression of the gene(s) for cephalosporinase production of Citrobacter freundii. Mol Gen Genet. 1983;190(1):85–91. doi: 10.1007/BF00330328. [DOI] [PubMed] [Google Scholar]
- Yang Y. J., Wu P. J., Livermore D. M. Biochemical characterization of a beta-lactamase that hydrolyzes penems and carbapenems from two Serratia marcescens isolates. Antimicrob Agents Chemother. 1990 May;34(5):755–758. doi: 10.1128/aac.34.5.755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Yotsuji A., Minami S., Inoue M., Mitsuhashi S. Properties of novel beta-lactamase produced by Bacteroides fragilis. Antimicrob Agents Chemother. 1983 Dec;24(6):925–929. doi: 10.1128/aac.24.6.925. [DOI] [PMC free article] [PubMed] [Google Scholar]