Summary
Hox transcription factors control many aspects of animal morphogenetic diversity. The segmental pattern of Drosophila larval muscles shows stereotyped variations along the anteroposterior body axis. Each muscle is seeded by a founder cell and the properties specific to each muscle reflect the expression by each founder cell of a specific combination of ‘identity’ transcription factors. Founder cells originate from asymmetric division of progenitor cells specified at fixed positions. Using the dorsal DA3 muscle lineage as a paradigm, we show here that Hox proteins play a decisive role in establishing the pattern of Drosophila muscles by controlling the expression of identity transcription factors, such as Nautilus and Collier (Col), at the progenitor stage. High-resolution analysis, using newly designed intron-containing reporter genes to detect primary transcripts, shows that the progenitor stage is the key step at which segment-specific information carried by Hox proteins is superimposed on intrasegmental positional information. Differential control of col transcription by the Antennapedia and Ultrabithorax/Abdominal-A paralogs is mediated by separate cis-regulatory modules (CRMs). Hox proteins also control the segment-specific number of myoblasts allocated to the DA3 muscle. We conclude that Hox proteins both regulate and contribute to the combinatorial code of transcription factors that specify muscle identity and act at several steps during the muscle-specification process to generate muscle diversity.
Keywords: Hox proteins, Cis-regulatory modules, Collier (Knot)/EBF, Nautilus/MyoD, Myogenesis, Drosophila
INTRODUCTION
Anatomy drawings illustrate the stereotyped patterns of skeletal muscles that are essential for coordinated movements. Each muscle has its own name/identity, reflecting its specific properties and function. The genetic and molecular bases of muscle identity remain, however, largely unknown. The rather simple pattern of Drosophila embryonic/larval skeletal muscles makes it an ideal model with which to study this process (Bate 1990; Bate 1993). Every (hemi)segment of the Drosophila larva contains ~30 different somatic muscles, each composed of a single multinucleate fiber. Whereas all muscles express the same myogenic program (e.g. of sarcomeric proteins), each has its own identity characterized by its specific position and orientation with respect to the dorsoventral (D/V) and anteroposterior (A/P) axes, size, sites of attachment on the epidermis and innervation (Bate 1993; Baylies et al., 1998; Knirr et al., 1999). Each syncytial muscle fiber is seeded by a ‘founder’ myoblast (founder cell, FC). FCs possess the unique property of being able to undergo multiple rounds of fusion with fusion-competent myoblasts (FCMs). The current view is that muscle identity reflects the expression by each FC of a specific combination of ‘identity’ transcription factors (iTFs), with Apterous, Collier (Col; Knot), Even-skipped (Eve), Krüppel, Ladybird, Nautilus (Nau) and Slouch (Slou; S59) being among the best characterized (Bourgouin et al., 1992; Ruiz-Gomez et al., 1997; Jagla et al., 1998; Crozatier and Vincent, 1999; Knirr et al., 1999; Balagopalan et al., 2001; Fujioka et al., 2005; Dubois et al., 2007). FCs originate from the asymmetric division of progenitor cells, which are themselves singled out from promuscular clusters (equivalence groups) by Notch (N)-mediated lateral inhibition (Carmena et al., 1995; Ruiz Gomez and Bate, 1997). Each muscle progenitor/FC is specified at a specific A/P and D/V position within the somatic mesoderm. This position determines the final location of the muscle(s) issued from this progenitor.
The abdominal A2-A7 segments present the same final muscle pattern. The patterns of the thoracic T2-T3 and abdominal A1 segments are variations on this pattern, whereas the first thoracic segment (T1) and eighth abdominal segment (A8) present fewer and more diversified muscles (Bate and Rushton, 1993). Although it is well established that the Hox transcription factors are major regulators for patterning of the animal body, our understanding of how each Hox protein specifies distinct morphological features and cellular identities within each body part is still fragmentary (Hueber and Lohmann, 2008; Mann et al., 2009). The segment-specific aspects of the Drosophila musculature represent an interesting paradigm with which to address this question (Greig and Akam, 1993; Michelson, 1994). Based on the expression pattern of Nau, which is the Drosophila ortholog of the mammalian bHLH myogenic factor MyoD, in different Hox conditions, Michelson (Michelson, 1994) suggested that segmental differences in the somatic muscle pattern reflect the regulation of muscle iTFs by Hox proteins. Subsequent studies of apterous expression in thoracic lateral muscles supported this notion and suggested that Hox activity could control the segment-specific variation in the number of myoblasts allocated to a specific group of muscles (Capovilla et al., 2001). As a whole, however, when and how homeotic genes act during the muscle-specification program remain to be established.
Here, we addressed this question, focusing on the dorsal acute DA3 muscle lineage, which depends upon the combinatorial activity of Nau and Col, which is the Drosophila ortholog of mammalian early B-cell factors (EBFs) (Michelson et al., 1990; Keller et al., 1997; Balagopalan et al., 2001). Our data show that Hox activity is superimposed on mesoderm-specific and positional information at the progenitor stage to establish the pattern of somatic muscles and acts at several independent steps during the muscle-specification process. Very few studies have addressed the role of Hox genes in controlling the fate of skeletal muscle precursors in vertebrates (Alvares et al., 2003). Our data provide a new framework for studying the integration of Hox information into the generic process of myogenesis and the generation of muscle diversity.
MATERIALS AND METHODS
Drosophila genetics
The strains used were w118 as wild-type (wt) reference (Bloomington Stock Center, IN, USA), Antp1 (Abbott and Kaufman, 1986), Ubx1 (Bloomington Stock Center), Antp25; Ubx1 (Ernesto Sanchez-Herrero, Madrid, Spain), rP298-lacZ (Nose et al., 1998), UAS-Antp (Heuer et al., 1995), UAS-Ubx, UAS-abdA (Michelson, 1994), UAS-col (Vervoort et al., 1999), UAS-nau (Keller et al., 1997), 24B-Gal4 (Brand and Perrimon, 1993), rP298-Gal4 (Menon and Chia, 2001) and sns-Gal4 (Kocherlakota et al., 2008). The mutant strains were balanced over marked (TM3 twist-lacZ) chromosomes. All Gal4-UAS crosses were performed at 25°C.
Plasmid constructions and transgenic lines
A BamHI-NaeI DNA fragment containing the attB site from the pUASTattB vector (Bischof et al., 2007) was substituted for the P-element-containing NsiI-AvrII fragment in the H-Pelican lacZ transformation vector (Barolo et al., 2000) (GenBank AF242361.2) to generate attB-inslacZ, in which lacZ is under the control of the minimal hsp70 promoter and flanked by gypsy insulator sequences. The 4_0.9, 2.6_0.9 and CRM276 col genomic fragments were inserted into the attB-inslacZ and/or attB-inslacZi vectors. Each attB construct was inserted at position 49D on the second chromosome by injection into nosC31NLS; Zh8 embryos (Bischof et al., 2007).
Construction of an intron-containing lacZi reporter gene
The unique intron of the D. virilis β-tubulin56D (βtub56D) gene (Dvir\GJ20466; FlyBase ID FBgn0207606) was inserted into the lacZ coding region between Asp119 and Val120 by standard PCR-based cloning. The 5′ to 3′ sequences of the created splice junctions are as follows: 5′-CGTGAGGT — AGGTCTCG-3′; the intron sequence is underlined. This resulted in a single C-to-G nucleotide change (bold) in the lacZ coding sequence, resulting in an Asp235-to-Glu substitution. The resulting intron-containing lacZ gene, denoted lacZi, was used to generate attB-inslacZi from attB-inslacZ. Mutagenesis of the consensus AbdA/Ubx binding site TAATTA (Ekker et al., 1994) to TGGGGA was by PCR.
Immunohistochemical staining and in situ hybridization
Embryos were fixed and processed for antibody staining and/or in situ hybridization as described (Crozatier et al., 1996). Primary antibodies were: mouse anti-Col (1/100) (Dubois et al., 2007); rabbit anti-Mef2 and anti-Nau (1/100) (provided by Eileen Furlong, Heidelberg, Germany and Bruce Paterson, Bethesda, MD, USA, respectively); mouse anti-β-galactosidase (Promega, 1/1000); and mouse anti-Ubx and anti-Antp (1/100, Developmental Studies Hybridoma Bank, IA, USA). Secondary antibodies were: Alexa Fluor 488-conjugated goat anti-rabbit and goat anti-mouse; Alexa Fluor 555-conjugated goat anti-rabbit and goat anti-mouse; Alexa Fluor 647-conjugated goat anti-mouse (all Molecular Probes, 1/300); and biotinylated goat anti-mouse (Vector Laboratories, 1/1000). Double fluorescence in situ hybridization and immunostaining using intronic probes and Col antibodies were as described (Dubois, 2007).
Sequence alignments and transcription factor binding sites
Pairwise sequence alignments of col upstream and CRM276 sequences from various Drosophila species (http://flybase.org/cgi-bin/gbrowse/) were performed using NCBI-BLAST (bl2seq), Genome Browser (University of California, Santa Cruz, CA, USA) and Evoprinter (NINDS, NIH, Bethesda, MD, USA) and manually edited by eye. The search for individual binding sites for transcription factors made use of Cis Analyst (http://rana.lbl.gov/cis-analyst/) and FlyEnhancer (http://opengenomics.org/) and manual inspection based on the matrices for binding sites of Twi, Tin, dTCF, Mad and Pnt (Philippakis et al., 2006). Access to the Tin and Twi in vivo binding sites (Sandmann et al., 2006; Sandmann et al., 2007; Zinzen et al., 2009) was via the E. Furlong lab site (http://furlonglab.embl.de/data/).
RESULTS
Segment-specific properties of the DA3 muscle
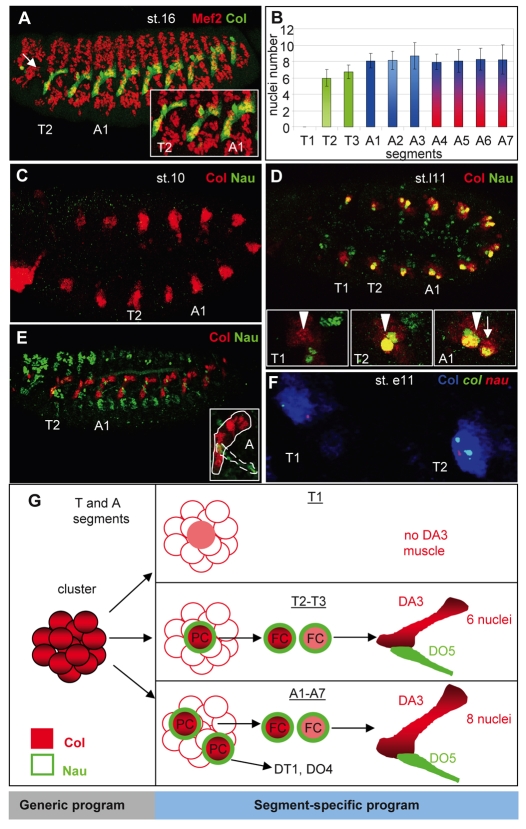
Col expression in stage 15 embryos showed that the DA3 muscle forms in the thoracic (T) T2-T3 segments and abdominal (A) A1-A7 segments, but not in the T1 segment and was thinner in T2-T3 than in A1-A7 (Fig. 1A). Since the size of Drosophila larval muscles correlates with their number of nuclei (Demontis and Perrimon, 2009), we counted the number of nuclei in the DA3 muscle. Fewer nuclei were present in T2 and T3 (six on average) than in A1-A7 (eight on average) (Fig. 1B), showing that this number is segment specific. Col expression in the somatic mesoderm is first detected at embryonic stage 10, in a cluster of cells at the same dorsal position in all trunk segments, including T1 (Fig. 1C). This cluster gives rise to the DA3/DO5 progenitor in T2 and T3 and to the DA3/DO5 and DO4/DT1 progenitors in A1-A7 (Fig. 1D). Following asymmetric division of the DA3/DO5 progenitor, col transcription is maintained in the DA3 FC but is repressed in the sibling DO5 FC, a repression mediated by N; it is also not maintained in the DO4 and DT1 FCs (Crozatier and Vincent, 1999). Nau is expressed in the same progenitors as Col (Fig. 1C,D). However, Nau and Col co-expression is only transient. col transcription is maintained in the DA3 FC and is activated in the nucleus of each FCM incorporated into the growing DA3 myofiber, whereas nau is transiently transcribed in the DA3 lineage and is activated in FCM nuclei incorporated into the DO5 myofiber (Dubois et al., 2007). This leads to a specific accumulation of Col and Nau in the DA3 and DO5 muscles, respectively (Fig. 1E). Thus, the progenitor/FC stage is the specific step in the DA3/D05 lineage at which Nau and Col are expressed together. One Col- and Nau-expressing progenitor is found in T2 and T3, two progenitors in A1-A7 and none in T1, correlating with the final muscle pattern (Bate and Rushton, 1993; Crozatier and Vincent, 1999). Transient co-transcription of col and nau was nevertheless observed in a cell issued from the Col-expressing cluster in T1, although at a very low level compared with other segments (Fig. 1F). This indicates that a positive input required to upregulate the expression of these two iTFs in progenitor cells is missing in T1. In summary, transient expression of Nau and Col at the progenitor stage is segment specific and foreshadows the segment-specific formation of DA3/DO5 and DO4/DT1 muscles (Fig. 1G).
Fig. 1.
Segment-specific formation and properties of the DA3 muscle. (A) Mef2 (red) and Col (green) expression in a stage (st.) 16 Drosophila embryo, showing the entire muscle pattern and DA3 muscle, respectively. The arrow indicates the absence of DA3 muscle in the T1 segment. The inset shows an enlarged view of the T2, T3 and A1 segments. Note that the DA3 muscle is smaller in T than in A segments. (B) The segment-specific number of nuclei in the DA3 muscle (number of segments counted=30). The mesodermal domain of expression of Antp is in green, Ubx in blue and AbdA in red (Bate,1993) (see Fig. S3 in the supplementary material). (C-E) Comparison of expression of Col (red) and Nau (green) in stage 10 (C), late stage 11 (st.l11) (D) and stage 15 (E) embryos. Insets in D show the Col-expressing cluster in T1 (arrowhead), the DA3/DO5 progenitor in T2, and DA3/DO5 (arrowhead) and the DT1/DO4 progenitors (arrow) in A1. The inset in E is an enlarged view of the abdominal DA3 (circled) and DO5 (dotted circle) muscles that express Col and Nau, respectively. (F) Double in situ hybridization for col (green) and nau (red) primary transcripts and immunostaining for Col (blue). The T1 and T2 segments of an early stage 11 embryo are shown. Two dots per nucleus reflect the two alleles. All embryos are oriented anterior to the left and dorsal to the top. (G) Col (red) and Nau (green) expression during the process of DA3 muscle formation in different trunk segments, highlighting the segment-specific aspects. PC, progenitor; FC, founder cell.
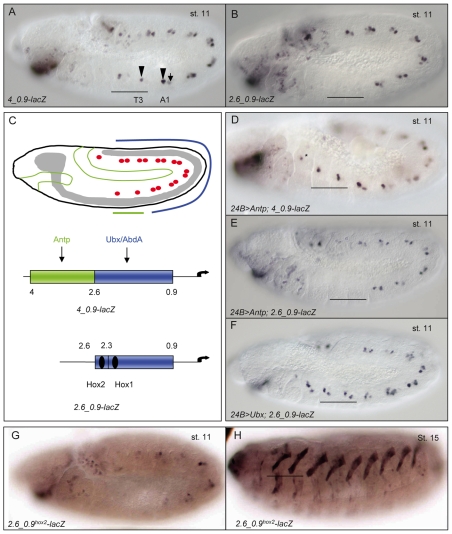
Two independent cis-regulatory modules (CRMs) control the different phases of col transcription in the DA3 muscle
A lacZ reporter gene containing 4 kb of the col upstream cis-regulatory region (4_0.9-lacZ; Fig. 2A) reproduces col expression in the DA3 FC and muscle, but not in promuscular clusters (Dubois et al., 2007). An enhancer that drives Eve expression in dorsal promuscular clusters at the origin of the DA1 and DO2 muscles (see Fig. S1 in the supplementary material) has been described (Halfon et al., 2000; Knirr and Frasch, 2001; Speicher et al., 2008). This enhancer integrates positional information issued from the ectoderm with mesoderm-intrinsic information via the binding of five different transcription factors: Mothers against dpp (Mad), dTCF (Pangolin), Pointed (Pnt), Twist (Twi) and Tinman (Tin) (Carmena et al., 1998; Halfon et al., 2000; Knirr and Frasch, 2001). Mad and dTCF are the downstream effectors of Decapentaplegic (Dpp) and Wingless (Wg) signaling, respectively, with these ectodermal signals defining a positional grid within each segment. Pnt is an effector of the Ras/MAPK pathway that contributes to the specification of equivalence groups within the mesoderm. Twi and Tin are mesoderm-specific transcription factors (Halfon et al., 2000).
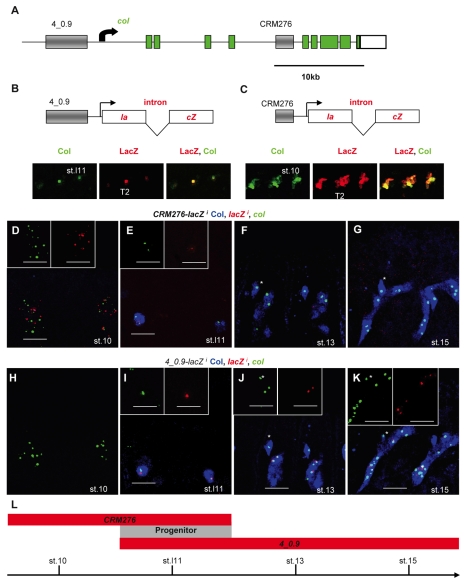
Fig. 2.
Two separate CRMs control col transcription at different steps during DA3 muscle specification. (A) The Drosophila col genomic region. Coding exons are in green. The transcription start site is indicated by an arrow. Gray boxes represent the muscle CRMs 4_0.9 and 276. (B,C) The 4_0.9-lacZi and CRM276-lacZi transgenes, in which the lacZ coding sequence (red) is split by the Drosophila virilis βtub56D intron. The transcription start site is indicated by an arrow. Beneath are shown (B) stage 11 4_0.9-lacZi and (C) stage 10 CRM276-lacZi embryos immunostained for Col (green) and β-galactosidase (red, LacZ), or both, as indicated. (D-K) Double in situ hybridization for col (green) and lacZ (red) primary transcripts and Col immunostaining (blue) on (D-G) CRM276-lacZi and (H-K) 4_0.9-lacZi embryos. Developmental stages are indicated in each panel. The T2 and T3 segments are shown. Insets show only lacZi or col transcripts. Note that the red and green dots corresponding to lacZi and col, respectively, do not always overlap because the two genes are located at separate chromosomal positions, with two dots per gene per nucleus reflecting the two alleles. The asterisk (F,G,J,K) indicates a Col-expressing multidendritic (md) neuron. CRM276-lacZi transcription is only active in promuscular clusters and in the corresponding progenitor (D,E). 4_0.9-lacZi transcription starts in the DA3/DO5 progenitor (I) and is active in DA3 nuclei until stage 15 (J,K). (L) The temporal windows of activity of CRM276 and 4_0.9, which specifically overlap at the progenitor stage (gray).
By comparing gene expression profiles for myoblasts of different genotypes representing perturbations of the Wg, Dpp, Ras and N pathways, Estrada et al. identified ~160 genes that were possibly regulated similarly to eve in the mesoderm, including col (Estrada et al., 2006). Using an in silico search, with the ModuleFinder protocol (Philippakis et al., 2006), we identified five non-coding sequences that were selectively enriched for combinations of Tin, Twi, Pnt, Mad and dTCF binding sites within the col gene. Each was individually tested by transgenic reporter analysis (data not shown). The intron-located sequence with the highest prediction score, despite the absence of a conserved Mad binding site (see Fig. S2 in the supplementary material), which is referred to below as CRM276 (Fig. 2A), drove lacZ expression in promuscular clusters. Double staining of stage 10-11 embryos showed a precise overlap between CRM276-lacZ and Col expression (data not shown).
The stability of lacZ mRNA and β-gal protein prevents, however, a precise determination of temporal aspects of the activity of CRMs, a central aspect of our study. To circumvent this problem, we introduced the Drosophila virilis βtub56D intron into the lacZ coding region (lacZi, see Materials and methods) and generated reporters in which lacZi is placed under control of the CRM276 and 4_0.9 col fragments (4_0.9-lacZi and CRM276_lacZi, respectively; Fig. 2B,C). The patterns of lacZ expression were identical when driven by intron-devoid (not shown) and intron-containing transgenes (Fig. 2B,C), demonstrating that the intron is efficiently spliced. In situ hybridization with a mixture of col and lacZi intronic probes confirmed at the primary transcript level that CRM276 precisely reproduces col activation and integrates the same A/P and D/V positional information (Fig. 2D). CRM276-lacZi transcription was subsequently restricted to the DA3/DO5 progenitor, identical to endogenous col, indicating that it is subject to repression by N (Fig. 2E). Unlike col, however, CRM276-lacZi transcription was very weak in progenitors and was not detected beyond that stage (Fig. 2F,G). Thus, CRM276 is an early mesodermal enhancer that imparts positional and mesodermal information to col activation in a specific promuscular cluster. Conversely, the 4_0.9 CRM was only active from the progenitor stage and was activated in the nuclei of FCMs that have fused with the DA3 FC (Fig. 2H-K). CRM276 and 4_0.9 CRM together account for all aspects of col transcription, including its repression by N signaling, during the successive steps of selection and asymmetric division of the DA3/D05 progenitor (Fig. 2I).
The precise determination of temporal windows of activity established that the activity of the position-specific CRM276 is transient and is relayed by another 4_0.9 CRM in progenitor cells. As it does not operate in T1, this relay is segment specific, providing evidence that it is at the progenitor stage that segment-specific information superimposes on positional information provided by segmentation and D/V patterning genes (Fig. 2L).
Hox proteins are required for DA3 muscle specification and allocate its number of nuclei
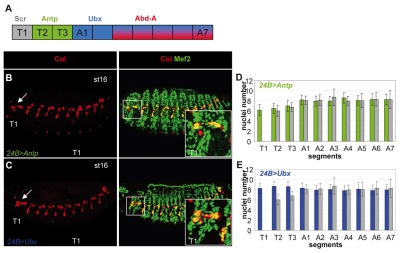
The register of mesodermal expression of the thoracic Hox proteins Sex combs reduced (Scr), Antennapedia (Antp), Ultrabithorax (Ubx) and Abdominal-A (AbdA) is schematized in Fig. 3A (Bate, 1993) (see Fig. S3 in the supplementary material; data not shown) (Bate and Rushton, 1993). Comparison between 4_0.9-lacZ and either Scr, Antp or Ubx expression in stage 11 embryos (see Fig. S3A,B in the supplementary material; data not shown) confirmed that the Col-expressing progenitor in T2 and T3 expresses Antp and that the two col-positive progenitors in A1-A7 express Ubx. Overlapping expression with the FC-specific marker, rP298-lacZ (Nose et al., 1998), in late stage 11 embryos showed that Antp and Ubx expression is maintained in T2-T3 and A1-A7 FCs, respectively (see Fig. S3C,D in the supplementary material). We looked at the effect of overexpressing either Antp or Ubx (Fig. 3B,C) or AbdA (not shown) throughout the mesoderm on the specification of the DA3 muscle. Pan-mesodermal expression of either Hox protein (Brand and Perrimon, 1993; Michelson, 1994) resulted in the formation of an ectopic Col-expressing muscle in T1, at the same position as in other segments (Fig. 3B,C). Thus, providing Antp, Ubx or AbdA activity is sufficient to convert the Col-expressing promuscular cluster in T1 into a DA3 muscle. We observed, however, that the size of this ectopic DA3 muscle varied depending on which Hox protein was expressed. When Antp was overexpressed, the number of nuclei in this muscle was identical to that in wild-type (wt) T2 and T3 (Fig. 3D). However, when either Ubx or AbdA was overexpressed, this number was converted to that in wt A segments (Fig. 3E and data not shown), a clear example of posterior prevalence (Duboule and Morata, 1994). These data show that Antp, Ubx and AbdA activities in the mesoderm control the formation of the DA3 muscle and regulate its number of nuclei.
Fig. 3.
Hox proteins are required for the formation of the DA3 muscle and allocate its number of nuclei. (A) A color-coded representation of Scr, Antp, Ubx and AbdA expression in the Drosophila trunk segments. (B,C) Col (red) and Mef2 (green) expression in stage 16 embryos expressing Antp (B) or Ubx (C) in all mesodermal cells (24B-Gal4 driver). Col staining alone is shown on the left. The arrow points to the DA3-like muscle that forms in T1. (D,E) The number of nuclei per DA3 muscle in each T1 to A7 segment. The genotypes are shown at the top, with the Hox color code as in Fig. 1. The gray bars show wild-type (wt) nuclei numbers. Number of segments counted=30.
Hox proteins control the number of muscle progenitors expressing Col and Nau
We then looked at when during the muscle-specification process expression of Col and Nau was modified by pan-mesodermal expression of Antp or Ubx. In stage 11 embryos, residual Col expression was detected in all segments in cells that do not become progenitors; high levels of Col and Nau co-localized in the DA3/DO5 progenitor in T2-T3 and in the DA3/DO5 and DO4/DT1 progenitors in A1-A7 (Fig. 1D,G; Fig. 4A). Pan-mesodermal expression of Antp resulted in high-level expression of Col and Nau in one dorsal progenitor in T1 (Fig. 4B), a pattern normally restricted to T2 and T3 (Fig. 4A). Pan-mesodermal expression of Ubx resulted in high-level expression of Col and Nau in two progenitors in all three T segments (Fig. 4C), a pattern typical of wt A1-A7 segments (see Fig. 1D). We conclude that Hox activity in the mesoderm controls the segment-specific number of progenitors expressing Col and Nau that emerge from a pre-defined promuscular cluster.
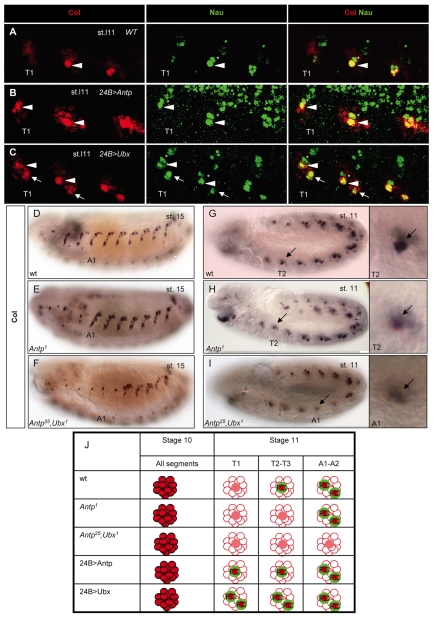
Fig. 4.
Hox proteins control the number of muscle progenitors expressing Col and Nau. (A-C) Col (red) and Nau (green) expression in T segments of late stage 11 Drosophila embryos. Overlap (yellow) is shown on the right. (A) One progenitor expressing high levels of Col and Nau is found in wt T2 (arrowhead) and T3 and in T1 upon expression of Antp (B) or Ubx (C). (C) Ubx induces a second progenitor (arrow) to express high levels of Col and Nau in all three T segments, similar to wt A segments (see Fig. 1D). The asterisk indicates another muscle progenitor that expresses Nau, but not Col, in wt T segments and sometimes low levels of Col upon Hox overexpression. (D-I) Col expression in stage 15 (D-F) and 11 (G-I) embryos. The DA3 muscle is lacking in T2-T3 and T2-A2 in Antp (E) and Antp; Ubx (F) mutants, respectively, correlating with the absence of high-level Col-expressing progenitors (H,I). The remaining Col-positive nucleus corresponds to that of an md neuron, as shown by its expression of pickpocket (Crozatier and Vincent, 2008). Enlarged views of the Col-expressing cluster/progenitor in T2 (G,H) and A1 (I) show that Col upregulation in progenitor cells does not occur in the absence of Hox protein. Residual Col protein indicates the position of the Col-expressing cluster. (J) Col (red) and Nau (green) expression in muscle progenitors in different trunk segments in Hox mutants and Hox-overexpression conditions.
In order to complement the Hox gain-of-function data, we followed Col expression and the DA3 muscle lineage in Hox loss-of-function mutant embryos, using null alleles (Fig. 4D-I). No DA3 muscle formed in T2 and T3 in Antp mutant embryos (Fig. 4E). In Ubx mutant embryos, the DA3 muscle pattern was similar to that of the wt (not shown), presumably owing to the derepression of Antp expression in the A1 and A2 segments (Hooper, 1986). Accordingly, we observed that Antp, Ubx double-mutant embryos lacked a DA3 muscle in all T and A1-A2 segments (Fig. 4F). Close examination of stage 11 Antp mutant embryos showed that one cell in each thoracic segment transiently accumulated more Col protein than the other cells of the cluster, similar to T1 in wt embryos. This transient, low-level Col accumulation confirms that the process of progenitor selection is initiated and prematurely aborts in the absence of Antp input (Fig. 4H). In Antp, Ubx double mutants, Col upregulation in progenitors was only observed in A3-A7 (Fig. 4I), further confirming that Hox-dependent upregulation and the final muscle pattern are linked (Fig. 4J).
Antp and Ubx/AbdA control col expression in muscle progenitors via distinct cis-regulatory elements
The loss of col expression in Hox mutant embryos and the pattern of 4_0.9-lacZ expression together indicated that the 4_0.9 CRM mediates col regulation by Hox proteins (Figs 2 and 4; Fig. 5A). The observation that a reporter construct containing only 2.6 kb of col upstream DNA (2.6_0.9-lacZ) was only active in A progenitors (Fig. 5B,C) further raised the possibility that differential control by the Antp and Ubx/AbdA paralogs could involve different cis elements. To investigate this, we compared 2.6_0.9-lacZ and 4_0.9-lacZ expression in stage 11 embryos when either Antp or Ubx was expressed throughout the entire mesoderm. Unlike col, 2.6_0.9-lacZ was not activated by Antp in T1, consistent with the fact that it is not expressed in T2 and T3 in wt embryos (Fig. 5E; see also Fig. 4). By contrast, 2.6_0.9-lacZ was activated by Ubx (or AbdA, not shown) in two progenitors in each segment, including all three T segments, thereby reproducing the pattern of col expression under these conditions (Fig. 5F). These results showed that upregulation of col expression by Ubx/AbdA, and not Antp, involves cis elements present within the 2.6_0.9 col upstream region. Conversely, 4_0.9-lacZ expression was upregulated by Antp in one progenitor in T1, therefore reproducing Col expression under the same conditions (Fig. 5D; Fig. 4B). These data show that Antp and Ubx/AbdA control col expression in muscle progenitors via distinct cis-regulatory elements (Fig. 5C).
Fig. 5.
Hox regulation of col transcription at the progenitor stage is modular. (A,B,D-H) Lateral views of stage 11 Drosophila embryos expressing lacZ in the DA3/DO5 (A, arrowhead) and DT1/DO4 (A, arrow) progenitors under control of either the 4_0.9 or 2.6_0.9 CRM. (A,B) lacZ expression in T2 and T3 (horizontal bar) in wt embryos requires the 4_2.6 CRM. (D-F) 2.6_0.9-lacZ expression is induced in T2 and T3 by Ubx but not by Antp; 4_0.9-lacZ expression is induced by Antp in all segments, including in T1 (asterisk in D). (G) Mutation of the Hox2 site abolishes 2.6_0.9-lacZ expression in progenitors. (H) lacZ expression in the DA3 muscle at stage 15 is independent of the 4_2.6 CRM and Hox2. (C) Schematic of a stage 11 embryo with the Col-expressing progenitors shown as red dots and domains of mesodermal Antp and Ubx/AbdA expression as green and blue lines, respectively. Beneath are the 4_0.9-lacZ and 2.6_0.9-lacZ constructs showing the Antp- and Ubx-responsive elements in green and blue, respectively. Position of the Hox1 and Hox2 sites is indicated (see Fig. S4 in the supplementary material).
Ubx/AbdA regulation of col transcription in muscle progenitors is direct
Since 2.6_0.9-lacZ is only expressed in A segments, we asked whether its regulation by Ubx/AbdA is direct. An in silico search for high-affinity Hox binding sites [TAATTA (Ekker et al., 1994; Affolter et al., 2008)] within the 2.6_0.9 CRM identified two sites that are conserved at the same relative position in several Drosophila species (Fig. 5C and see Fig. S4 in the supplementary material). Each of these sites, named Hox1 and Hox2, was individually mutated (TAAT to GGGG) within the 2.6_0.9-lacZ construct, giving rise to the 2.6_0.9Hox1-lacZ and 2.6_0.9Hox2-lacZ transgenes. 2.6_0.9Hox2-lacZ expression was no longer detected in progenitors (Fig. 5G), whereas the Hox1 mutation had little if any effect. Mutation of both Hox1 and Hox2 sites had no additional effect over the Hox2 mutation alone (data not shown), indicating that Hox2 is required for col regulation by Ubx/AbdA in abdominal muscle progenitors. Expression of 2.6_0.9Hox2-lacZ recovered later during DA3 muscle development (Fig. 5H), consistent with our previous finding that cis elements responsible for col activation in the DA3-recruited FCM are all contained within the 2.3_1.6 interval, which does not cover the Hox2 site (Dubois et al., 2007). We conclude that a single Hox binding site mediates the specific upregulation of col expression by Ubx/AbdA in A progenitors (Fig. 5C).
Forced expression of Nau plus Col bypasses the need for Hox activity in forming a DA3 muscle, but not in allocating its nuclei number
Our finding that Hox activity is required in progenitors for prolonged expression of Nau and Col and for implementation of the muscle differentiation process suggested that forced expression of these iTFs in progenitor cells might bypass the need for Hox activity. We therefore repeated the 24B-Gal4×UAS-overexpression experiments, but using both Col and Nau in place of a Hox protein. To visualize the DA3 muscle, we used the 2.6_0.9-lacZ reporter (Fig. 6A). As previously shown (Dubois et al., 2007), 24B-Gal4-driven expression of Nau alone does not affect 2.6_0.9-lacZ expression, whereas expression of Col results in strong activation in the DA2 and VL1-2 muscles in T2 to A7, in DT1 in A segments, plus, more sporadically, in a few other muscles (Fig. 6B,C) (Dubois et al., 2007). The same pattern was observed upon expression of Col plus Nau, with the notable exception of additional activation in a muscle in T1. This ectopic muscle is at the same position and shows the same orientation and attachment sites as the ectopic DA3 muscle induced by Hox proteins (see Fig. 3). We infer from this observation that high levels of Nau plus Col can bypass the need for a Hox protein in implementing the DA3 muscle formation process. It strengthens our conclusion that the upregulation of expression of a combination of iTFs by Hox proteins plays a decisive role in converting positional information within the mesoderm into a specific muscle pattern.
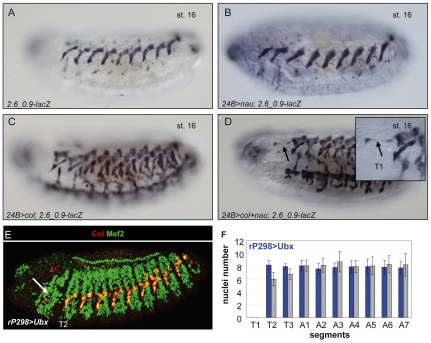
Fig. 6.
Expression of Nau and Col can bypass the Hox requirement for DA3 muscle specification. (A-D) lacZ expression in stage 16 2.6_0.9-lacZ Drosophila embryos upon pan-mesodermal expression of Nau (B), Col (C), Col plus Nau (D) or neither protein (A). Only co-expression of Col plus Nau results in the formation of a DA3-like muscle in T1 (arrow). This ectopic muscle has only two nuclei (inset in D). (E) Double immunostaining of stage 15 embryos for Mef2 (green) and Col (red) showing Col expression upon expression of Ubx in all FCs. (F) The number of nuclei per DA3 muscle in each segment of rP298>Ubx embryos is in blue. Wt values are in gray. Number of segments counted=30.
However, the ectopic DA3 muscle that formed in T1 upon forced expression of Col and Nau displayed very few nuclei (two to three), indicating that Hox activity is essential for allocating this muscle a normal number of nuclei. To better define in which myoblasts Hox was acting, we repeated the Ubx-overexpression experiments using two different drivers, rP298-Gal4 and sns-Gal4, which are specific for FCs and muscle precursors and for FCMs, respectively (Menon and Chia, 2001; Kocherlakota et al., 2008). We found that the number of nuclei in the T2 and T3 segments was converted to that in A segments upon Ubx expression in FCs (Fig. 6E,F) but not in FCMs. These results show that Hox factors act to regulate the size of each muscle. Ubx expression in FCs did not, however, promote the formation of an ectopic DA3 muscle in T1 (Fig. 6E), strengthening our conclusion that Hox activity is required at the progenitor stage to implement the muscle differentiation process.
A model for the transcriptional history of a Drosophila muscle
Segment-specific upregulation of col and nau expression by Hox proteins suggests the following model for the transcriptional history of the DA3 muscle (Fig. 7). The first step is the activation of col transcription in a cluster of mesodermal cells in all T and A segments. This is controlled by the early-acting CRM276 enhancer, which integrates positional information issued from the ectoderm, mesoderm-intrinsic cues and repression by N signaling in non-progenitor cells. In a second step, maintenance of Col and Nau expression in the DA3/DO5 progenitor relies upon Hox factors and is mediated by another CRM, the 4_0.9 CRM, which contains separate elements for regulation by Antp and Ubx/AbdA. The 4_0.9 CRM is also subject to repression by N, leading to restriction of col transcription to the DA3 FC. In a third step, implementation of the DA3 muscle differentiation process requires positive and direct Col autoregulation, which converts all of the DA3 muscle nuclei to the same transcriptional program (Dubois et al., 2007). In essence, Col accumulation in the DA3/DO5 progenitor is required to maintain its own expression in the DA3 FC and muscle precursor, thus representing a case of forward autoregulation. Finally, Hox proteins collaborate with iTFs to control the number of myoblasts assigned to the DA3 muscle. Central to our model is the switch in regulation from the early to late muscle CRM that occurs at the progenitor stage and requires Hox activity, thereby linking positional information along the A/P axis to muscle diversity.
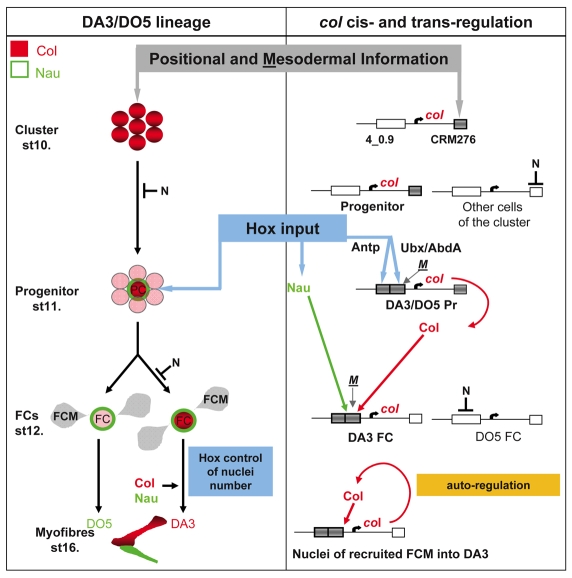
Fig. 7.
Transcriptional control of Drosophila muscle identity: the central role of Hox proteins. Expression of Col (red) and Nau (green) in the DA3 muscle lineage is shown on the left, with the corresponding cis control of col expression represented on the right. col transcription is activated in one promuscular cluster per trunk segment under control of the early-acting CRM276, which integrates A/P and D/V positional information, mesoderm-intrinsic cues and repression by N signaling. This results in Col expression in one (T segments) or two (A1-A7 segments) dorsal progenitors. Upregulation of col (red) and nau (green) transcription in the DA3/DO5 progenitor at stage 11 is controlled by Antp and Ubx/AbdA and is mediated by two separate elements of the 4_0.9 CRM. This is the key step, when segment-specific variations are imposed on the segmental muscle pattern. Following progenitor division, restriction of col transcription to the DA3 FC requires positive regulation by Col and Nau and repression by N. From this stage on, Col is required for activating its own transcription in all FCM nuclei incorporated into the DA3 myofiber. Hox activity in FCs controls the number of myoblasts allocated to the DA3 muscle, independently of the control of nau and col. The key regulatory steps are shaded and centered: early positional information is relayed by segment-specific Hox information at the progenitor stage. The active and inactive CRMs are indicated by gray and white boxes, respectively. Binding of mesodermal transcription factors (M), such as Twi and Mef2, to CRM 276 and 4_0.9 is suggested by phylogenomic footprinting (see Figs S3 and S4 in the supplementary material) and ChIP-on-chip data (Sandmann et al., 2006; Sandmann et al., 2007).
DISCUSSION
Cis-reading of positional information: intersecting computational predictions and ChIP-on-chip data
Eve expression in the DA1 muscle lineage provided the first paradigm for studying the early steps of muscle specification. Detailed characterization of an eve muscle CRM showed that positional and tissue-specific information were directly integrated at the level of CRMs via the binding of multiple transcription factors, including dTCF, Mad, Pnt, Tin and Twi (Carmena et al., 1998; Halfon et al., 2000; Knirr and Frasch, 2001). Based on this transcription factor code and using the ModuleFinder computational approach (Philippakis et al., 2006), we have identified a CRM, CRM276, that precisely reproduces the early phase of col transcription. This CRM also drove expression in cells of the lymph gland, another organ that is issued from the dorsal mesoderm where col is expressed (Crozatier et al., 2004). Parallel to our study, two col genomic fragments were selectively retrieved in chromatin immunoprecipitation (ChIP-on-chip) experiments designed to identify in vivo binding sites for Twi, Tin or Mef2 in early embryos (Sandmann et al., 2007; Liu et al., 2009). One fragment overlaps with CRM276. Based on this overlap and interspecies sequence conservation, we tested a 1.4 kb subfragment of CRM276 that retained most of the transcription factor binding sites identified by ModuleFinder and found that it specifically reproduced promuscular col expression (M. de Taffin, personal communication). This in vivo validation shows that intersecting computational predictions and ChIP-on-chip data should provide a very efficient approach to identify functional CRMs on a genome-wide scale (Zinzen et al., 2009).
The eve and col early mesodermal CRMs are activated at distinct A/P and D/V positions. We are now in a position to undertake a comparison of these two CRMs, in terms of the number and relative spacing of common activator and repressor sites and their expanded combinatorial code, in order to understand how different mesodermal cis elements perform a specific interpretation of positional information.
The Hox code relays positional information in a segment-specific manner
A progenitor is selected from the Col promuscular cluster in T2 and T3 but not T1. One cell issued from the Col-expressing promuscular cluster in T1 nevertheless shows transiently enhanced Col expression, suggesting that the generic process of progenitor selection is correctly initiated in T1. This process aborts, however, in the absence of a Hox input, as shown by the loss of progenitor Col expression and DA3 muscle in specific segments in Hox mutants. The similar changes in Nau and Col expression observed under Hox gain-of-function conditions allow us to conclude that the expression of iTFs is regulated by Hox factors at the progenitor stage. The superimposition of Hox information onto the intrasegmental information thereby implements the iTF code in a segment-specific manner and establishes the final muscle pattern. Unlike DA3, a number of specific muscles are found in both T1 and T2-A7 (Bate and Rushton, 1993), such as the Eve-expressing DA1 muscle; other muscles form in either abdominal or thoracic segments, as illustrated by the pattern of Nau expression in stage 16 embryos (Michelson, 1994) (Fig. 1E). This diversity in segment-specific patterns indicates that Hox regulation of iTF expression is iTF and/or progenitor specific.
Modular cis regulation of col transcription by Hox proteins
As early as 1994, Hox proteins were proposed to regulate the segment-specific expression of iTFs. Seven years later, Capovilla et al. characterized an apterous mesodermal enhancer (apME680) active in the LT1-4 muscles and proposed that regulation by Antp was direct (Capovilla et al., 2001). However, mutation of the predicted Antp binding sites present in apME680 abolished its activity also in A segments, suggesting that some of the same sites were bound by Ubx/AbdA. We now have evidence that the regulation of col expression by Ubx/AbdA in muscle progenitors is direct and involves a single Hox binding site. However, regulation by Antp does require other cis elements. It remains to be seen whether regulation by Antp is also direct. Since Antp, Ubx and AbdA display indistinguishable DNA-binding preferences in vitro (Ekker, 1994), the modular regulation of col expression by different Hox paralogs suggests that other cis elements and/or Hox collaborators contribute to Hox specificity (Mann et al., 2009). Direct regulation of col by Ubx has previously been documented in another cellular context, that of the larval imaginal haltere disc, via a wing-specific enhancer (Hersh and Carroll, 2005). In this case, Ubx directly represses col expression by binding to several sites, contrasting with col-positive regulation via a single site in muscle progenitors. This is the second example, in addition to CG13222 regulation in the haltere disc, of direct positive regulation by Ubx via a single binding site (Hersh et al., 2007). Hox ‘selector’ proteins collaborate on some cis elements with ‘effector’ transcription factors that are downstream of cell-cell signaling pathways (Grienenberger et al., 2003; Walsh and Carroll, 2007; Mann et al., 2009). In the DA3 lineage, it seems that Dpp, Wg and Ras signaling act on one col cis element and the Hox proteins on others. The regulation of col expression by Hox proteins in different tissues via different CRMs provides a new paradigm to decipher how different Hox paralogs cooperate and/or collaborate with tissue- and lineage-specific factors to specify cellular identity (Brodu et al., 2004; Gebelein et al., 2004; Walsh and Carroll, 2007; Stobe et al., 2009).
Hox proteins control the number of myoblasts allocated to each muscle
The DA3 muscle displays fewer nuclei in T2 and T3 than in A1-A7, an opposite situation to that described for an aggregate of the four LT1-4 muscles. Capovilla et al. proposed that the variation in the number of LT1-4 nuclei was controlled by Hox proteins (Capovilla et al., 2001). Our studies of the DA3 muscle extend this conclusion by showing that the variations due to Hox control are specific to each muscle and are exerted at the level of FCs. Since the number of nuclei is both muscle- and segment-specific, Hox proteins must cooperate and/or collaborate with various iTFs to differentially regulate the nucleus-counting process. As such, Hox proteins contribute to the combinatorial code of muscle identity. Identifying the nature of the cellular events and genes that act downstream of the iTF/Hox combinatorial code and that are involved in the nucleus-counting process represents a new challenge.
Supplementary Material
Acknowledgements
We thank the Bloomington Stock Center, S. Abmayr, S. Menon, E. Sanchez-Herrero and F. Karch for fly stocks; B. Patterson and E. Furlong for antibodies; F. Karch and members of our laboratory for critical reading of the manuscript; and E. Furlong for communicating unpublished data. We acknowledge the help of the Toulouse RIO Imaging Platform and B. Ronsin and A. Leru for confocal microscopy. This work was supported by CNRS and Ministère de la Recherche et de la Technologie (MRT), Université Paul Sabatier, Association Française contre les Myopathies (AFM) and the US National Institutes of Health. J.E. was supported by fellowships from MRT and AFM and H.B. by a fellowship from MRT. Deposited in PMC for release after 12 months.
Footnotes
Competing interests statement
The authors declare no competing financial interests.
Supplementary material
Supplementary material for this article is available at http://dev.biologists.org/lookup/suppl/doi:10.1242/dev.045286/-/DC1
References
- Abbott M. K. and Kaufman, T. C. (1986). The relationship between the functional complexity and the molecular organization of the Antennapedia locus of Drosophila melanogaster. Genetics 114, 919-942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Affolter M., Slattery, M. and Mann, R. S. (2008). A lexicon for homeodomain-DNA recognition. Cell 133, 1133-1135. [DOI] [PubMed] [Google Scholar]
- Alvares L. E., Schubert, F. R., Thorpe, C., Mootoosamy, R. C., Cheng, L., Parkyn, G., Lumsden, A. and Dietrich, S. (2003). Intrinsic, Hox-dependent cues determine the fate of skeletal muscle precursors. Dev. Cell 5, 379-390. [DOI] [PubMed] [Google Scholar]
- Balagopalan L., Keller, C. A. and Abmayr, S. M. (2001). Loss-of-function mutations reveal that the Drosophila nautilus gene is not essential for embryonic myogenesis or viability. Dev. Biol. 231, 374-382. [DOI] [PubMed] [Google Scholar]
- Barolo S., Carver, L. A. and Posakony, J. W. (2000). GFP and beta-galactosidase transformation vectors for promoter/enhancer analysis in Drosophila. Biotechniques 29, 726, 728, 730, 732. [DOI] [PubMed] [Google Scholar]
- Bate M. (1990). The embryonic development of larval muscles in Drosophila. Development 110, 791-804. [DOI] [PubMed] [Google Scholar]
- Bate M. (1993). The mesoderm and its derivatives. In The Development of Drosophila melanogaster, vol. 2 (ed. M. Bate and A. Martinez-Arias), p. 1013-1090. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press. [Google Scholar]
- Bate M. and Rushton, E. (1993). Myogenesis and muscle patterning in Drosophila. C. R. Acad. Sci. III 316, 1047-1061. [PubMed] [Google Scholar]
- Baylies M. K., Bate, M. and Ruiz Gomez, M. (1998). Myogenesis: a view from Drosophila. Cell 93, 921-927. [DOI] [PubMed] [Google Scholar]
- Bischof J., Maeda, R. K., Hediger, M., Karch, F. and Basler, K. (2007). An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc. Natl. Acad. Sci. USA 104, 3312-3317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourgouin C., Lundgren, S. E. and Thomas, J. B. (1992). Apterous is a Drosophila LIM domain gene required for the development of a subset of embryonic muscles. Neuron 9, 549-561. [DOI] [PubMed] [Google Scholar]
- Brand A. H. and Perrimon, N. (1993). Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401-415. [DOI] [PubMed] [Google Scholar]
- Brodu V., Elstob, P. R. and Gould, A. P. (2004). EGF receptor signaling regulates pulses of cell delamination from the Drosophila ectoderm. Dev. Cell 7, 885-895. [DOI] [PubMed] [Google Scholar]
- Capovilla M., Kambris, Z. and Botas, J. (2001). Direct regulation of the muscle-identity gene apterous by a Hox protein in the somatic mesoderm. Development 128, 1221-1230. [DOI] [PubMed] [Google Scholar]
- Carmena A., Bate, M. and Jimenez, F. (1995). Lethal of scute, a proneural gene, participates in the specification of muscle progenitors during Drosophila embryogenesis. Genes Dev. 9, 2373-2383. [DOI] [PubMed] [Google Scholar]
- Carmena A., Gisselbrecht, S., Harrison, J., Jimenez, F. and Michelson, A. M. (1998). Combinatorial signaling codes for the progressive determination of cell fates in the Drosophila embryonic mesoderm. Genes Dev. 12, 3910-3922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crozatier M. and Vincent, A. (1999). Requirement for the Drosophila COE transcription factor Collier in formation of an embryonic muscle: transcriptional response to notch signalling. Development 126, 1495-1504. [DOI] [PubMed] [Google Scholar]
- Crozatier M. and Vincent, A. (2008). Control of multidendritic neuron differentiation in Drosophila: the role of Collier. Dev. Biol. 315, 232-242. [DOI] [PubMed] [Google Scholar]
- Crozatier M., Valle, D., Dubois, L., Ibnsouda, S. and Vincent, A. (1996). collier, a novel regulator of Drosophila head-development, is expressed in a single mitotic domain. Curr. Biol. 6, 707-718. [DOI] [PubMed] [Google Scholar]
- Crozatier M., Ubeda, J. M., Vincent, A. and Meister, M. (2004). Cellular immune response to parasitization in Drosophila requires the EBF orthologue collier. PLoS Biol. 2, E196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demontis F. and Perrimon, N. (2009). Integration of Insulin receptor/Foxo signaling and dMyc activity during muscle growth regulates body size in Drosophila. Development 136, 983-993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubois L., Enriquez, J., Daburon, V., Crozet, F., Lebreton, G., Crozatier, M. and Vincent, A. (2007). Collier transcription in a single Drosophila muscle lineage: the combinatorial control of muscle identity. Development 134, 4347-4355. [DOI] [PubMed] [Google Scholar]
- Duboule D. and Morata, G. (1994). Colinearity and functional hierarchy among genes of the homeotic complexes. Trends Genet. 10, 358-364. [DOI] [PubMed] [Google Scholar]
- Ekker S. C., Jackson, D. G., von Kessler, D. P., Sun, B. I., Young, K. E. and Beachy, P. A. (1994). The degree of variation in DNA sequence recognition among four Drosophila homeotic proteins. EMBO J. 13, 3551-3560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Estrada B., Choe, S. E., Gisselbrecht, S. S., Michaud, S., Raj, L., Busser, B. W., Halfon, M. S., Church, G. M. and Michelson, A. M. (2006). An integrated strategy for analyzing the unique developmental programs of different myoblast subtypes. PLoS Genet. 2, e16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujioka M., Wessells, R. J., Han, Z., Liu, J., Fitzgerald, K., Yusibova, G. L., Zamora, M., Ruiz-Lozano, P., Bodmer, R. and Jaynes, J. B. (2005). Embryonic even skipped-dependent muscle and heart cell fates are required for normal adult activity, heart function, and lifespan. Circ. Res. 97, 1108-1114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gebelein B., McKay, D. J. and Mann, R. S. (2004). Direct integration of Hox and segmentation gene inputs during Drosophila development. Nature 431, 653-659. [DOI] [PubMed] [Google Scholar]
- Greig S. and Akam, M. (1993). Homeotic genes autonomously specify one aspect of pattern in the Drosophila mesoderm. Nature 362, 630-632. [DOI] [PubMed] [Google Scholar]
- Grienenberger A., Merabet, S., Manak, J., Iltis, I., Fabre, A., Berenger, H., Scott, M. P., Pradel, J. and Graba, Y. (2003). Tgfbeta signaling acts on a Hox response element to confer specificity and diversity to Hox protein function. Development 130, 5445-5455. [DOI] [PubMed] [Google Scholar]
- Halfon M. S., Carmena, A., Gisselbrecht, S., Sackerson, C. M., Jimenez, F., Baylies, M. K. and Michelson, A. M. (2000). Ras pathway specificity is determined by the integration of multiple signal-activated and tissue-restricted transcription factors. Cell 103, 63-74. [DOI] [PubMed] [Google Scholar]
- Hersh B. M. and Carroll, S. B. (2005). Direct regulation of knot gene expression by Ultrabithorax and the evolution of cis-regulatory elements in Drosophila. Development 132, 1567-1577. [DOI] [PubMed] [Google Scholar]
- Hersh B. M., Nelson, C. E., Stoll, S. J., Norton, J. E., Albert, T. J. and Carroll, S. B. (2007). The UBX-regulated network in the haltere imaginal disc of D. melanogaster. Dev. Biol. 302, 717-727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heuer J. G., Li, K. and Kaufman, T. C. (1995). The Drosophila homeotic target gene centrosomin (cnn) encodes a novel centrosomal protein with leucine zippers and maps to a genomic region required for midgut morphogenesis. Development 121, 3861-3876. [DOI] [PubMed] [Google Scholar]
- Hooper J. E. (1986). Homeotic gene function in the muscles of Drosophila larvae. EMBO J. 5, 2321-2329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hueber S. D. and Lohmann, I. (2008). Shaping segments: Hox gene function in the genomic age. BioEssays 30, 965-979. [DOI] [PubMed] [Google Scholar]
- Jagla T., Bellard, F., Lutz, Y., Dretzen, G., Bellard, M. and Jagla, K. (1998). ladybird determines cell fate decisions during diversification of Drosophila somatic muscles. Development 125, 3699-3708. [DOI] [PubMed] [Google Scholar]
- Keller C. A., Erickson, M. S. and Abmayr, S. M. (1997). Misexpression of nautilus induces myogenesis in cardioblasts and alters the pattern of somatic muscle fibers. Dev. Biol. 181, 197-212. [DOI] [PubMed] [Google Scholar]
- Knirr S. and Frasch, M. (2001). Molecular integration of inductive and mesoderm-intrinsic inputs governs even-skipped enhancer activity in a subset of pericardial and dorsal muscle progenitors. Dev. Biol. 238, 13-26. [DOI] [PubMed] [Google Scholar]
- Knirr S., Azpiazu, N. and Frasch, M. (1999). The role of the NK-homeobox gene slouch (S59) in somatic muscle patterning. Development 126, 4525-4535. [DOI] [PubMed] [Google Scholar]
- Kocherlakota K. S., Wu, J. M., McDermott, J. and Abmayr, S. M. (2008). Analysis of the cell adhesion molecule sticks-and-stones reveals multiple redundant functional domains, protein-interaction motifs and phosphorylated tyrosines that direct myoblast fusion in Drosophila melanogaster. Genetics 178, 1371-1383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y. H., Jakobsen, J. S., Valentin, G., Amarantos, I., Gilmour, D. T. and Furlong, E. E. (2009). A systematic analysis of Tinman function reveals Eya and JAK-STAT signaling as essential regulators of muscle development. Dev. Cell 16, 280-291. [DOI] [PubMed] [Google Scholar]
- Mann R. S., Lelli, K. M. and Joshi, R. (2009). Hox specificity unique roles for cofactors and collaborators. Curr. Top. Dev. Biol. 88, 63-101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Menon S. D. and Chia, W. (2001). Drosophila rolling pebbles: a multidomain protein required for myoblast fusion that recruits D-Titin in response to the myoblast attractant Dumbfounded. Dev. Cell 1, 691-703. [DOI] [PubMed] [Google Scholar]
- Michelson A. M. (1994). Muscle pattern diversification in Drosophila is determined by the autonomous function of homeotic genes in the embryonic mesoderm. Development 120, 755-768. [DOI] [PubMed] [Google Scholar]
- Michelson A. M., Abmayr, S. M., Bate, M., Arias, A. M. and Maniatis, T. (1990). Expression of a MyoD family member prefigures muscle pattern in Drosophila embryos. Genes Dev. 4, 2086-2097. [DOI] [PubMed] [Google Scholar]
- Nose A., Isshiki, T. and Takeichi, M. (1998). Regional specification of muscle progenitors in Drosophila: the role of the msh homeobox gene. Development 125, 215-223. [DOI] [PubMed] [Google Scholar]
- Philippakis A. A., Busser, B. W., Gisselbrecht, S. S., He, F. S., Estrada, B., Michelson, A. M. and Bulyk, M. L. (2006). Expression-guided in silico evaluation of candidate cis regulatory codes for Drosophila muscle founder cells. PLoS Comput. Biol. 2, e53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz Gomez M. and Bate, M. (1997). Segregation of myogenic lineages in Drosophila requires numb. Development 124, 4857-4866. [DOI] [PubMed] [Google Scholar]
- Ruiz-Gomez M., Romani, S., Hartmann, C., Jackle, H. and Bate, M. (1997). Specific muscle identities are regulated by Kruppel during Drosophila embryogenesis. Development 124, 3407-3414. [DOI] [PubMed] [Google Scholar]
- Sandmann T., Jensen, L. J., Jakobsen, J. S., Karzynski, M. M., Eichenlaub, M. P., Bork, P. and Furlong, E. E. (2006). A temporal map of transcription factor activity: mef2 directly regulates target genes at all stages of muscle development. Dev. Cell 10, 797-807. [DOI] [PubMed] [Google Scholar]
- Sandmann T., Girardot, C., Brehme, M., Tongprasit, W., Stolc, V. and Furlong, E. E. (2007). A core transcriptional network for early mesoderm development in Drosophila melanogaster. Genes Dev. 21, 436-449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Speicher S., Fischer, A., Knoblich, J. and Carmena, A. (2008). The PDZ protein Canoe regulates the asymmetric division of Drosophila neuroblasts and muscle progenitors. Curr. Biol. 18, 831-837. [DOI] [PubMed] [Google Scholar]
- Stobe P., Stein, M. A., Habring-Muller, A., Bezdan, D., Fuchs, A. L., Hueber, S. D., Wu, H. and Lohmann, I. (2009). Multifactorial regulation of a hox target gene. PLoS Genet. 5, e1000412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vervoort M., Crozatier, M., Valle, D. and Vincent, A. (1999). The COE transcription factor Collier is a mediator of short-range Hedgehog-induced patterning of the Drosophila wing. Curr. Biol. 9, 632-639. [DOI] [PubMed] [Google Scholar]
- Walsh C. M. and Carroll, S. B. (2007). Collaboration between Smads and a Hox protein in target gene repression. Development 134, 3585-3592. [DOI] [PubMed] [Google Scholar]
- Zinzen R. P., Girardot, C., Gagneur, J., Braun, M. and Furlong, E. E. (2009). Combinatorial binding predicts spatio-temporal cis-regulatory activity. Nature 462, 65-70. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.