Abstract
The multiprotein replisome complex that replicates DNA, has been extensively characterized in vitro, but its composition and architecture in vivo is unknown. Using millisecond single molecule fluorescence microscopy in living cells expressing YPet derivatives of replisome components, we have examined replisome stoichiometry and architecture. Active Escherichia coli replisomes contain three molecules of the replicative polymerase, rather than the historically accepted two. These are associated with three molecules of τ, a clamp loader component that trimerizes polymerase. Only two of the three sliding clamps are always associated with the core replisome. Single strand binding protein has a broader spatial distribution than the core components, with five to eleven tetramers per replisome. This in vivo technique could provide single molecule insight into other molecular machines.
Replisomes are dynamic multiprotein machines that replicate DNA by copying the leading strand template continuously and the lagging strand template discontinuously. In E. colithe replisome couples activities of more than 11 proteins during genome replication (1, 2). The DnaB helicase, loaded onto the lagging strand template, separates the two templates that are subsequently copied by PolIII polymerase ([αεθ). PolIII processivity results from binding to a sliding clamp (β) encircling duplex DNA; sliding clamps are added and removed by a clamp loader ([τ/γ]3δδ′ψχ) whose τ component oligomerizes PolIII. Unwound DNA on the lagging template strand is bound by single strand binding protein (Ssb) tetramers that remove DNA secondary structure and protect against nucleases. Primase binds to helicase during cycles of priming and DNA synthesis on the lagging strand template.
In vitro studies of the replisome have yielded details of replisome organization and replication mechanism, but have not revealed how replication is organized within living cells: additional copies of known replisome components may be present at replication forks, while additional cellular factors, absent from in vitro assays, may modify the composition of the replisome and act in DNA processing. Furthermore, the techniques used to determine stoichiometry and architecture in vitro favor strong interactions, potentially biasing estimates on numbers and interactions, and are subject to complications if any component is proteolytically sensitive. We thus investigated active replisome architecture in living cells by using a fluorescence microscopy protocol with single molecule sensitivity and millisecond temporal resolution. Using fully functional fluorescent C- or N-terminal YPet derivatives of E. coli replisome components expressed from their endogenous promoters, we showed previously that the two spatially separable sister replisomes derived from a single initiation event at the replication origin, oriC, track independently along DNA (3). Using these and additional fusions, 10 components of individual replisomes were analyzed using ‘slimfield’ fluorescence microscopy, which uses a compact Gaussian laser excitation field (~30 μm2) that encompasses single cells with an excitation intensity ~100 times greater than in widefield fluorescence (4, 5). This imaging allows quantitative detection of single fluorescent molecules at 3 ms capture rates (Fig. 1A, B). The high laser excitation intensity does not abolish DNA replication, as judged by a 2-fold difference in turnover on DNA of Ssb-YPet in replicating and non-replicating cells exposed to an excitation intensity and duration similar to that of slimfield (fig. S1), consistent with the demonstration that similar exposures do not inhibit flagellar rotation (4).
Fig. 1.
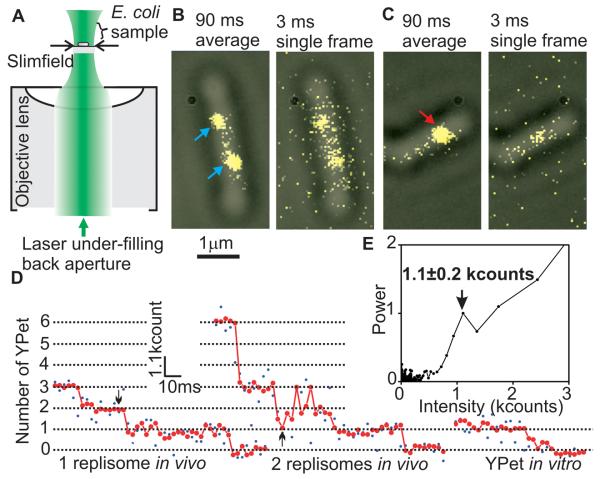
Slimfield microscopy and photobleach analysis. (A) Slimfield schematic; a laser under-fills the back aperture of an objective lens generating an intense Gaussian field at the sample large enough to image single E. coli. (B) Overlaid brightfield (gray) and 90 ms frame-averaged fluorescence images (yellow) of ε-YPet strain, arrows indicating spots with a stoichiometry of ~3 (cyan) and (C) ~6 (red) ε-YPet molecules, with corresponding single 3 ms frame taken after 45 ms showing that stochastic photobleaching generates different brightnesses. (D) Raw intensity (blue) and filtered (red) for a putative single (left panel) and double (right panel) replisome spot, 45 ms point indicated (arrows), with surface-immobilized YPet in vitro. (E) Fourier spectral analysis for a photobleach trace of the ε-YPet strain with mean±SD peak indicated for brightness of a single YPet.
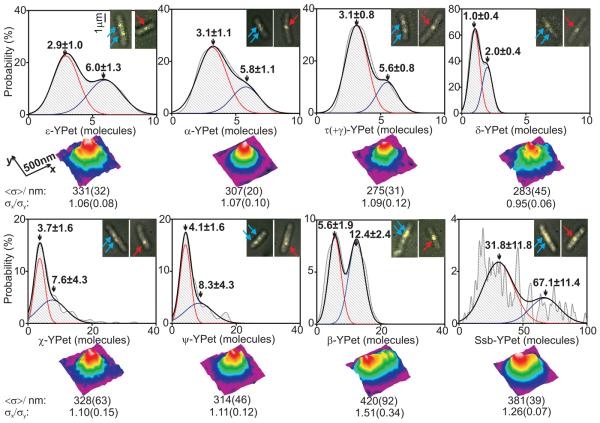
For estimating stoichiometry, image frames were averaged over 90 ms to define ‘regions of interest’ that are hotspots for localization of a given YPet fusion protein. The position, size, shape and intensity of the spots were measured automatically for each individual image frame, generating step-like intensity traces as photobleaching occurred (Fig. 1D, E). We measured the step spacing using an edge-preserving filter (6, 7) combined with Fourier spectral analysis and compared these with intensity traces from purified YPet in vitro (8-10), indicating that the in vivo steps were approximately integer multiples of the in vitro intensity of a single YPet molecule (4, 5, 8-13). Thus intensities prior to photobleaching enabled single molecule stoichiometry determination (5). For most replisome components, we observed bimodal 1:2 distributions of stoichiometries, reflecting the observation that ~75% of cells contain two spatially separated replisomes, each associated with independent forks, while ~25% have sister replisomes separated by a distance smaller than the diffraction limit of our system (~250 nm), thus observed as a single spot (3). Lower stoichiometries were always associated with spots from cells containing two spots and two-fold higher stoichiometries from cells with a single spot (Fig. 2). With the ε-YPet strain we observed a bimodal distribution with two Gaussians, peaks centered on 2.9 ± 1.0 (mean ± SD, SEM ~0.2) and 6.0 ± 1.3 molecules, consistent with a model in which a single fork contains three copies of ε, the proofreading exonuclease of the core polymerase. The PolIII catalytic subunit, α, had a bimodal distribution with peaks centered on 3.1 ± 1.1 and 5.8 ± 1.1 molecules. Similarly, the clamp loader component, τ, which also oligomerizes the polymerase, had peaks at 3.1 ± 0.8 and 5.6 ± 0.8 molecules. δ, an essential component of the clamp loader, expected to be present in one copy, had peaks at 1.0 ± 0.4 and 2.0 ± 0.4 molecules. Furthermore, we observed a stoichiometry of ~6 and ~12 molecules for DnaB (fig. S2), as expected for the hexameric helicase.
Fig. 2.
Stoichiometries of replisome components and spatial distributions. Stoichiometry distributions per spot (upper panels) using unbiased kernel density estimation for different E. coli strains, N=27-51 cells used in each dataset. Insets indicate examples of overlaid brightfield (gray) and single 3ms fluorescence images (yellow) for each, arrows indicating foci in cells containing two (cyan) and one (red) replisome. 2-Gaussian fits (black) with contributing single Gaussian curves (red and blue) and mean±SD of Gaussian peaks indicated. Lower panels indicate false-color contour plots for 2D averaged spatial distributions for each strain, N=42-151 spots in each dataset. Estimates for mean FWHM <σ> of a symmetrical 2D Gaussian fit and the ratio σx/σy of the FWHM for the 1D Gaussian fits through the mean spot parallel to the x and y axes are indicated, SD errors in brackets.
Non-essential χ and ψ, which heterodimerize, have been reported to be present as one copy per replisome, due to ψ binding to γ/τ in the clamp loader (1, 14). Instead, we found a mean of ~4 copies per single replisome spot (Fig. 2), with single spot cells having ~8 molecules per spot, with a broader distribution of stoichiometries than for other low copy components. We propose this results from one χψ heterodimer being tethered to the clamp loader, while the other χψ dimers bind available C-terminal tails of the same or different Ssb tetramers by a characterized interaction through χ (14). The intensity of ψ-YPet foci is greatly reduced when χ is absent, but χ focus intensity is unchanged when ψ is absent, supporting this hypothesis (fig. S3).
dnaX expresses comparable amounts of τ and a truncated form, γ, formed by a programmed frameshift (15). τ and γ can interchangeably be clamp loader constituents, but only τ oligomerizes PolIII (1). Our demonstration that three copies of τ are associated with the replisome, suggests that γ is not associated with the single clamp loader in most replisomes. To test this, we both constructed a strain, dnaX(γ−), that did not express γ, because the frameshift was abrogated, and a strain, dnaX(γ-YPet), that expressed γ-YPet. The strain that failed to express γ grew well, confirming that γ is non-essential (16), while the strain expressing γ–YPet showed fluorescent replisome-associated foci (fig. S4A). Since γ can interact with χψ, we considered whether γ might be Ssb-associated via a linking interaction with the χψ heterodimers not associated with the clamp loader (Fig. 3). Targeted proteolysis of degron-tagged ψ, or deletion of χ or ψ, led to a loss of replisome-associated γ–YPet foci, but not ε-YPet or τ-YPet foci; remaining γ–YPet foci were of reduced intensity (fig. S4B). Thus the single replicative clamp loader in each replisome contains τ but not γ, and the clamp loader-independent copies of χψ associated with Ssb are likely to recruit γ to Ssb, in addition to acting in primase release (17). We propose that γ may replace τ as a clamp loader component in post-replication repair-associated events at the replication fork. Comparison of τ stoichiometry in dnaX and dnaX(γ−) strains, showed a ~30% increase when γ was absent (fig. S5), indicating that γ and τ can compete for binding to Ssb.
Fig. 3.
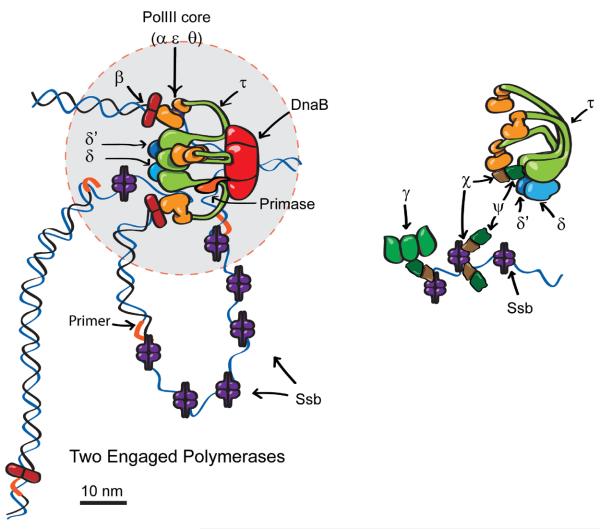
Schematic model for replisome components. (A) Two engaged polymerases, and one of the three β clamps at a distance from the core replisome (circle of diameter 50 nm indicated in gray). The data indicate that ~75% of replisomes have this organization, while ~25% have all three β clamps associated with the core replisome and potentially associated with active PolIII. (B) Expanded view of clamp loader (τ3δδ′ψχ) and three additional molecules of χψ interacting with Ssb tails. The χψ heterodimer bound to the clamp loader may also contact Ssb (14). γ (shown as a trimer, but the stoichiometry is unknown) then interacts with Ssb-associated χψ.
In contrast to the structural skeleton components of the replisome, Ssb showed a broad distribution of stoichiometries (Fig. 2) with a periodicity of ~4 molecules (fig. S6) and a mean of 31.8 ± 11.1 molecules per spot for cells containing two spots per cell, consistent with 8 ± 3 Ssb tetramers per replication fork. Single spot cells had a stoichiometry larger by a factor of roughly two at ~70 molecules per spot. The number of Ssb molecules bound at the replication fork is expected to be proportional to the length of ssDNA. The average stoichiometry of Ssb within replisomes approximately doubled in cells treated with hydroxyurea (5), as expected, since ssDNA accumulates at the fork when replication is stalled (18). The ssDNA associated with each Ssb tetramer is either 35 or 65 nt in vitro; (Ssb)35 forms compact filaments, while (Ssb)65 forms dimers of tetramers covering 170 nt (19). The contribution of each binding mode in vivo is unknown. Assuming the stretch of ssDNA at the replication fork equals the Okazaki fragment length (~650 nt at 22 °C (19)) then the presence of (Ssb)65 would give an occupancy of ~8 tetramers, close to our mean stoichiometry estimate for single replisome spots (Fig. 2).
Structural investigations of replisome components suggest that their cumulative volume is contained in a sphere of maximum diameter ~50 nm (20-23). To investigate this, we studied size and shape of images from each strain (Fig. 2, lower panels). Strains expressing fluorescent α, ε, τ, δ, χ and ψ all produced spots with a circularly symmetrical shape and a mean full-width at half maximum (FWHM) across the cells strains of 305 ± 30 nm (5). There was no significant difference to the individual FWHM measurements for each strain (p=0.05; Student t-test), but all were significantly larger than the FWHM of surface-immobilized YPet in vitro of ~250 nm (fig. S7). In contrast, β and Ssb produced spots spread further over the long axis of the cell, Ssb foci extending up to ~200 nm, with ~55% of the tetramers being present within a 50 nm diameter. Although the ~4 χψ heterodimers may interact with Ssb tails, we note that the spatial distribution of χψ is that of the core replisome-clamp loader rather than that of Ssb. Furthermore, the stoichiometry of χψ is much lower than that of Ssb. Taken together these results suggest that χψ is associated preferentially with one or a few Ssb molecules in the vicinity of the core replisome. The stoichiometry of the dimeric β sliding clamp was expected to be partly the result of its interaction with PolIII and the clamp loader, and partly due to association with DNA at the 3′ end of Okazaki fragments. Nevertheless, its bimodal distribution peaked at ~3 and ~6 dimers (Fig. 2), which raised the possibility that it might be continually associated with PolIII. Analysis of β-YPet spots indicated two subpopulations with different spatial intensity distributions (fig. S8). The first subgroup (~27% of spots) had circular distributions that support a scheme in which three β dimers are associated with active PolIII or with the clamp loader. In the second subgroup, the spots were more extended (circularity >1.2), consistent with at least one of the sliding clamps being localized at a distance >50 nm from the replisome core. Historically, the replisome has been considered to have two opposing yet coordinated polymerases connected to the rest of the replisome (24). Instead, we provide strong evidence for an in vivo core replisome containing three PolIIIs associated with a clamp loader whose three copies of τ also trimerize PolIII. The results are consistent with two of the three PolIIIs being dedicated for use during lagging strand synthesis, while one is the highly processive leading strand polymerase. In a minority of replisomes, all three PolIIIs may be associated with sliding clamps, with two being potentially simultaneously active on the lagging strand, supporting suggestions from in vitro studies of E. coli and phage T4 replisomes (2, 25). Nevertheless, the majority of replisomes appear to have only two of the polymerases associated with a sliding clamp, suggesting that the third polymerase is waiting to be loaded onto the next lagging strand primer (Fig. 3). We measured the number of non-replisome Ssb tetramers to be 330±105 per cell (table S1), in agreement to estimates using quantitative westerns (5). Similarly, 0.5-5% of other replisome molecules in a cell were associated with each replisome (table S1), showing that the measured stoichiometries reflect biologically relevant complexes, and are not the consequence of the replisome's component being rate limiting. The level of replisome molecules is sufficient to support more than 10 cellular replication forks associated with the chromosome, plasmids or phage. Slimfield microscopy provides a powerful non-invasive in vivo analytical tool that extends previous analyses of the assembly, and action of molecular machines (10, 26, 27). In combination with degron-targeted proteolysis of specific proteins, it has provided unanticipated insight into replisome architecture. In combination with partial pre-bleaching and stochastic photoactivation and photoswitching techniques, the methodology may provide new insight into biological systems that contain substantially higher numbers of freely diffusing fluorescent proteins than were investigated here.
Supplementary Material
Acknowledgements
The work was supported by the Wellcome Trust and the Royal Society. M.L. is a Royal Society University Fellow and research fellow of Hertford College Oxford. R. R-L is a research fellow of New College Oxford and was supported by Conacyt and a Clarendon post-graduate award. We thank C. Possoz and C. Lesterlin for stimulating discussions and acknowledge NBRP (NIG, Japan): E. coli for the provision of the holC and holD deletion strains.
Footnotes
This manuscript has been accepted for publication in Science. This version has not undergone final editing. Please refer to the complete version of record at http://www.sciencemag.org/. The manuscript may not be reproduced or used in any manner that does not fall within the fair use provisions of the Copyright Act without the prior, written permission of AAAS.
References and notes
- 1.Johnson A, O'Donnell M. Annu Rev Biochem. 2005;74:283. doi: 10.1146/annurev.biochem.73.011303.073859. [DOI] [PubMed] [Google Scholar]
- 2.McInerney P, Johnson A, Katz F, O'Donnell M. Mol Cell. 2007;27:527. doi: 10.1016/j.molcel.2007.06.019. [DOI] [PubMed] [Google Scholar]
- 3.Reyes-Lamothe R, Possoz C, Danilova O, Sherratt DJ. Cell. 2008;133:90. doi: 10.1016/j.cell.2008.01.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Plank M, Wadhams GH, Leake MC. Integr. Biol. 2009;1:602. doi: 10.1039/b907837a. [DOI] [PubMed] [Google Scholar]
- 5.Materials and methods are available as supporting material on Science Online.
- 6.Chung SH, Kennedy RA. J. Neurosci. Methods. 1991;40:71. doi: 10.1016/0165-0270(91)90118-j. [DOI] [PubMed] [Google Scholar]
- 7.Smith DA. Phil. Trans. R. Soc. Lond. B. 1998;353:1969. doi: 10.1098/rstb.1998.0348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Leake MC, Wilson D, Bullard B, Simmons RM. FEBS Lett. 2003;535:55. doi: 10.1016/s0014-5793(02)03857-7. [DOI] [PubMed] [Google Scholar]
- 9.Leake MC, Wilson D, Gautel M, Simmons RM. Biophys J. 2004;87:1112. doi: 10.1529/biophysj.103.033571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Leake MC, et al. Nature. 2006;443:355. doi: 10.1038/nature05135. [DOI] [PubMed] [Google Scholar]
- 11.Leake MC, et al. Proc Natl Acad Sci U S A. 2008;105:15376. doi: 10.1073/pnas.0806338105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lenn T, Leake MC, Mullineaux CW. Mol Microbiol. 2008;70:1397. doi: 10.1111/j.1365-2958.2008.06486.x. [DOI] [PubMed] [Google Scholar]
- 13.Lenn T, Leake MC, Mullineaux CW. Biochem Soc Trans. 2008;36:1032. doi: 10.1042/BST0361032. [DOI] [PubMed] [Google Scholar]
- 14.Simonetta KR, et al. Cell. 2009;137:659. doi: 10.1016/j.cell.2009.03.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Blinkova AL, Walker JR. Nucleic Acids Res. 1990;18:1725. doi: 10.1093/nar/18.7.1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Blinkova A, et al. J Bacteriol. 1993;175:6018. doi: 10.1128/jb.175.18.6018-6027.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yuzhakov A, Kelman Z, O'Donnell M. Cell. 1999;96:153. doi: 10.1016/s0092-8674(00)80968-x. [DOI] [PubMed] [Google Scholar]
- 18.McInerney P, O'Donnell M. J Biol Chem. 2007;282:25903. doi: 10.1074/jbc.M703777200. [DOI] [PubMed] [Google Scholar]
- 19.Lohman TM, Ferrari ME. Annu Rev Biochem. 1994;63:527. doi: 10.1146/annurev.bi.63.070194.002523. [DOI] [PubMed] [Google Scholar]
- 20.Kazmirski SL, Podobnik M, Weitze TF, O'Donnell M, Kuriyan J. Proc Natl Acad Sci U S A. 2004;101:16750. doi: 10.1073/pnas.0407904101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lamers MH, Georgescu RE, Lee SG, O'Donnell M, Kuriyan J. Cell. 2006;126:881. doi: 10.1016/j.cell.2006.07.028. [DOI] [PubMed] [Google Scholar]
- 22.Georgescu RE, et al. Cell. 2008;132:43. doi: 10.1016/j.cell.2007.11.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bailey S, Eliason WK, Steitz TA. Science. 2007;318:459. doi: 10.1126/science.1147353. [DOI] [PubMed] [Google Scholar]
- 24.Alberts BM, et al. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):655. doi: 10.1101/sqb.1983.047.01.077. [DOI] [PubMed] [Google Scholar]
- 25.Nossal NG, Makhov AM, Chastain PD, 2nd, Jones CE, Griffith JD. J Biol Chem. 2007;282:1098. doi: 10.1074/jbc.M606772200. [DOI] [PubMed] [Google Scholar]
- 26.Xie XS, Yu J, Yang WY. Science. 2006;312:228. doi: 10.1126/science.1127566. [DOI] [PubMed] [Google Scholar]
- 27.Wu JQ, Pollard TD. Science. 2005;310:310. doi: 10.1126/science.1113230. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.