Abstract
Persons of African ancestry, on average, have lower white blood cell (WBC) counts than those of European descent (ethnic leukopenia), but whether this impacts negatively on HIV-1 disease course remains unknown. Here, in a large natural history cohort of HIV-infected subjects, we show that, although leukopenia (< 4000 WBC/mm3 during infection) was associated with an accelerated HIV disease course, this effect was more prominent in leukopenic subjects of European than African ancestry. The African-specific −46C/C genotype of Duffy Antigen Receptor for Chemokines (DARC) confers the malaria-resisting, Duffy-null phenotype, and we found that the recently described association of this genotype with ethnic leukopenia extends to HIV-infected African Americans (AAs). The association of Duffy-null status with HIV disease course differed according to WBC but not CD4+ T-cell counts, such that leukopenic but not nonleukopenic HIV+ AAs with DARC −46C/C had a survival advantage compared with all Duffy-positive subjects. This survival advantage became increasingly pronounced in those with progressively lower WBC counts. These data highlight that the interaction between DARC genotype and the cellular milieu defined by WBC counts may influence HIV disease course, and this may provide a partial explanation of why ethnic leukopenia remains benign in HIV-infected AAs, despite immunodeficiency.
Introduction
Leukopenia is observed frequently during HIV-1 infection.1–3 For example, evaluation of a large number of incarcerated adults revealed that leukopenia correlated strongly with HIV seropositivity, independent of other variables, such as sex with an HIV-positive (HIV+) partner, injection drug use, ethnicity, and presence of sexually transmitted diseases.4,5 Interestingly, in these studies, the risk for HIV seropositivity associated with leukopenia and a history of sex with an HIV+ partner were similarly high,4 underscoring that leukopenia is strongly associated with HIV positivity in at-risk persons and is commonly observed soon after HIV infection. Of note, although the sine qua non feature of infection with HIV is progressive immunodeficiency related to CD4+ T-cell lymphopenia,6 CD4+ T-cell counts contribute minimally in quantitative terms to the overall white blood cell (WBC) count and their decline during the course HIV infection contributes minimally to this observed leukopenia, which is mainly attributable to neutropenia.
However, the impact of leukopenia on HIV disease course is largely undefined. Most of the studies that have examined the frequency or HIV disease-influencing effects of cytopenias have used cross-sectional study designs (eg, HIV+ vs HIV−1 or HIV+ neutropenics vs HIV+ nonneutropenics7). These studies have documented that the prevalence of cytopenias is higher in advanced disease. For example, neutropenia ranges from 0.8% to 13.4% when CD4+ counts are less than 250 cells/mm3 and from 13% to 44% in those with AIDS.1,8 By contrast, there are no prospective studies in natural history cohorts that have examined whether leukopenia impacts on HIV disease course, independent of the known strong relationship between immunodeficiency (reflected by low CD4+ counts or high viral loads) and either leukopenia or neutropenia.1,2,9
To address this, we determined whether leukopenia impacts on the HIV disease course of subjects in a natural history cohort.10–13 The large representation of both European Americans (EAs) and African Americans (AAs) in the study population allowed us to examine whether the impact of leukopenia on disease course differed according to race. This possibility is of particular interest because persons of African ancestry, on average, have significantly lower WBC counts, secondary to lower neutrophil cell counts, than persons of European descent.14–17 In otherwise healthy persons, low WBC counts in persons of African ancestry are thought to be both genetically determined and benign, as they have not been associated withan increased incidence of bacterial infection.14 This has resulted in the designation of this condition as “benign ethnic leukopenia or neutropenia.”14,16,17
In light of the preceding discussion, we considered whether differences in WBC counts also result in racial differences in HIV disease outcome. Such an analysis could be confounded by social factors (eg, access to health care, socioeconomic status). However, our studies were conducted in a well-characterized natural history cohort of HIV infection in which several factors that may confound assessment of race-specific differences in disease progression were minimized (eg, equal access to health care, similar living standards, and minimal loss to follow-up). In our study cohort, the HIV+ AAs had a slight survival advantage compared with HIV+ EAs.10 This finding was consistent with the survival advantage observed in the HIV-infected AAs in a much larger segment of the same cohort of HIV+ subjects, of which the group that we studied was a subset.12 In view of the epidemiologic observation that ethnic leukopenia is “benign” in otherwise healthy AAs, a parsimonious explanation for the survival advantage observed in the overall group of HIV-infected AAs compared with EAs was that it may be the result of a subset of leukopenic AAs who had a benign disease course compared with their leukopenic HIV+ EA counterparts.
The results of our studies conducted herein affirmed this possibility. This finding and the previously ascribed genotype-phenotype relationships observed for Duffy Antigen Receptor for Chemokines (DARC), a decoy chemokine receptor,18 prompted us to examine whether the associations for DARC genotype in HIV-infected subjects may explain in part why leukopenic AAs had a survival advantage compared with leukopenic EAs. DARC is expressed mainly on the surface of red blood cells (RBCs) and endothelial cells, where it binds multiple chemokines that have relevance to both HIV pathogenesis (eg, RANTES and MCP-1) and neutrophil biology (eg, GROα).18,19 The −46T→C polymorphism is specific to persons of African ancestry,10,20–22 and homozygosity for the −46C allele (−46C/C genotype) abrogates DARC expression on erythrocytes.18–20 Because DARC on RBC serves as a receptor for the malarial protozoa Plasmodium vivax,19,20 Duffy-null status (DARC −46C/C genotype) on RBC is associated with a selective advantage against this form of malaria. Recent studies revealed 2 phenotypes associated with the DARC −46C/C genotype that had relevance to both leukopenia and HIV pathogenesis. First, admixture mapping and other genetic studies in large nonimmunosuppressed cohorts of EAs and AAs demonstrate that, over and above its strong link with African ancestry, DARC −46C/C is the main genetic basis for ethnic leukopenia21 and neutropenia22; other observational studies substantiate this relationship between DARC −46C/C genotype and leukopenia or neutropenia.23,24 Second, in our study population of HIV+ AAs, those with the DARC −46C/C genotype had a slower rate of HIV disease progression than those with DARC −T/T or C/T.10 The latter 2 findings provided the rationale for investigating further the relationships among leukopenia, DARC genotype, and HIV disease course. The results of these studies highlight that an interaction between DARC genotype and the endogenous environment as defined by WBC counts impacts on HIV disease course.
Methods
Study cohort
We used DARC genotyping data, which we reported on recently,10 derived from HIV+ subjects from the Department of Defense HIV Natural History Study cohort followed originally at Wilford Hall Medical Center and more recently at the Brooke Army Medical Center. The studied population is the local (San Antonio, TX) site of a prospective multisite observational cohort from the United States Military's HIV Natural History Study. Extensive details of the Wilford Hall Medical Center cohort have been provided elsewhere, and all subjects were followed prospectively from the early stages of their infection.10,11 The time from diagnosis to entry into the cohort was similar for subjects from different ethnic groups. The genetic and clinical data came from 1132 subjects representing a total of 7125.15 person-years of follow-up, in which a total of 440 (38.9%) deaths occurred. The cohort included 515 (45.5%) subjects with documented seroconversion, and these subjects are referred to here as seroconverters. The proportion of seroconverters across the major ethnic groups was similar (42.1% of EA, 48.4% of AA, and 42.0% of other ethnic groups). The clinical characteristics of the subjects are shown in supplemental Table 1 (available on the Blood website; see the Supplemental Materials link at the top of the online article). The research was approved by the Institutional Review Boards of Wilford Hall Medical Center and the University of Texas Health Science Center, San Antonio, TX.
Study outcomes
We classified the study subjects as those with leukopenia, that is, a low WBC count (WBClow) and those with a high WBC count (WBChigh) according to whether they maintained an average WBC count during HIV disease of less than or more than or equal to 4000 cells/mm3, respectively. There were a total of 26585 WBC count measurements on the study subjects during the HIV disease course (16032 measurements in EAs at a mean of 25.7 measurements per patient; 8266 measurements in AAs at a mean of 20.8 measurements per patient and 2287 measurements in other ethnicities at a mean of 22.9 measurements per patient). To assess the influence of genetic determinants on WBC counts during disease, we determined the mean of the total WBC counts measured during disease for each patient as that patient's average WBC count measured during disease. The frequency of assessment of WBC counts did not differ significantly according to racial/ethnic group or DARC −46C/C genotype (data not shown). Mixed effects models revealed that the time trends of WBC counts during the HIV disease course remained steady in the racial/ethnic group and according to DARC −46C/C genotype (supplemental Figure 1), and for this reason we also defined race/ethnicity-specific tertiles for the average WBC counts. To assess the influence of genotypes on survival, we used time to death as the endpoint.
Genotyping
A single-nucleotide polymorphism in DARC T-46C (rs2814778) was genotyped using a TaqMan allelic discrimination assay as described previously.10
Statistical analyses
All continuous variables were compared across 2 groups using the nonparametric Mann-Whitney test. For survival analyses, we used Kaplan-Meier plots to compare graphically the survival outcomes. Statistical significance of these differences was assessed by log-rank test as well as univariate and multivariate Cox proportional hazards models after testing the assumption of proportionality of hazards using the Schoenfield residuals. To control for the potential effect of varying lead times, seroprevalent status, and antiretroviral therapy (categorized as described previously11), we conducted stratified multivariate logistic and Cox regression analyses using the seroconversion status and receipt of antiretoriviral therapy as a stratifying variable. We used generalized linear mixed models to examine the time trends of the WBC counts during disease. This was done because the WBC count measurements were irregularly spaced and not all subjects had the same length of follow-up. Statistical significance was assessed using a 2-tailed P value at a type I error rate of .05. Statistical analyses were conducted using the Stata 7.0 statistical software package (Stata Corp).
Results
WBC counts and HIV disease course
There was a very high degree of correlation between the initial WBC count at diagnosis (ie, at entry into cohort) and the average WBC count maintained during HIV disease course (Spearman rho = 0.7035, P < .001, supplemental Figure 2). The time to death or AIDS in HIV-infected persons was significantly shorter in those with WBClow than those with WBChigh (Figure 1A; and data not shown). Because the results were similar whether we used AIDS or death as endpoints, the data presented are only for time to death and average WBC counts.
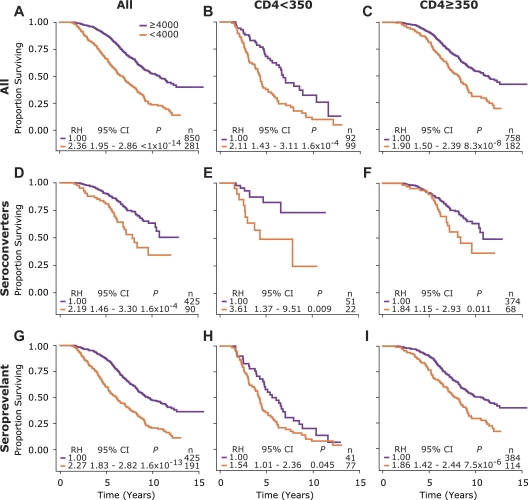
Figure 1.
Association of low WBC counts during disease with survival in HIV-infected subjects. Each panel shows Kaplan-Meier (KM) survival plots for subjects with an average WBC count during disease that was < (orange) or ≥ 4000 cells/mm3 (purple). The KM plots are for all subjects (left column), and those with baseline CD4+ T-cell counts of < 350 (middle column) ≥ 350 (right column) cells/mm3 in the entire cohort (top row), and subjects categorized as seroconverter (middle row) and seroprevalent (bottom row) component of the cohort. RH indicates relative hazards (reference RH = 1 represents those with WBC counts of ≥ 4000 cells/mm3); CI, confidence interval; P, significance value estimated using Cox proportional hazards modeling; and n, number of subjects.
As the average WBC count during infection was strongly positively correlated with the baseline CD4+ T-cell count (Spearman rho = 0.4340, P < .001), one possibility was that WBClow simply tracked the disease-accelerating effects associated with a low CD4+ count. To evaluate this, we investigated whether WBClow versus WBChigh had differential impacts on HIV disease course in subjects with a low (< 350 cells/mm3) or high (≥ 350 cells/mm3) baseline/initial CD4+ T-cell count, CD4+ count thresholds that in this study population were very strong predictors of rapid versus slow disease course, respectively.11,25 These analyses revealed that, in subjects who presented with low or high baseline CD4+ counts, WBClow was associated with a significantly shorter survival time than in those with WBChigh (Figure 1B-C). This suggested that low WBC counts are associated with HIV disease progression independent of the CD4+ T-cell count.
Because WBC counts were associated with an accelerated disease course (Figure 1A-C), it could be surmised that a low WBC count merely reflected a more advanced disease stage in those subjects for whom the date of seroconversion was not known. In this instance, the differential survival outcomes observed for WBClow versus WBChigh (Figure 1A-C) would simply be a reflection of the differences in the disease course of subjects categorized as seroconverters versus seroprevalent persons. However, this was unlikely to be the case because, compared with those subjects with WBChigh, those with WBClow had a faster disease course regardless of whether they were categorized as seroconverting or seroprevalent, or whether the baseline CD4+ count was low or high (Figure 1D-I).
To test the robustness of these observations, we conducted multivariate Cox proportional hazards regression analyses and found that WBClow was a strong predictor of shorter survival times, independent of baseline CD4+ counts and steady-state viral load that in this study population are strong predictors of disease progression11,25 (Table 1). For example, in the seroconverting subjects, the hazard ratio for time to death for those with WBClow was 1.80 (Table 1). Collectively, these findings (Figure 1; Table 1) indicated that a low WBC count is an independent determinant of an accelerated HIV disease course.
Table 1.
Association of low WBC counts during disease with rapid rate of HIV disease progression, independent of baseline CD4+ T-cell count, and steady-state viral load in the entire study population and in subjects categorized as seroconverters or seroprevalent
| All subjects | Seroconverters | Seroprevalent | |
|---|---|---|---|
| Average WBC count < 4000 cells/mm3 | 1.70 (1.35-2.14) | 1.80 (1.17-2.76) | 1.64 (1.24-2.17) |
| P | < .001 | .007 | < .001 |
| Baseline CD4+ T-cell count < 350 cells/mm3 | 2.50 (1.90-3.30) | 2.08 (1.20-3.59) | 2.68 (1.94-3.70) |
| P | < .001 | .009 | < .001 |
| Steady-state viral load ≥ 20000 copies/mL | 4.54 (3.43-6.01) | 4.27 (2.70-6.75) | 4.64 (3.24-6.65) |
| P | < .001 | < .001 | < .001 |
Results are from full multivariate Cox proportional hazards models showing relative hazards (95% confidence interval) for time to death and P value.
Race and leukopenia and HIV infection
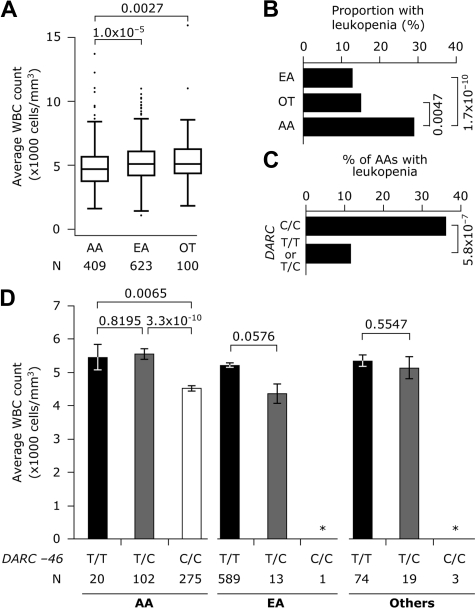
As the prevalence of leukopenia is higher in persons of African than European ancestry, we next determined whether this phenotype exists in HIV+ persons. The average WBC counts during disease (Figure 2A) were significantly lower in HIV-infected AAs (median, 4694 cells/mm3) than EAs (median, 5100 cells/mm3) or other ethnicities (median, 5093 cells/mm3); similarly, the initial WBC count at diagnosis was lower in HIV+ AAs (supplemental Figure 3A). The prevalence (percentage) of leukopenia at time of diagnosis was significantly higher in HIV+ AAs (28%) than in EAs (15%, P < .005) or HIV+ persons of other ethnicities (13%, P < .001) that were represented in the study cohort (Figure 2B). These data indicated that ethnic leukopenia present in healthy AAs was also present in the setting of HIV infection.
Figure 2.
WBC counts according to race/ethnicity and DARC −46C/C genotype. (A) Box-and-whisker plots depicting the distribution of the average of the total WBC counts during disease, in the major ethnic groups represented in the study population. AA indicates African American; EA indicates European American. OT indicates other ethnicities. Numbers at the top are P values obtained by Mann-Whiney tests for the indicated comparisons. N indicates number of subjects. (B) The proportion of subjects with leukopenia (initial WBC count < 4000 cells/mm3) in the indicated racial/ethnic groups. Vertically oriented numbers are significance values obtained by Pearson χ2 test. (C) The proportion of AA (n = 397) possessing the indicated DARC T-46C genotype (122 subjects with T/C or T/T and 275 subjects with C/C) who were leukopenic at presentation. The significance value for the comparison was obtained using the Pearson χ2 test. (D) The bar charts represent the mean (vertical bars) and 95% confidence intervals (error bars) for the average WBC counts during disease in the indicated racial/ethnic groups according to their DARC T-46C genotype. Numbers at the top are P values obtained by Mann-Whiney tests for the indicated comparisons. N indicates number of subjects. *Too few subjects (1 EA and 3 others), for whom the vertical bars are not shown.
Race and leukopenia and HIV disease
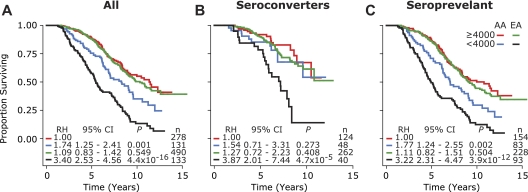
Because ethnic leukopenia is associated with a benign phenotype in HIV-uninfected AAs, we investigated whether this benign phenotype was reflected in a more benign HIV disease course in HIV-infected AAs by evaluating whether the disease course in AAs with WBClow (AA-WBClow) was more similar to those of EAs with WBClow (EA-WBClow) or nonleukopenic AAs (AA-WBChigh) and EAs (EA-WBChigh). In the overall cohort, before (Figure 3A) and after stratifying subjects according to their seroconversion status (Figure 3B-C), the disease course of AA-WBClow and EA-WBClow was dissimilar, whereas the disease course of AA-WBChigh and EA-WBChigh was similar. Among AA seroconverters, those with WBClow and WBChigh had similar rates of disease progression (Figure 3B). By contrast, among EA seroconverters, those with WBClow had a significantly faster disease course than those with WBChigh (Figure 3B). Among subjects categorized to the seroprevalent group, although AAs with WBClow progressed faster than AAs with WBChigh (compare red vs blue survival curves; Figure 3C), their disease course was slower than EAs with WBClow (compare blue vs black survival curves; Figure 3C). Hence, the hierarchy of disease progression according to ethnicity and WBC status revealed that AAs with WBClow had a survival advantage relative to EAs with WBClow, and this hierarchical relationship was consistent among seroconverters (AA-WBChigh = EA-WBChigh = AA-WBClow > EA-WBClow) and seroprevalent (AA-WBChigh = EA-WBChigh > AA-WBClow > EA-WBClow) subjects (Figure 3).
Figure 3.
Differential effects of low WBC counts on HIV disease course in HIV-infected EA and AA. KM plots depict survival curves computed according to race/ethnicity and average WBC count in the entire cohort (A), and HIV+ subjects categorized as seroconverter (B), and seroprevalent (C). RH indicates relative hazards (reference RH = 1 represents AAs with WBC counts ≥ 4000 cells/mm3); CI, confidence interval; P, significance values obtained by Cox proportional hazards modeling; and n, number of subjects.
We undertook further analyses and found that leukopenic EAs had a survival disadvantage compared with leukopenic AAs leukopenia whether leukopenia was defined by contemporary (< 4000 cells/mm3) versus population-specific cutoffs observed in the study cohort (supplemental Figure 4).
Duffy status, race, and leukopenia
The aforementioned data suggested that ethnic leukopenia existed in HIV+ AAs and that leukopenic HIV+ AAs had a more benign disease course relative to leukopenic EAs. To investigate whether there was a genetic basis for this, we determined whether there was any relationship between the aforementioned findings and the recent observation demonstrating that DARC −46C/C is a strong genetic basis for benign ethnic leukopenia, attributable to neutropenia, in HIV-negative subjects.21,22
In our study population, DARC −46C/C was present nearly exclusively in AAs (69.1%) compared with EA (0.2%).10 Among the hematologic parameters available for assessment, DARC T-46C genotype (T/T, T/C, and C/C), but not race, was a strong determinant of the variability in the WBC counts during disease (Table 2) and at entry into the cohort (P < .001). Furthermore, among HIV+ AAs, leukopenia at time of diagnosis was 3 times more prevalent in those with −46C/C (36%) than those with DARC −46 C/T or −46 T/T (11.5%; Figure 2C). Concordantly, WBC counts were lower in those AAs bearing DARC −46C/C than other DARC genotypes (Figures 2D, S3B). Taken together (Table 2; Figure 2), our results in HIV+ patients are concordant with those recent findings that the DARC genotype, and not race, was a strong predictor of variability in WBC/neutrophil counts in non-HIV cohorts.21,22
Table 2.
Two-way analyses of variance of hematologic parameters as outcomes predicted by race/ethnicity and DARC T-46C genotypes
| Race/ethnicity |
DARC T-46C |
|||
|---|---|---|---|---|
| F | P | F | P | |
| Average WBC count | 1.08 | .372 | 17.43 | < .001 |
| Hemoglobin | 0.72 | .607 | 0.24 | .790 |
| Platelet count | 1.18 | .316 | 0.77 | .465 |
| Lymphocyte count | 0.83 | .529 | 0.10 | .909 |
| CD4+ T-cell count | 0.81 | .544 | 0.80 | .452 |
| CD8+ T-cell count | 0.50 | .777 | 0.39 | .678 |
F indicates Snedecor F estimated using analysis of variance; and P, significance value.
DARC-46C/C, leukopenia, and disease course
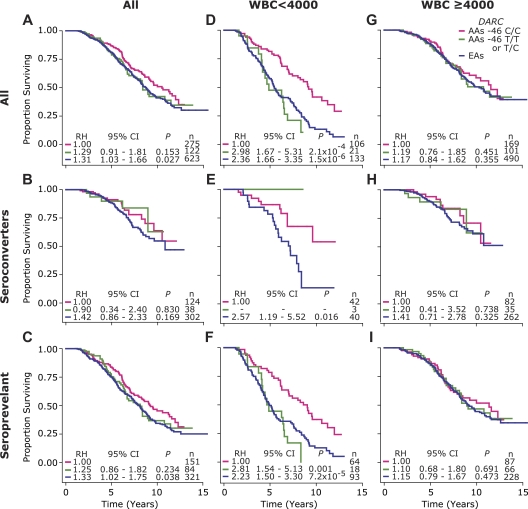
Consistent with our previous data10 in the overall study population and, therein the seroconverting and seroprevalent populations, there was a trend toward a survival advantage for HIV+ AAs with −46 C/C compared with HIV+ EAs, or AAs with DARC −46 C/T or T/T (Figure 4A-C). However, the survival advantage associated with −46C/C was highly dependent on WBC counts as this association was greatly magnified in subjects with WBClow (Figure 4D-F) and muted in those with WBChigh (Figure 4G-I).
Figure 4.
Rate of HIV disease progression in HIV+ EAs, and HIV+ AAs who are Duffy-null (DARC −46C/C) or -positive (DARC −46C/T or T/T). Each panel shows KM survival plots for 3 groups of HIV-infected subjects: EA (blue), AAs possessing DARC −46 C/C (pink), and AAs possessing DARC −46 C/T or T/T (green). The KM plots are for all subjects (left column) and those with baseline CD4+ T-cell counts of < 4000 (middle column) or ≥ 4000 (right column) cells/mm3 in the entire cohort (top row), and those subjects categorized as seroconverter (middle row) and seroprevalent (bottom row). RH indicates relative hazards (reference RH = 1 represents AAs with DARC −46C/C); CI, confidence interval; P, significance value estimated using Cox proportional hazards modeling, n, number of subjects. Whether DARC −46C/T or T/T was associated with disease acceleration in leukopenic seroconverting AAs could not be evaluated as in this group; there were only 4 subjects who did not have −46C/C, and no death events had occurred (green survival curve in panel E).
To investigate whether the accentuation of the DARC genotype-phenotype associations with leukopenia was meaningful and not artifactual, we determined whether disease course differed according to DARC genotype in subjects categorized according to their baseline CD4+ T-cell count (< 350 or > 350 cells/mm3; supplemental Figure 5). These analyses revealed that the disease-influencing phenotype associated with DARC −46C/C differed substantially according to the WBC (Figure 4) but not CD4+ counts (supplemental Figure 5).
We further explored the relationships between DARC genotype, leukopenia, and HIV disease course in HIV+ AAs using univariate, multivariate, and other statistical approaches. Univariate models revealed that, among those with WBClow, the hazard ratio for survival associated with DARC −46C/C was one-third (relative hazard [RH] = 0.33) of that associated with DARC −46 C/T or T/T (RH = 1; Table 3). In a multivariate model stratified for seroconversion status and receipt of antiretroviral therapy, DARC −46C/C was associated with a nearly 80% (RH = 0.20) lower risk of progressive disease, independent of other covariates that are strong predictors of disease progression11 as well as a genetic parameter designated as admixture score10 (Table 3). Substantiating this, in stepwise Cox regression modeling, the DARC −46C/C genotype was retained as a significant predictor of disease progression, but only in subjects with a low WBC count (Table 3). In these analyses, an association between DARC −46C/C and survival was not observed in subjects with WBChigh (Table 3). These data indicate that DARC −46C/C is an independent determinant of disease course, but mainly in those with a low WBC count.
Table 3.
Association of DARC −46C/C with disease progression in HIV+ African Americans
| Univariate | Full model | Final model | |
|---|---|---|---|
| All subjects | 0.77 (0.55-1.09) | 0.66 (0.40-1.09) | — |
| P | .146 | .102 | |
| Average WBC < 4000 | 0.33 (0.19-0.60) | 0.20 (0.07-0.60) | 0.22 (0.08-0.64) 0.005 |
| P | < .001 | .004 | |
| Average WBC ≥ 4000 | 0.84 (0.54-1.31) | 1.18 (0.62-2.27) | — |
| P | .441 | .617 |
Values are relative hazards (95% confidence interval) time-to-death and P value. Covariates for the full multivariate model are baseline CD4+ T-cell count; cumulative CD4+ T-cell count (cCD4), a parameter that reflects the changes in CD4+ counts over disease course and a very strong predictor of disease progression11; steady-state viral load; and an admixture score derived from 11 genetic markers that was highly predictive of self-reported ethnicity (> 98% accuracy)10 and was also strongly correlated with the degree of admixture as assessed by the first principal component (obtained by EIGENSOFT program) of 96 ancestry informative markers.26 Both full model and final stepwise model have been adjusted for seroconversion and receipt of antiretroviral therapy, as defined previously.11 RH = 1 represents those with DARC −46C/T or T/T.
— indicates not retained in model.
DARC −46C/C, magnitude of leukopenia, and disease course
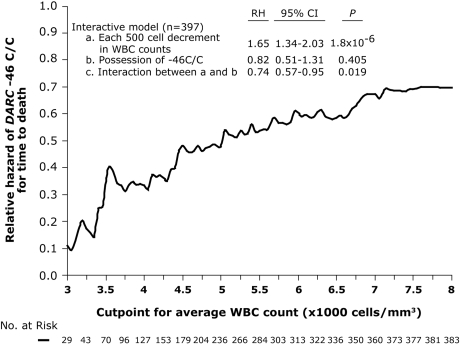
The aforementioned genotype-phenotype analyses were conducted with a fixed WBC cutoff of 4000 cells/mm3. However, it was plausible that the association between DARC genotype with disease progression was quantitatively dependent on the magnitude of leukopenia. We therefore evaluated the hazard ratio for survival associated with DARC −46C/C in HIV+ AA subjects at different thresholds of average WBC counts (Figure 5). This analysis revealed that, with each stepwise decrement in the average WBC count, the hazard ratios for time to death associated with DARC −46C/C progressively improved (Figure 5). This suggested that it was the interaction between DARC genotype and WBC counts that actually influenced HIV disease course. This inference was consistent with the results of the interactive logistic regression model that we used to investigate whether (1) leukopenia, (2) DARC −46C/C, and (3) the interaction between leukopenia and DARC genotype were each associated with independent disease-influencing effects (Figure 5). In this model, among HIV+ AAs, each 500-cell decrement in the average WBC count during disease was associated with a significantly increased risk of progressive HIV disease (RH = 1.65; P < .001; Figure 5 inset). Notably, this hazard ratio of 1.65 is consistent with that observed for leukopenic AAs (Figure 3A, RH = 1.74 for blue plot). However, although leukopenia per se was associated with disease acceleration in HIV+ AAs, the interaction term between DARC −46C/C and WBC counts was significantly associated with a reduced risk of progressive disease (RH = 0.74; P = .019; Figure 5 inset), rendering the association of DARC −46C/C in itself not significant (RH = 0.82; P = .405, Figure 5 inset).
Figure 5.
Relationship between decrements in the average WBC counts and the hazard ratios for time to death associated with the DARC −46C/C genotype in HIV-infected AAs. The plot represents the hazard ratios (y-axis) for rate of progression to death estimated using Cox proportional hazards modeling in subjects with average WBC counts below the indicated cutpoints (x-axis). The hazard ratios were estimated over the range of WBC counts shown on the x-axis for every 50-cell count change. The reference group is those subjects who possess DARC −46T/T or T/C (RH = 1). Inset represents data showing an interactive effect of low total WBC count on the survival advantage associated with the DARC −46C/C genotype in HIV+ AAs. RH indicates relative hazards (reference RH = 1 represents AAs with DARC −46T/T or T/C); CI, confidence interval; and P, significance value estimated using Cox proportional hazards modeling with indicated parameters (a, b, and c).
Discussion
In our study population, we observed the following relationships for WBC counts during HIV infection. First, a low WBC count during disease was associated with a faster disease progression, independent of known predictors of AIDS development, such as the CD4+ T-cell count and viral load. Second, although twice as many HIV+ AAs presented with leukopenia than EAs, leukopenic AAs had a slower disease course than leukopenic EAs, whereas the disease course in nonleukopenic EAs and nonleukopenic AAs was similar. This suggested that ethnic leukopenia in HIV-infected AAs may be associated with a benign phenotype, despite HIV-induced immunodeficiency. In addition, we found that, over and above its strong link to African ancestry, DARC genotype is a strong predictor of WBC variability during HIV disease. Furthermore, there was a significant survival benefit in DARC −46C/C-bearing HIV+ AAs with low WBCs but not high WBCs. This survival advantage became increasingly pronounced in those with progressively lower WBC counts.
Highlighting that ethnic leukopenia associated with DARC −46C/C persists during HIV infection, we found that, among HIV+ AAs who presented with leukopenia, 36% possessed DARC −46C/C versus 12% who had −46T/C or T/T. The levels of total lymphocytes or CD4+ and CD8+ T cells were similar among AAs with different DARC genotypes. Hence, concordant with published reports,21–24 the lower WBC levels found in −46C/C-bearing HIV+ AAs we suspect are because of a reduction in neutrophil and, possibly, monocyte cell counts.
Our previous data showed that DARC −46C/C associated with a slow HIV disease course in the overall AA study population.10 We now show that this association is evident mainly in those AAs who maintained a low WBC count during disease, suggesting that the genotype-phenotype relationships for the DARC-null state are highly dependent on the cellular milieu. These findings may also help explain the observation that neutropenia, which is found commonly in HIV-infected patients,2,3 was not associated with decreased survival in a large study of HIV-infected African American women.2
The basis for the association between DARC −46C/C and survival advantage during HIV disease remains unknown, but we offer the following possible explanations. First, it can be argued that the association of DARC −46C/C with survival advantage in leukopenic HIV+ AAs represents a tautology. As DARC −46C/C is African-specific and associated with leukopenia, it follows that the association between DARC −46C/C and disease protection in leukopenic AAs was merely a reflection of the association between −46C/C with a low WBC count in AAs. Although this argument has merit, it does not account for the following observations, which make this line of reasoning implausible: (1) not all leukopenic HIV+ AAs possessed DARC −46C/C, and conversely, not all subjects with DARC −46C/C were leukopenic; and (2) leukopenic AAs with DARC −46C/T or -T/T had an accelerated disease course that was similar to the disease course of leukopenic EAs. This suggested that DARC −46C/C may offset the detrimental effects associated with leukopenia in AAs. Supporting the latter inference were the results of statistical analyses showing that among HIV+ AAs it was the interaction between DARC −46C/C and WBC counts, and not DARC genotype alone, that was associated with a survival advantage (Figure 5 inset). This dependence of DARC genotype on WBC counts for full expression of its phenotypic effects on disease was also highlighted by the observation that the survival advantage associated with DARC −46C/C was augmented with increasing degrees of leukopenia.
A second possibility is that DARC −46C/C may be merely a surrogate for other unknown genotypes that influence HIV disease course in the setting of leukopenia. Because WBC counts are a genetically related phenotype, it follows that subjects with leukopenia may be enriched for polymorphisms in other genes that confer a low WBC count trait. In this setting, the cellular milieu defined by genetically determined leukocyte count profiles and/or genes that track with leukopenia (which may include DARC −46C/C) may serve as a modifier of HIV disease. Although we cannot exclude this possibility, we outline in the following paragraphs several observations that together favor a role for DARC in HIV pathogenesis.
Why does the DARC −46C/C genotype specifically associate with a survival advantage in those HIV-infected AAs who have a low WBC count? Indeed, that leukopenia could be “good” for a patient appears on the surface to be counterintuitive. We speculate that what we have observed in the context of HIV infection may reflect an evolutionarily adaptive mechanism to limit both inflammatory response and cell entry, to an ancestral pathogen such as P vivax. The DARC locus shows complex genetic signatures of positive natural selection,27,28 and it has been suggested that the −46C allele has arisen under the selective pressure of an ancestral form of malaria that also used DARC but was more lethal than P vivax, which is generally not associated with mortality and is no longer very common in Africa.20 Interestingly, neutropenia is commonly observed during malaria and is more common in infection with P vivax than P falciparum,29,30 and there is clinical and experimental evidence that neutrophils contribute to the adverse outcomes after malarial infection.31–35 Furthermore, a genome-wide analysis of the host response to malaria also implicates neutrophil-mediated pathways in underlying malarial pathogenesis.36 Hence, one possibility is that the protection against neutrophil-mediated inflammation during infection by a lethal ancestral form of malaria may have also promoted the selection of the Duffy-null trait.
Studies suggest that ethnic leukopenia/neutropenia may be related to a reduced ability to mobilize neutrophils from the marginated granulocyte pool.37 Supporting this, studies in mouse models show that DARC expression on both RBC and endothelial cells influences neutrophil migration.38,39 In addition, DARC −46C/C may associate with impaired hematopoiesis because some of the chemokines that bind to DARC (eg, GROα40) regulate many hematopoietic responses, including survival, proliferation, migration, and homing.41 Supporting this possibility, healthy subjects of African ancestry have reduced numbers of bone marrow progenitor cells than Caucasians.42 Although the precise mechanism underlying ethnic leukopenia is unknown, given the importance of neutrophils in inflammation, the reduced numbers of circulating neutrophils associated with DARC −46C/C may lead to dampened inflammatory responses, and this may explain in part the “benign” nature of ethnic leukopenia. In contrast, the HIV-associated leukopenia/neutropenia observed in EAs or AAs who bear at least one DARC −46T allele, is associated with a less favorable outcome possibly because this leukopenia occurs within the context of the Duffy-positive state and hence originates from a different mechanism that is not associated with a “benign” leukopenia.
Additional data suggest a link between DARC −46C/C and inflammatory responses relevant to HIV pathogenesis. Studies demonstrate a key role for circulating lipopolysaccharide (LPS), a marker of microbial translocation, and inflammation in the pathogenesis of HIV infection, especially acute infection.43 In this respect, it is notable that, after administration of LPS (endotoxin) to healthy persons, compared with Duffy-positive Caucasians, Duffy-null persons of African descent had significantly lower levels of total blood chemokine levels (derived from plasma levels, RBC- and neutrophil-associated MCP-1, GROα, and interleukin [IL]-8 ligands of DARC).24 This may have relevance to HIV pathogenesis because prior studies had found that plasma levels of MCP-1, a chemokine that associates with higher HIV viral load in vivo and increased viral replication in vitro,44 are lower in Duffy-null subjects.45 In addition, after administration of LPS, compared with Duffy-positive Caucasians, Duffy-null subjects had significantly lower levels of 3 biomarkers of in vivo thrombin generation (TAT, F1 + 2, and D-dimer), suggesting that the Duffy-null state associates with reduced tissue factor-triggered activation of coagulation pathways.46 This may also be relevant to HIV pathogenesis, as Kuller et al demonstrated recently that the degree of HIV-induced activation of coagulation pathways, as assessed by D-dimer levels, has an adverse effect on all-cause mortality among HIV+ patients with relatively preserved CD4+ counts.47 Hence, future studies are warranted to determine whether the increased survival in HIV-infected AAs associated with the Duffy-null/low WBC profile is a result of attenuated coagulation responses and chemokine- and neutrophil-mediated inflammation.
The importance of neutrophil-mediated inflammation has been highlighted in simian models of simian immunodeficiency virus (SIV) infection. For example, Elbim et al48 showed that the degree of neutropenia during acute infection correlates with viral load, and the differential sensitivity of neutrophils to apoptosis may distinguish pathogenic versus nonpathogenic SIV infections of nonhuman primates. Further affirming a possible role of neutrophils during the acute phases of lentiviral infection are the studies of Li et al,49 who used a vaginal transmission model of SIV infection in rhesus macaques and found that, after viral inoculation of SIV, infected cells in the cervix were generally in areas of inflammation containing IL-8–positive cells.49 Notably, IL-8 is a potent chemoattractant of neutrophils and application of glycerol monolaurate, an inhibitor of IL-8, protected these animals from infection.49
There may be additional mechanisms by which DARC influences HIV pathogenesis. In a previous study,10 we found that, in addition to binding to HIV-suppressive chemokines, HIV also binds to erythrocyte DARC in vitro and that RBC-bound HIV can be displaced by chemokines and transferred in trans to CD4+ T cells.10 Thus, we had speculated that the binding of fewer HIV particles to DARC-null RBCs may lead to a slower disease course.
In conclusion, prior results associated the DARC-null state with malaria resistance, and the present data suggest that an intricate interaction between DARC −46C/C genotype and leukopenia associates with survival advantage in HIV-infected AAs. Thus, further evaluation of the intersection of DARC genotype, the cellular milieu, as reflected by the WBC count, and HIV disease outcome may provide new insights into pathogenesis. Of broad relevance, accounting for the modifier effects of the cellular environment (eg, as reflected in the present study by genetically determined leukocyte parameters) may provide greater insights into genotype-phenotype relationships for DARC and other genes during HIV infection as well as other diseases.
Acknowledgments
The authors thank Duane Hospenthal, Brian Agan, and the anonymous reviewers for their critical feedback.
This work was supported by the Veterans Administration Center on AIDS and HIV infection of the South Texas Veterans Health Care System, and a MERIT (R37046326) and other awards (AI043279 and MH069270) from the National Institutes of Health (S.K.A.). S.K.A. is also supported by a Veterans Administration MERIT award and is a recipient of the Elizabeth Glaser Scientist Award, the Burroughs Wellcome Clinical Scientist Award in Translational Research, and the Doris Duke Distinguished Clinical Scientist Award. Support for the Department of Defense HIV Natural History Study cohort and staff involved in this work was provided by the Infectious Disease Clinical Research Program of the Uniformed Services University of the Health Sciences, of which the HIV Natural History Study is a component. The Infectious Disease Clinical Research Program is a Department of Defense tri-service program executed through Uniformed Services University of the Health Sciences and the Henry M. Jackson Foundation for the Advancement of Military Medicine, in collaboration with Department of Health and Human Services/National Institutes of Health/National Institute of Allergy and Infectious Diseases/Division of Clinical Research through Interagency Agreement Y1-AI-5072.
The content of this publication is the sole responsibility of the authors and does not necessarily reflect the views or policies of the National Institutes of Health or the Department of Health and Human Services, the Department of Defense or the Departments of the Army, Navy or Air Force. Mention of trade names, commercial products, or organizations does not imply endorsement by the US Government.
Footnotes
An Inside Blood analysis of this article appears at the front of this issue.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Authorship
Contribution: S.K.A., H.K., and M.J.D. conceptualized the research, analyzed the data, and wrote the manuscript; V.C.M., M.L.L., J.F.O., J.D., J.C., and M.J.D. contributed substantially to clinical cohort development, conceptual ideas, data analyses, and preparation of the manuscript; and W.H., D.K., S.S.A., E.J.W., R.A.W., and R.A.C. contributed conceptual ideas and immensely to manuscript preparation.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Sunil K. Ahuja, Department of Medicine, University of Texas Health Science Center at San Antonio, 7703 Floyd Curl Dr, San Antonio, TX 78229; e-mail: ahujas@uthscsa.edu.
References
- 1.Kaslow RA, Phair JP, Friedman HB, et al. Infection with the human immunodeficiency virus: clinical manifestations and their relationship to immune deficiency. A report from the Multicenter AIDS Cohort Study. Ann Intern Med. 1987;107:474–480. doi: 10.7326/0003-4819-107-4-474. [DOI] [PubMed] [Google Scholar]
- 2.Levine AM, Karim R, Mack W, et al. Neutropenia in human immunodeficiency virus infection: data from the women's interagency HIV study. Arch Intern Med. 2006;166:405–410. doi: 10.1001/archinte.166.4.405. [DOI] [PubMed] [Google Scholar]
- 3.Farber BF, Lesser M, Kaplan MH, Woltmann J, Napolitano B, Armellino D. Clinical significance of neutropenia in patients with human immunodeficiency virus infection. Infect Control Hosp Epidemiol. 1991;12:429–434. doi: 10.1086/646373. [DOI] [PubMed] [Google Scholar]
- 4.D'Angelo LJ, Getson PR, Luban NL, Gayle HD. Human immunodeficiency virus infection in urban adolescents: can we predict who is at risk? Pediatrics. 1991;88:982–986. [PubMed] [Google Scholar]
- 5.Altice FL, Marinovich A, Khoshnood K, Blankenship KM, Springer SA, Selwyn PA. Correlates of HIV infection among incarcerated women: implications for improving detection of HIV infection. J Urban Health. 2005;82:312–326. doi: 10.1093/jurban/jti055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Henry WK, Tebas P, Lane HC. Explaining, predicting, and treating HIV-associated CD4 cell loss: after 25 years still a puzzle. JAMA. 2006;296:1523–1525. doi: 10.1001/jama.296.12.1523. [DOI] [PubMed] [Google Scholar]
- 7.Hermans P, Sommereijns B, Van Cutsem N, Clumeck N. Neutropenia in patients with HIV infection: a case control study in a cohort of 1403 patients between 1982 and 1993. J Hematother Stem Cell Res. 1999;8(suppl 1):S23–S32. doi: 10.1089/152581699319894. [DOI] [PubMed] [Google Scholar]
- 8.Murphy MF, Metcalfe P, Waters AH, et al. Incidence and mechanism of neutropenia and thrombocytopenia in patients with human immunodeficiency virus infection. Br J Haematol. 1987;66:337–340. doi: 10.1111/j.1365-2141.1987.tb06920.x. [DOI] [PubMed] [Google Scholar]
- 9.Servais J, Nkoghe D, Schmit JC, et al. HIV-associated hematologic disorders are correlated with plasma viral load and improve under highly active antiretroviral therapy. J Acquir Immune Defic Syndr. 2001;28:221–225. doi: 10.1097/00042560-200111010-00003. [DOI] [PubMed] [Google Scholar]
- 10.He W, Neil S, Kulkarni H, et al. Duffy antigen receptor for chemokines mediates trans-infection of HIV-1 from red blood cells to target cells and affects HIV-AIDS susceptibility. Cell Host Microbe. 2008;4:52–62. doi: 10.1016/j.chom.2008.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dolan MJ, Kulkarni H, Camargo JF, et al. CCL3L1 and CCR5 influence cell-mediated immunity and affect HIV-AIDS pathogenesis via viral entry-independent mechanisms. Nat Immunol. 2007;8:1324–1336. doi: 10.1038/ni1521. [DOI] [PubMed] [Google Scholar]
- 12.Silverberg MJ, Wegner SA, Milazzo MJ, et al. Effectiveness of highly-active antiretroviral therapy by race/ethnicity. AIDS. 2006;20:1531–1538. doi: 10.1097/01.aids.0000237369.41617.0f. [DOI] [PubMed] [Google Scholar]
- 13.Crum-Cianflone N, Hullsiek KH, Marconi V, et al. Trends in the incidence of cancers among HIV-infected persons and the impact of antiretroviral therapy: a 20-year cohort study. AIDS. 2009;23:41–50. doi: 10.1097/QAD.0b013e328317cc2d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Haddy TB, Rana SR, Castro O. Benign ethnic neutropenia: what is a normal absolute neutrophil count? J Lab Clin Med. 1999;133:15–22. doi: 10.1053/lc.1999.v133.a94931. [DOI] [PubMed] [Google Scholar]
- 15.Freedman DS, Gates L, Flanders WD, et al. Black/white differences in leukocyte subpopulations in men. Int J Epidemiol. 1997;26:757–764. doi: 10.1093/ije/26.4.757. [DOI] [PubMed] [Google Scholar]
- 16.Grann VR, Bowman N, Joseph C, et al. Neutropenia in 6 ethnic groups from the Caribbean and the U.S. Cancer. 2008;113:854–860. doi: 10.1002/cncr.23614. [DOI] [PubMed] [Google Scholar]
- 17.Hsieh MM, Everhart JE, Byrd-Holt DD, Tisdale JF, Rodgers GP. Prevalence of neutropenia in the U.S. population: age, sex, smoking status, and ethnic differences. Ann Intern Med. 2007;146:486–492. doi: 10.7326/0003-4819-146-7-200704030-00004. [DOI] [PubMed] [Google Scholar]
- 18.Rot A. Contribution of Duffy antigen to chemokine function. Cytokine Growth Factor Rev. 2005;16:687–694. doi: 10.1016/j.cytogfr.2005.05.011. [DOI] [PubMed] [Google Scholar]
- 19.Hadley TJ, Peiper SC. From malaria to chemokine receptor: the emerging physiologic role of the Duffy blood group antigen. Blood. 1997;89:3077–3091. [PubMed] [Google Scholar]
- 20.Langhi DM, Jr, Bordin JO. Duffy blood group and malaria. Hematology. 2006;11:389–398. doi: 10.1080/10245330500469841. [DOI] [PubMed] [Google Scholar]
- 21.Nalls MA, Wilson JG, Patterson NJ, et al. Admixture mapping of white cell count: genetic locus responsible for lower white blood cell count in the Health ABC and Jackson Heart studies. Am J Hum Genet. 2008;82:81–87. doi: 10.1016/j.ajhg.2007.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Reich D, Nalls MA, Kao WH, et al. Reduced neutrophil count in people of African descent is due to a regulatory variant in the Duffy antigen receptor for chemokines gene. PLoS Genet. 2009;5:e1000360. doi: 10.1371/journal.pgen.1000360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Grann VR, Ziv E, Joseph CK, et al. Duffy (Fy), DARC, and neutropenia among women from the United States, Europe and the Caribbean. Br J Haematol. 2009;145:266–267. doi: 10.1111/j.1365-2141.2009.07588.x. [DOI] [PubMed] [Google Scholar]
- 24.Mayr FB, Spiel AO, Leitner JM, et al. Duffy antigen modifies the chemokine response in human endotoxemia. Crit Care Med. 2008;36:159–165. doi: 10.1097/01.CCM.0000297875.55969.DB. [DOI] [PubMed] [Google Scholar]
- 25.Kulkarni H, Agan BK, Marconi VC, et al. CCL3L1-CCR5 genotype improves the assessment of AIDS Risk in HIV-1-infected individuals. PLoS ONE. 2008;3:e3165. doi: 10.1371/journal.pone.0003165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.He W, Marconi VC, Castiblanco J, et al. Response: association of Duffy antigen genotype with HIV-AIDS susceptibility. Cell Host Microbe. 2009;5:418–419. [Google Scholar]
- 27.Hamblin MT, Thompson EE, Di Rienzo A. Complex signatures of natural selection at the Duffy blood group locus. Am J Hum Genet. 2002;70:369–383. doi: 10.1086/338628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tournamille C, Blancher A, Le Van Kim C, et al. Sequence, evolution and ligand binding properties of mammalian Duffy antigen/receptor for chemokines. Immunogenetics. 2004;55:682–694. doi: 10.1007/s00251-003-0633-2. [DOI] [PubMed] [Google Scholar]
- 29.Rodriguez-Morales AJ, Sanchez E, Arria M, et al. White blood cell counts in Plasmodium vivax malaria. J Infect Dis. 2005;192:1675–1676. doi: 10.1086/496993. author reply 1676–1677. [DOI] [PubMed] [Google Scholar]
- 30.Jadhav UM, Singhvi R, Shah R. Prognostic implications of white cell differential count and white cell morphology in Malaria. J Postgrad Med. 2003;49:218–220. discussion 221. [PubMed] [Google Scholar]
- 31.Chen L, Zhang Z, Sendo F. Neutrophils play a critical role in the pathogenesis of experimental cerebral malaria. Clin Exp Immunol. 2000;120:125–133. doi: 10.1046/j.1365-2249.2000.01196.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Senaldi G, Vesin C, Chang R, Grau GE, Piguet PF. Role of polymorphonuclear neutrophil leukocytes and their integrin CD11a (LFA-1) in the pathogenesis of severe murine malaria. Infect Immun. 1994;62:1144–1149. doi: 10.1128/iai.62.4.1144-1149.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mohammed AO, Elghazali G, Mohammed HB, et al. Human neutrophil lipocalin: a specific marker for neutrophil activation in severe Plasmodium falciparum malaria. Acta Trop. 2003;87:279–285. doi: 10.1016/s0001-706x(03)00116-5. [DOI] [PubMed] [Google Scholar]
- 34.Hemmer CJ, Vogt A, Unverricht M, Krause R, Lademann M, Reisinger EC. Malaria and bacterial sepsis: similar mechanisms of endothelial apoptosis and its prevention in vitro. Crit Care Med. 2008;36:2562–2568. doi: 10.1097/CCM.0b013e31818441ee. [DOI] [PubMed] [Google Scholar]
- 35.Huy NT, Trang DT, Kariu T, et al. Leukocyte activation by malarial pigment. Parasitol Int. 2006;55:75–81. doi: 10.1016/j.parint.2005.10.003. [DOI] [PubMed] [Google Scholar]
- 36.Griffiths MJ, Shafi MJ, Popper SJ, et al. Genomewide analysis of the host response to malaria in Kenyan children. J Infect Dis. 2005;191:1599–1611. doi: 10.1086/429297. [DOI] [PubMed] [Google Scholar]
- 37.Phillips D, Rezvani K, Bain BJ. Exercise induced mobilisation of the marginated granulocyte pool in the investigation of ethnic neutropenia. J Clin Pathol. 2000;53:481–483. doi: 10.1136/jcp.53.6.481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lee JS, Wurfel MM, Matute-Bello G, et al. The Duffy antigen modifies systemic and local tissue chemokine responses following lipopolysaccharide stimulation. J Immunol. 2006;177:8086–8094. doi: 10.4049/jimmunol.177.11.8086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pruenster M, Mudde L, Bombosi P, et al. The Duffy antigen receptor for chemokines transports chemokines and supports their promigratory activity. Nat Immunol. 2009;10:101–108. doi: 10.1038/ni.1675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Fukuda S, Bian H, King AG, Pelus LM. The chemokine GRObeta mobilizes early hematopoietic stem cells characterized by enhanced homing and engraftment. Blood. 2007;110:860–869. doi: 10.1182/blood-2006-06-031401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Broxmeyer HE. Chemokines in hematopoiesis. Curr Opin Hematol. 2008;15:49–58. doi: 10.1097/MOH.0b013e3282f29012. [DOI] [PubMed] [Google Scholar]
- 42.Rezvani K, Flanagan AM, Sarma U, Constantinovici N, Bain BJ. Investigation of ethnic neutropenia by assessment of bone marrow colony-forming cells. Acta Haematol. 2001;105:32–37. doi: 10.1159/000046530. [DOI] [PubMed] [Google Scholar]
- 43.Douek D. HIV disease progression: immune activation, microbes, and a leaky gut. Top HIV Med. 2007;15:114–117. [PubMed] [Google Scholar]
- 44.Weiss L, Si-Mohamed A, Giral P, et al. Plasma levels of monocyte chemoattractant protein-1 but not those of macrophage inhibitory protein-1alpha and RANTES correlate with virus load in human immunodeficiency virus infection. J Infect Dis. 1997;176:1621–1624. doi: 10.1086/517341. [DOI] [PubMed] [Google Scholar]
- 45.Jilma-Stohlawetz P, Homoncik M, Drucker C, et al. Fy phenotype and gender determine plasma levels of monocyte chemotactic protein. Transfusion. 2001;41:378–381. doi: 10.1046/j.1537-2995.2001.41030378.x. [DOI] [PubMed] [Google Scholar]
- 46.Mayr FB, Spiel AO, Leitner JM, et al. Racial differences in endotoxin-induced tissue factor-triggered coagulation. J Thromb Haemost. 2009;7:634–640. doi: 10.1111/j.1538-7836.2009.03307.x. [DOI] [PubMed] [Google Scholar]
- 47.Kuller LH, Tracy R, Belloso W, et al. Inflammatory and coagulation biomarkers and mortality in patients with HIV infection. PLoS Med. 2008;5:e203. doi: 10.1371/journal.pmed.0050203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Elbim C, Monceaux V, Mueller YM, et al. Early divergence in neutrophil apoptosis between pathogenic and nonpathogenic simian immunodeficiency virus infections of nonhuman primates. J Immunol. 2008;181:8613–8623. doi: 10.4049/jimmunol.181.12.8613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Li Q, Estes JD, Schlievert PM, et al. Glycerol monolaurate prevents mucosal SIV transmission. Nature. 2009;458:1034–1038. doi: 10.1038/nature07831. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.