Abstract
An extensive family-based study of linkage disequilibrium (LD) in the killer cell immunoglobulin-like receptors (KIR) cluster was performed. We aimed to describe the LD structure in the KIR gene cluster using a sample of 418 founder haplotypes identified by segregation in a group of 106 families from Northern Ireland. The LD was studied at two levels of polymorphism: the structural level (presence or absence of KIR genes) and the allelic level (between alleles of KIR genes). LD was further assessed using the predictive value of one KIR polymorphism for another one in order to provide an interpretative framework for the LD effect in association studies. In line with previous research, distinct patterns of KIR genetic diversity within the genomic region centromeric to KIR2DL4 (excluding KIR2DL4) and within the telomeric region including KIR2DL4 were found. In a comprehensive PPV/NPV-based LD analysis within the KIR cluster, robust tag markers were found that can be used to identify which genes are concomitantly present or absent and to further identify groups of associated KIR alleles. Several extended KIR haplotypes in the study population were identified (KIR2DS2*POS–KIR2DL2*001–KIR2DL5B*002–KIR2DS3*00103–KIR2DL1*00401; KIR2DL4*011–KIR3DL1/S1*005–KIR2DS4*003–KIR3DL2*003; KIR2DL4*00802–KIR3DL1/S1*004–KIR2DS4*006–KIR3DL2*005; KIR2DL4*00801–KIR3DL1/S1*00101–KIR2DS4*003–KIR3DL2*001; KIR2DL4*00103–KIR3DL1/S1*008–KIR2DS4*003–KIR3DL2*009; KIR2DL4*00102–KIR3DL1/S1*01502/*002–KIR2DS4*00101–KIR3DL2*002; KIR2DL4*00501–KIR3DL1/S1*013–KIR2DL5A*001–KIR2DS5*002–KIR2DS1*002–KIR3DL2*007). The present study provides a rationale for analyzing associations of KIR polymorphisms by taking into account the complex LD structure of the KIR region.
Electronic supplementary material
The online version of this article (doi:10.1007/s00251-010-0478-4) contains supplementary material, which is available to authorized users.
Keywords: KIR, Linkage disequilibrium, Tagging, Association studies
Introduction
Natural killer (NK) cells are key components of the innate immune response. According to the missing-self model, NK cells integrate activating and inhibitory signals and modulate the targeting of MHC Class I-deficient cells, in particular virus-infected and transformed malignant cells (Bashirova et al. 2006). NK cells maintain wide-ranging interactions with other immune cells, such as macrophages and dendritic cells, resulting in numerous effects on the immune response as a whole through the stimulation of cytokine production and induction of cytotoxicity (Carrington and Martin 2006). NK cells may provide benefit or act to the detriment of to the host: They play a central role not only in viral clearance (Khakoo et al. 2004; Martin et al. 2007), cancer (Middleton et al. 2009; Verheyden and Demanet 2008), and hematopoietic stem cell transplantation (Velardi 2008) but also in the development of autoimmune disorders (Lowe et al. 2009; Ploski et al. 2006) and less common conditions such as pre-eclampsia (Hiby et al. 2008).
The NK cell receptors consist of two distinct families, C-type lectin-like group (CD94/NKG2) mapping to chromosome 12q1.3-13.4 and the immunoglobulin-like super family consisting of the killer cell immunoglobulin-like receptors (KIR), leukocyte immunoglobulin-like receptors, and the leukocyte-associated immunoglobulin-like receptors mapping to chromosome 19q13.4 (Bashirova et al. 2006; Carrington and Martin 2006). The KIR gene cluster has generated great interest due to its complex genetic variability (Bashirova et al. 2006; Carrington and Martin 2006; Middleton et al. 2007; Parham 2005).
KIR genes may be either present or absent, leading to substantial variation in gene content across individuals and populations. The deletion and/or duplication of KIR genes occur frequently, creating great structural variation and generating a variable number of copies of KIR genes relative to a reference sequence (i.e., the most frequent one; Hsu et al. 2002). When present, individual KIR genes show high allelic polymorphisms. Their frequencies vary across populations, some being common, others rare. The patterns of gene presence or absence and allelic polymorphism combine to generate a high degree of KIR heterogeneity between individuals (Middleton and Gonzelez 2010). The polymorphism of genes in the KIR cluster may be assessed at two levels: the structural variation (also known as copy number variation, CNV) of genes and the allelic variation of each gene.
Linkage disequilibrium (LD) is the nonrandom association of alleles at two or more neighboring loci. Furthermore, because KIR genes are arrayed in tandem over more than 150 kb (Uhrberg et al. 1997), extensive LD implicates both gene content and allelic variation (Shilling et al. 2002). Such complex LD patterns, already complicated in pedigree analyses, may lead to ambiguous interpretation in population association studies because nonindependent associations between KIR genes or alleles may result in potential synergistic or antagonist functional effects. This complexity may explain the difficulty encountered in replication of disease CNV genetic association studies. The KIR cluster can be considered as a model region for CNV genomics, because it is the most important region outside the MHC establishing the functional role for variation in the number of genes. The complexity of KIR genetics increases the risk for false-positive findings and misinterpretation of genetic associations, particularly in studies with small sample size, requiring the development of improved analytical methods and better characterization of control populations. In the present study using predictive values, we provide a statistical description of LD that aids in the interpretation of associations between KIR polymorphisms.
The LD tagging approach identifies a single genetic position that marks a block of positions in LD with one another (Gu et al. 2008; Johnson et al. 2001). This approach helps to deal with the complexity of the haplotype structure by measuring whether the different polymorphisms (genes and/or alleles) carry different or redundant information. The tagging concept applies LD between KIR genes in order to help understand the complex associations observed in KIR haplotypes.
LD studies require haplotype frequency data. To supplement the lack of phase information in standard genotyping, haplotypes can be experimentally deduced from segregation analysis of pedigrees. Thus far, only numerous-offspring or multigenerational pedigree studies have been able to appropriately address both phase information and dose ambiguity (Martin et al. 2008; Middleton et al. 2007). However, KIR typing of families requires a lot of resources, and such homogeneous samples of families typed for KIR are limited (Martin et al. 2008). Describing the LD structure in the KIR gene cluster using founder haplotypes from existing family data would provide an interesting analytical tool for in-depth KIR analysis in nonfamily cases, such as genetic association analyses.
In the present study, we performed an extensive family-based study of LD in the KIR cluster, with the aim of describing the LD structure in the KIR gene cluster using founder haplotypes (Middleton et al. 2007). LD was sequentially studied at two levels: the structural level of LD (between presence or absence of KIR genes) and the allelic level of LD (between alleles of KIR genes). The KIR cluster LD was analyzed using the tagging concept outlined above and predictive values.
Dataset and methods
For this study, 106 families from Northern Ireland with numerous offspring and complex pedigrees were recruited. Three parents were available in one family. Genotyping DNA from this set of families allowed the identification of 426 founder haplotypes, which are the actual statistical unit of the study. The typing protocols used were the same as in a previous publication in which 77 of the 106 families were reported (Middleton et al. 2007). Because DNA specimens were lacking for some individuals, eight haplotypes with only partial typing at the allelic level were excluded. Thus, a total of 418 haplotypes were included in this analysis. Informed consent was obtained prior to sample collection. Only anonymous data, which protect the privacy rights of the persons involved and their family, were used in the analyses.
The following KIR genes were studied (listed in consensus order from the most centromeric to the most telomeric): KIR3DL3, KIR2DS2, KIR2DL3, KIR2DL2, KIR2DL5B, KIR2DS3, KIR2DL1, KIR2DL4, KIR3DL1, KIR3DS1, KIR2DL5A, KIR2DS5, KIR2DS1, KIR2DS4, and KIR3DL2. The physical gene position is a consensus map, and another order may better reflect the reality of a given haplotype. Although LD computations do not use the physical positions of the genes and their alleles, LD measures are not necessarily independent of gene order. For example, this is the case when one considers the possibility of KIR2DS3 and KIR2DS5 being present on either side or both sides of KIR2DL4. Each gene was typed to the allelic level, except KIR2DS2 and KIR3DL3. All parents were positive for KIR3DL3, although it is not possible to ensure that two copies are present; KIR3DL3 was considered to be always present. Allele typing together with segregation analysis allows the dose ambiguity in all other situations to be ascertained (Middleton et al. 2007).
The two historical KIR haplotype arrangements, A and B, were identified in relation to the typed KIR genes according the definition of (Middleton et al. 2007). Briefly, the basis of each A or B haplotype consists of four framework genes: KIR2DL4, KIR3DL2, KIR3DL3, and KIR3DP1. The A haplotype contains eight genes, those of this framework plus KIR2DL1, KIR2DL3, KIR2DS4, and KIR3DL1. The B haplotype is defined by the presence of one or more of the activating KIR genes in addition to the framework: KIR2DS1/2/3/5 genes, KIR3DS1, and the inhibitory genes KIR2DL5A/B and KIR2DL2. Several genes (KIR2DL1, KIR2DL3, KIR2DS4, and KIR3DL1) normally associated with the A haplotype can be found in the B haplotype. Only when all four are present was the haplotype considered to be the A haplotype. Among the 418 haplotypes studied, 230 (55.02%, with a 95% confidence interval (CI) of 50.11–59.86) were A haplotypes and 188 (44.98%, 95% CI of 40.14–49.89) were B haplotypes.
First, at the structural variation level, the LD is between the presence or absence of KIR genes, where KIR genes are taken as biallelic markers. KIR3DL3 and KIR3DL2 were assumed to be always present, preventing the assessment of LD between these genes. In the segregation analysis and allele typing, KIR2DL4 was shown to be missing in two haplotypes. This made it possible to compute LD for presence/absence between KIR2DL4 and the other KIR genes that may be present or absent. Second, LD between alleles of KIR genes was studied. KIR3DL2, which was always present in the studied sample, was not considered in LD computations. However, alleles of KIR3DL2 were considered in the second part of the analysis.
Haplotype frequencies were computed by gene counting. The total number of haplotypes used in the estimation is 418. It remained the same throughout the LD analyses. Furthermore, to better address which KIR genes are preferentially associated, positive predictive value (PPV) and negative predictive value (NPV) were used. PPV and NPV are computed using one allele/gene as the tagging marker and another gene/allele as the tagged/targeted marker. PPV is defined as the probability of occurrence of the targeted marker given the presence of the tagging marker. NPV is defined as the probability of absence of the targeted marker given the absence of the tagging marker. Each parameter is calculated as the marginal probability of the corresponding 2 × 2 table. PPV and NPV are commonly used in epidemiology and have been used to better describe polymorphism associations in the MHC region (Foissac et al. 2001). As classically performed for insertion/deletion (indel) polymorphisms, the absence of a KIR gene is considered as an allele and, therefore, can serve as the tagging marker of alleles at other loci. In order to avoid associations that may occur due to sampling fluctuations, a minimal frequency of 5% (20 occurrences) was applied. The computational and graphical display of matrix of D′ LD measures was performed in the Haploview software package (a SNP-oriented software (Barrett et al. 2005) implementing tagging algorithms) as well as the Gold software (Abecasis and Cookson 2000). Both are publicly available. The PPV/NPV-based LD analysis was performed with the STATA ver. 10 software (Stata Corp LP, College Station, TX, USA).
Results
First, we analyzed the distribution of KIR genes as copy number variants in the familial dataset (presence or absence of each KIR gene) and their associations. A parsimonious tagging strategy of KIR haplotype structural diversity was derived, leading to the quantification of the tagging ability of KIR genes toward KIR haplotype structure. Second, we analyzed those alleles of KIR genes that are in LD.
Figure 1 displays D′ pairwise LD metrics between the presence and absence of 13 KIR genes (all but KIR3DL2 and KIR3DL3, which are always present). In this approach to assess KIR LD, the KIR cluster genetic polymorphism is reduced to presence (POS) or absence (NEG) of KIR genes (Fig. 1). Higher D′ values appear toward the darker end of the gray scale and lower D′ values appear toward the lighter end. LD in KIR splits around the KIR2DL4 gene, clearly demarcating two regions, one centromeric (KIR3DL3, KIR2DS2, KIR2DL3, KIR2DL2, KIR2DL5B, KIR2DS3, and KIR2DL1) and one telomeric (KIR2DL4, KIR3DL1, KIR3DS1, KIR2DL5A, KIR2DS5, KIR2DS1, KIR2DS4, and KIR3DL2). In the centromeric region, KIR2DL5B and KIR2DS3 display moderate LD with genes of the telomeric region, with the exception of KIR2DS5. In the telomeric region, KIR2DL4 is in moderate LD with the genes of the centromeric region, in particular KIR2DL3. Hereafter, the subdivision of the KIR cluster into two regions was adopted as a suitable way to describe LD in the KIR region.
Fig. 1.
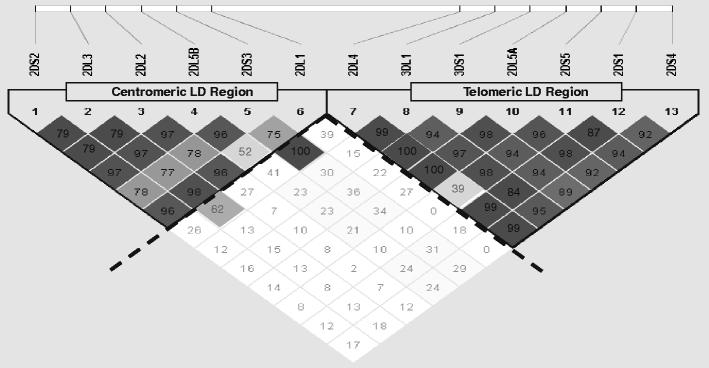
Pairwise linkage disequilibrium between the presence or absence of 13 KIR genes. Lewontin’s pairwise standardized metrics describe linkage disequilibrium (LD) between all KIR genes but KIR3DL2 and KIR3DL3 which were thought to be always present. KIR genes are considered at the structural variation level (the presence or absence level; copy number variant). Computations were double-checked using Haploview software (a SNP-oriented software) as well as the combined use of Gold software for graphical display and Stata ver. 10.0 SE for computing the LD metrics. KIR3DL1 and KIR3DS1 are considered as two different genes. The physical gene position is a consensus map, and another order may better reflect the reality of a given haplotype. Although LD computations do not use the physical positions of the genes and their alleles, LD measures are not necessarily independent of gene order. For example, this is the case when one considers the possibility of 2DS3 and 2DS5 being present on either side or both sides of 2DL4
As shown in Fig. 2, the population distribution of KIR gene haplotype structural diversity can be visually represented in a compact manner. Haplotypes are read following the thin (frequency between 0.3% and 10 %) and thick (frequency greater than 10%) connecting lines between the centromeric and telomeric regions. Haplotypes whose frequencies are less than 0.3% are not connected because the sample size does not allow the distinction of preferential associations from a rare pattern combination. As suggested by the branching of connecting lines, multiple events of recombination between KIR2DL1 and KIR2DL4 are needed to generate the observed structural diversity of these haplotypes of the centromeric and telomeric regions. The structural diversity of the haplotypes in both regions suggests multiple events of duplication and/or deletion of the genes as well as the selection and/or genetic drift events that occurred in the Northern European population. In the centromeric region, three gene structural haplotypes represent 93.8% of the haplotypes observed in our sample. In the telomeric region, two gene structural haplotypes represent 92.4%. The low LD (0.36) between regions suggests that multiple ancestral recombination events occurred between KIR2DL4 and KIR2DL1. Individuals bearing the haplotypes generated by such events do not seem to have been eliminated by selection or genetic drift. As expected, the structural diversity of B haplotypes is much greater than that of A haplotypes. The figure illustrates that the A or B haplotype definition depends on how the various structural haplotypes of both regions combine.
Fig. 2.
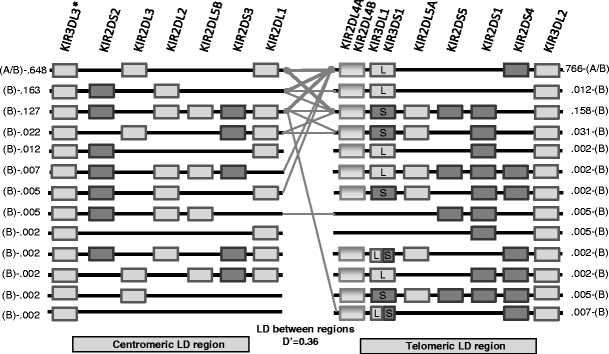
Compact illustration of the KIR genes framework haplotype using centromeric and telomeric regions. Lines indicate associations of telomeric and centromeric regions. Thin lines connect haplotypes whose frequency is between 0.3% and 10%. Thick lines connect haplotype whose frequency is greater than 10%. Rectangles indicate presence of the KIR gene, and lines represent absence of the KIR gene. All haplotype structures within the sample are displayed. This figure also presents a specific D′: the Hedrick’s multiallelic D′, which represents the degree of LD between two regions, treating each haplotype within a region as an allele. The display is adapted from the Haploview software haplotype blocks display. KIR3DL2 is always present at the telomeric end. (*) KIR3DL3 is assumed to be always present but was not typed at the allelic level in the families. All founders and offspring were positive for KIR3DL3. Because KIR3DL3 and KIR3DL2 appear to be always present, it is not possible to study LD for these genes in terms of their presence or absence. The physical gene position is a consensus map, and another order may better reflect the reality of a given haplotype. Although LD computations do not use the physical positions of the genes and their alleles, LD measures are not necessarily independent of gene order. For example, this is the case when one considers the possibility of 2DS3 and 2DS5 being present on either side or both sides of 2DL4. Frequencies and membership in the A or B haplotype are displayed on the side of the haplotypes. For example, in the centromeric region, the haplotype KIR3DL3–KIR2DL3–KIR2DL1 can belong to both A and B haplotypes and its frequency in our data is 0.648 (64.8%), it combines in more than 10% of the cases with KIR2DL4–KIR3DS1–KIR2DL5A–KIR2DS1–KIR3DL2 telomeric haplotype, which is found in B haplotypes and whose frequency is 15.8%
Subdividing the KIR cluster into two regions of LD, as displayed in Fig. 2, simplifies the description of the KIR genes’ haplotype structure. As commonly performed for SNPs, a tagging strategy was applied to the gene structural variation level of KIR haplotypes. The analysis revealed limited structural variation patterns (number of instances of presence or absence of KIR genes that are always associated). Table 1 presents the tagging approach derived from haplotype frequencies. Studying six genes is sufficient to account for the haplotype structural diversity observed in the 13 member genes. These associations of presence and/or absence of KIR genes introduce a structural framework for further KIR allelic diversity. The definition of the presence or absence of these six genes is sufficient to determine the presence or absence of all other genes in a single haplotype (r 2 = 0.8). That is, the status (present or absent) of the 13 KIR genes in a haplotype can be deduced from the presence or absence of six of them. It is worth noting that most of the tagging is achieved within each LD region (centromeric or telomeric). Interestingly, a nonpairwise LD component test (test 7, Table 1) has to be used to identify KIR2DL5B using the simultaneous presence of KIR2DL2–KIRD2S3 as a tagging marker, meaning that KIR2DL5B is present when both KIR2DL2 and KIR2DS3 are present. Unlike the other tagging genes, the absence (not the presence) of KIR3DL1 is taken into account. Similarly, unlike other tagged genes, KIR2DS4 and KIR2DL3 are deduced to be absent instead of present. With regard to tests 4 and 5, even if KIR2DS3 and KIR2DL4 are in complete LD with one another, they are embedded in different haplotypes and need to be tested separately. In addition, with an r 2 = 0.87 (not shown), the absence of KIR3DL1 tagging the absence of KIR2DS4 is the least efficient tagging relationship. For studies in which not all KIR genes are typed, such a table can help extend the possible interpretation to other untyped KIR genes.
Table 1.
Tagging strategy of KIR genes using LD between genes to minimize the number of KIR genes to study
| Test | KIR genes | KIR genes captured through LD |
|---|---|---|
| 1 | Absence of KIR3DL1 | KIR2DL5A; KIR3DS1; KIR2DS1; absence of KIR2DS4 |
| 2 | KIR2DL2 | KIR2DS2; absence of KIR2DL3 |
| 3 | KIR2DS5 | |
| 4 | KIR2DS3 | |
| 5 | KIR2DL4 | |
| 6 | KIR2DL1 | |
| 7 | KIR2DL2–KIRD2S3 | KIR2DL5B |
According to the haplotype display in Fig. 2, a parsimonious strategy of testing for the presence or absence of KIR genes can be drawn. The 13 genes can be tested using six genes and seven tests. In the tagging approach, KIR2DL3, KIR3DL1, and KIR2DS4 differ from the other genes because it is not the genes themselves that are taken into account but their absence. Test 7 corresponds to the simultaneous presence of KIR2DL2 and KIR2DS3. Test 7 cannot be directly inferred from the data in Table 2 and must consider KIR2DL2–KIR2DS3 or KIR2DS2–KIR2DS3 structural haplotypes to capture the KIR2DL5B gene (Fig. 2)
Association studies more conveniently handle pairwise relationships among genes. For example, it is commonly noted that KIR2DL2 and KIR2DS2 are “preferentially associated” with one another. To both quantify and generalize such a statement to any pair of KIR genes, in addition to Table 1, Supplementary Tables S1 and S2 for the centromeric region and S3 and S4 for the telomeric region present PPVs and NPVs between the presence or absence of all KIR genes. PPVs quantify associations due to both LD and high frequency of the reference haplotype. For example, when KIR2DS2 is absent, there is a 100% chance that KIR2DL2 is absent as well and an 82.39% chance that KIR2DS4 is present (PPVs). Similarly for NPVs, when KIR2DS4 is NOT present, there is a 39.33% chance that KIR2DL2 is present and a 94.38% chance that KIR2DS1 is present.
We now present results for LD at the allelic level (preferential association of alleles). KIR3DL1 and KIR3DS1 are considered as mutually exclusive alleles of the same gene (the “KIR3DL1/S1” locus): the simultaneous presence in cis of the KIR3DL1 and KIR3DS1 alleles can occur and is considered as a specific KIR3DL1/S1 allele. It includes haplotypes that have two copies of one gene (Middleton et al. 2007). This analysis includes KIR3DL2, which is always present but shows allelic variation. Data on LD between genes were computed using allelic information. Tables 2 and 3 present, respectively, the highest LD components between KIR alleles in the centromeric LD region (KIR2DL3, KIR2DL2, KIR2DL5B, KIR2DS3, KIR2DL1) and in the telomeric LD region (KIR2DL4, KIR3DL1/KIR3DS1, KIR2DL5A, KIR2DS5, KIR2DS1, KIR2DS4, KIR3DL2). Only tagging relations that have a minimal PPV of 65% are displayed. For robust estimation, only haplotypes with a frequency over 5% were studied.
Table 2.
KIR gene alleles of the centromeric LD region tagging other KIR gene alleles
| KIR tagging allele | KIR tagged allele | PPV | NPV | Haplotype frequency | D' (*) | |
|---|---|---|---|---|---|---|
| KIR2DL3 | *002 | KIR2DL1*002 | 94.30% | 99.70% | 23.90% | 0.45 |
| *001 | KIR2DL1*00302 | 94.80% | 96.60% | 35.20% | 0.45 | |
| KIR2DL2 | *003 | KIR2DS2*POS | 100.00% | 77.60% | 12.40% | 0.79 |
| *001 | KIR2DS2*POS | 100.00% | 83.30% | 18.40% | 0.79 | |
| *001 | KIR2DL5B*002 | 71.40% | 99.40% | 13.20% | 0.95 | |
| KIR2DL5B | *002 | KIR2DS2*POS | 98.20% | 78.40% | 13.40% | 0.97 |
| *002 | KIR2DL1*00401 | 78.90% | 98.10% | 10.80% | 0.82 | |
| *002 | KIR2DS3*00103 | 84.20% | 98.60% | 11.50% | 0.84 | |
| *002 | KIR2DL2*001 | 96.50% | 93.90% | 13.20% | 0.95 | |
| KIR2DS3 | *00103 | KIR2DS2*POS | 90.60% | 76.40% | 11.50% | 0.79 |
| *00103 | KIR2DL1*00401 | 69.80% | 95.90% | 8.90% | 0.77 | |
| *00103 | KIR2DL5B*002 | 90.60% | 97.50% | 11.50% | 0.84 | |
| *00103 | KIR2DL2*001 | 86.80% | 91.50% | 11.00% | 0.79 | |
| KIR2DL1 | *00401 | KIR2DS2*POS | 100.00% | 77.60% | 12.40% | 0.75 |
| *00401 | KIR2DL5B*002 | 86.50% | 96.70% | 10.80% | 0.82 | |
| *00401 | KIR2DS3*00103 | 71.20% | 95.60% | 8.90% | 0.77 | |
| *00401 | KIR2DL2*001 | 84.60% | 91.00% | 10.50% | 0.74 | |
| *00302 | KIR2DL3*001 | 94.20% | 96.90% | 35.20% | 0.45 | |
| *002 | KIR2DL3*002 | 99.00% | 98.10% | 23.90% | 0.45 | |
Only alleles of the previously defined centromeric region are displayed. Other constraints were applied: tagging must have a minimal PPV of 65%. For robustness of the estimation, only haplotypes of more than 5% were studied; haplotypes without KIR gene at another locus were not studied. (*) PPV and NPV drastically differ from D′. PPV and NPV are measures of association at the allelic level, whereas D′ is a locus-level average measure of LD. D′ stands for the pairwise standardized Hedrick’s measure of LD between multiallelic genes. Confidence intervals can be computed because frequencies were estimated by gene counting. The total number of haplotypes is 418 (~209 founders of the families). For example (2nd row), the probability of having KIR2DL1*00302 given the presence of KIR2DL3*001 is 94.8%, the probability of not having KIR2DL1*00302 given the absence of KIR2DL3*001 is 96.6%, the KIR2DL3*001–KIR2DL1*00302 haplotype frequency is 35.2%, and D′ between KIR2DL3 and KIR2DL1 is 0.45
PPV positive predictive value, the probability of the tagged KIR allele given the presence of the tagging KIR allele; NPV negative predictive value, the probability of the absence of the tagged KIR allele given the absence of the tagging KIR allele
Table 3.
KIR gene alleles of the telomeric LD region tagging other KIR genes alleles
| KIR tagging allele | KIR tagged allele | PPV | NPV | Haplotype Frequency | D′ (*) | |
|---|---|---|---|---|---|---|
| KIR2DL4 | *011 | KIR3DL1/S1*005 | 100.00% | 99.70% | 13.60% | 0.21 |
| *011 | KIR2DS4*003 | 100.00% | 72.30% | 13.60% | 0.49 | |
| *011 | KIR3DL2*001 | 77.20% | 85.90% | 10.50% | 0.27 | |
| *00802 | KIR2DS4*006 | 98.60% | 99.40% | 16.70% | 0.49 | |
| *00802 | KIR3DL1/S1*004 | 95.80% | 99.70% | 16.30% | 0.21 | |
| *00801 | KIR2DS4*003 | 94.90% | 75.60% | 17.70% | 0.49 | |
| *00801 | KIR3DL1/S1*00101 | 89.70% | 99.70% | 16.70% | 0.21 | |
| *00501 | KIR3DL1/S1*013 | 93.80% | 99.70% | 18.20% | 0.21 | |
| *00501 | KIR3DL2*007 | 95.10% | 97.60% | 18.40% | 0.27 | |
| *00501 | KIR2DS1*002 | 100.00% | 97.60% | 19.40% | 0.71 | |
| *00501 | KIR2DS5*002 | 82.70% | 98.80% | 16.00% | 0.86 | |
| *00501 | KIR2DL5A*001 | 79.00% | 99.40% | 15.30% | 0.69 | |
| *00103 | KIR3DL2*009 | 81.30% | 99.20% | 6.20% | 0.27 | |
| *00103 | KIR3DL1/S1*008 | 75.00% | 100.00% | 5.70% | 0.21 | |
| *00103 | KIR2DS4*003 | 71.90% | 65.30% | 5.50% | 0.49 | |
| *00102 | KIR3DL2*002 | 100.00% | 99.10% | 17.50% | 0.27 | |
| *00102 | KIR2DS4*00101 | 98.60% | 96.80% | 17.20% | 0.49 | |
| KIR3DL1/S1 | *01502 | KIR3DL2*002 | 100.00% | 89.30% | 8.40% | 0.25 |
| *01502 | KIR2DL4*00102 | 100.00% | 90.10% | 8.40% | 0.21 | |
| *01502 | KIR2DS4*00101 | 97.10% | 87.20% | 8.10% | 0.47 | |
| *013 | KIR3DL2*007 | 94.80% | 96.50% | 17.50% | 0.25 | |
| *013 | KIR2DL4*00501 | 98.70% | 98.50% | 18.20% | 0.21 | |
| *013 | KIR2DS1*002 | 100.00% | 96.50% | 18.40% | 0.71 | |
| *013 | KIR2DS5*002 | 81.80% | 97.70% | 15.10% | 0.84 | |
| *013 | KIR2DL5A*001 | 77.90% | 98.20% | 14.40% | 0.69 | |
| *008 | KIR2DS4*003 | 95.80% | 66.00% | 5.50% | 0.47 | |
| *008 | KIR2DL4*00103 | 100.00% | 98.00% | 5.70% | 0.21 | |
| *005 | KIR2DL4*011 | 98.30% | 100.00% | 13.60% | 0.21 | |
| *005 | KIR2DS4*003 | 98.30% | 72.20% | 13.60% | 0.47 | |
| *005 | KIR3DL2*001 | 75.90% | 85.80% | 10.50% | 0.25 | |
| *004 | KIR2DL4*00802 | 98.60% | 99.10% | 16.30% | 0.21 | |
| *004 | KIR2DS4*006 | 98.60% | 98.90% | 16.30% | 0.47 | |
| *002 | KIR3DL2*002 | 100.00% | 89.50% | 8.60% | 0.25 | |
| *002 | KIR2DL4*00102 | 100.00% | 90.30% | 8.60% | 0.21 | |
| *002 | KIR2DS4*00101 | 100.00% | 87.70% | 8.60% | 0.47 | |
| *00101 | KIR2DL4*00801 | 98.60% | 97.70% | 16.70% | 0.21 | |
| *00101 | KIR2DS4*003 | 95.80% | 74.40% | 16.30% | 0.47 | |
| KIR2DL5A | *001 | KIR3DL1/S1*013 | 90.90% | 95.20% | 14.40% | 0.69 |
| *001 | KIR3DL2*007 | 95.50% | 93.80% | 15.10% | 0.8 | |
| *001 | KIR2DL4*00501 | 97.00% | 95.20% | 15.30% | 0.69 | |
| *001 | KIR2DS1*002 | 100.00% | 93.50% | 15.80% | 0.85 | |
| *001 | KIR2DS5*002 | 100.00% | 98.60% | 15.80% | 0.83 | |
| KIR2DS5 | *002 | KIR3DL1/S1*013 | 88.70% | 96.00% | 15.10% | 0.84 |
| *002 | KIR3DL2*007 | 95.80% | 95.10% | 16.30% | 0.84 | |
| *002 | KIR2DL4*00501 | 94.40% | 96.00% | 16.00% | 0.86 | |
| *002 | KIR2DS1*002 | 100.00% | 94.80% | 17.00% | 0.87 | |
| *002 | KIR2DL5A*001 | 93.00% | 100.00% | 15.80% | 0.83 | |
| KIR2DS1 | *002 | KIR3DL1/S1*013 | 86.50% | 100.00% | 18.40% | 0.71 |
| *002 | KIR3DL2*007 | 92.10% | 99.10% | 19.60% | 0.86 | |
| *002 | KIR2DL4*00501 | 91.00% | 100.00% | 19.40% | 0.71 | |
| *002 | KIR2DS5*002 | 79.80% | 100.00% | 17.00% | 0.87 | |
| *002 | KIR2DL5A*001 | 74.20% | 100.00% | 15.80% | 0.85 | |
| KIR2DS4 | *006 | KIR2DL4*00802 | 97.20% | 99.70% | 16.70% | 0.49 |
| *006 | KIR3DL1/S1*004 | 94.40% | 99.70% | 16.30% | 0.47 | |
| *00101 | KIR3DL2*002 | 89.20% | 99.40% | 17.70% | 0.62 | |
| *00101 | KIR2DL4*00102 | 86.70% | 99.70% | 17.20% | 0.49 | |
| KIR3DL2 | *011 | KIR2DL4*00801 | 95.70% | 85.80% | 5.30% | 0.27 |
| *011 | KIR2DS4*003 | 91.30% | 65.60% | 5.00% | 0.62 | |
| *009 | KIR2DL4*00103 | 89.70% | 98.50% | 6.20% | 0.27 | |
| *007 | KIR3DL1/S1*013 | 85.90% | 98.80% | 17.50% | 0.25 | |
| *007 | KIR2DL4*00501 | 90.60% | 98.80% | 18.40% | 0.27 | |
| *007 | KIR2DS1*002 | 96.50% | 97.90% | 19.60% | 0.86 | |
| *007 | KIR2DS5*002 | 80.00% | 99.10% | 16.30% | 0.84 | |
| *007 | KIR2DL5A*001 | 74.10% | 99.10% | 15.10% | 0.8 | |
| *005 | KIR2DL4*00802 | 92.10% | 90.50% | 8.40% | 0.27 | |
| *005 | KIR2DS4*006 | 94.70% | 90.50% | 8.60% | 0.62 | |
| *005 | KIR3DL1/S1*004 | 86.80% | 90.50% | 7.90% | 0.25 | |
| *003 | KIR2DL4*00802 | 93.90% | 89.60% | 7.40% | 0.27 | |
| *003 | KIR2DS4*006 | 97.00% | 89.60% | 7.70% | 0.62 | |
| *003 | KIR3DL1/S1*004 | 93.90% | 90.10% | 7.40% | 0.25 | |
| *002 | KIR2DL4*00102 | 96.10% | 100.00% | 17.50% | 0.27 | |
| *002 | KIR2DS4*00101 | 97.40% | 97.40% | 17.70% | 0.62 | |
| *001 | KIR2DS4*003 | 95.80% | 79.60% | 21.80% | 0.62 | |
Only alleles of the previously defined central telomeric region are displayed. Other constraints were applied: Tagging must have a minimal PPV of 65%. For robustness of the estimation, only haplotypes of more than 5% were studied; haplotypewithout the KIR gene at another locus were not studied. (*) PPV and NPV drastically differ from D′. PPV and NPV are measures of association at the allelic level, whereas D′ is a locus-level average measure of LD. Confidence intervals can be computed because frequencies were estimated by gene counting. The total number of haplotype is 418 (~209 founders of the families). For example, (1st row) the probability of having KIR3DL1/S1*005 given the presence of KIR2DL4*011 is 100.0%, the probability of not having KIR3DL1/S1*005 given that KIR2DL4*011 is absent in 99.7%, the KIR3DL1/S1*005–KIR2DL4*0011 haplotype frequency is 13.6%, and D′ between KIR2DL4 and KIR3DL1/S1 is 0.21
PPV positive predictive value, the probability of the tagged KIR allele given the presence of the tagging KIR allele; NPV negative predictive value, the probability of the absence of the tagged KIR allele given the absence of the tagging KIR allele
As expected, strong LD between KIR genes (as measured by D′) is driven by specific allelic associations (high PPV, high NPV, and high D′; Table 2). For example, while KIR2DS3 and KIR2DL1 are in strong LD (D′ computed with alleles = 0.77), KIR2DS3*00103 specifically associates with KIR2DL1*00401: KIR2DL1*00401 is present in 69.8% of the cases when KIR2DS3*00103 is present (i.e., PPV = 69.8%). Interestingly, several alleles of KIR genes that are not in strong LD were found to tag each other (high PPV, high NPV at allelic level, and low D′ at averaged level between loci). That is the case for two allelic associations between KIR2DL1 and KIR2DL3 (D′ computed with alleles = 0.45): KIR2DL1*00302 tags KIR2DL3*001 (PPV = 94.2%, NPV = 96.9%) and KIR2DL1*002 tags KIR2DL3*002 (PPV = 99.0%, NPV = 98.1%). In addition to the balance between KIR2DS2 and KIR2DL2*001 associated with KIR2DL5B*002, KIR2DS3*00103 and KIR2DL1*00401 may have a specific role in tuning the NK cell response. The data in Table 2 thus suggest the existence of very strong LD spanning the centromeric KIR region: KIR2DS2*POS–KIR2DL2*001–KIR2DL5B*002–KIR2DS3*00103–KIR2DL1*00401 (as listed in Table 4), as each of these genes/alleles has a high PPV.
Table 4.
Haplotypes exhibiting the strongest KIR allele associations in the KIR gene cluster
| Centromeric haplotype |
| KIR2DS2*POS–KIR2DL2*001–KIR2DL5B*002–KIR2DS3*00103–KIR2DL1*00401 |
| Telomeric haplotypes |
| KIR2DL4*011–KIR3DL1*005–KIR2DS4*003–KIR3DL2*003 |
| KIR2DL4*00802–KIR3DL1*004–KIR2DS4*006– KIR3DL2*005 |
| KIR2DL4*00801–KIR3DL1*00101–KIR2DS4*003–KIR3DL2*001 |
| KIR2DL4*00103–KIR3DL1*008–KIR2DS4*003–KIR3DL2*009 |
| KIR2DL4*00102–KIR3DL1*01502/*002–KIR2DS4*00101–KIR3DL2*002 |
| KIR2DL4*00801–KIR2DS4*003–KIR3DL2*011 |
| KIR2DL4*00501–KIR3DS1*013–KIR2DL5A*001–KIR2DS5*002–KIR2DS1*002–KIR3DL2*007 |
The telomeric LD regions show more allele diversity as well as a greater number of tagging-like relationships between KIR alleles (Table 3). As observed in the centromeric region, strong LD between KIR genes is driven by specific allelic association (high PPV, high NPV, and high D′). For example, KIR3DL1/S1*013 tags KIR2DS1*002, KIR2DS5*002, and KIR2DL5A*001, and KIR2DL4*00501 tags KIR2DS1*002, KIR2DS5*002, and KIR2DL5A*001. KIR2DL4, KIR3DL1/S1, and KIR3DL2 have a particularly high number of alleles included in haplotypes in strong LD (Table 3). Interestingly, these associations can extend across relatively low pairwise LD between loci (high PPV, high NPV, and low D′). For example, KIR3DL1/S1*013 tags KIR2DL4*00501 and KIR3DL2*007 and KIR2DL4*00501 tags KIR3DL1/KIR3DS1*013 and KIR3DL2*007 (Table 4).
More haplotypes in high LD are found in the telomeric region (see allele association in Table 3 and haplotype in Table 4). Most of them involve KIR2DL4, KIR3DL1, and KIR3DL2. Seven telomeric haplotypes explain most of the 2 × 2 associations observed (Table 4). Two additional extended haplotypes involved different genes (Table 4). One points to the potential specificity of the association between KIR3DL2*011 and KIR2DS4*003 in the context of KIR2DL4*00801, and the other extended haplotype involves a KIR3DS1 allele (KIR2DL4*00501–KIR3DL1/S1*013–KIR2DL5A*001–KIR2DS5*002–KIR2DS1*002–KIR3DL2*007). Table 4 provides a list of extended haplotypes in KIR genes.
Discussion
In this study, LD was analyzed at two levels: the structural presence or absence of KIR genes in haplotypes and the allelic variation of KIR genes. A tagging approach demonstrated that the strongest LD, at both the structural and allelic levels, is found within the centromeric region from KIR3DL3 to KIR2DL1 and within the telomeric region from KIR2DL4 to KIR3DL2. This LD pattern was used to identify the tagging properties of specific genes and alleles, and a series of tagging associations were presented. These patterns reduced the number of markers that must be considered in the association analysis. Using predictive values to assess LD in KIR polymorphisms is complementary to classical r 2 and D′ and aids in efforts to evaluate the genes or alleles expected to be found together due to LD based on specific KIR associations with diseases.
To our knowledge, this study is the largest one using a familial pedigree design to define KIR haplotypes. Moreover, it was performed using one of the most homogeneous European populations, as all Northern Irish families are of European ancestry, with very little history of immigration. Compared to a previous report on Irish families (Middleton et al. 2007), the number of haplotypes studied increased more than 33% (from 308 to 416 founder haplotypes), providing a more accurate dataset for a more representative and more detailed description of LD. In previous family-based studies of LD in the KIR region, the sample sizes, heterogeneity of the families, and lower typing resolution did not permit a comprehensive quantification of LD (Martin et al. 2008; Whang et al. 2005). The present results were obtained using a systematic approach. They reflect the major characteristics of KIR LD in populations of Northern European ancestry. Although not typed to the allelic level, the analysis of LD by Whang et al. (2005) in families of Korean ancestry suggests that LD in KIR is affected by the variation of the proportions of A and B haplotypes in both the telomeric and centromeric regions.
Regarding LD between genes, the use of the tag approach confirms previous classical LD findings (Norman et al. 2004; Single et al. 2008) as well as nonpairwise associations such as KIR2DL2–KIR2DS3–KIR2DL5B (Ordonez et al. 2008). Such associations can be used to detect genotyping error when unexpected associations are found. At the structural level, we observed the same LD pattern for KIR3DL1 and KIR3DS1 as that of Martin et al. (2008). KIR3DL1 and KIR3DS1 were presented as the two alleles of the KIR3DL1/S1 locus because allele typing was performed in the present study. To clarify the multiple features of LD in KIR, LD was formulated in terms of positive and negative predictive value. Strong LD between KIR genes (as measured by D′) is driven by specific allelic associations. The present data considerably update the previous allelic associations described by Shilling et al. (Shilling et al. 2002), as the improvement of typing techniques and the greater number of families studied allowed for a more comprehensive analysis. Although LD between alleles of KIR2DL3, KIR2DL1, KIR3DL1, and KIR3DL2 was studied by Shilling et al., many more associations are presented here. Consequently, very few direct comparisons between the results can be made. The most significant (p < 0.0001) positive LD components reported by Shilling et al. (2002) are found among our results: KIR3DL2*003 tags KIR3DL1*004, KIR3DL2*005 tags KIR3DL1*004, KIR3DL2*007 tags KIR3DS1 (KIR3DL1/S1*013 in present study), and KIR2DL1*003 (refined to be *00302 here) tags KIR2DL3*001. In addition, because haplotypes were experimentally deduced, no “blank” alleles had to be defined in the present study. By looking only at the most frequent haplotypes, LD associations are more robust because a minimal frequency of 5% corresponds to a minimal number of haplotypes of more than 20. In addition, the associations were quantified rather than qualitatively evaluated for their statistical significance (H 0: D′ ≠ 0). Such approaches allow the identification of extended KIR allele haplotypes that are in strong LD. These are the KIR equivalents of the extended or conserved haplotypes of HLA alleles (most-frequent-most-in-LD haplotypes, also termed “ancestral haplotypes” and “complotypes” (Alper et al. 1992; Dawkins et al. 1983)).
For association studies, analysis at the DNA level of the presence or absence of KIR genes, or structural variation such as CNV, should take into account unexpressed and/or untranscribed alleles, as well as the genetic structure within these complex loci, to make sense of observed associations with diseases. Furthermore, the same approach can be used for various subgroups of KIR haplotypes, such as the A and B haplotypes separately, the presence or absence of KIR2DS4, high and low expression KIR3DL1 alleles, or expressed and unexpressed KIR2DS4 alleles. Such analyses would provide an alternative view of the LD associations between KIR alleles conditioned by the characteristics of the subgroup of haplotypes. When computed between polymorphic loci, LD appears to be much smaller in A haplotypes than in B haplotypes. In B haplotypes, an unrevealed LD component between KIR2DS2 (and KIR2DL2) and telomeric genes is observed. KIR2DS1 and KIR2DS2 are strongly associated in B haplotypes (D′ = 0.96, data not shown (Gourraud et al. 2008)). Strong LD components are also found in KIR2DS4* negative haplotypes. Because PPV and NPV also can be computed in any subset of haplotypes in populations of various ancestries, the PPV/NPV-based approach of LD can be applied to any subset of haplotypes. Dedicated software is needed to make such analyses easier to achieve.
The numerous statistics presented here are relevant for association studies. They provide a simplified, working approach and an overview of the associations between KIR polymorphisms. For example, two studies suggested that KIR2DS1 is associated with psoriasis vulgaris (Luszczek et al. 2004; Ploski et al. 2006), but another paper suggested that both KIR2DS1 and KIR2DL5 are associated with this disease (Suzuki et al. 2004). The data in Tables S1 to S4 shed light on these findings, revealing a strong association between the presence of KIR2DS1 and the presence of KIR2DL5A (PPV = 96.5%, NPV = 99.7%) and between the presence of KIR2DS1with the absence of KIR2DL5B (PPV = 73.3%), although KIR2DL5B can be present together with KIR2DS1 (NPV = 10.8%). As hypothesized for extended HLA haplotypes, these KIR haplotypes might have specific immunopathological properties that should be further studied as potentially coexpressed receptors. PPVs and NPVs account for both allele frequencies and LD structure. In addition, PPVs and NPVs describe LD in ways that are easier to express in natural language than r 2 and D′. Classically, high LD can be created by genetic drift, selection, and the founder effect. Although our sample of KIR families is the largest ever published, the sample size may be not sufficient to observe all possible haplotypes. Due to the sampling effect, this can result in an underestimation of the KIR haplotype diversity and an overestimation of LD.
The associations between KIR revealed in the present genetic study also can be of specific relevance for expression and functional studies. For example, Pando et al. (2003) suggested that the KIR3DL1*004 protein is poorly expressed at cell surfaces due to substitution at position 86 in Ig domain 0 and position 182 in Ig domain 1. The present data indicated that the KIR3DL1/S1*004 allele is embedded in the haplotype KIR2DL4*00802–KIR3DL1/S1*004–KIR2DS4*006–KIR3DL2*005, suggesting that KIR3DL2*005 and/or KIR2DL4*00802 might play a more significant role in compensating for the poorly expressed KIR3DL1 protein. Because the absence of a KIR gene must be taken as a null allele at a given locus, it is less important to define a KIR gene than to perform typing at the allelic level. Mathematically and for geneticists, the absence of a gene may be treated formally as an allele. With regard to immunological relevance, the absence of a gene should be considered as equivalent to alleles with no or poor expression.
Family approaches require great effort, and samples of unrelated individuals also may be of interest in studying LD in KIR. Similar analyses performed with estimated haplotype frequencies offer more accessible opportunities to challenge LD findings in non-Caucasian populations using designs based on families and/or unrelated individuals with (Single et al. 2008; Yoo et al. 2007) or without (Gourraud et al. 2007) a priori assumptions on the possible haplotypes. These techniques are much more appropriate than the use of Arlequin (Excoffier et al. 2005) for KIR, as performed when PYPOP, HAPLO-IHP, ESTIHAPLO software were not available (Shilling et al. 2002). The LD data can provide a framework of the plausible gene structural haplotypes of the KIR region. Assuming that all the gene structural haplotypes of the centromeric and telomeric regions could be combined, the extent of structural variation would reach a maximum of 169 haplotypes. Bringing allelic diversity to these theoretically possible gene structural haplotypes would greatly help software to handle the possible haplotype diversity. In addition, starting values for expectation–maximization algorithms can adequately reflect the distribution of KIR haplotype diversity of the three most frequent gene structural haplotypes in the centromeric region with the two most frequent gene structural haplotypes in the telomeric region. The 418 haplotypes experimentally deduced in the present study highlights the great structural diversity of the KIR cluster, which can be underestimated in studies with smaller sample sizes.
Our description of LD in the KIR region suggests that meiotic recombination combines with an ancestral core diversity to create new KIR phenotypes upon which natural selection and genetic drift regenerate LD. Our findings shed light on the biological implications of KIR gene associations in terms of the potential balance between inhibitory and activating effects. We hypothesize that selection would favor specific combinations of activating and inhibitory KIR genes and alleles. Our findings also suggest that the distinction between genes and alleles is artificial, as recently observed by Norman et al. (2009). The results clearly show that using the notion of blocks to describe the complex structure of LD is a good first step. For the sake of simplicity, we describe KIR LD using the two regions where the strongest associations occur, but this does not exclude other LD associations between these two regions. For example, at the allelic level, we observed that KIR2DL4*011 is in LD with KIR2DL1*00302 (PPV = 65%, NPV = 67%, D′ = 0.44; data not shown) in addition to a low LD averaged over all allele pairs between KIR2DL4 and KIR2DL1 (D′ = 0.22).
LD refers to the complex nonrandom association of genetic polymorphisms. The various dimensions of LD in such a complex gene cluster as the KIR region is difficult to summarize. LD may be quantified in order to adequately interpret statistical associations between genetic polymorphisms in genotypes. The PPV/NPV-based LD analysis presented here may be used to supplement classical pairwise analyses by identifying the extended haplotypes that are the best candidates for conferring differential genetic risks to human diseases. In turn, such an approach may provide an evolutionary overview of the genetic control of the activating/inhibitory balance of NK cell activity in the immune response.
Electronic supplementary material
Below is the link to the electronic supplementary material.
KIR2DS2, KIR2DL3, and KIR2DL2 genes whose presence or absence in the centromeric LD region tags the presence or absence of other KIR genes (DOC 100 kb)
KIR2DL5B, KIR2DS3, and KIR2DL1 genes whose presence or absence in the centromeric LD region tags the presence or absence of other KIR genes (DOC 93 kb)
KIR2DL4, KIR3DL1, KIR3DS1, and KIR2DL5A genes whose presence or absence in the telomeric LD region tags the presence or absence of other KIR genes (DOC 93 kb)
KIR2DS2, KIR2DS1, and KIR2DS4 genes whose presence or absence in the telomeric LD region tags the presence or absence of others KIR genes (DOC 81 kb)
Acknowledgments
We thank the families that participated in this study. We also thank Jason D. Barbour, Jorge R. Oksenberg, and Jill Hollenbach for insightful discussions.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Footnotes
Contributions
Ashley Meenagh performed the genotyping. Pierre-Antoine Gourraud set up the analytical strategy. Anne Cambon-Thomsen contributed to the discussion and the manuscript. Derek Middleton and Pierre-Antoine Gourraud conceived the research and wrote the paper.
References
- Abecasis GR, Cookson WO. GOLD–graphical overview of linkage disequilibrium. Bioinformatics. 2000;16:182–183. doi: 10.1093/bioinformatics/16.2.182. [DOI] [PubMed] [Google Scholar]
- Alper CA, Awdeh Z, Yunis EJ. Conserved, extended MHC haplotypes. Exp Clin Immunogenet. 1992;9:58–71. [PubMed] [Google Scholar]
- Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–265. doi: 10.1093/bioinformatics/bth457. [DOI] [PubMed] [Google Scholar]
- Bashirova AA, Martin MP, McVicar DW, Carrington M. The killer immunoglobulin-like receptor gene cluster: tuning the genome for defense. Annu Rev Genomics Hum Genet. 2006;7:277–300. doi: 10.1146/annurev.genom.7.080505.115726. [DOI] [PubMed] [Google Scholar]
- Carrington M, Martin MP. The impact of variation at the KIR gene cluster on human disease. Curr Top Microbiol Immunol. 2006;298:225–257. doi: 10.1007/3-540-27743-9_12. [DOI] [PubMed] [Google Scholar]
- Dawkins RL, Christiansen FT, Kay PH, Garlepp M, McCluskey J, Hollingsworth PN, Zilko PJ. Disease associations with complotypes, supratypes and haplotypes. Immunol Rev. 1983;70:1–22. doi: 10.1111/j.1600-065X.1983.tb00707.x. [DOI] [PubMed] [Google Scholar]
- Excoffier L, Laval G, Schneider S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online. 2005;1:47–50. [PMC free article] [PubMed] [Google Scholar]
- Foissac A, Fort M, Clayton J, Abbal M, Raffoux C, Moine A, Bensa JC, Bignon JD, Mercier P, Cambon-Thomsen A. Microsatellites in the HLA region: HLA prediction and strategies for bone marrow donor registries. Transplant Proc. 2001;33:491–492. doi: 10.1016/S0041-1345(00)02107-2. [DOI] [PubMed] [Google Scholar]
- Gourraud PA, Gagne K, Bignon JD, Cambon-Thomsen A, Middleton D. Preliminary analysis of a KIR haplotype estimation algorithm: a simulation study. Tissue Antigens. 2007;69(Suppl 1):96–100. doi: 10.1111/j.1399-0039.2006.762_4.x. [DOI] [PubMed] [Google Scholar]
- Gourraud PA, Meenagh A, Cambon-Thomsen A, Middleton D. Linkage disequilibrium patterns in the KIR cluster: several point of view on the same non independent association of KIR genes polymorphisms. Tissue Antigens. 2008;72:299. [Google Scholar]
- Gu CC, Yu K, Ketkar S, Templeton AR, Rao DC. On transferability of genome-wide tagSNPs. Genet Epidemiol. 2008;32:89–97. doi: 10.1002/gepi.20269. [DOI] [PubMed] [Google Scholar]
- Hiby SE, Regan L, Lo W, Farrell L, Carrington M, Moffett A. Association of maternal killer-cell immunoglobulin-like receptors and parental HLA-C genotypes with recurrent miscarriage. Hum Reprod. 2008;23:972–976. doi: 10.1093/humrep/den011. [DOI] [PubMed] [Google Scholar]
- Hsu KC, Chida S, Geraghty DE, Dupont B. The killer cell immunoglobulin-like receptor (KIR) genomic region: gene-order, haplotypes and allelic polymorphism. Immunol Rev. 2002;190:40–52. doi: 10.1034/j.1600-065X.2002.19004.x. [DOI] [PubMed] [Google Scholar]
- Johnson GC, Esposito L, Barratt BJ, Smith AN, Heward J, Di Genova G, Ueda H, Cordell HJ, Eaves IA, Dudbridge F, Twells RC, Payne F, Hughes W, Nutland S, Stevens H, Carr P, Tuomilehto-Wolf E, Tuomilehto J, Gough SC, Clayton DG, Todd JA. Haplotype tagging for the identification of common disease genes. Nat Genet. 2001;29:233–237. doi: 10.1038/ng1001-233. [DOI] [PubMed] [Google Scholar]
- Khakoo SI, Thio CL, Martin MP, Brooks CR, Gao X, Astemborski J, Cheng J, Goedert JJ, Vlahov D, Hilgartner M, Cox S, Little AM, Alexander GJ, Cramp ME, O'Brien SJ, Rosenberg WM, Thomas DL, Carrington M. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science. 2004;305:872–874. doi: 10.1126/science.1097670. [DOI] [PubMed] [Google Scholar]
- Lowe DP, Cook MA, Bowman SJ, Briggs DC. Association of killer cell immunoglobulin-like receptors with primary Sjogren's syndrome. Rheumatology (Oxford) 2009;48:359–362. doi: 10.1093/rheumatology/ken503. [DOI] [PubMed] [Google Scholar]
- Luszczek W, Manczak M, Cislo M, Nockowski P, Wisniewski A, Jasek M, Kusnierczyk P. Gene for the activating natural killer cell receptor, KIR2DS1, is associated with susceptibility to psoriasis vulgaris. Hum Immunol. 2004;65:758–766. doi: 10.1016/j.humimm.2004.05.008. [DOI] [PubMed] [Google Scholar]
- Martin MP, Qi Y, Gao X, Yamada E, Martin JN, Pereyra F, Colombo S, Brown EE, Shupert WL, Phair J, Goedert JJ, Buchbinder S, Kirk GD, Telenti A, Connors M, O'Brien SJ, Walker BD, Parham P, Deeks SG, McVicar DW, Carrington M. Innate partnership of HLA-B and KIR3DL1 subtypes against HIV-1. Nat Genet. 2007;39:733–740. doi: 10.1038/ng2035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin MP, Single RM, Wilson MJ, Trowsdale J, Carrington M. KIR haplotypes defined by segregation analysis in 59 Centre d'Etude Polymorphisme Humain (CEPH) families. Immunogenetics. 2008;60:767–774. doi: 10.1007/s00251-008-0334-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Middleton D, Gonzelez F. The extensive polymorphism of KIR genes. Immunology. 2010;129:8–19. doi: 10.1111/j.1365-2567.2009.03208.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Middleton D, Meenagh A, Gourraud PA. KIR haplotype content at the allele level in 77 Northern Irish families. Immunogenetics. 2007;59:145–158. doi: 10.1007/s00251-006-0181-7. [DOI] [PubMed] [Google Scholar]
- Middleton D, Diler AS, Meenagh A, Sleator C, Gourraud PA. Killer immunoglobulin-like receptors (KIR2DL2 and/or KIR2DS2) in presence of their ligand (HLA-C1 group) protect against chronic myeloid leukaemia. Tissue Antigens. 2009;73:553–560. doi: 10.1111/j.1399-0039.2009.01235.x. [DOI] [PubMed] [Google Scholar]
- Norman PJ, Cook MA, Carey BS, Carrington CV, Verity DH, Hameed K, Ramdath DD, Chandanayingyong D, Leppert M, Stephens HA, Vaughan RW. SNP haplotypes and allele frequencies show evidence for disruptive and balancing selection in the human leukocyte receptor complex. Immunogenetics. 2004;56:225–237. doi: 10.1007/s00251-004-0674-1. [DOI] [PubMed] [Google Scholar]
- Norman PJ, Abi-Rached L, Gendzekhadze K, Hammond JA, Moesta AK, Sharma D, Graef T, McQueen KL, Guethlein LA, Carrington CV, Chandanayingyong D, Chang YH, Crespi C, Saruhan-Direskeneli G, Hameed K, Kamkamidze G, Koram KA, Layrisse Z, Matamoros N, Mila J, Park MH, Pitchappan RM, Ramdath DD, Shiau MY, Stephens HA, Struik S, Tyan D, Verity DH, Vaughan RW, Davis RW, Fraser PA, Riley EM, Ronaghi M, Parham P. Meiotic recombination generates rich diversity in NK cell receptor genes, alleles, and haplotypes. Genome Res. 2009;19:757–769. doi: 10.1101/gr.085738.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ordonez D, Meenagh A, Gomez-Lozano N, Castano J, Middleton D, Vilches C. Duplication, mutation and recombination of the human orphan gene KIR2DS3 contribute to the diversity of KIR haplotypes. Genes Immun. 2008;9:431–437. doi: 10.1038/gene.2008.34. [DOI] [PubMed] [Google Scholar]
- Pando MJ, Gardiner CM, Gleimer M, McQueen KL, Parham P (2003) The protein made from a common allele of KIR3DL1 (3DL1*004) is poorly expressed at cell surfaces due to substitution at positions 86 in Ig domain 0 and 182 in Ig domain 1. J Immunol 171:6640–6649 [DOI] [PubMed]
- Parham P. Influence of KIR diversity on human immunity. Adv Exp Med Biol. 2005;560:47–50. doi: 10.1007/0-387-24180-9_6. [DOI] [PubMed] [Google Scholar]
- Ploski R, Luszczek W, Kusnierczyk P, Nockowski P, Cislo M, Krajewski P, Malejczyk J. A role for KIR gene variants other than KIR2DS1 in conferring susceptibility to psoriasis. Hum Immunol. 2006;67:521–526. doi: 10.1016/j.humimm.2006.04.001. [DOI] [PubMed] [Google Scholar]
- Shilling HG, Guethlein LA, Cheng NW, Gardiner CM, Rodriguez R, Tyan D, Parham P. Allelic polymorphism synergizes with variable gene content to individualize human KIR genotype. J Immunol. 2002;168:2307–2315. doi: 10.4049/jimmunol.168.5.2307. [DOI] [PubMed] [Google Scholar]
- Single RM, Martin MP, Meyer D, Gao X, Carrington M. Methods for assessing gene content diversity of KIR with examples from a global set of populations. Immunogenetics. 2008;60:711–725. doi: 10.1007/s00251-008-0331-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki Y, Hamamoto Y, Ogasawara Y, Ishikawa K, Yoshikawa Y, Sasazuki T, Muto M. Genetic polymorphisms of killer cell immunoglobulin-like receptors are associated with susceptibility to psoriasis vulgaris. J Invest Dermatol. 2004;122:1133–1136. doi: 10.1111/j.0022-202X.2004.22517.x. [DOI] [PubMed] [Google Scholar]
- Uhrberg M, Valiante NM, Shum BP, Shilling HG, Lienert-Weidenbach K, Corliss B, Tyan D, Lanier LL, Parham P. Human diversity in killer cell inhibitory receptor genes. Immunity. 1997;7:753–763. doi: 10.1016/S1074-7613(00)80394-5. [DOI] [PubMed] [Google Scholar]
- Velardi A. Role of KIRs and KIR ligands in hematopoietic transplantation. Curr Opin Immunol. 2008;20:581–587. doi: 10.1016/j.coi.2008.07.004. [DOI] [PubMed] [Google Scholar]
- Verheyden S, Demanet C. NK cell receptors and their ligands in leukemia. Leukemia. 2008;22:249–257. doi: 10.1038/sj.leu.2405040. [DOI] [PubMed] [Google Scholar]
- Whang DH, Park H, Yoon JA, Park MH. Haplotype analysis of killer cell immunoglobulin-like receptor genes in 77 Korean families. Hum Immunol. 2005;66:146–154. doi: 10.1016/j.humimm.2004.10.013. [DOI] [PubMed] [Google Scholar]
- Yoo YJ, Tang J, Kaslow RA, Zhang K. Haplotype inference for present-absent genotype data using previously identified haplotypes and haplotype patterns. Bioinformatics. 2007;23:2399–2406. doi: 10.1093/bioinformatics/btm371. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
KIR2DS2, KIR2DL3, and KIR2DL2 genes whose presence or absence in the centromeric LD region tags the presence or absence of other KIR genes (DOC 100 kb)
KIR2DL5B, KIR2DS3, and KIR2DL1 genes whose presence or absence in the centromeric LD region tags the presence or absence of other KIR genes (DOC 93 kb)
KIR2DL4, KIR3DL1, KIR3DS1, and KIR2DL5A genes whose presence or absence in the telomeric LD region tags the presence or absence of other KIR genes (DOC 93 kb)
KIR2DS2, KIR2DS1, and KIR2DS4 genes whose presence or absence in the telomeric LD region tags the presence or absence of others KIR genes (DOC 81 kb)


