Abstract
Chromosome segregation and cell division are essential, highly ordered processes that depend on numerous protein complexes. Results from recent RNA interference (RNAi) screens indicate that the identity and composition of these protein complexes is incompletely understood. Using gene tagging on bacterial artificial chromosomes, protein localization and tandem affinity purification-mass spectrometry, the MitoCheck consortium has analyzed about 100 human protein complexes, many of which had not or only incompletely been characterized. This work has led to the discovery of previously unknown, evolutionarily conserved subunits of the anaphase-promoting complex (APC/C) and the γ-tubulin ring complex (γ-TuRC), large complexes which are essential for spindle assembly and chromosome segregation. The approaches we describe here are generally applicable to high throughput follow-up analyses of phenotypic screens in mammalian cells.
Phenotypic screens using random mutagenesis, systematic gene deletion or RNA interference (RNAi) are powerful techniques for cataloguing gene function. To interpret the resulting genotype-phenotype relationships, detailed molecular analyses are required, among which protein localization and identification of protein interactions are particularly informative. In yeast, the modification of most genes at their endogenous loci with tag-coding sequences has been valuable for systems-wide analyses of protein function (1-4). In mammalian cells, however, large-scale localization and interaction studies of proteins expressed under control of their own regulatory sequences have so far lagged behind phenotypic analysis. The MitoCheck consortium (www.mitocheck.org) has therefore used recombineering techniques (5) to develop a fast and reliable procedure for the introduction of genes tagged in bacterial artificial chromosomes (BACs) into human tissue culture cells. This technique allows the stable expression of genes under their own promoters at near-physiological levels (6). Here we have combined this “BAC TransgeneOmics” technology with large-scale protein localization and interaction experiments to characterize about 100 mitotic protein complexes (Fig. 1A). By using this combined approach we discovered previously unknown subunits of the γ-TuRC and the APC/C, complexes that are essential for spindle assembly and chromosome segregation, respectively (7, 8).
Fig. 1.
Use of BAC TransgeneOmics for identification and characterization of mitotic protein complexes in human cells.
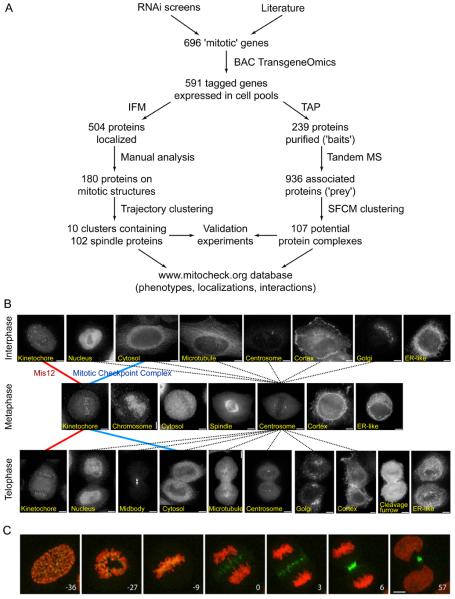
(A) Schematic outline of the workflow established for BAC tagging, GFP localization and tandem affinity purification (TAP)-mass spectrometry (MS) of mitotic proteins. IFM, immunofluorescence microscopy.
(B) Representative images of fixed HeLa cells, obtained by immunofluorescence microscopy with antibodies to GFP. Examples of cells are shown that stably express LAP-tagged proteins that have different locations in interphase, metaphase or telophase, as indicated in the images. Images are connected by lines that represent a subset of the localization trajectories observed for different proteins. The red and blue lines represent the trajectories of MIS12 and MCC, respectively. Dotted lines represent all observed trajectories that include centrosome localization in metaphase. Size bar, 10 μm.
(C) Time-lapse images of a mitotic HeLa cell stably expressing H2B-mCherry (red) and the CPC subunit INCENP-LAP (green). Numbers indicate time (minutes) before and after anaphase onset. Size bar, 10 μm.
Generation of a library of HeLa cell pools stably expressing GFP tagged BACs
We chose to characterize proteins required for mitosis because this process is essential for eukaryotic life, is of relevance for tumor biology, and is known to depend on numerous protein complexes. Many of these had been characterized before, providing prior knowledge that we could use to control our approaches and to draw hypotheses for unknown genes. RNAi screens performed in C. elegans, Drosophila and mammalian cells as well as proteomic studies have furthermore identified numerous uncharacterized proteins required for mitosis (9-17). In addition, the MitoCheck consortium carried out a genome-wide RNAi screen by time-lapse imaging of chromosome segregation in live cells that provided detailed phenotypic information for the majority of human proteins (16).
From these screens and the literature, we selected 696 proteins (table S1) for C-terminal tagging with a combined localization affinity-purification (LAP) tag (18), using high-throughput BAC recombineering in Escherichia coli (6). In most cases we tagged mouse genes and expressed them in human cells because this allows functional testing of the tagged proteins by RNAi-mediated depletion of their endogenous counterparts (19). N-terminal tags were introduced if C-terminal tagging failed, or in some cases to validate data obtained with the C-terminal tag. We were able to tag and stably express 591 (89%) of the selected proteins in HeLa cells (fig. S1). For each gene we obtained at least one non-clonal pool of stably-expressing cells, resulting in a library of 657 pools. Using antibodies to the GFP moiety of the LAP tag, we could detect the tagged proteins in 559 pools (85%), corresponding to 504 unique proteins (77%), by immunofluorescence microscopy (table S1 and fig. S1C).
Localization of mitotic proteins
First we analyzed all cell pools in which a GFP signal could be detected for the intracellular localization of tagged proteins in interphase, metaphase and telophase, using fixed cells stained with antibodies to GFP and α-tubulin, and 4′,6-diamidino-2-phenylindole (DAPI) to visualize DNA (Fig. 1B). In mitotic cells we observed specific association with centrosomes, spindles, kinetochores, chromosomes, cleavage furrows, midbodies or cortical structures for 180 proteins, of which 25 had not been characterized in mitosis, and 54 not at all (fig. S1C). For 14 proteins we confirmed our fixed-cell data by time-lapse imaging of living cells (Fig. 1C and fig. S2).
To identify proteins with potential roles in spindle assembly we localized in more detail 102 proteins that showed mitotic centrosome or spindle association. Of these, 23 had not been characterized in mitosis and 9 not at all. Immunofluorescence images were classified into 87 staining patterns at five different mitotic stages, resulting in specific localization trajectories (Fig. 1B, fig. S3). The frequency distribution of different patterns was scored manually, and the 102 proteins were clustered into 10 groups according to these scores (Fig. 2). Localization analysis with this resolution allowed separation even of complexes with very similar localization, e.g. the kinetochore complex MIS12 and the mitotic checkpoint complex (MCC), into separate localization trajectories (see fig. S3). In all cases, subunits of known complexes were recovered in the same clusters (chromosomal passenger complex [CPC], centralspindlin, MIS12, Aurora A-targeting protein for Xklp2 [TPX2], γ-TuRC), although prior knowledge about the existence of these complexes had not been used to “train” the cluster algorithm. This suggests that clustering of localization trajectories can be used to formulate hypotheses about functions and physical interactions of uncharacterized proteins. For example, centrosomal protein of 120 kDa (CEP120) clustered with proteins required for centriole duplication, suggesting that CEP120 may have a role in this process. RNAi experiments indicated that this is indeed the case (fig. S4).
Fig. 2.
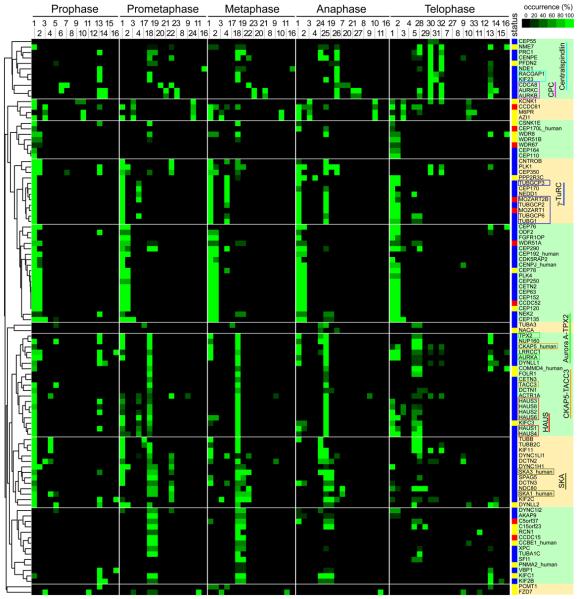
Heat map showing hierarchical clustering of the localization trajectories of 102 spindle and centrosome proteins.
Immunofluorescence microscopy images of fixed HeLa cells expressing LAP-tagged proteins (listed on the y-axis) in five mitotic phases were classified into 33 staining patterns (mouse ortholog proteins were imaged unless suffixed as “_human”). The frequency with which each staining pattern was observed for each protein in each phase is represented by a square with color ranging from black (0% occurrence) to bright green (100% occurrence). The number codes for the staining patterns are as follows: 1: centrosome (PCM-like); 2: centrosome (fine dots); 3: dots near centrosome; 4: centrosome and partial microtubule (MT); 5: microtubule; 6: chromosome; 7: kinetochore; 8: cortex (whole); 9: cortex (partial); 10: dots in cytoplasm; 11: ER-like meshwork; 12: nuclear envelope; 13: nucleus; 14: dots in nucleus; 15: nucleolus; 16: whole cell; 17: spindle MT (whole); 18: spindle MT (K-fiber); 19: spindle matrix-like; 20: chromosome (axis); 21: chromosome (periphery); 22: kinetochore (sisters); 23: kinetochore (inner); 24: microtubule (whole); 25: microtubule (K-fiber); 26: spindle midzone; 27: cleavage furrow; 28: microtubule next to midbody; 29: microtubule (unclear); 30: midbody; 31: midbody ring; 32: midbody next to ring; 33: Golgi-like. The ‘status’ column shows the characterization status of each gene, as defined by literature searching in PubMed, according to the following color scheme: blue, reported to be involved in mitosis; yellow, some function reported but not in the context of mitosis; red, functionally uncharacterized. The dendrogram on the left indicates the relative similarities of the trajectories of individual proteins to the entire trajectory pattern. The clustered heat map was divided into ten subclusters based on a visual inspection of clustered localization patterns (subclusters indicated by alternating green and yellow shading). The names of subunits of previously identified complexes are boxed in different colors (Centralspindlin, light blue; CPC, magenta; γ-TuRC, dark blue; Aurora A-TPX2, bright green; chTOG-TACC3, orange; Augmin/HAUS, red; SKA, olive-green).
Identification of mitotic protein complexes
To characterize mitotic protein complexes, we isolated LAP tagged proteins from cells arrested in mitosis, using tandem-affinity purification (fig. S5)(6). Samples were analyzed by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) followed by silver staining, and by in-solution trypsinization and tandem mass spectrometry. Initially, proteins were selected based on localization identified by GFP imaging or on reported mitotic functions. Once interaction partners had been identified, interaction mapping was performed iteratively by producing new LAP-tagged cell pools to validate a subset of the interactions through reciprocal analyses. In total, cell pools containing 254 different tagged genes were analyzed. In 239 cases (94%), the “bait” proteins could be identified. These interacted with a total of 936 “prey” proteins which were present in specific samples, corresponding to 2011 unique pair-wise interactions (20). Other proteins, which were found in more than 4.5% of all samples or in “mock” purifications, were excluded from further analyses because these proteins might represent contaminants (tables S2 and S3). For a complete presentation of all data, see http://www.mitocheck.org [username: members; password: con_priv. Public access will be granted upon publication of this manuscript]. Additional information on tagged BACs can be found at http://hymanlab.mpi-cbg.de/BACE.
We analyzed baits from 11 previously described reference complexes which, according to the literature, contain 74 subunits (fig. S6A). Our experiments identified 70 of these, indicating a low false-negative detection rate (fig. S6B). In 175 cases, our experiments revealed interactions between two proteins, both of which had been tagged and used as baits. Of these interactions, 94 (54%) could be detected with both baits. This frequency of reciprocal interactions is higher than in previous studies performed in yeast (15% in (3); 8% in (4)). These results suggest that the number of false-positive interactions in our dataset is relatively low. However, we cannot exclude that some false-positive interactions were detected.
To identify previously unknown complexes we analyzed the dataset of all interactions for the presence of proteins that are densely connected with each other (fig. S7), using a clustering algorithm that we call spectral fuzzy C-means (SFCM). We identified 35 singletons (cases where only the bait had been found), 107 clusters which contain between two and 20 proteins, and 13 clusters with more than 20 components (Fig. 3, fig. S7, fig. S8 and table S4). The 13 large clusters contain sets of loosely connected proteins, which presumably had been grouped together because the density of the interaction network was not high enough to separate these proteins into smaller, more meaningful clusters. However, among the 107 small clusters, 11 matched the reference complexes with an average precision of 59% (the fraction of cluster members that belong to the same reference complex) and an average recall of 89% (the fraction of the reference complex subunits assigned to the same cluster). These values indicate that many of the small clusters represent bona fide protein complexes, or groups of closely related complexes (note, for example, that different isoforms of cohesin complexes clustered together; table S4). As an example, Fig. 3 shows a graphical representation of 10 of the 107 small clusters and how they compare to reference complexes described in the literature. The entire interaction network can be visualized in the Cytoscape session file S1.
Fig. 3.
Combined interaction and localization map for a selection of 10 out of 107 small clusters (cluster size 2 to 20 proteins). Protein interaction data obtained by tandem affinity purification-mass spectrometry were analyzed by SFCM clustering. The resulting clusters (enclosed by orange dashed lines) were manually annotated based on literature knowledge (29) (for composition of reference complexes as described in the literature see fig. S6). The resulting curated clusters are shown as gray ellipses: complexes containing reciprocal interactions are enclosed by solid gray lines, while those without reciprocal interactions are denoted by dashed gray lines. Inter- and intra-cluster interactions are represented by dashed and solid blue lines, respectively, with the number of interactions corresponding to the thickness of these lines. Localization as determined by GFP imaging is shown in the color codes as indicated. Note that cluster and protein complexes can have interactions with several other clusters/complexes. For example, MCC co-purified and therefore co-clustered both with APC/C and with the MIS12/NDC80 cluster. The former association reflects binding of MCC to APC/C in prometaphase cells (30), from which these proteins were purified, whereas MCC association with MIS12/NDC80 may reflect recruitment of MCC components to unattached kinetochores in prometaphase (31). For an expanded ‘master map’, integrating localization and interaction data for more proteins, see fig. S8.
Identification and characterization of mitotic protein complexes by combined interaction and localization studies
To test whether co-purifying proteins interact in vivo we analyzed in how many cases similar localization patterns had been obtained for interacting proteins. We manually annotated a subset of 728 interactions and found that 49% of all pair-wise interacting bait and prey proteins had similar localizations. This frequency was even higher (79%) when only reciprocally confirmed interactions were considered (fig. S9). For example, we observed that CEP120 both co-localized and physically interacted with coiled coil domain containing 52 (CCDC52), suggesting that these proteins form a complex (Figs. 2 and 3). Similarly, we observed that eight proteins, which had not been characterized when we performed our experiments, interacted reciprocally with each other and were all located on mitotic spindles (fig. S10). These proteins are subunits of the Augmin/HAUS complex which has recently been shown to be essential for spindle function (21-23). Combining localization and interaction data can also identify unknown interactions between well characterized proteins and complexes, as is illustrated by our finding that Polo-like kinase 1-interacting checkpoint helicase (PICH, also known as excision repair cross-complementing rodent repair deficiency, complementation group 6-like [ERCC6L]) interacts and colocalizes on chromosome bridges with subunits of the RTR complex (RecQ helicase [BLM]-topoisomerase III [TOP3A]-RecQ mediated genome instability 1 [RMI1] complex; (24); fig. S11).
Identification of C10orf104 as a subunit of the APC/C
We further characterized chromosome 10 open reading frame 104 (C10orf104) because this 11.7-kDa protein co-purified with seven different APC/C subunits (fig. S12A), but not with any other bait, and thus co-clustered with the APC/C in our SFCM analysis (Fig. 3). The APC/C is a 1.5-MDa ubiquitin ligase complex essential for chromosome segregation and mitotic exit (8). Because APC/C has been characterized in detail it was surprising that a previously uncharacterized protein co-purified with APC/C subunits. However, also when C10orf104 was used as bait, several APC/C subunits were detected (fig. S12A), and antibodies raised against C10orf104 immunoprecipitated the entire APC/C and its associated cyclin B ubiquitylation activity (Fig. 4, A and B). Immunoblot experiments showed that C10orf104 is present throughout the cell cycle (fig. S12, B and C), and density gradient centrifugation experiments indicated that most of C10orf104 is associated with the APC/C (Fig. 4C). These observations indicate that C10orf104 is a constitutive subunit of the APC/C and not a substrate or a transiently associating regulatory protein. Electron microscopic analysis of APC/C labeled with C10orf104 antibodies suggested that C10orf104 is located at the top of APC/C's “arc lamp” domain, in the vicinity of the subunit CDC27 (Fig. 4D and fig. S12, D, E and F). The amino acid sequence of C10orf104 is highly conserved among vertebrates (95% identical between human and zebrafish), suggesting that, despite its small size, this protein performs an important function within the APC/C. Related sequences also exist in invertebrates (fig. S12G), although we could not yet identify homologous sequences in yeast. These observations indicate that C10orf104 is an evolutionarily-conserved APC/C subunit, which we propose to call APC16 (gene symbol ANAPC16). We suspect that APC16 has previously escaped detection in protein gels or by mass spectrometry due to its small size.
Fig. 4.
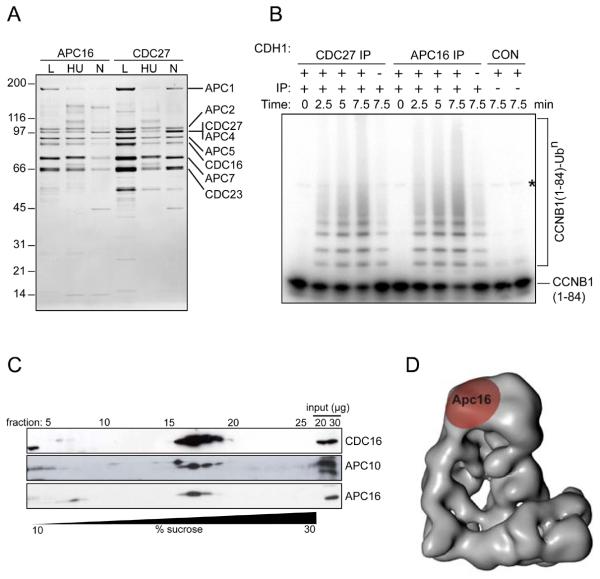
Characterization of APC16, a previously unknown subunit of the APC/C.
(A) Silver-stained SDS-PAGE gel showing proteins immunoprecipitated using APC16 or CDC27 antibodies from extracts of HeLa cells, cultured under logarithmic growth conditions (L), or arrested in S phase by treatment for 18 hr with hydroxyurea (HU) or arrested in prometaphase by treatment for 18 hr with nocodazole (N). Numbers on the left indicate the molecular masses of reference proteins.
(B) Phosphorimage showing the ubiquitylation of Cyclin B1 (CCNB1), catalyzed by CDC27 and APC16 immunoprecipitates. [125I]-labelled human CCNB1 fragment (amino acids 1 to 84) was incubated with CDC27 or APC16 immunoprecipitates from logarithmically growing HeLa cells, plus E1 and E2 enzymes, ubiquitin and ATP, with or without the recombinant co-activator protein CDH1, for the times indicated, then analyzed by SDS-PAGE and phosphorimaging. CON indicates empty protein-A beads (left) and a condensin antibody immunoprecipitate (right). The asterisk marks a contaminating band present in the CCNB1 sample.
(C) Immunoblots showing the co-sedimentation of APC16 with core APC/C subunits CDC16 and APC10, following density gradient centrifugation. An extract of logarithmically-growing HeLa cells was subjected to centrifugation through a 10-30% sucrose density gradient. 28 fractions were collected and analyzed by SDS-PAGE and immunoblotting with the antibodies indicated.
(D) Three-dimensional model of the human APC/C obtained by electron microscopy (30), showing the location of APC16, as determined by antibody labeling.
Identification of proteins interacting with the γ-TuRC
We also characterized chromosome 13 open reading frame 37 (C13orf37) and two closely-related proteins, called family with sequence similarity 128, member A and member B (FAM128A and FAM128B), because these proteins co-purified with three different subunits of the γ-TuRC. This complex is located at centrosomes and mediates the formation of bipolar spindles in mitosis (7). When C13orf37 and FAM128B were used as baits, all six known γ-TuRC subunits were identified (Fig. 3, Fig. 5, A and B). Sucrose density gradient centrifugation experiments confirmed that C13orf37 and FAM128B are associated with the γ-TuRC component tubulin, gamma 1 (TUBG1; fig S13A). Like γ-TuRC subunits, C13orf37 and FAM128B were located on centrosomes throughout the cell cycle, and to a lesser extent on mitotic spindles (Fig. 2, Fig. 5, C and D, and fig. S13B). Proteins homologous to C13orf37 and FAM128A/B are predicted to exist in many eukaryotes, including, in the case of C13orf37, the fission yeast Schizosaccharomyces pombe (fig. S13, C and D). However, the corresponding genomic sequences have not been annotated as genes in all organisms, possibly because C13orf37 and FAM128A/B are small proteins of 8.5 and 16.2 kDa, respectively. C13orf37 and FAM128A/B may thus be evolutionarily conserved γ-TuRC subunits that previously may not have been detected due to their small size. However, unlike the known subunits of γ-TuRC (tubulin, gamma complex associated proteins, TUBGCP2-6), C13orf37 and FAM128A/B do not contain the conserved “Spc97_Spc98” GCP domain (25). The TUBGCP nomenclature can therefore not be applied to C13orf37, FAM128A and FAM128B. Instead, we propose to call these proteins mitotic-spindle organizing proteins associated with a ring of gamma-tubulin, MOZART1, MOZART2A and MOZART2B, respectively.
Fig. 5.
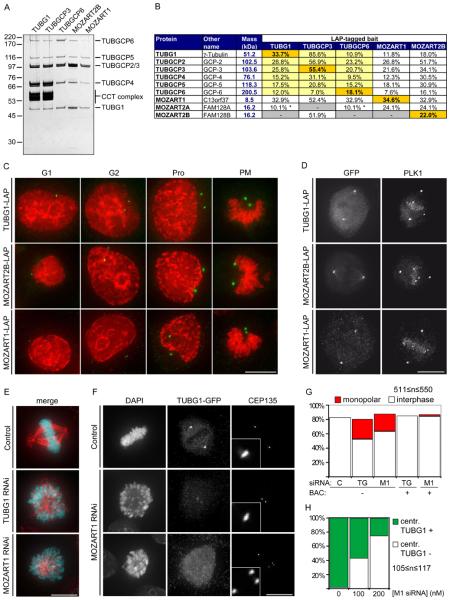
Characterization of the γ-TuRC interacting proteins MOZART1 and MOZART2B.
(A) Silver-stained SDS-PAGE gel of LAP-purified TUBG1, TUBGCP3, TUBGCP6, MOZART2B and MOZART1. Numbers on the left indicate the molecular masses of reference proteins. Proteins annotated based on expected electrophoretic mobility are listed on the right. CCT, chaperonin containing T-complex.
(B) Mass spectrometry results obtained from the samples in (A), showing identification of γ-TuRC subunits plus MOZART1, MOZART2A and MOZART2B. Entries highlighted in orange are bait proteins, yellow indicates proteins whose interaction with the bait has previously been reported, a white background indicates that the interaction was previously unknown. Asterisks indicate proteins identified by a single unique peptide.
(C) Immunofluorescence microscopy images showing centrosomal localization of TUBG1, MOZART2B and MOZART1 through the cell cycle (G1-phase, G2-phase, prophase (Pro) or prometaphase (PM)). HeLa cells expressing the indicated proteins were fixed and stained with antibodies to GFP (green) and with DAPI (red).
(D) Immunofluorescence microscopy images showing localization of TUBG1, MOZART2B and MOZART1 to centrosomes and the mitotic spindle in metaphase in fixed LAP-cells stained with antibodies to GFP and PLK1. All images in this figure are maximum-intensity projections of DeltaVision stacks. Size bar, 10 μm.
(E) Immunofluorescence microscopy images of HeLa cells treated for 72 hr with 100 nM siRNA specific to either TUBG1 or MOZART1. Cells were fixed and stained with DAPI (cyan) and TUBA1A antibodies (red). Size bar, 10 μm.
(F) Immunofluorescence microscopy images of TUBG1-LAP HeLa cells treated for 72 hr with 100 nM siRNA against TUBG1 to deplete the endogenous TUBG1 and, simultaneously, transfected either with luciferase siRNA (control siRNA) or MOZART1 siRNA. Cells were fixed and stained with DAPI, GFP antibodies (to visualize TUBG1-LAP) and CEP135 (to stain centrioles). Two distinct phenotypes were observed: CEP135 foci in close proximity to each other that colocalize with faint TUBG1-GFP foci (middle row) and CEP135 foci dispersed randomly dispersed throughout the cell with no detectable TUBG1-GFP foci (bottom row). Insets in the third column are magnified five-fold to show one centrosome. Size bar, 10 μm.
(G) Histogram showing quantification of the experiment described in (E). In addition, cells containing the mouse TUBG1-LAP or the mouse MOZART1-LAP (BAC +) were treated with 100 nM siRNA specific for TUBG1 or MOZART1, respectively, and fixed and stained as in E. C, control, TG, TUBG1 siRNA, M1, MOZART1 siRNA.
(H) Histogram showing quantification of the experiment described in (F). The number of cells with GFP positive (TUBG1 +) or negative (TUBG1 −) centrosomes was counted. The GFP-negative cells are a combination of both MOZART1-depletion phenotypes shown in (F).
MOZART1, an evolutionarily conserved protein essential for γ-TuRC function
To test if the MOZARTs are important for γ-TuRC function, we performed RNAi experiments in HeLa cells. Transfection of MOZART2A/B siRNA did not result in detectable mitotic phenotypes, but we cannot exclude that this was due to incomplete depletion of these proteins. In contrast, depletion of either MOZART1 or TUBG1 led to the accumulation of prometaphase cells with mono-polar spindles and closely spaced centrosome pairs (Fig. 5, E to G). These phenotypes were fully reverted by stable integration of the corresponding LAP-tagged mouse genes on BACs (Fig. 5G), ruling out off-target RNAi effects and showing that the LAP-tagged proteins used for localization and interaction mapping are functional.
Monopolar spindle phenotypes have been observed after depletion of γ-TuRC, but also after inactivation of PLK1 (26) or Aurora A kinase (27). We therefore tested if MOZART1 depletion could interfere with spindle assembly directly by preventing γ-tubulin recruitment to centrosomes, or indirectly by decreasing PLK1 or Aurora A activity. In immunofluorescence microscopy experiments MOZART1 depleted cells were stained equally well as control cells with antibodies specific for phospho-epitopes generated by PLK1 or Aurora A (fig S13E and F), suggesting that MOZART1 is not required for the activation of these kinases. However, depletion of MOZART1 did strongly reduce TUBG1-LAP staining at centrosomes in 70% of the cells (Fig. 5, F and H). These observations indicate that MOZART1 is required for γ-TuRC recruitment to centrosomes. Because orthologs of MOZART1 exist in lower eukaryotes, including fission yeast, it is possible that this function has been highly conserved during evolution.
www.mitocheck.org, a human functional-genomics database
The data obtained in this study have enabled us to identify previously unknown protein complexes (CEP120-CCDC52; Augmin/HAUS), new subunits of well-studied protein complexes such as the APC/C (APC16) and γ-TuRC (the MOZARTs), and unknown interactions between known proteins and complexes (PICH-RTR). However, the majority of our data has not yet been used for follow-up experiments. We suspect that such experiments will lead to additional important discoveries about the functions of human protein complexes in mitosis (28). This notion is supported by the observation that most of the protein interactions detected in our experiments have not been reported previously. For example, for 60 of the 107 small SFCM clusters none of the interactions have been reported in seven major public interaction databases (fig. S7A). Many of these clusters may therefore represent uncharacterized protein complexes. To enable the exploitation of these data by the scientific community we have generated a human genome-wide database (www.mitocheck.org) that contains all data generated by the MitoCheck consortium (fig. S14). These include information on tagged BACs, immunofluorescence images obtained by GFP localization, silver-stained SDS-PAGE gels of all protein samples obtained by tandem affinity purification, and all protein interaction lists obtained by in-solution trypsinization-tandem mass spectrometry. In addition, this database contains movies from the MitoCheck RNAi screen in which mitosis has been analyzed by live imaging of cells in which all human proteins have been targeted by siRNAs (16). The database also provides information about gene synonyms used in the literature, orthologs in other species and protein interactions reported in public databases. This collection of localization, interaction and phenotypic data will be a useful resource for understanding the functions of human proteins.
Conclusion
The widespread application of RNAi for phenotypic screens has not been accompanied by the development of approaches to rapidly study protein function. This means that it is difficult to characterize the results of such screens. Similar problems apply to the results of human genetic and genomic studies, which often identify many uncharacterized proteins potentially associated with disease. The combined use of BAC tagging, protein localization and interaction mapping techniques which we describe here for mitotic proteins helps to overcome this limitation by allowing systems-scale approaches to studying protein function. These systematic non-genetic approaches represent a valuable counterpart to RNAi screens, in which limited penetrance and off-target effects can result in ambiguity in identifying gene function. Rather than relying on phenotypic screens, hypotheses can be generated and tested from analysis of the protein complexes and localization of uncharacterized proteins.
Supplementary Material
Acknowledgments
We are grateful to the following colleagues for their excellent assistance: E. Kreidl, M. Mazanek, M. Madalinski, G. Mitulović, M. Novatchkova, C. Stingl, Y. Sun (IMP-IMBA, Vienna); A. Bird, K. Kozak, D. Krastev, Z. Maliga, D. Richter, M. Theis, M. Toyoda (MPI, Dresden); P. Dube (MPI, Göttingen); and N. Kraut (Boehringer Ingelheim, Vienna). This work was funded in the most part by the European Commission via the Sixth Framework Programme Integrated Project ‘MitoCheck’ (LSHG-CT-2004-503464). Work in the laboratories of J.-M.P. and K.M. received support from Boehringer Ingelheim, the Vienna Spots of Excellence Programme, the Austrian Science Fund Special Research Programme “Chromosome Dynamics” and the Genome Research in Austria Programme. Work in the laboratory of L.P. was supported by operating grants from the Natural Science and Engineering Research Council of Canada (RGPIN-355644-2008), the National Cancer Institute of Canada (019562) and the Human Frontier Science Program (CDA0044/200). L.P. holds a Canada Research Chair in Centrosome Biogenesis and Function. Y.T. was supported by a Postdoctoral Fellowship for Research Abroad from the Japan Society for the Promotion of Science (JSPS).
Footnotes
Materials and methods
Supplementary figures S1 to S14
Tables S1 to S4
Cytoscape session file S1
Supplementary references
References and notes
- 1.Ghaemmaghami S, et al. Nature. 2003 Oct 16;425:737. doi: 10.1038/nature02046. [DOI] [PubMed] [Google Scholar]
- 2.Huh WK, et al. Nature. 2003 Oct 16;425:686. doi: 10.1038/nature02026. [DOI] [PubMed] [Google Scholar]
- 3.Gavin AC, et al. Nature. 2006 Mar 30;440:631. doi: 10.1038/nature04532. [DOI] [PubMed] [Google Scholar]
- 4.Krogan NJ, et al. Nature. 2006 Mar 30;440:637. doi: 10.1038/nature04670. [DOI] [PubMed] [Google Scholar]
- 5.Zhang Y, Buchholz F, Muyrers JP, Stewart AF. Nat Genet. 1998 Oct;20:123. doi: 10.1038/2417. [DOI] [PubMed] [Google Scholar]
- 6.Poser I, et al. Nat Methods. 2008 May;5:409. doi: 10.1038/nmeth.1199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Patel U, Stearns T. Curr Biol. 2002 Jun 25;12:R408. doi: 10.1016/s0960-9822(02)00908-9. [DOI] [PubMed] [Google Scholar]
- 8.Peters JM. Nat Rev Mol Cell Biol. 2006 Sep;7:644. doi: 10.1038/nrm1988. [DOI] [PubMed] [Google Scholar]
- 9.Gönczy P, et al. Nature. 2000 Nov 16;408:331. doi: 10.1038/35042526. [DOI] [PubMed] [Google Scholar]
- 10.Andersen JS, et al. Nature. 2003 Dec 4;426:570. doi: 10.1038/nature02166. [DOI] [PubMed] [Google Scholar]
- 11.Kamath RS, et al. Nature. 2003 Jan 16;421:231. doi: 10.1038/nature01278. [DOI] [PubMed] [Google Scholar]
- 12.Sönnichsen B, et al. Nature. 2005 Mar 24;434:462. doi: 10.1038/nature03353. [DOI] [PubMed] [Google Scholar]
- 13.Goshima G, et al. Science. 2007 Apr 20;316:417. doi: 10.1126/science.1141314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kittler R, et al. Nat Cell Biol. 2007 Dec;9:1401. doi: 10.1038/ncb1659. [DOI] [PubMed] [Google Scholar]
- 15.Somma MP, et al. PLoS Genet. 2008 Jul;4:e1000126. doi: 10.1371/journal.pgen.1000126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Neumann B. 2010 e. al., Manuscript submitted. [Google Scholar]
- 17.Theis M, et al. Embo J. 2009 May 20;28:1453. doi: 10.1038/emboj.2009.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cheeseman IM, Desai A. Sci STKE. 2005 Jan 11;2005:pl1. doi: 10.1126/stke.2662005pl1. [DOI] [PubMed] [Google Scholar]
- 19.Kittler R, et al. Proc Natl Acad Sci U S A. 2005 Feb 15;102:2396. doi: 10.1073/pnas.0409861102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.The protein interactions from this publication have been submitted to the International Molecular Exchange (IMEx) Consortium (http://imex.sf.net) through IntAct (http://www.ebi.ac.uk/intact), and assigned the identifier IM-11719.
- 21.Goshima G, Mayer M, Zhang N, Stuurman N, Vale RD. J Cell Biol. 2008 May 5;181:421. doi: 10.1083/jcb.200711053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lawo S, et al. Curr Biol. 2009 May 26;19:816. doi: 10.1016/j.cub.2009.04.033. [DOI] [PubMed] [Google Scholar]
- 23.Uehara R, et al. Proc Natl Acad Sci U S A. 2009 Apr 28;106:6998. doi: 10.1073/pnas.0901587106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mankouri HW, Hickson ID. Trends Biochem Sci. 2007 Dec;32:538. doi: 10.1016/j.tibs.2007.09.009. [DOI] [PubMed] [Google Scholar]
- 25.Finn RD, et al. Nucleic Acids Res. 2010 Jan;38:D211. doi: 10.1093/nar/gkp985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lénárt P, et al. Curr Biol. 2007 Feb 20;17:304. doi: 10.1016/j.cub.2006.12.046. [DOI] [PubMed] [Google Scholar]
- 27.Hannak E, Kirkham M, Hyman AA, Oegema K. J Cell Biol. 2001 Dec 24;155:1109. doi: 10.1083/jcb.200108051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kittler R, Pelletier L, Buchholz F. Cell Cycle. 2008 Jul 15;7:2123. doi: 10.4161/cc.7.14.6322. [DOI] [PubMed] [Google Scholar]
- 29.For details see ‘Materials and methods’ section in supporting online material.
- 30.Herzog F, et al. Science. 2009 Mar 13;323:1477. doi: 10.1126/science.1163300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Musacchio A, Salmon ED. Nat Rev Mol Cell Biol. 2007 May;8:379. doi: 10.1038/nrm2163. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.