Abstract
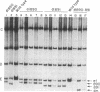
Repair of heteroduplex DNA, generated between two interacting DNA molecules during homologous recombination, has been implicated as a contributing factor in the process of gene conversion. To assess patterns of heteroduplex repair in mammalian cells, we constructed 13 different heteroduplexes from simian virus 40 wild-type and deletion mutant DNAs. Each heteroduplex contained one or multiple single-stranded loops in the intron of the gene for large tumor antigen, which is not essential during lytic infection. After transfection into cultured monkey cells, cellular repair was evaluated by restriction analysis of the amplified viral progeny from 1123 individual plaques, each representing the clonal expansion of a single repair event. Single-stranded loops were corrected prior to replication with an overall efficiency of 90%. At the position of a loop, one of the two heteroduplex strands served as a template for accurate repair 98% of the time. Repair of single-stranded loops was biased nearly 2 to 1 in favor of the strand without the loop. The efficiency, accuracy, and strand bias of repair were unaffected by loop size within the tested range, which was 25-247 nucleotides. The excision tract associated with repair of single-stranded loops rarely exceeds 200-400 nucleotides in length.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abastado J. P., Cami B., Dinh T. H., Igolen J., Kourilsky P. Processing of complex heteroduplexes in Escherichia coli and Cos-1 monkey cells. Proc Natl Acad Sci U S A. 1984 Sep;81(18):5792–5796. doi: 10.1073/pnas.81.18.5792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bianchi M. E., Radding C. M. Insertions, deletions and mismatches in heteroduplex DNA made by recA protein. Cell. 1983 Dec;35(2 Pt 1):511–520. doi: 10.1016/0092-8674(83)90185-x. [DOI] [PubMed] [Google Scholar]
- Chakrabarti S., Seidman M. M. Intramolecular recombination between transfected repeated sequences in mammalian cells is nonconservative. Mol Cell Biol. 1986 Jul;6(7):2520–2526. doi: 10.1128/mcb.6.7.2520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dohet C., Wagner R., Radman M. Repair of defined single base-pair mismatches in Escherichia coli. Proc Natl Acad Sci U S A. 1985 Jan;82(2):503–505. doi: 10.1073/pnas.82.2.503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fishel R. A., Kolodner R. An Escherichia coli cell-free system that catalyzes the repair of symmetrically methylated heteroduplex DNA. Cold Spring Harb Symp Quant Biol. 1984;49:603–609. doi: 10.1101/sqb.1984.049.01.068. [DOI] [PubMed] [Google Scholar]
- Fishel R. A., Siegel E. C., Kolodner R. Gene conversion in Escherichia coli. Resolution of heteroallelic mismatched nucleotides by co-repair. J Mol Biol. 1986 Mar 20;188(2):147–157. doi: 10.1016/0022-2836(86)90300-1. [DOI] [PubMed] [Google Scholar]
- Folger K. R., Thomas K., Capecchi M. R. Efficient correction of mismatched bases in plasmid heteroduplexes injected into cultured mammalian cell nuclei. Mol Cell Biol. 1985 Jan;5(1):70–74. doi: 10.1128/mcb.5.1.70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamza H., Kalogeropoulos A., Nicolas A., Rossignol J. L. Two mechanisms for directional gene conversion. Proc Natl Acad Sci U S A. 1986 Oct;83(19):7386–7390. doi: 10.1073/pnas.83.19.7386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hare J. T., Taylor J. H. One role for DNA methylation in vertebrate cells is strand discrimination in mismatch repair. Proc Natl Acad Sci U S A. 1985 Nov;82(21):7350–7354. doi: 10.1073/pnas.82.21.7350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967 Jun 14;26(2):365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- Lichten M., Fox M. S. Effects of nonhomology on bacteriophage lambda recombination. Genetics. 1983 Jan;103(1):5–22. doi: 10.1093/genetics/103.1.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lichten M., Fox M. S. Evidence for inclusion of regions of nonhomology in heteroduplex products of bacteriophage lambda recombination. Proc Natl Acad Sci U S A. 1984 Nov;81(22):7180–7184. doi: 10.1073/pnas.81.22.7180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liskay R. M., Stachelek J. L. Information transfer between duplicated chromosomal sequences in mammalian cells involves contiguous regions of DNA. Proc Natl Acad Sci U S A. 1986 Mar;83(6):1802–1806. doi: 10.1073/pnas.83.6.1802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meselson M. S., Radding C. M. A general model for genetic recombination. Proc Natl Acad Sci U S A. 1975 Jan;72(1):358–361. doi: 10.1073/pnas.72.1.358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller L. K., Cooke B. E., Fried M. Fate of mismatched base-pair regions in polyoma heteroduplex DNA during infection of mouse cells. Proc Natl Acad Sci U S A. 1976 Sep;73(9):3073–3077. doi: 10.1073/pnas.73.9.3073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norkin L. C. Marker-specific effects in genetic recombination. J Mol Biol. 1970 Aug;51(3):633–655. doi: 10.1016/0022-2836(70)90013-6. [DOI] [PubMed] [Google Scholar]
- Pukkila P. J., Peterson J., Herman G., Modrich P., Meselson M. Effects of high levels of DNA adenine methylation on methyl-directed mismatch repair in Escherichia coli. Genetics. 1983 Aug;104(4):571–582. doi: 10.1093/genetics/104.4.571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roth D. B., Porter T. N., Wilson J. H. Mechanisms of nonhomologous recombination in mammalian cells. Mol Cell Biol. 1985 Oct;5(10):2599–2607. doi: 10.1128/mcb.5.10.2599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarkar S., Dasgupta U. B., Summers W. C. Error-prone mutagenesis detected in mammalian cells by a shuttle vector containing the supF gene of Escherichia coli. Mol Cell Biol. 1984 Oct;4(10):2227–2230. doi: 10.1128/mcb.4.10.2227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seidman M. M., Dixon K., Razzaque A., Zagursky R. J., Berman M. L. A shuttle vector plasmid for studying carcinogen-induced point mutations in mammalian cells. Gene. 1985;38(1-3):233–237. doi: 10.1016/0378-1119(85)90222-7. [DOI] [PubMed] [Google Scholar]
- Shortle D. A genetic system for analysis of staphylococcal nuclease. Gene. 1983 May-Jun;22(2-3):181–189. doi: 10.1016/0378-1119(83)90102-6. [DOI] [PubMed] [Google Scholar]
- Szostak J. W., Orr-Weaver T. L., Rothstein R. J., Stahl F. W. The double-strand-break repair model for recombination. Cell. 1983 May;33(1):25–35. doi: 10.1016/0092-8674(83)90331-8. [DOI] [PubMed] [Google Scholar]
- Thimmappaya B., Shenk T. Nucleotide sequence analysis of viable deletion mutants lacking segments of the simian virus 40 genome coding for small t antigen. J Virol. 1979 Jun;30(3):668–673. doi: 10.1128/jvi.30.3.668-673.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas K. R., Capecchi M. R. Introduction of homologous DNA sequences into mammalian cells induces mutations in the cognate gene. Nature. 1986 Nov 6;324(6092):34–38. doi: 10.1038/324034a0. [DOI] [PubMed] [Google Scholar]
- Wagner R., Dohet C., Jones M., Doutriaux M. P., Hutchinson F., Radman M. Involvement of Escherichia coli mismatch repair in DNA replication and recombination. Cold Spring Harb Symp Quant Biol. 1984;49:611–615. doi: 10.1101/sqb.1984.049.01.069. [DOI] [PubMed] [Google Scholar]
- Wagner R., Jr, Meselson M. Repair tracts in mismatched DNA heteroduplexes. Proc Natl Acad Sci U S A. 1976 Nov;73(11):4135–4139. doi: 10.1073/pnas.73.11.4135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wake C. T., Gudewicz T., Porter T., White A., Wilson J. H. How damaged is the biologically active subpopulation of transfected DNA? Mol Cell Biol. 1984 Mar;4(3):387–398. doi: 10.1128/mcb.4.3.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wake C. T., Wilson J. H. Defined oligomeric SV40 DNA: a sensitive probe of general recombination in somatic cells. Cell. 1980 Aug;21(1):141–148. doi: 10.1016/0092-8674(80)90121-x. [DOI] [PubMed] [Google Scholar]
- White J. H., Lusnak K., Fogel S. Mismatch-specific post-meiotic segregation frequency in yeast suggests a heteroduplex recombination intermediate. Nature. 1985 May 23;315(6017):350–352. doi: 10.1038/315350a0. [DOI] [PubMed] [Google Scholar]
- White R. L., Fox M. S. On the molecular basis of high negative interference. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1544–1548. doi: 10.1073/pnas.71.4.1544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wildenberg J., Meselson M. Mismatch repair in heteroduplex DNA. Proc Natl Acad Sci U S A. 1975 Jun;72(6):2202–2206. doi: 10.1073/pnas.72.6.2202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson J. H., Berget P. B., Pipas J. M. Somatic cells efficiently join unrelated DNA segments end-to-end. Mol Cell Biol. 1982 Oct;2(10):1258–1269. doi: 10.1128/mcb.2.10.1258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson J. H. Genetic analysis of host range mutant viruses suggests an uncoating defect in simian virus 40-resistant monkey cells. Proc Natl Acad Sci U S A. 1977 Aug;74(8):3503–3507. doi: 10.1073/pnas.74.8.3503. [DOI] [PMC free article] [PubMed] [Google Scholar]