Abstract
Background
It has become increasingly clear that molecular and neural mechanisms underlying learning and memory and drug addiction are largely shared. To confirm and extend these findings, we analyzed ethanol-responsive behaviors of a collection of Drosophila long-term memory mutants.
Methods
For each mutant, sensitivity to the acute uncoordinating effects of ethanol was quantified using the inebriometer. Additionally, 2 distinct forms of ethanol tolerance were measured: rapid tolerance, which develops in response to a single brief exposure to a high concentration of ethanol vapor; and chronic tolerance, which develops following a sustained low-level exposure.
Results
Several mutants were identified with altered sensitivity, rapid or chronic tolerance, while a number of mutants exhibited multiple defects.
Conclusions
The corresponding genes in these mutants represent areas of potential overlap between learning and memory and behavioral responses to alcohol. These genes also define components shared between different ethanol behavioral responses.
Keywords: Alcohol, Sensitivity, Tolerance, Drosophila, Learning/Memory
Alcohol is one of the most used and abused drugs in the world, yet the mechanisms by which it exerts its effects upon behavior are incompletely understood. Better understanding of the genes and mechanisms underlying behavioral responses to alcohol will facilitate the development of earlier, more accurate and effective treatments for alcohol abuse and dependence. In humans, the degree of initial ataxic response to alcohol is genetically influenced, and inversely correlated with the risk of alcohol abuse or dependence (Heath et al., 1999; Schuckit, 1994; Schuckit and Smith, 1996; Schuckit et al., 2004). The molecular underpinnings of this correlation, and the identities of the genes involved, largely remain to be elucidated.
Model organisms, such as the fruit fly Drosophila melanogaster, offer powerful genetic methodologies with which to dissect genes and molecular pathways underlying behaviors of interest. Flies exhibit many complex behaviors and, moreover, share with mammals many features of the behavioral response to ethanol [reviewed (Heberlein et al., 2004)]. Behavioral screens for altered ethanol responses in flies have pinpointed several underlying genes and mechanisms (Dzitoyeva et al., 2003; Moore et al., 1998; Scholz et al., 2000, 2005). In several cases, genes identified in flies have also been implicated in mammalian ethanol responses (Thiele et al., 2000; Wand et al., 2001; Wen et al., 2005; Zaleski et al., 2001).
Recent investigations have highlighted similarities between genes and pathways utilized in learning and memory and behavioral responses to drugs of abuse [reviewed (Hyman et al., 2006; Nestler, 2002)]. In Drosophila, several learning and/or memory mutants have been found to display altered responses to ethanol. For example, the mutant cheapdate, identified by increased ethanol sensitivity, was found to be allelic to amnesiac [amn, encoding a PACAP-like neuropeptide regulating cAMP signaling (Feany and Quinn, 1995; Moore et al., 1998)]. Additionally, mutants of the learning/memory gene rutabaga (rut, encoding calcium/calmodulin-sensitive adenylate cyclase) or Pka-C1 (encoding protein kinase A catalytic subunit) displayed increased ethanol sensitivity (Lane and Kalderon, 1993; Levin et al., 1992; Livingstone et al., 1984; Moore et al., 1998). In contrast, mutation of the Pka-RII gene, encoding a regulatory subunit of cAMP-dependent PKA, caused reduced sensitivity (Park et al., 2000). Lastly, fasciclin II (NCAM) mutants displayed both defective learning and increased ethanol sensitivity (Cheng et al., 2001), and flies lacking synapsin, while essentially normal in brain and synapse morphology and by electrophysiological criteria, showed alterations in both learning and memory and ethanol tolerance (Godenschwege et al., 2004).
Recently, a collection of 60 Drosophila long-term memory mutants, representing 52 genetic loci, was described (Dubnau et al., 2003). These mutants exhibited reduced memory following spaced training in a classical olfactory aversive conditioning paradigm, and therefore comprised a large number of potential new components of long-term memory (Tully et al., 1994). To further explore the connection between learning and memory and ethanol-induced behaviors, we quantified ethanol responses in these mutants. In the present study, we used the inebriometer (Moore et al., 1998; Weber, 1988; Weber and Diggins, 1990) to measure ethanol sensitivity and 2 different forms of ethanol tolerance. Our findings reinforce the notion of overlapping biological functions in learning and memory and behavioral responses to ethanol, and suggest specific genes functioning in both of these processes.
Materials and Methods
Fly Strains and Genetics
Flies were grown and maintained at 25°C and 70% relative humidity in constant light on standard cornmeal/molasses/yeast/agar media. The collection of fly long-term memory mutants has been described (Dubnau et al., 2003). Briefly, these mutants were generated by P-element transposon mutagenesis, with each mutant harboring a single homozygous, adult-viable transposon insertion somewhere on the second or third (autosomal) chromosome (Dubnau et al., 2003). Prior to ethanol behavioral testing, all mutants were crossed to [w1118(iso CJ1)] (Dubnau et al., 2003) flies for 5 generations and then re-homozygosed, so that the genetic background of all lines was identical. Each of the mutants described in Dubnau et al., 2003 was assayed for ethanol sensitivity and tolerance with the following exceptions: tungus, valiet-2, and mampus, which could not be recovered as homozygous viable stocks following backcrossing, and milkah-2, beluy-2, and the 3 homozygous lethal mutants (beluy-1, mirta, and lis), which were not included as part of the original, pre-publication collection. In all, 53 lines (52 mutants plus an isogenic control, Ctl) were tested. For some mutants (100% of those checked, namely box, iks, ikar, jack, joy, krasavietz, milord-1, milord-2, murashka-1, murashka-2, premjera, rogdi, ruslan, valiet-1, valiet-3, and zolotistuy), P-element insertion positions were verified to be those previously reported (Dubnau et al., 2003), using inverse PCR and DNA sequencing (http://www.fruitfly.org/about/methods/inverse.pcr.html). Additionally, the position of the P-element in the control strain was determined (Ctl, Table 1); this strain exhibited wild-type inebriometer sensitivity, rapid, and chronic tolerance, and was used as a behavioral control [we did not choose the w-parental line as our behavioral control, as w encodes an ABC-transporter recently shown to affect anesthetic sensitivity (Campbell and Nash, 2001)].
Table 1. Ethanol Sensitivity and Tolerance of Fly Memory Mutants.
| Mutant name | Line | Ethanol phenotypes | MET, min. (SEM) | Rapid tolerance, min.(SEM) | Chronic tolerance, min.(SEM) | P-element position |
|---|---|---|---|---|---|---|
| Ctl | – | 20.9 (0.2) | 10.2 (0.9) | 6.8 (1.3) | ∼800 bp 5′ of path (amino acid transporter)≫ ∼800 bp 3′ of CG3408 (LRR repeat) | |
| arap | E4294 | – | 21.5 (0.8) | 9.4 (1.0) | 8.5 (1.9) | 2,728 bp 5′ of ced-6 ⪡ 4,637 bp 5′ of CG13952 (novel) |
| arleekin | C0141 | – | 21.6 (0.9) | 9.3 (1.5) | 4.3 (1.0) | ⪡ tolkin (BMP-1 homolog) |
| avgust | D0361 | – | 19.2 (0.9) | 6.8 (1.5) | 4.6 (0.6) | 365 bp 3′ of CycB (cyclin) ≫ 973 bp 3′ of blw (mitochondrial ATP synthase) |
| baikal | E1715 | LCT | 18.7 (0.8) | 8.4 (0.8) | 3.1 (0.7); 3.2 (0.8) | ⪡paxillinB (Hh/smo signaling; neurogenesis) |
| barbos-1 | C0015 | S | 16.6 (0.5); 16.3 (1.1) | 10.0 (1.4) | 4.8 (0.6) | 1,782 bp 5′ of CG6044 (novel) ⪡ 371 bp 5′ of CG3624 (Ig repeat) |
| barbos-2 | E2740 | – | 21.0 (1.1) | 7.6 (1.0) | 7.1 (0.2) | same as barbos-1 |
| beck-1 | D0077 | – | 18.6 (1.6) | 11.1 (1.4) | 6.8 (1.4) | ≫lightoid (Rab-RP1) (rab GTPase; eye pigment granule biogenesis) |
| beck-2 | D0091 | – | 21.7 (0.9) | 6.9 (0.6) | 6.1 (1.0) | same as beck-1 |
| box | E1847 | S | 15.4 (0.2); 13.9 (0.8) | 10.7 (0.6) | 8.8 (1.7) | ⪡Gclm (glutathione biosynthesis) |
| chingis khan | E1023 | – | 21.4 (0.9) | 6.4 (1.0) | 5.7 (1.0) | ≫Src64B (src oncogene family) |
| chyorny | E1654 | – | 18.7 (0.7) | 12.3 (1.6) | 4.6 (1.4) | 20,978 bp 5′ of CG9922 (novel) ⪡ 64 bp 5′ of foxo |
| diana | E0391 | LCT | 18.9 (1.0) | 7.2 (0.4) | 3.2 (1.2); 3.7 (0.7) | 250 bp 5′ of CG14696 (novel) ≫ 463 bp 5′ of CG4674 (novel) |
| dikar | E3145 | – | 17.3 (0.4) | 10.3 (1.0) | 6.1 (1.4) | ≫CG32393 (novel) |
| drujok | E3309 | – | 18.1 (1.2) | 7.4 (0.9) | 4.7 (1.0) | 6,623 bp 3′ of Tsp42Ee ⪡ 25 bp 5′ of Tsp42Ef |
| gryzun | E0627 | – | 17.4 (1.4) | 9.9 (1.0) | 3.7 (1.2) | ≫CG17569 (novel) |
| ikar | C0108 | S; LRT | 11.1 (0.8) | 5.5 (0.8); 6.6 (0.4) | 3.8 (0.8) | 4,158 bp 3′ of CG6930 (zinc finger TF) ⪡ 37 bp 5′ of Lk6 (kinase) |
| iks | A0544 | S; LCT | 12.2 (0.3) | 11.8 (0.7) | 3.3 (1.0); 2.7 (0.6) | ≫rhomboid (EGFR-activating peptidase) |
| jack | E3548 | LCT | 19.4 (1.0) | 12.2 (1.3) | 1.5 (0.9); 5.0 (1.0) | 30 bp 5′ of CG2765 (novel) ⪡ 443 bp 3′ of CG30424 (novel) |
| john | E4526 | HCT | 20.2 (2.0) | 9.7 (1.8) | 11.2 (1.0); 8.3 (0.4) | 37 bp 5′ of CG5384 (ubiquitin-specific protease) ≫107 bp 5′ of CG4972 (Zn-dependent exopeptidase) |
| joy | E3947 | S | 16.7 (0.8); 16.3 (0.5) | 10.1 (0.8) | 4.3 (1.5) | ≫cheerio (actin-binding) |
| jurka | E3505 | – | 22.4 (0.7) | 5.8 (0.8); 10.7 (0.2) | 9.0 (1.5) | ≫Mob1 (cell cycle) |
| krasavietz | C0150 | R; LRT; LCT | 25.2 (0.6); 26.8 (0.4) | 0.56 (0.48) | 2.9 (1.1); 2.7 (0.6) | ⪡exba (elF-5C) (translational regulation; axon guidance) |
| laska | C0029 | – | 19.0 (1.2) | 7.6 (2.2) | 3.9 (0.4) | 122 bp 3′ of CG6272 (transcription factor) ⪡ 767 bp 5′ of CG6279 (nitroreductase) |
| lyadi | E4490 | – | 18.2 (0.7) | 8.2 (0.3) | 4.0 (0.4) | ≫CG7897 (gp120, transporter) |
| martik | E3914 | – | 19.5 (1.4) | 12.3 (2.0) | 7.5 (1.6) | ≫CG3361 (novel) |
| milkah-1 | D0283 | – | 20.0 (0.9) | 5.2 (0.6); 10.8 (0.5) | 4.0 (1.4) | ∼40,000 bp 5′ of CG12426 (novel) ≫ 6,409 bp 5′ of CG5017 (nucleosome assembly chaperone) |
| milord-1 | E0469 | S; LRT | 16.8 (0.5); 15.8 (0.9) | 3.8 (0.4) | 4.1 (0.6) | ≫pumilio (translational regulator) |
| milord-2 | E2655 | S | 17.4 (0.6); 14.0 (0.7) | 12.9 (0.7); 8.7 (1.4) | 7.9 (0.7) | ⪡pumilio (translational regulator) |
| moladietz | E1877 | – | 19.0 (0.1) | 6.7 (0.4) | 3.8 (0.6) | ⪡CG4482 (novel) |
| murashka-1 | D575B | – | 21.4 (0.6) | 8.0 (1.0) | 9.0 (1.6) | ⪡CG9381 (RING finger) |
| murashka-2 | E3803 | R | 30.0 (1.5) | 15.5 (2.0); 9.8 (1.6) | 7.1 (1.0) | ⪡CG9381 (RING finger) |
| nord | E3272 | – | 20.7 (1.2) | 11.4 (0.5) | 12.2 (1.2); 5.4 (0.7) | ⪡CG3372 (novel) |
| norka | D0067 | – | 19.8 (0.9) | 8.6 (1.7) | 6.8 (1.6) | 624 bp 5′ of CG11962 (novel) ≫ 3,265 bp 5′ of oskar (mRNA localization) |
| novichok | D0940 | – | 16.2 (0.9); 18.2 (0.6) | 6.9 (0.4) | 5.7 (1.4) | 346 bp 5′ of CG10646 (WD40 repeats) ≫ 109 bp 5′ of vihar (cyclin-selective ubiquitin ligase) |
| pastrel | C0167 | – | 21.0 (3.2) | 13.2 (1.7) | 8.1 (1.1) | ≫Hsp70Aa |
| pingiel | D0991 | – | 18.1 (0.3) | 9.5 (1.3) | 5.2 (0.9) | ⪡slmb (ubiquitin protein ligase) |
| premjera | E3945 | S | 15.8 (0.9); 15.0 (1.0) | 9.3 (0.5) | 3.6 (0.4) | ≫CG1910 (novel) |
| rafael | D0264 | S; LCT | 14.3 (1.1) | 8.0 (0.9) | 3.4 (0.6); 2.2 (0.6) | 1,151 bp 3′ of CG5208 (novel) ⪡ 6,075 bp 5′ of Dad (negative regulator of TGF-β |
| rex | D0753 | – | 23.0 (1.2) | 6.8 (1.0) | 4.1 (1.0) | 1,212 bp 5′ of EP2237/cabut (Zn finger C2H2) ≫ 32,932 bp 5′ of usher (steroid transcription factor) |
| rijiy I | E0389 | – | 18.3 (1.3) | 8.9 (1.0) | 4.6 (0.8) | 46,606 bp 5′ of CG5620 (transcription factor) ≫ 776 bp 5′ of CG10429 (novel) |
| rijiy II | E4309 | – | 21.4 (1.8) | 13.4 (1.1) | 4.9 (1.6) | ≫fasciclin3 (neurogenesis) |
| rogdi | C0113 | LRT | 23.8 (0.9) | 5.4 (1.4); 9.2 (0.8) | 6.8 (1.6) | ⪡CG7725 (novel) |
| rosa | E0602 | – | 20.6 (0.7) | 7.8 (0.6) | 5.4 (0.4) | 51,287 bp 5′ of CG6559 (novel) ⪡ 48,903 bp 5′ of CG12362 (ubiquitin ligase) |
| ruslan | D0417 | S; LRT; LCT | 14.6 (0.2) | 6.1 (1.9); 5.5 (0.6) | 3.4 (0.5); 2.5 (0.2) | ⪡klingon (Ig superfamily; neurogenesis) |
| toi | D0177 | – | 20.5 (1.6) | 11.8 (2.1) | 6.0 (1.4) | 13,243 bp 3′ of CG13299 (novel) ≫ 669 bp 5′ of LanA (laminin) |
| trezor | E3503 | – | 17.1 (0.8) | 11.1 (2.0) | 8.2 (1.0) | ≫Mob1 (cell cycle) |
| umnitza | A0563 | – | 25.8 (1.1); 16.9 (1.4) | 12.6 (0.4) | 9.2 (2.0) | ≫schnurri (transcription factor; dpp signaling) |
| valiet-1 | A0023 | LRT | 25.2 (0.7); 20.7 (0.7) | 8.4 (0.8); 7.1 (0.3) | 4.6 (0.3) | ⪡RluA-1 (pseudouridine synthase; RNA metabolism) |
| valiet-3 | C0085 | LRT | 25.6 (1.8); 20.2 (0.4) | 4.5 (1.2); 2.9 (0.8) | 8.9 (2.0) | same as valiet-1 |
| visgun | E4203 | LCT | 20.2 (1.3) | 9.3 (1.1) | 2.4 (0.5); 4.8 (0.6) | ⪡CG16707(endolyn homolog) |
| zloday | E4299 | – | 21.9 (1.1) | 6.8 (0.9) | 4.7 (0.9) | ∼20,000 bp 5′ of SP1070 ⪡ 2921 bp 5′ of CG13776 (chaperone J-domain) |
| zolotistuy | E0511 | S; LRT | 14.2 (1.4) | 6.2 (1.5); 7.4 (0.6) | 3.9 (1.3) | ⪡formin3 (actin assembly promoting) |
Mutant names, line designations, and P-element insertion positions are from Dubnau et al., 2003. The learning and memory defects of the mutants have been described (Dubnau et al., 2003). P-element insertion positions were verified for 16 of 16 selected mutants by iPCR and DNA sequencing (Materials and Methods). Gene annotations were updated based on Flybase (http://flybase.bio.indiana.edu). For insertions between 2 genes, the distance from each is given in base pairs (bp). Arrowheads indicate orientation of the P-element with respect to adjacent gene(s). Ethanol sensitivity, rapid, and chronic tolerance were quantified in the inebriometer (Materials and Methods). All values are mean ± SEM. For each genotype, n ≥ 3 for ethanol sensitivity and rapid tolerance; n ≥ 4 for chronic tolerance and for sensitivity and tolerance retests; values for retests, if performed, are listed after original values. S, increased ethanol sensitivity; R, decreased ethanol sensitivity; LRT, low rapid tolerance; HRT, high rapid tolerance; LCT, low chronic tolerance; HCT, high chronic tolerance; –, not significantly different from Ctl. Our behavioral control (Ctl) consisted of a strain harboring a P-element insertion of the PlacZ type, the same as that of the “E” lines from the memory mutants, in the isogenic background strain, 2202U [(w; iso(CJ1) (Dubnau et al., 2003)]. Ethanol sensitivity and tolerance of this strain appeared wild-type compared with other control strains used in our laboratory (data not shown).
All experiments used adult male flies which were collected 0 to 3 days following eclosion (day 13 after egg-laying). Flies were kept in large vials on standard fly food at 25°C and 70% relative humidity for an additional 2 days prior to experiments to minimize any effects of CO2 anesthesia on behavior.
Inebriometer Assay for Ethanol Sensitivity
The inebriometer assay was used to measure ethanol sensitivity essentially as previously described (Moore et al., 1998; Singh and Heberlein, 2000). Briefly, for each assay, ∼120 flies of a single genotype were introduced into the top of an inebriometer column, a ∼4-ft high glass column containing multiple, uniformly distributed oblique mesh baffles throughout which ethanol vapor is circulated. Columns were pre-equilibrated at a temperature of 20°C with a mixture of ethanol vapor and humidified air at a ratio of ∼55/45 ethanol/air (E/A). Over time, flies' internal ethanol concentration increases and they lose the ability to stand on the mesh baffles, eventually falling (eluting) from the column. Flies eluting were counted at 3-minute intervals using a Drosophila Activity Monitoring System (TriKinetics Inc., Waltham, MA). The mean elution time (MET) for each sample population of ∼120 flies was calculated as the sum of the number of flies eluting at a given time (minute) multiplied by the time of elution divided by the total number of flies and used as a measure of sensitivity to ethanol intoxication. Those mutants that were most sensitive in the inebriometer (ikar, iks, rafael, ruslan, and zolotistuy) were additionally tested for baseline ability to stay in the inebriometer column in the absence of ethanol vapor. In 30 minutes, which is the inebriometer MET of the most resistant line, essentially no flies fell from the inebriometer for even these most sensitive mutants. On this basis, most if not all of the differences in ethanol sensitivity observed between the mutants were based on different levels of response to ethanol, rather than gross differences in motor coordination or ability to cling to the inebriometer column.
Rapid Tolerance Assay
Inebriometer assays measuring rapid tolerance were performed essentially as previously described (Berger et al., 2004). Briefly, for each assay, duplicate samples of 120 flies each were pre-exposed for 30 minutes in perforated 50-mL Falcon tubes without food, to either a mixture of ethanol vapor and humidified air at a relative flow rate of 60/40 E/A, or to an equivalent total flow rate of humidified air alone (treatment control). Following this pre-exposure, both sets of flies were allowed to recover for 3.5 hours at 25°C and 70% relative humidity and assayed in the inebriometer. Tolerance values were calculated as the increase in MET of flies pre-exposed to ethanol vapor relative to those pre-exposed to humidified air, using the formula METeth − METair. To obtain both sensitivity and rapid tolerance data simultaneously, we used the METair values obtained for the control-treated flies to quantify ethanol sensitivity; in separate tests, air pre-exposure did not have any significant effect on inebriometer MET (data not shown).
Chronic Tolerance Assay
Inebriometer measurement of chronic tolerance was performed essentially as previously described (Berger et al., 2004). For each assay, duplicate samples of 120 flies each were placed into perforated tubes containing 15 mL standard fly food, and were pre-exposed for 20 to 28 hours to a mixture of ethanol vapor and humidified air at a relative flow rate of 10/80 E/A, or to an equivalent flow rate of humidified air alone (treatment control). Immediately following this pre-exposure, flies were assayed in the inebriometer, and tolerance was calculated using the formula METeth − METair.
Retests for Sensitivity, Rapid, and Chronic Tolerance
Initial statistical analyses were carried out by one-way ANOVA and post hoc Holm–Sidak tests comparing all 52 mutants to control, resulting in only a few strains that were significantly different. To increase statistical sensitivity, a subset of mutants was selected for small-group retests versus control. For ethanol sensitivity, mutants were retested in additional inebriometer assays if their METs differed by more than 4 minutes from the control strain MET of 20.9 minutes (>24.9 or <16.9 minutes), a cutoff used in previous inebriometer screens (Singh and Heberlein, 2000; D. Guarnieri, unpublished data). For rapid tolerance, strains were selected for retesting if their rapid tolerance differed by more than 3.9 minutes from the control value of 10.2 minutes (>14.1 or <6.3 minutes); this cutoff was chosen to be roughly equivalent by one-way ANOVA to the inebriometer MET cutoff used for sensitivity retests (data not shown). For chronic tolerance, candidate mutants were retested if their tolerance was more than 3.3 minutes different from the control value, 6.8 minutes (>10.1 or <3.5 minutes). This cutoff led to retesting approximately the same number of mutants as for sensitivity and rapid tolerance.
Ethanol Absorption Assay
Fly internal ethanol concentrations were determined from whole-fly homogenates of 25 flies per sample. Flies were exposed 8 samples at a time in perforated 50-mL Falcon tubes. The ethanol exposure regimen used for mutants was based on their altered ethanol phenotype(s). For ethanol sensitivity mutants, flies were exposed for 30 minutes to a mixture of humidified air and ethanol vapor at a relative flow rate of 60/40 or 55/45 E/A (60/40 E/A was used in earlier assays, while 55/45 E/A was used subsequently to match the inebriometer settings). For rapid tolerance, flies were pre-exposed for 30 minutes to ethanol vapor at 60/40 E/A or to an equivalent flow of humidified air alone (control), allowed to recover 3.5 hours, and then exposed to 55/45 E/A for 30 minutes. For chronic tolerance, flies were pre-exposed for 18 to 24 hours to 10/80 E/A or to an equivalent flow of humidified air alone, then immediately exposed to 55/45 E/A for 30 minutes. In all cases, flies were snap-frozen on dry ice following the final exposure to ethanol, and either stored at −80°C or homogenized immediately at 4°C in 250 μL of 50 mM Tris–HCl, pH 7.5. Homogenates were centrifuged at 15,000 × g for 20 minutes at 4°C and ethanol concentrations in supernatants were measured using an alcohol dehydrogenase-based spectrophotometric assay (Ethanol Assay Kit #229-29; Diagnostics Chemicals Limited, Oxford, CT). To calculate fly internal ethanol concentration, the volume of 1 fly was estimated to be ∼2 μL; there were no obvious strain-based differences in fly volumes (sizes).
Calculations and Statistics
Ethanol sensitivity was quantified based on the average inebriometer MET of sample populations of ∼120 male flies, as described above and previously (Moore et al., 1998; Singh and Heberlein, 2000). Rapid and chronic tolerance were calculated as the difference in MET between ethanol and air pre-exposed flies, using the equation (METeth − METair) to express tolerance as a difference value. From each assay, a value for tolerance was calculated from each value of METeth and METair, and averages and standard errors for tolerance were calculated within each genotype and set of experiments based on these individual tolerance values. Statistical significance was assessed by ANOVA and post hoc Holm–Sidak tests using SigmaStat (version 3.1) statistical software (Systat Software Inc., San Jose, CA). In all experiments, n refers to the number of samples, not the number of flies.
Results
Ethanol Sensitivity
To measure sensitivity of the collection of memory mutants to the uncoordinating effects of ethanol, each line was assayed in the inebriometer (Moore et al., 1998; Weber, 1988; Weber and Diggins, 1990). When the 52 mutants were tested, the resulting METs ranged widely, from a low of 11.1 ± 0.8 minutes (ikar) to a high of 30.0 ± 1.5 minutes (murashka-2) (Fig. 1 and Table 1). Initial analysis identified 8 mutants with altered ethanol sensitivity, 7 with increased sensitivity and 1 with decreased sensitivity (Fig. 1B). Inebriometer elution profiles are shown for one of the increased sensitivity mutants, ikar, and the decreased sensitivity mutant, murashka-2 (Fig. 1C and D).
Fig. 1.

Ethanol sensitivity of the Drosophila memory mutants. (A) Histogram illustrates frequency distribution of inebriometer mean elution times (MET, min; 3-minute bins) for 52 memory mutants. Solid line indicates mutant population mean MET, 19.5 minutes, and dashed line the control strain MET, 20.9 ± 0.2 minutes. (B) Bar graph represents inebriometer METs (±SEM) of the 8 mutants showing altered ethanol sensitivity in first-round analysis (Ctl, control; *denotes p < 0.001 for each mutant vs. Ctl; F52,198 = 10.45, p < 0.001). (C and D) Individual inebriometer elution profiles for the most ethanol-sensitive mutant, ikar, versus control (C) and for the ethanol-resistant mutant, murashka-2, versus control (D); n = 4 (here and elsewhere, n represents the number of samples, not the number of flies; in all figures, error bars represent standard errors of the mean).
Initially, all the ethanol sensitivity data were compared by one-way ANOVA and adjusted post hoc for multiple comparisons across 53 strains (52 mutants plus control). Thus, this initial analysis was of fairly high statistical stringency, and few mutants were significantly different from control. To increase statistical sensitivity and ensure mutants were not being missed, some strains were selected to retest in small groups. Strains were retested in the inebriometer if their MET was at least 4 minutes different from the control strain (<16.9 or >24.9 minutes). This cutoff was chosen based on previous inebriometer behavioral screens performed in our laboratory (Singh and Heberlein, 2000; D. Guarnieri, unpublished data). Using this cutoff, 8 strains, 4 candidates for increased sensitivity (barbos-1, joy, novichok, and milord-1) and 4 for reduced sensitivity (krasavietz, umnitza, valiet-1, and valiet-3), were selected for retesting. Additionally, the strain milord-2 was retested because it was allelic to milord-1, 1 of the candidates for increased sensitivity, and moreover was close to the 16.9-minute retest cutoff.
Strains were assayed in small groups of 1 to 3 strains plus the control (Fig. 2; Table 1). In these assays, 3 of the 4 candidates for increased sensitivity behaved consistent with their earlier performances and were significantly more sensitive to ethanol than control (barbos-1, joy, and milord-1); milord-2 also exhibited increased ethanol sensitivity (Fig. 2 and Table 1). In contrast, only 1 of the 4 candidates for decreased sensitivity differed significantly from control upon retesting (krasavietz; Fig. 2 and Table 1). It was possible that the other 3 candidates for decreased sensitivity might have accumulated one or more modifier mutations that suppressed their original phenotype during the time elapsed between initial assays and retesting. Accordingly, the mutants umnitza, valiet-1, and valiet-3 were re-outcrossed to the parental line, and newly homozygous lines were re-derived and tested immediately. Ethanol sensitivity of the newly backcrossed lines did not differ significantly from control (data not shown). In summary, of the 52 memory mutants tested, 11 showed significant alterations in ethanol sensitivity, 9 with increased sensitivity (barbos-1, box, ikar, iks, joy, milord-1, milord-2, premjera, rafael, ruslan, and zolotistuy) and 2 with decreased sensitivity (krasavietz and murashka-2).
Fig. 2.

Additional ethanol sensitivity mutants identified by retesting. Inebriometer assays were performed on small groups of 1 to 3 candidate mutants plus control (Ctl); p values for each mutant versus Ctl are denoted by asterisks: *p < 0.05; **p < 0.001; n = 4. (A) Only barbos-1 was significantly different from control (F3,15 = 4.99, p = 0.018; barbos-1 vs. Ctl, p = 0.013). (B) Neither novichok nor valiet-3 was significantly different from control (F2,11 = 5.22, p = 0.03; novichok vs. Ctl, p = 0.24; valiet-3 vs. Ctl, p = 0.08). (C) Both joy and milord-1 showed increased ethanol sensitivity (F2,11 = 32.17, p < 0.001), as did milord-2 (D) (F1,7 = 47.7, p < 0.001), while krasavietz (E) exhibited significantly reduced sensitivity (F1,7 = 48.05, p < 0.001).
Rapid Ethanol Tolerance
Tolerance, defined as a decreased response to repeated drug exposures, is a hallmark of alcohol abuse and addiction, and represents a form of behavioral plasticity in response to alcohol (Fadda and Rossetti, 1998). Multiple forms of tolerance have been defined operationally, based on the regimen used to establish tolerance (Kalant et al., 1971). In procedures used to study rapid tolerance in flies as well as in mammals, a single intoxicating dose of ethanol is allowed time to be fully metabolized and then assessed for effects on behavioral response to a second ethanol exposure (Scholz et al., 2000; Tabakoff et al., 1986). In rodents, rapid tolerance is traditionally measured 24 hours after the first exposure (e.g., Crabbe et al., 1979), whereas in flies, rapid tolerance is most readily assessed at its peak levels 2 to 4 hours after initial exposure, a time when the ethanol absorbed during the first exposure is completely metabolized (Scholz et al., 2000).
To quantify rapid tolerance, flies were pre-exposed to an intoxicating level of ethanol vapor, given sufficient time (3.5 hours) to recover and metabolize this ethanol, and then assayed in the inebriometer. A duplicate sample was pre-exposed to humidified air and assayed as a treatment control in parallel for each experiment. METs were determined for the ethanol and air pre-exposed flies (denoted METeth and METair, respectively). To evaluate tolerance of these mutants, of which a large fraction exhibited substantially altered sensitivity to ethanol, tolerance values were calculated as the increase in MET of flies pre-exposed to ethanol relative to control (air pre-exposed) flies.
When the memory mutants were measured for development of rapid tolerance, values ranged from a low of 0.6 ± 0.5 minutes (krasavietz) to a high of 15.5 ± 2.0 minutes (murashka-2) (Fig. 3). The mean rapid tolerance value of the collection was 8.8 minutes and rapid tolerance of the control strain was 10.2 ± 0.9 minutes. Initial analysis of data across the entire collection identified 3 mutants, each exhibiting reduced rapid tolerance: krasavietz, milord-1, and valiet-3 (Fig. 3). To explore the possibility of additional mutants, strains were selected for retesting if their rapid tolerance differed by >3.9 minutes from control (tolerance >14.1 or <6.3 minutes). Using this cutoff, 7 mutants were selected for retesting: 6 candidates for reduced tolerance (ikar, jurka, milkah, rogdi, ruslan, and zolotistuy) and 1 for increased tolerance (murashka-2). Additionally, we retested milord-2 and valiet-1, which were allelic to the milord-1 and valiet-3 rapid tolerance mutants previously identified. Lastly, we also retested valiet-3, to ascertain whether this mutant's reduced rapid tolerance phenotype was stable despite its loss of decreased ethanol resistance (described above; see Fig. 2).
Fig. 3.

Rapid ethanol tolerance in the memory mutants. Flies were pre-exposed for 30 minutes to ethanol vapor (at a relative flow rate of 60/40 E/A, a sedating dose) or to humidified air at an equivalent total flow rate as a control, allowed to recover for 3.5 hours and assayed in the inebriometer. (A) Histogram represents frequency distribution of values for rapid tolerance (minutes, min.), using 3 minute bins. Vertical solid line denotes mutant population mean tolerance (8.8 minutes), and dashed line rapid tolerance of the control strain (Ctl, 10.2 ± 0.9 minutes). (B) MET and rapid tolerance values for krasavietz, milord-1, and valiet-3, the 3 reduced rapid tolerance mutants identified in first-round analysis (F52,198 = 5.83, p < 0.001; *denotes p < 0.001 for each mutant vs. Ctl; n = 4). For MET values, dark gray bars denote flies pre-exposed to ethanol vapor, and light gray bars, humidified air (treatment control). (C and D) Average inebriometer elution profiles for Ctl flies (C) and the krasavietz mutant (D). “Ethanol” denotes flies pre-exposed to ethanol vapor, and “Air” the treatment control, air pre-exposed flies.
Upon retesting, 5 additional mutants, all of which showed reduced rapid tolerance, were identified: ikar, rogdi, ruslan, valiet-1, and zolotistuy (Fig. 4). Similar to valiet-1, the valiet-3 mutant again exhibited reduced rapid tolerance along with wild-type ethanol sensitivity (Table 1). Unlike milord-1, milord-2 exhibited rapid tolerance that was not significantly reduced compared with control (Fig. 4C). The mutant murashka-2, which was the only candidate for increased rapid tolerance, displayed wild-type tolerance upon retesting (Fig. 4D). In all, 8 mutants (ikar, krasavietz, milord-1, rogdi, ruslan, valiet-1, valiet-3, and zolotistuy) exhibited reduced rapid tolerance, while no mutants exhibited increased rapid tolerance.
Fig. 4.

Additional rapid tolerance mutants identified by retesting. Inebriometer assays measuring tolerance were performed on small groups of 1 to 3 candidate mutants plus control (Ctl). Dark gray bars denote MET values for flies pre-exposed to ethanol vapor, and light gray bars, humidified air; p values for tolerance of each mutant compared with Ctl are denoted by asterisks: *p < 0.01; **p < 0.001; n = 4–6. The mutants valiet-1 (A) (F1,7 = 41.38, p < 0.001) and rogdi (B) (F3,17 = 5.53, p = 0.01; rogdi vs. Ctl, p = 0.0013) showed reduced tolerance, while milord-2 was not significantly different from Ctl (C) (F1,7 = 0.9; p = 0.38). (D) The mutants ikar, ruslan and zolotistuy also exhibited significant reductions in rapid tolerance (F4,25 = 6.96, p < 0.001).
Chronic Ethanol Tolerance
Lastly, the memory mutant collection was evaluated in a second ethanol tolerance paradigm, which we have termed chronic tolerance. Wild-type flies have previously been shown to develop tolerance following a continuous, prolonged (24 or 48 hours) exposure to a low concentration of ethanol vapor, at internal ethanol concentrations that never reached intoxicating levels (Berger et al., 2004). This form of tolerance could be at least partially distinguished genetically and mechanistically from rapid tolerance, and was specifically occluded by prefeeding flies a protein synthesis inhibitor (Berger et al., 2004). Each of the 52 mutants was evaluated for chronic tolerance compared with the control strain (Fig. 5). Values ranged from a low of 1.5 ± 0.9 minutes (jack) to a high of 12.2 ± 1.2 minutes (nord); mean chronic tolerance of the mutant collection was 5.6 minutes, and chronic tolerance of the control strain was 6.8 ± 1.3 minutes.
Fig. 5.

Chronic ethanol tolerance in the memory mutants. Flies were pre-exposed overnight (20 to 28 hours) to a nonsedating concentration of ethanol vapor (10/80 E/A) or to humidified air as a control and assayed in the inebriometer. (A) Histogram represents frequency distribution of values for chronic tolerance (minutes, min.), using 2-minute bins. Vertical solid line denotes mutant population mean chronic tolerance (5.6 minutes), and dashed line denotes tolerance of the control strain (Ctl, 6.8 ± 1.3 minutes). (B) MET and chronic tolerance values for the control strain (Ctl) and the mutant with the lowest chronic tolerance, jack. Dark gray bars denote MET values for flies pre-exposed to ethanol, and light gray bars, humidified air. In first-round analysis across the entire collection, no mutants exhibited chronic tolerance significantly different from Ctl (F52,272 = 4.27; p < 0.001; n = 4–10). (C and D) Average inebriometer elution profiles for Ctl (C) or jack (D) flies pre-exposed overnight to low-level ethanol vapor (Ethanol) or to an equivalent flow rate of humidified air (Air).
In first-round data analysis, no mutants were significantly different from control (Fig. 5). Mutants were selected for subsequent retesting using a cutoff of >3.3 minutes from the control value of 6.8 minutes (<3.5 or >10.1 minutes). Using this criterion, 8 mutants (listed below) were candidates for reduced chronic tolerance and 2 mutants (john and nord) were candidates for enhanced chronic tolerance. These strains were retested for chronic tolerance in small groups of 2 or 3 candidate mutants plus the control strain (Fig. 6). The mutant john, but not the mutant nord, displayed significantly increased chronic tolerance (Fig. 6A), while each of the 8 candidates for reduced chronic tolerance exhibited significantly reduced tolerance (Fig. 6B–D). In all, 8 mutants, baikal, diana, iks, jack, krasavietz, rafael, ruslan, and visgun, displayed reduced chronic tolerance and the mutant john displayed enhanced chronic tolerance.
Fig. 6.

Chronic tolerance mutants identified by small-group retesting. Inebriometer assays measuring tolerance were performed on small groups of 1 to 3 candidate mutants plus control (Ctl). (A–D) Dark gray bars denote MET values for flies pre-exposed to ethanol, and light gray bars, humidified air; p values for tolerance of each mutant versus Ctl are denoted by asterisks: *p < 0.05; **p < 0.01; ***p < 0.001; n = 5–9. (A) The mutant john exhibited significantly increased chronic tolerance (F2,21 = 10.0, p < 0.001), while (B–D) a total of 8 mutants exhibited significantly reduced tolerance [(B) F3,26 = 6.77, p = 0.002; (C) F2,15 = 6.45, p = 0.011; (D) F3,22 = 7.42, p = 0.002)].
Ethanol Pharmacokinetics
To assess whether altered ethanol pharmacokinetics might underlie the ethanol phenotypes of any of the mutants, each mutant showing altered sensitivity or tolerance was exposed to ethanol vapor and then processed to determine ethanol content (Fig. 7). Of the 13 lines displaying altered sensitivity, none displayed a significant alteration in ethanol content following an ethanol vapor exposure comparable with that used in the inebriometer assay (Fig. 7A and B). Of the strains that displayed altered rapid tolerance, one, milord-1, showed reduced ethanol content compared with control; this reduction was independent of whether flies were pre-exposed to ethanol vapor or humidified air (Fig. 7C). Finally, among the chronic tolerance mutants, 2 mutants, rafael and ruslan, showed increased ethanol content (Fig. 7D). In the case of rafael, this increase was independent of pre-exposure condition. For ruslan, and also for the mutants jack and john, flies pre-exposed to ethanol showed reduced ethanol content relative to isogenic (treatment control) flies pre-exposed to humidified air. The reason for this difference is unknown; however, it was in a direction opposite to that which would explain the decreased chronic tolerance of these mutants. In summary, the mechanisms underlying most if not all of the altered mutant ethanol phenotypes appeared to be functional, rather than dispositional, in nature.
Fig. 7.
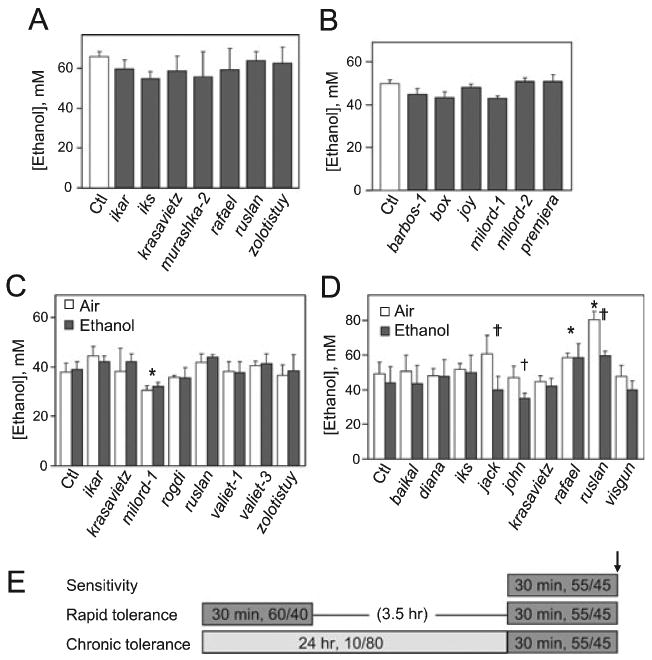
Ethanol pharmacokinetics of ethanol sensitivity and tolerance mutants. (A and B) Mutants exhibiting altered ethanol sensitivity were exposed to ethanol vapor at a relative flow rate of 60/40 (A) or 55/45 E/A (B) for 30 minutes and processed immediately. No significant difference in ethanol content was seen for any of the altered sensitivity mutants [(A), F7,31 = 1.02, p = 0.44; (B) F6,29 = 2.38, p = 0.06; n = 4–6)]. (C) Mutants exhibiting reduced rapid tolerance were pretreated by a 30-minute exposure to 60/40 E/A (Ethanol), or to humidified air at an equivalent flow rate (Air). Following a 3.5-hour recovery, all samples were exposed for 30 minutes to 55/45 E/A and processed. In two-way ANOVA, a significant main effect of genotype was seen (F8,79 = 6.78, p < 0.001), with the mutant milord-1 exhibiting a significant decrease in ethanol content compared with Ctl (p < 0.001); no significant effect of pre-exposure treatment was observed (F1,79 = 0.50, p = 0.48), and there was no significant interaction between genotype and pre-exposure treatment (F8,79 = 0.38, p = 0.93). n = 4–8. (D) Mutants exhibiting altered chronic tolerance were pretreated by ∼24-hour exposure to 10/80 E/A (Ethanol) or to humidified air at an equivalent flow rate (Air), then all samples were exposed for 30 minutes to 55/45 E/A and processed. In two-way ANOVA, a significant effect of genotype was detected (F9,89 = 12.03, p < 0.001), with the mutants rafael and ruslan significantly different from Ctl (*p < 0.001). A significant effect of pre-exposure condition was also detected (F1,89 = 26.2, p < 0.001), and there was a significant interaction between genotype and pre-exposure condition (F9,89 = 2.56, p < 0.013) for the mutants jack (p < 0.001), john (p = 0.017) and ruslan (p < 0.001; significance of interaction is denoted by daggers: †p < 0.05; ††p < 0.001; n = 3–8). (E) Line diagram illustrates exposure regimens for sensitivity and tolerance mutants; arrow indicates point at which flies were processed to determine ethanol content.
Discussion
Of the 52 mutants examined, 21 (∼40%) exhibited alterations in ethanol sensitivity and/or tolerance. While direct comparisons to unbiased screens previously conducted by others in our laboratory are challenged by differences such as genetic background and screening methodology, such screens have typically identified mutants at a frequency of 1% or less [altered ethanol sensitivity, 12 mutants of 5,000 screened (Moore et al., 1998) or 23 mutants of nearly 30,000 screened (Singh and Heberlein, 2000); altered ethanol tolerance, 2 mutants of 404 screened (Scholz et al., 2005)]. Conservatively, then, the substantially higher frequency with which we detected ethanol mutants among the memory mutants is at least consistent with the notion of shared pathways for learning and memory and ethanol behavioral responses.
In rodents, a number of studies have implicated common pathways, and in some cases specific genes, either directly or indirectly in both ethanol behavioral responses and learning and memory. Ethanol tolerance in particular likely involves adaptive changes in synaptic function, and may be described as a form of synaptic plasticity (Chandler et al., 1998). Earlier studies in fact suggest a learning component for multiple forms of tolerance (e.g. Khanna et al., 1994; Le and Kalant, 1992). Proteins implicated in both learning and memory and ethanol behaviors include neurotransmitters and neuromodulators, as well as other components of intracellular signaling and/or synaptic plasticity such as cAMP-dependent PKA, protein kinase C, and cAMP-response element-binding protein (see Crabbe et al., 2006; Hyman et al., 2006 for recent reviews). Here and elsewhere it is important to keep in mind the procedural differences in establishing tolerance in flies versus rodents, principally the different time frames involved, which may correspond to underlying mechanistic differences. For example, in rats a requirement for new protein synthesis was shown for rapid tolerance (measured at 24 hours after first exposure), while in flies such a requirement was not observed (with rapid tolerance measured 3.5 hours after first exposure) (Berger et al., 2004; Bitran and Kalant, 1993).
In the current study, while 9 mutants exhibited increased sensitivity, only 2 exhibited decreased sensitivity (Table 2). Similarly, mutants exhibiting reduced tolerance, either rapid or chronic, were much more frequent (15 total) than mutants exhibiting enhanced tolerance (only 1 mutant). This is perhaps not surprising, given that P-element insertions will most often disrupt expression of the nearby gene, and are therefore generally more likely to reduce rather than enhance function of the gene. Interestingly, however, a recent analysis of ethanol sensitivity and tolerance of 20 fly lines with P-element insertions in or near genes of interest (genes that showed expression changes following ethanol exposure) identified 7 lines exhibiting decreased sensitivity, and only 1 with increased sensitivity (Morozova et al., 2006). This same study also identified more strains with enhanced ethanol tolerance (7) than reduced tolerance (4). Although these studies were performed in a different genetic background than ours [Canton S, vs. w1118(iso CJ1)], a more obvious difference seems to be that the mutants tested here were preselected, having been previously identified as defective for long-term (24 hours) memory (Dubnau et al., 2003).
Table 2. Summary of Ethanol Sensitivity and Tolerance Mutants.
| Ethanol phenotype | ||
|---|---|---|
| Mutant name (line) | Other ethanol phenotypes | Gene(s) implicated based on P-element position |
| Increased sensitivity | ||
| barbos-1 (C0015) | – | 1,782 bp 5′ of CG6044 (novel) ⪡ 371 bp 5′ of CG3624 (Ig repeat) |
| box (E1847) | – | ⪡Gclm (glutathione biosynthesis) |
| ikar (C0108) | LRT | 4,158 bp 3′ of CG6930 (zinc finger TF)⪡37 bp 5′ of Lk6 (kinase) |
| iks (A0544) | LCT | ≫rhomboid (EGFR-activating peptidase) |
| joy (E3947) | – | ≫cheerio (actin-binding) |
| milord-1 (E0469) | LRT | ≫pumilio (translational regulator) |
| milord-2 (E2655) | – | ⪡pumilio (translational regulator) |
| premjera (E3945) | – | ≫CG1910 (novel) |
| rafael (D0264) | LCT | 1,151 bp 3′ of CG5208 ⪡ 6,075 bp 5′ of Dad (negative regulator of TGF-β) |
| ruslan (D0417) | LRT, LCT | ⪡klingon (Ig superfamily; neurogenesis) |
| zolotistuy (E0511) | LRT | ⪡formin3 (actin assembly promoting) |
| Decreased sensitivity | ||
| krasavietz (C0150) | LRT, LCT | ⪡exba (eIF-5C) (translational regulation; axon guidance) |
| murashka-2 (E3803) | – | ⪡CG9381 (RING finger) |
| Reduced rapid tolerance | ||
| ikar (C0108) | S | 4,158 bp 3′ of CG6930 (zinc finger TF) ⪡ 37 bp 5′ of Lk6 (kinase) |
| krasavietz (C0150) | R, LCT | ⪡exba (eIF-5C) (translational regulation; axon guidance) |
| milord-1 (E0469) | S | ≫pumilio (translational regulator) |
| rogdi (C0113) | – | ⪡CG7725 (novel) |
| ruslan (D0417) | S, LCT | ⪡klingon (Ig superfamily; neurogenesis) |
| valiet-1 (A0023) | – | ⪡RluA-1 (pseudouridine synthase; RNA metabolism) |
| valiet-3 (C0085) | – | ⪡RluA-1: same as valiet-1 |
| zolotistuy (E0511) | S | ⪡formin3 (actin assembly promoting) |
| Reduced chronic tolerance | ||
| baikal (E1715) | – | ⪡paxillinB (Hh/smo signaling; neurogenesis) |
| diana (E0391) | – | 250 bp 5′ of CG14696 (novel) ≫ 463 bp 5′ of CG4674 (novel) |
| iks (A0544) | S | ≫rhomboid (EGFR-activating peptidase) |
| jack (E3548) | – | 30 bp 5′ of CG2765 (novel) ⪡ 443 bp 3′ of CG30424 (novel) |
| krasavietz (C0150) | R, LRT | ⪡exba (eIF-5C) (translational regulation; axon guidance) |
| rafael (D0264) | S | 1,151 bp 3′ of CG5208 ⪡ 6,075 bp 5′ of Dad (negative regulator of TGF-β) |
| ruslan (D0417) | S, LRT | ⪡klingon (Ig superfamily; neurogenesis) |
| visgun (E4203) | – | ⪡CG16707 (endolyn homolog) |
| Enhanced chronic tolerance | ||
| john (E4526) | – | 37 bp 5′ of CG5384 (ubiquitin-specific protease) ≫ 107 bp 5′ of CG4972 (Zn-dependent exopeptidase) |
We compared the previously published strain mean values for the memory deficits of the mutants (ibid.) with their levels of ethanol sensitivity and tolerance, and did not observe any significant overall genetic correlations (sensitivity vs. memory, r = 0.04, p = 0.77; rapid tolerance vs. memory, r = 0.05, p = 0.70; chronic tolerance vs. memory, r = 0.16, p = 0.24). We also examined whether there were overall relationships between different ethanol phenotypes in the mutants. There was a modest but significant positive overall correlation between rapid tolerance and chronic tolerance (r = 0.3, p = 0.03), but no significant overall correlation between rapid tolerance and sensitivity, nor chronic tolerance and sensitivity (r = −0.04, p = 0.78; r = 0.23, p = 0.10, respectively).
A third form of tolerance not yet mentioned here is acute (within-session) tolerance, tolerance which is acquired and manifested during a single ethanol exposure (Kalant et al., 1971). While we did not explicitly measure acute tolerance, the time spent by flies in the inebriometer during a typical experiment (from 10 to 30 minutes) may well permit its development, so that inebriometer MET would reflect acute tolerance as well as initial sensitivity to the onset of intoxication. In rodents, acute tolerance appears to contribute differentially to various behavioral measures of ethanol sensitivity, and acute tolerance development may offset some sensitivity measures (Crabbe et al., 2005). Strain-based differences in acute tolerance might also impact upon inebriometer measurements of rapid and chronic tolerance. The potential contribution of acute tolerance to our findings combined with the large number of ethanol mutants identified may suggest overlap between this form of tolerance and altered learning/memory.
Despite the lack of a significant overall correlation, a number of mutants displayed alterations in both sensitivity and rapid or chronic tolerance (Table 2; this number was not significantly greater than expected based on the frequencies with which either phenotype was observed alone—Fisher's exact test, p = 0.44). Supporting the notion of at least partial overlap in underlying mechanisms, homer mutant flies showed both increased sensitivity and reduced tolerance (Urizar et al., 2007). Likewise, among the 20 mutants in genes with ethanol-dependent altered transcription mentioned above, those with decreased sensitivity to ethanol most often also showed increased tolerance (Morozova et al., 2006). Conversely, here and in this group's study, there were a number of other mutants whose defects were specific to sensitivity or tolerance, indicating that these processes are partially distinct. Similarly, flies lacking synapsin showed impairments in multiple learning paradigms and no change in ethanol sensitivity, yet showed enhanced ethanol tolerance (Godenschwege et al., 2004).
In rodents, studies addressing the relationship between sensitivity and tolerance have yielded mixed results. Some studies have suggested that lines showing greater initial sensitivity also show greater tolerance (specifically, acute tolerance); however, outcomes have varied depending on the populations or genotypes compared and the behavioral assays used [e.g. (Crabbe et al., 2005); (Bennett et al., 2007) and references therein]. A recent study of inbred mouse lines suggested that rapid tolerance shared genetic determinants principally with acute tolerance, rather than with initial sensitivity (Radcliffe et al., 2006). The relationship between different forms of tolerance in rodents also remains fairly open, with some evidence to suggest rapid and chronic tolerance share at least partial mechanistic overlap (Kalant, 1996; Khanna et al., 1991). Despite the positive overall correlation seen here between levels of rapid and chronic tolerance, most of the individual mutants appeared defective specifically for one or the other, consistent with previous findings distinguishing these forms of tolerance genetically and mechanistically (Berger et al., 2004). The krasavietz mutant, an exception, was defective for both, implying that the gene affected may encode a component shared by both forms of tolerance. In addition to reduced tolerance, the krasavietz mutant showed decreased sensitivity. Thus, without specifying an underlying mechanism, the ethanol-naïve krasavietz flies might be “pretolerant,” i.e. altered by the mutation in baseline levels of a normal tolerance component. Conversely, a number of mutants exhibited both increased sensitivity and reduced tolerance: either rapid (ikar, milord-1, and zolotistuy), chronic (iks and rafael), or both (ruslan). Based on this, decreased sensitivity and reduced development of tolerance were not necessarily correlated, and in fact enhanced sensitivity to ethanol was more often associated with reduced tolerance [as for the homer mutant, mentioned above (Urizar et al., 2007)]. Lastly, the murashka-2 mutant showed the greatest reduction in initial sensitivity, yet no impairment in either form of tolerance; thus, a lower initial behavioral response did not necessarily correlate with impaired tolerance development. In sum, the relationship between ethanol sensitivity and tolerance appears complex, and elucidation of underlying mechanisms will require analysis of the roles of specific genes and identifying relevant brain regions.
Previous studies have made progress towards determining particular regions of the fly brain important for ethanol sensitivity (Rodan et al., 2002) and tolerance (Scholz et al., 2000). In the fly brain, the structures known as mushroom bodies (MBs) are of central importance for olfactory learning and memory [reviewed (Heisenberg, 2003)]. For many of the memory mutants, including several that showed altered ethanol sensitivity (8 of the 13, compared with 32 of 52 mutants total; represented at the same frequency), P-element enhancer trap expression was detected in the MB (Dubnau et al., 2003), as well as elsewhere in the adult brain (images available online at http://dart.cshl.edu/DART/public/scripts/browse_GEPlib.pl). However, a previous study indicated that the MB were dispensable for the regulation of ethanol sensitivity (Rodan et al., 2002), implying other brain regions are most likely involved. Acute inhibition of PKA signaling in specific fly brain regions implicated a relatively small number of neurons, including possible neurosecretory cells in the pars intercerebralis (PI), in regulation of ethanol sensitivity (Rodan et al., 2002); subsequently, the function of these PI cells in the release of insulin-like peptides was implicated in the behavioral response to ethanol (Corl et al., 2005). As recently described, the expression of another gene affecting ethanol sensitivity, white rabbit (whir), is widespread in the adult brain, including the MB, PI neurons, and parts of the central complex including the ellipsoid and fan-shaped bodies (Rothenfluh et al., 2006). Future investigations of other genes may further implicate these cells and regions in additional pathways regulating ethanol sensitivity; alternatively, different underlying mechanisms might involve additional brain regions.
In rapid tolerance, a previous analysis implicated the central brain region known as the ellipsoid body (EB), and in particular a subset of small field neurons connecting the EB with other structures of the central complex (Scholz et al., 2000). Analysis of the homer mutant, which showed increased sensitivity and reduced tolerance, implicated a small group of EB neurons in both phenotypes (Urizar et al., 2007). The hangover mutant, defective for a component of rapid ethanol tolerance that overlaps with other cell stress responses, encodes a nuclear zinc finger protein broadly expressed in the adult fly brain (Scholz et al., 2005). A potential role for the MB in rapid tolerance has neither conclusively been proved nor ruled out (Scholz et al., 2000). Of the mutants defective in tolerance analyzed here, P-element enhancer trap expression in the MB was detected for 7 of 8 of the reduced rapid tolerance mutants, but for only 3 of 9 chronic tolerance mutants (these numbers of rapid and chronic tolerance mutants showing MB expression were not significantly different from each other; p = 0.42, Fisher's exact test). A detailed analysis of the fly brain regions involved in chronic tolerance has not yet been performed.
While we undertook this study with the broad goal of examining the overlap between altered memory and ethanol behaviors, we were also interested in the identities of genes potentially affected in the mutants. From the 52 memory mutants we screened, some ethanol mutants were identified by multiple alleles. The 2 mutants valiet-1 and valiet-3 correspond to the same P-element insertion position (Dubnau et al., 2003) (in the gene RluA-1, discussed below), and exhibited very similar ethanol-related behaviors: both showed significant reductions in rapid tolerance and normal ethanol sensitivity (despite both being early candidates for decreased sensitivity, as described above). The mutants milord-1 and milord-2 showed increased sensitivity to ethanol, but only milord-1 showed reduced rapid tolerance. The corresponding pLacW insertions, while in the same gene at nearly the identical position (in the first large intron of the gene pumilio, discussed below), are in the opposite orientation (Dubnau et al., 2003 and our unpublished data). Similarly, both of the P-element insertions in the mutants murashka-1 and murashka-2 lie in the gene CG9381, encoding an uncharacterized RING finger domain protein, but only murashka-2 exhibited altered ethanol behavior (strongly reduced sensitivity). These 2 mutants showed different P-element enhancer trap expression (Dubnau et al., 2003), and the P-elements in them are not the same (pGawB in murashka-1 and pLacW in murashka-2), although the insertion sites are similar [in the 5′-end of CG9381, <100 bp apart; (Dubnau et al., 2003) and our unpublished data]. Likewise, barbos-1 and barbos-2 mutations correspond to different P-elements (pGawB, barbos-1; pLacW, barbos-2) in the same insertion site, lying between and upstream of 2 genes: CG6044, encoding a novel protein, and CG3624, encoding an otherwise uncharacterized immunoglobulin repeat protein (Dubnau et al., 2003; Table 1); only barbos-1 showed increased ethanol sensitivity. The memory mutants were characterized for ethanol sensitivity and tolerance after 5 generations of outcrossing to the wild-type genetic background strain (Materials and Methods), greatly increasing the likelihood that the P-elements in the mutants, rather than unlinked second-site mutations, are responsible for the altered ethanol phenotypes. Nevertheless, definitively linking the mutant ethanol phenotypes with specific genes requires more stringent genetic and/or molecular analysis. For this reason, in the absence of molecular or genetic evidence other than P-element location to link mutations with particular genes, we refer to mutants by their original names throughout (Dubnau et al., 2003).
With the above caveat, the identities of the genes potentially implicated in the sensitivity and tolerance mutants may suggest underlying pathways or mechanisms (Table 2). Altered regulation of actin dynamics has previously been associated with altered ethanol sedation in the whir mutant mentioned above [whir encodes a Rho-type GTPase that interacts genetically with Rho1 and Rac (Rothenfluh et al., 2006)], as well as the altered ethanol sensitivity of Eps8 mutants in both flies and mice (Eps8 encodes a regulator of actin dynamics implicated in modulating NMDA receptor responses to ethanol) (Offenhauser et al., 2006). A role for appropriate regulation of the actin cytoskeleton in ethanol sensitivity is further suggested by the genes implicated in 2 of the increased sensitivity mutants [cheerio, encoding a filamin homolog (Sokol and Cooley, 1999), in the mutant joy, and formin3, whose product promotes actin assembly (Tanaka et al., 2004), in the mutant zolotistuy].
Three other mutants displaying altered ethanol sensitivity potentially affect kinase signaling pathways [ikar—P-element just 5′ of Lk6, encoding a kinase homologous to mammalian Mnk1 and Mnk2 (Arquier et al., 2005; Reiling et al., 2005); iks, insertion in rhomboid, encoding a positive regulator of epidermal growth factor receptor-mediated signaling (Tsruya et al., 2007; Urban et al., 2002); and rafael, in which the closest gene is Dad, encoding a negative regulator of transforming growth factor-β (TGF-β) signaling (Tsuneizumi et al., 1997)]. Other mutants hint at roles for oxidative stress response (box, in which the corresponding P-element insertion lies in the 5′-end of Gclm) encoding a modifier subunit of glutamate-cysteine ligase involved in glutathione homeostasis (Fraser et al., 2003, 2002; Reed, 2004), and neuronal excitability [milord-1 and milord-2, in the gene pumilio (Dubnau et al., 2003; Murata and Wharton, 1995; Schweers et al., 2002; Weston and Baines, 2007)].
The P-element locations in other mutants may imply a role for RNA localization and/or translational regulation in rapid tolerance [krasavietz—P-element in exba, recently shown to play a role in axon guidance and to act as a translational repressor (Lee et al., 2007); milord-1—pumilio (discussed above); and valiet-1 and valiet-3, each corresponding to a P-element in RluA-1, encoding a pseudouridine synthase (isomerizes uridine to pseudouridine in tRNAs) capable of binding 3′-untranslated regions of a subset of mRNAs (Ofengand et al., 2001; Zhu et al., 2007)]. These mutants potentially represent specific links between ethanol tolerance, a form of behavioral plasticity in response to ethanol, and the targeted regulation of mRNA localization and translation at synapses, a recently intensively studied form of synaptic plasticity [reviewed (Schuman et al., 2006; Sutton and Schuman, 2006)]. While the krasavietz mutant showed both reduced rapid and chronic tolerance, the milord-1, valiet-1, and valiet-3 mutants were reduced specifically for rapid tolerance. This was somewhat surprising, given that chronic but not rapid tolerance has been shown to exhibit a dependence on new protein synthesis (Berger et al., 2004). An explanation is unclear; potentially, the milord-1 and valiet mutants might have developmental compensations masking a role in chronic tolerance. As far as other pathways or mechanisms suggested in chronic tolerance, the broadly defined category of signaling appeared again, represented by the mutants iks and rafael (potentially affecting the genes rhomboid and Dad, as described above), and baikal [insertion in paxillinB (pxb), a modulator of Hedgehog signaling (Inaki et al., 2002)]. More detailed studies of specific mutants will be required to elucidate the potential activities of gene products underlying the altered ethanol behaviors.
In summary, the high frequency with which strains showing one or more altered behavioral responses to ethanol were identified among the memory mutants further supports the notion of overlap between genes and mechanisms required for these processes. Additionally, the identities of the genes that may be affected will provide targets for more detailed future investigations.
Acknowledgments
The authors thank Doug Guarnieri for advice with inebriometer experiments and Chris Kliethermes for advice with statistical analysis and manuscript review. We are grateful to Vilma Roldan and Willa Abrone for preparing fly food. We thank Jody Barditch (CSHL, NY) for DNA sequence information corresponding to the rafael insertion site. We express our appreciation for reviewers' insightful comments and suggestions.
This work was supported by funds provided by the state of California for medical research on alcohol and substance abuse through the University of California, San Francisco (KHB, MSM, UH), and by the Department of the Army Grant DAMD 17-01-1-0798 (KHB, MSM).
References
- Arquier N, Bourouis M, Colombani J, Leopold P. Drosophila Lk6 kinase controls phosphorylation of eukaryotic translation initiation factor 4E and promotes normal growth and development. Curr Biol. 2005;15(1):19–23. doi: 10.1016/j.cub.2004.12.037. [DOI] [PubMed] [Google Scholar]
- Bennett B, Downing C, Carosone-Link P, Ponicsan H, Ruf C, Johnson TE. Quantitative trait locus mapping for acute functional tolerance to ethanol in the L × S recombinant inbred panel. Alcohol Clin Exp Res. 2007;31(2):200–208. doi: 10.1111/j.1530-0277.2006.00296.x. [DOI] [PubMed] [Google Scholar]
- Berger KH, Heberlein U, Moore MS. Rapid and chronic: two distinct forms of ethanol tolerance in Drosophila. Alcohol Clin Exp Res. 2004;28(10):1469–1480. doi: 10.1097/01.alc.0000141817.15993.98. [DOI] [PubMed] [Google Scholar]
- Bitran M, Kalant H. Effect of anisomycin on the development of rapid tolerance to ethanol-induced motor impairment. Pharmacol Biochem Behav. 1993;45(1):225–228. doi: 10.1016/0091-3057(93)90109-7. [DOI] [PubMed] [Google Scholar]
- Campbell JL, Nash HA. Volatile general anesthetics reveal a neurobiological role for the white and brown genes of Drosophila melanogaster. J Neurobiol. 2001;49(4):339–349. doi: 10.1002/neu.10009. [DOI] [PubMed] [Google Scholar]
- Chandler LJ, Harris RA, Crews FT. Ethanol tolerance and synaptic plasticity. Trends Pharmacol Sci. 1998;19(12):491–495. doi: 10.1016/s0165-6147(98)01268-1. [DOI] [PubMed] [Google Scholar]
- Cheng Y, Endo K, Wu K, Rodan AR, Heberlein U, Davis RL. Drosophila fasciclinII is required for the formation of odor memories and for normal sensitivity to alcohol. Cell. 2001;105:757–768. doi: 10.1016/s0092-8674(01)00386-5. [DOI] [PubMed] [Google Scholar]
- Corl AB, Rodan AR, Heberlein U. Insulin signaling in the nervous system regulates ethanol intoxication in Drosophila melanogaster. Nat Neurosci. 2005;8(1):18–19. doi: 10.1038/nn1363. [DOI] [PubMed] [Google Scholar]
- Crabbe JC, Metten P, Cameron AJ, Wahlsten D. An analysis of the genetics of alcohol intoxication in inbred mice. Neurosci Biobehav Rev. 2005;28(8):785–802. doi: 10.1016/j.neubiorev.2004.08.002. [DOI] [PubMed] [Google Scholar]
- Crabbe JC, Phillips TJ, Harris RA, Arends MA, Koob GF. Alcohol-related genes: contributions from studies with genetically engineered mice. Addict Biol. 2006;11(3-4):195–269. doi: 10.1111/j.1369-1600.2006.00038.x. [DOI] [PubMed] [Google Scholar]
- Crabbe JC, Rigter H, Uijlen J, Strijbos C. Rapid development of tolerance to the hypothermic effect of ethanol in mice. J Pharmacol Exp Ther. 1979;208(1):128–133. [PubMed] [Google Scholar]
- Dubnau J, Chiang AS, Grady L, Barditch J, Gossweiler S, McNeil J, Smith P, Buldoc F, Scott R, Certa U, Broger C, Tully T. The staufen/pumilio pathway is involved in Drosophila long-term memory. Curr Biol. 2003;13(4):286–296. doi: 10.1016/s0960-9822(03)00064-2. [DOI] [PubMed] [Google Scholar]
- Dzitoyeva S, Dimitrijevic N, Manev H. Gamma-aminobutyric acid B receptor 1 mediates behavior-impairing actions of alcohol in Drosophila: adult RNA interference and pharmacological evidence. Proc Natl Acad Sci USA. 2003;100(9):5485–5490. doi: 10.1073/pnas.0830111100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fadda F, Rossetti ZL. Chronic ethanol consumption: from neuroadaptation to neurodegeneration. Prog Neurobiol. 1998;56:385–431. doi: 10.1016/s0301-0082(98)00032-x. [DOI] [PubMed] [Google Scholar]
- Feany MB, Quinn WG. A neuropeptide gene defined by the Drosophila memory mutant amnesiac. Science. 1995;268(5212):869–873. doi: 10.1126/science.7754370. [DOI] [PubMed] [Google Scholar]
- Fraser JA, Kansagra P, Kotecki C, Saunders RD, McLellan LI. The modifier subunit of Drosophila glutamate-cysteine ligase regulates catalytic activity by covalent and noncovalent interactions and influences glutathione homeostasis in vivo. J Biol Chem. 2003;278(47):46369–46377. doi: 10.1074/jbc.M308035200. [DOI] [PubMed] [Google Scholar]
- Fraser JA, Saunders RD, McLellan LI. Drosophila melanogaster glutamate-cysteine ligase activity is regulated by a modifier subunit with a mechanism of action similar to that of the mammalian form. J Biol Chem. 2002;277(2):1158–1165. doi: 10.1074/jbc.M106683200. [DOI] [PubMed] [Google Scholar]
- Godenschwege TA, Reisch D, Diegelmann S, Eberle K, Funk N, Heisenberg M, Hoppe V, Hoppe J, Klagges BRE, Martin JR, Nikitina EA, Putz G, Reifegerste R, Reisch N, Rister J, Schaupp M, Scholz H, Schwarzel M, Werner U, Zars TD, Buchner S, Buchner E. Flies lacking all synapsins are unexpectedly healthy but are impaired in complex behaviour. Eur J Neurosci. 2004;20(3):611–622. doi: 10.1111/j.1460-9568.2004.03527.x. [DOI] [PubMed] [Google Scholar]
- Heath AC, Madden PA, Bucholz KK, Dinwiddie SH, Slutske WS, Bierut LJ, Rohrbaugh JW, Statham DJ, Dunne MP, Whitfield JB, Martin NG. Genetic differences in alcohol sensitivity and the inheritance of alcoholism risk. Psychol Med. 1999;29(5):1069–1081. doi: 10.1017/s0033291799008909. [DOI] [PubMed] [Google Scholar]
- Heberlein U, Wolf FW, Rothenfluh A, Guarnieri DJ. Molecular genetic analysis of ethanol intoxication in Drosophila melanogaster. Integr Comp Biol. 2004;44:269–274. doi: 10.1093/icb/44.4.269. [DOI] [PubMed] [Google Scholar]
- Heisenberg M. Mushroom body memoir: from maps to models. Nat Rev Neurosci. 2003;4(4):266–275. doi: 10.1038/nrn1074. [DOI] [PubMed] [Google Scholar]
- Hyman SE, Malenka RC, Nestler EJ. Neural mechanisms of addiction: the role of reward-related learning and memory. Annu Rev Neurosci. 2006;29:565–598. doi: 10.1146/annurev.neuro.29.051605.113009. [DOI] [PubMed] [Google Scholar]
- Inaki M, Kojima T, Ueda R, Saigo K. Requirements of high levels of Hedgehog signaling activity for medial-region cell fate determination in Drosophila legs: identification of pxb, a putative Hedgehog signaling attenuator gene repressed along the anterior-posterior compartment boundary. Mech Dev. 2002;116(1-2):3–18. doi: 10.1016/s0925-4773(02)00119-3. [DOI] [PubMed] [Google Scholar]
- Kalant H. Current state of knowledge about the mechanisms of alcohol tolerance. Addict Biol. 1996;1:133–141. doi: 10.1080/1355621961000124756. [DOI] [PubMed] [Google Scholar]
- Kalant H, LeBlanc AE, Gibbins RJ. Tolerance to, and dependence on, some non-opiate psychotropic drugs. Pharmacol Rev. 1971;23(3):135–191. [PubMed] [Google Scholar]
- Khanna JM, Kalant H, Shah G, Weiner J. Rapid tolerance as an index of chronic tolerance. Pharmacol Biochem Behav. 1991;38(2):427–432. doi: 10.1016/0091-3057(91)90302-i. [DOI] [PubMed] [Google Scholar]
- Khanna JM, Morato GS, Chau A, Shah G, Kalant H. Effect of NMDA antagonists on rapid and chronic tolerance to ethanol: importance of intoxicated practice. Pharmacol Biochem Behav. 1994;48(3):755–763. doi: 10.1016/0091-3057(94)90343-3. [DOI] [PubMed] [Google Scholar]
- Lane ME, Kalderon D. Genetic investigation of cAMP-dependent protein kinase function in Drosophila development. Genes Dev. 1993;7(7A):1229–1243. doi: 10.1101/gad.7.7a.1229. [DOI] [PubMed] [Google Scholar]
- Le AD, Kalant H. Influence of intoxicated practice on the development of acute tolerance to the motor impairment effect of ethanol. Psychopharmacology (Berl) 1992;106(4):572–576. doi: 10.1007/BF02244833. [DOI] [PubMed] [Google Scholar]
- Lee S, Nahm M, Lee M, Kwon M, Kim E, Zadeh AD, Cao H, Kim HJ, Lee ZH, Oh SB, Yim J, Kolodziej PA, Lee S. The F-actin-microtubule crosslinker shot is a platform for Krasavietz-mediated translational regulation of midline axon repulsion. Development. 2007;134(9):1767–1777. doi: 10.1242/dev.02842. [DOI] [PubMed] [Google Scholar]
- Levin LR, Han PL, Hwang PM, Feinstein PG, Davis RL, Reed RR. The Drosophila learning and memory gene rutabaga encodes a Ca2+/calmodulin-responsive adenylyl cyclase. Cell. 1992;68(3):479–489. doi: 10.1016/0092-8674(92)90185-f. [DOI] [PubMed] [Google Scholar]
- Livingstone MS, Sziber PP, Quinn WG. Loss of calcium/calmodulin responsiveness in adenylate cyclase of rutabaga, a Drosophila learning mutant. Cell. 1984;37(1):205–215. doi: 10.1016/0092-8674(84)90316-7. [DOI] [PubMed] [Google Scholar]
- Moore MS, DeZazzo J, Luk AY, Tully T, Singh CM, Heberlein U. Ethanol intoxication in Drosophila: genetic and pharmacological evidence for regulation by the cAMP signaling pathway. Cell. 1998;93:997–1007. doi: 10.1016/s0092-8674(00)81205-2. [DOI] [PubMed] [Google Scholar]
- Morozova TV, Anholt RRH, Mackay TFC. Transcriptional response to alcohol exposure in Drosophila melanogaster. Genome Biol. 2006;7(10):R95. doi: 10.1186/gb-2006-7-10-r95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murata Y, Wharton RP. Binding of pumilio to maternal hunchback mRNA is required for posterior patterning in Drosophila embryos. Cell. 1995;80(5):747–756. doi: 10.1016/0092-8674(95)90353-4. [DOI] [PubMed] [Google Scholar]
- Nestler EJ. Common molecular and cellular substrates of addiction and memory. Neurobiol Learn Mem. 2002;78:637–647. doi: 10.1006/nlme.2002.4084. [DOI] [PubMed] [Google Scholar]
- Ofengand J, Malhotra A, Remme J, Gutgsell NS, Del Campo M, Jean-Charles S, Peil L, Kaya Y. Pseudouridines and pseudouridine synthases of the ribosome. Cold Spring Harb Symp Quant Biol. 2001;66(1):147–160. doi: 10.1101/sqb.2001.66.147. [DOI] [PubMed] [Google Scholar]
- Offenhauser N, Castelletti D, Mapelli L, Soppo BE, Regondi MC, Rossi P, D'Angelo E, Frassoni C, Amadeo A, Tocchetti A, Pozzi B, Disanza A, Guarnieri D, Betsholtz C, Scita G, Heberlein U, Di Fiore PP. Increased ethanol resistance and consumption in Eps8 knockout mice correlates with altered actin dynamics. Cell. 2006;127(1):213–226. doi: 10.1016/j.cell.2006.09.011. [DOI] [PubMed] [Google Scholar]
- Park SK, Sedore SA, Cronmiller C, Hirsh J. Type II cAMP-dependent protein kinase-deficient Drosophila are viable but show developmental, circadian, and drug response phenotypes. J Biol Chem. 2000;275(27):20588–20596. doi: 10.1074/jbc.M002460200. [DOI] [PubMed] [Google Scholar]
- Radcliffe RA, Floyd KL, Lee MJ. Rapid ethanol tolerance mediated by adaptations in acute tolerance in inbred mouse strains. Pharmacol Biochem Behav. 2006;84(3):524–534. doi: 10.1016/j.pbb.2006.06.018. [DOI] [PubMed] [Google Scholar]
- Reed DJ. Mitochondrial glutathione and chemically induced stress including ethanol. Drug Metab Rev. 2004;36(3-4):569–582. doi: 10.1081/dmr-200033449. [DOI] [PubMed] [Google Scholar]
- Reiling JH, Doepfner KT, Hafen E, Stocker H. Diet-dependent effects of the Drosophila Mnk1/Mnk2 homolog Lk6 on growth via eIF4E. Curr Biol. 2005;15(1):24–30. doi: 10.1016/j.cub.2004.12.034. [DOI] [PubMed] [Google Scholar]
- Rodan AR, Kiger JA, Jr, Heberlein U. Functional dissection of neuroanatomical loci regulating ethanol sensitivity in Drosophila. J Neurosci. 2002;22(21):9490–9501. doi: 10.1523/JNEUROSCI.22-21-09490.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothenfluh A, Threlkeld RJ, Bainton RJ, Tsai LT, Lasek AW, Heberlein U. Distinct behavioral responses to ethanol are regulated by alternate RhoGAP18B isoforms. Cell. 2006;127(1):199–211. doi: 10.1016/j.cell.2006.09.010. [DOI] [PubMed] [Google Scholar]
- Scholz H, Franz M, Heberlein U. The hangover gene defines a stress pathway required for ethanol tolerance development. Nature. 2005;436(7052):845–847. doi: 10.1038/nature03864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scholz H, Ramond J, Singh CM, Heberlein U. Functional ethanol tolerance in Drosophila. Neuron. 2000;28:261–271. doi: 10.1016/s0896-6273(00)00101-x. [DOI] [PubMed] [Google Scholar]
- Schuckit MA. Low level of response to alcohol as a predictor of future alcoholism. Am J Psychiatry. 1994;151(2):184–189. doi: 10.1176/ajp.151.2.184. [DOI] [PubMed] [Google Scholar]
- Schuckit MA, Smith TL. An 8-year follow-up of 450 sons of alcoholic and control subjects. Arch Gen Psychiatry. 1996;53(3):202–210. doi: 10.1001/archpsyc.1996.01830030020005. [DOI] [PubMed] [Google Scholar]
- Schuckit MA, Smith TL, Kalmijn J. The search for genes contributing to the low level of response to alcohol: patterns of findings across studies. Alcohol Clin Exp Res. 2004;28(10):1449–1458. doi: 10.1097/01.alc.0000141637.01925.f6. [DOI] [PubMed] [Google Scholar]
- Schuman EM, Dynes JL, Steward O. Synaptic regulation of translation of dendritic mRNAs. J Neurosci. 2006;26(27):7143–7146. doi: 10.1523/JNEUROSCI.1796-06.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schweers BA, Walters KJ, Stern M. The Drosophila melanogaster translational repressor pumilio regulates neuronal excitability. Genetics. 2002;161(3):1177–1185. doi: 10.1093/genetics/161.3.1177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh CM, Heberlein U. Genetic control of acute ethanol-induced behaviors in Drosophila. Alcohol Clin Exp Res. 2000;24(8):1127–1136. [PubMed] [Google Scholar]
- Sokol NS, Cooley L. Drosophila filamin encoded by the cheerio locus is a component of ovarian ring canals. Curr Biol. 1999;9(21):1221–1230. doi: 10.1016/s0960-9822(99)80502-8. [DOI] [PubMed] [Google Scholar]
- Sutton MA, Schuman EM. Dendritic protein synthesis, synaptic plasticity, and memory. Cell. 2006;127(1):49–58. doi: 10.1016/j.cell.2006.09.014. [DOI] [PubMed] [Google Scholar]
- Tabakoff B, Cornell N, Hoffman PL. Alcohol tolerance. Ann Emerg Med. 1986;15(9):1005–1012. doi: 10.1016/s0196-0644(86)80119-6. [DOI] [PubMed] [Google Scholar]
- Tanaka H, Takasu E, Aigaki T, Kato K, Hayashi S, Nose A. Formin3 is required for assembly of the F-actin structure that mediates tracheal fusion in Drosophila. Dev Biol. 2004;274(2):413–425. doi: 10.1016/j.ydbio.2004.07.035. [DOI] [PubMed] [Google Scholar]
- Thiele TE, Willis B, Stadler J, Reynolds JG, Bernstein IL, McKnight GS. High ethanol consumption and low sensitivity to ethanol-induced sedation in protein kinase A-mutant mice. J Neurosci. 2000;20(10):RC75. doi: 10.1523/JNEUROSCI.20-10-j0003.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsruya R, Wojtalla A, Carmon S, Yogev S, Reich A, Bibi E, Merdes G, Schejter E, Shilo BZ. Rhomboid cleaves Star to regulate the levels of secreted Spitz. EMBO J. 2007;26(5):1211–1220. doi: 10.1038/sj.emboj.7601581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuneizumi K, Nakayama T, Kamoshida Y, Kornberg TB, Christian JL, Tabata T. Daughters against dpp modulates dpp organizing activity in Drosophila wing development. Nature. 1997;389(6651):627–631. doi: 10.1038/39362. [DOI] [PubMed] [Google Scholar]
- Tully T, Preat T, Boynton SC, Del Vecchio M. Genetic dissection of consolidated memory in Drosophila. Cell. 1994;79(1):35–47. doi: 10.1016/0092-8674(94)90398-0. [DOI] [PubMed] [Google Scholar]
- Urban S, Lee JR, Freeman M. A family of Rhomboid intramembrane proteases activates all Drosophila membrane-tethered EGF ligands. EMBO J. 2002;21(16):4277–4286. doi: 10.1093/emboj/cdf434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urizar NL, Yang Z, Edenberg HJ, Davis RL. Drosophila homer is required in a small set of neurons including the ellipsoid body for normal ethanol sensitivity and tolerance. J Neurosci. 2007;27(17):4541–4551. doi: 10.1523/JNEUROSCI.0305-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wand G, Levine M, Zweifel L, Schwindinger W, Abel T. The cAMP-protein kinase A signal transduction pathway modulates ethanol consumption and sedative effects of ethanol. J Neurosci. 2001;21(14):5297–5303. doi: 10.1523/JNEUROSCI.21-14-05297.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber KE. An apparatus for measurement of resistance to gas-phase reagents. Drosoph Inf Serv. 1988;67:91–93. [Google Scholar]
- Weber KE, Diggins LT. Increased selection response in larger populations. II. Selection for ethanol vapor resistance in Drosophila melanogaster at two population sizes. Genetics. 1990;125(3):585–597. doi: 10.1093/genetics/125.3.585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen T, Parrish CA, Xu D, Wu Q, Shen P. Drosophila neuropeptide F and its receptor, NPFR1, define a signaling pathway that acutely modulates alcohol sensitivity. Proc Natl Acad Sci USA. 2005;102(6):2141–2146. doi: 10.1073/pnas.0406814102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weston AJ, Baines RA. Translational regulation of neuronal electrical properties. Invert Neurosci. 2007;7(2):75–86. doi: 10.1007/s10158-006-0037-8. [DOI] [PubMed] [Google Scholar]
- Zaleski MJ, Nunes Filho JR, Lemos T, Morato GS. GABA(B) receptors play a role in the development of tolerance to ethanol in mice. Psychopharmacology (Berl) 2001;153(4):415–424. doi: 10.1007/s002130000581. [DOI] [PubMed] [Google Scholar]
- Zhu J, Gopinath K, Murali A, Yi G, Hayward SD, Zhu H, Kao C. RNA-binding proteins that inhibit RNA virus infection. Proc Natl Acad Sci USA. 2007;104(9):3129–3134. doi: 10.1073/pnas.0611617104. [DOI] [PMC free article] [PubMed] [Google Scholar]


