Abstract
Introduction
This paper describes the design, implementation, and potential use of a comparative anatomy information system (CAIS) for querying on similarities and differences between homologous anatomical structures across species, the knowledge base it operates upon, the method it uses for determining the answers to the queries, and the user interface it employs to present the results. The relevant informatics contributions of our work include (1) the development and application of the structural difference method, a formalism for symbolically representing anatomical similarities and differences across species; (2) the design of the structure of a mapping between the anatomical models of two different species and its application to information about specific structures in humans, mice, and rats; and (3) the design of the internal syntax and semantics of the query language. These contributions provide the foundation for the development of a working system that allows users to submit queries about the similarities and differences between mouse, rat, and human anatomy; delivers result sets that describe those similarities and differences in symbolic terms; and serves as a prototype for the extension of the knowledge base to any number of species. Additionally, we expanded the domain knowledge by identifying medically relevant structural questions for the human, the mouse, and the rat, and made an initial foray into the validation of the application and its content by means of user questionnaires, software testing, and other feedback.
Methods
The anatomical structures of the species to be compared, as well as the mappings between species, are modeled on templates from the Foundational Model of Anatomy knowledge base, and compared using graph-matching techniques. A graphical user interface allows users to issue queries that retrieve information concerning similarities and differences between structures in the species being examined. Queries from diverse information sources, including domain experts, peer-reviewed articles, and reference books, have been used to test the system and to illustrate its potential use in comparative anatomy studies.
Results
157 test queries were submitted to the CAIS system, and all of them were correctly answered. The interface was evaluated in terms of clarity and ease of use. This testing determined that the application works well, and is fairly intuitive to use, but users want to see more clarification of the meaning of the different types of possible queries. Some of the interface issues will naturally be resolved as we refine our conceptual model to deal with partial and complex homologies in the content.
Conclusions
The CAIS system and its associated methods are expected to be useful to biologists and translational medicine researchers. Possible applications range from supporting theoretical work in clarifying and modeling ontogenetic, physiological, pathological, and evolutionary transformations, to concrete techniques for improving the analysis of genotype–phenotype relationships among various animal models in support of a wide array of clinical and scientific initiatives.
Keywords: Ontology, Anatomy, Comparative anatomy, Knowledge base, Protégé, Homology, Graph matching, Graph similarity, Isomorphism, Foundational Model of Anatomy
1. Introduction
The amount of anatomical and associated medical information emerging from animal modeling in comparative medicine (the study of health, disease, and treatment in one species through comparison with similar conditions in other model species) and comparative genomics (the study of the genome in one species through comparison with the genomes in other model species and their evolutionary relationships) is increasing rapidly [1,2], and consequently, innovative techniques for evaluating, organizing, and managing that information for researchers and clinicians are called for. The increasing need for extrapolating information from one species to another has been highlighted by contemporary research in bioinformatics, genomics, proteomics, and animal models of human disease, as well as other fields [3]. Additionally, the urgency of finding ways to organize and manage the volume of data has been remarked upon by many observers, especially in light of the identification and characterization of genomic sequences across species [4]. Information systems have been and continue to be an important tool in this task.
At the same time that the amount of information generated is increasing so rapidly, traditional barriers between scientific domains are being blurred. As medical research becomes more interdisciplinary, researchers from traditional biomedical disciplines (e.g., anatomy, embryology) join forces with scientists from newer disciplines (e.g., molecular biology, genomics) and clinicians in the attempt to translate discoveries from bench science into clinical applications that can realize effective treatments for patients. Accordingly, the audience for information has expanded to include, among others, patients and policy makers [5]. Information systems dealing with this type of data must be flexible enough to accommodate the various needs of these different groups of users. Therefore, in addition to rigorous attention to the quality of the anatomical information involved, such a system must be flexible and extensible enough to accommodate different information views, depending on the needs of the user, whether a bench scientist, a clinician, a student, or a patient.
We have developed a cross-species model that provides a formalized ontological framework for the analysis of structural phenotype comparison, as well as application of the model’s foundational principles to real-life queries on animal models. Such a model will support formal reasoning about the comparisons of structural phenotypes involved [6] and provides a structure on which the quantity of information involved can be organized. The possibility of establishing and validating structural correspondences between different structural phenotypes has tremendous potential for addressing both issues, thereby improving the quality, management, and dissemination of information about animal models of human disease and genomics. While phenotypes of traits based on physiological processes are outside the scope of this application, the principles underlying their comparison remain the same, and we anticipate that the methods of defining and comparing phenotypes used for structure will be extensible to physiological and pathological phenotypes as well.
Some preliminary work in these areas has already been carried out. Cook et al. have associated qualitative and quantitative values with spatial and non-spatial physical properties of anatomical entities. This association has permitted them to instantiate instances of the canonical Foundational Model of Anatomy (FMA) and the related Foundational Model of Physiology (FMP) in order to create physiological simulations [7]. In response to the need of the Virtual Soldier Project for reasoning about traumatic injuries and prediction of their outcomes, Rosse et al. developed the Ontology of Biomedical Reality (OBR), which supports the representation of variant anatomical structures in addition to canonical ones [8]. Smith et al. extended OBR to support reasoning about carcinomas as representative pathological entities [9]. These initial efforts appear to provisionally support our hypothesis that CAIS may be similarly expanded to variations in physiologic and pathologic phenotypes, as well as to indicate directions in which further research in this area may be pursued.
A comparative anatomy information system is a computer system that allows users to compare canonical phenotypes of corresponding anatomical structures across medically relevant species at varying levels of granularity and detail and returns responses to queries about those comparisons. The need for such a system arises out of the importance of animal models in comparative medicine and genomics, as well as out of the explosion in th quantity of data to be managed. We have developed an information system that is an initial attempt to address some of the informatics issues involved in meeting that need.
The primary subject matter for our comparative anatomy information system, CAIS, consists of a subset of the cancer sites identified by the Mouse Models of Human Cancer Consortium (MMHCC) as medically important. Due to their importance to cancer researchers and clinicians, their structural complexity, and their specific similarities and differences with human structures, we selected five of these sites (prostate, breast/mammary gland, lung, ovary, and cervix) to model. We built on our foundational work in rodent mammary gland and prostate symbolic model development and comparison [3], to continue development of rodent anatomical models, including leveraging the work on mouse structures as templates for the corresponding rat structures. Our research design involved close collaboration with colleagues in biological structure and structural informatics, computer science, and comparative vertebrate embryology. These colleagues contributed domain content, assisted in development of the system, and evaluated its usefulness and accuracy.
This paper describes the design, implementation, and potential use of CAIS. The system is based on the structural difference method (SDM) formalism for symbolically representing the similarities and differences between homologous anatomical structures across dif ferent species [3]. The anatomical structures of the species to be compared, as well as the mappings between species, are modeled on templates from the FMA knowledge base, and implemented using frames in the Protégé-2000 ontology and knowledge-base editor [10]. A graphical user interface (GUI) allows users to issue queries that retrieve information concerning the similarities and differences between the species being examined. Queries from diverse information sources, including domain experts, peerreviewed articles, and reference books, have been used to test the system and to illustrate its potential use in comparative anatomy studies.
2. Background
This research is concerned with the design and implementation of CAIS. The work spans several fields including knowledge representation, information systems, and graph-matching algorithms, as well as symbolic modeling of humans and other species. Since the modeling part of our work is mainly directed at mouse anatomy, we will first discuss related work on mouse modeling and related databases. Next we will discuss the FMA, which is an integral part of our work. We will then describe some related works on graph matching and model management, both of which lead into our own structural difference methodology.
2.1. Mouse modeling and databases
Determining genotype–phenotype correlations is the basis for creating integrated systems for biological applications [11], and integrating diverse types of model organism data is crucial to the usefulness of these efforts [12]. The development of phenotypic standards – an essential component of rationalizing these correlations – is an area where CAIS has the potential to make a solid contribution. Some smaller, more specific efforts have already been undertaken along these lines.
Bao et al. have integrated behavioral and neurological genotype–phenotype relations from the Mammalian Phenotype Ontology Database curated by the Jackson Laboratory [13]. MitoP2, the mitochondrial proteome database, now contains data for mice as well as humans and yeast [14]. At the single nucleotide polymorphism (SNP) level, Agrafioti and Stumpf collected mouse, dog, rat, and chicken SNPs, as well as all inferrable human ones [15]. A database of mouse mutant strains that affect biological responses to DNA damage has been developed at the University of Texas-Southwestern [16].
The Mouse Phenome Database at the Jackson Laboratory contains data on the phenotypes and genotypes of commonly used strains of experimental mice [17,18], and the Jackson Laboratory’s Mouse Genome Database incorporates the Gene Ontology (GO), the Mammalian Phenotype Ontology, and the Anatomical Dictionary for Mouse Development and the Adult Anatomy [19–22]. Similarly, the Rat Genome Database at the Medical College of Wisconsin contains annotations for a phenotype ontology [23]. Lussier has addressed the challenge of the volume of data by using natural-language processing and data mining in order to semi-automatically assign a phenotypic context (PhenoGO) to the gene ontology annotations [24].
In addition to the numerous mouse genome resources available, there is also a large body of work on the representation of mouse anatomy. One of the most significant resources available is the Adult Mouse Anatomical Dictionary, from the Jackson Laboratory [25]. Despite the term “Dictionary” in the name, it is actually an ontology, organizing anatomical structures for postnatal mice by is-a and part-of relationships. The Adult Mouse Anatomical Dictionary is intended to integrate biological data of various types, including gene expression and phenotype data, and to this end, Hayamizu et al. argue – as we do – that anatomy is essential as the foundation for integrating these various types of processes and phenotypic observations [25].
Besides the adult mouse ontology, the Jackson Laboratory collaborates on a larger project, the Mouse Anatomical Dictionary Browser [26], with the Edinburgh Mouse Atlas Project (EMAP) [27]. EMAP develops the anatomical ontologies for the embryonic stages of the mouse; it and the Adult Mouse Anatomical Dictionary are components of the larger composite Mouse Anatomical Dictionary Browser.
A web-based resource for the visualization, searching and downloading of standard operating procedures and other documents, the European Mouse Phenotyping Resource for Standardized Screens (EMPReSS) has been developed by the Mammalian Genetics Unit at Oxford [28], and the German Mouse Clinic is an open-access platform for standardized phenotyping [29]. Pathbase is a database that stores images of the abnormal histology associated with spontaneous and induced mutations of both embryonic and adult mice, including those produced by transgenesis, targeted mutagenesis, and chemical mutagenesis [30].
The systems described above were developed to meet specific needs of researchers working with models within a species, although some of them have taken first steps to including different species. Additionally, while work at the molecular biology level is well-represented, and some steps have been taken to address disease phenotypes, there is currently no systematic basis for classifying normal anatomical phenotypes as a reference. In order to meet the larger goal of correlating genotype and phenotype across multiple different species, these systems still need detailed specifications about what the canonical phenotypes for different species are, and a normalization or correlation of the relevant terminologies.
CAIS has the potential to meet both needs. By systematically categorizing anatomical morphology in a manner that is species-independent, CAIS provides a generalized mechanism which makes possible phenotypic comparisons between any two species at a time, opening the door to multiple comparisons in an additive fashion. Its emphasis on biological realism and on entities, rather than concepts, provides a mechanism for solving thorny terminological challenges and confounds resulting from the separate and parallel histories of anatomical observation in different species. The phenotypic classifications generated by CAIS can be further developed with cladistic analysis (objective, quantitative analysis of phenotypic traits of organisms based on phylogenetic relationships, established by DNA and RNA sequencing) to approach the genotypic end of the phenotype–genotype correlation. Finally, the ability to export the CAIS knowledge base in XML contributes to the necessary interoperability to synthesize data from heterogeneous datasets [31] by syntactically supporting the exchange of data across those datasets, in order to provide new juxtapositions and visualizations of the data for hypothesis generation and discovery and to come closer to the goal of the Human Phenome Project [32].
The biology community is moving toward the ontology library known as Open Biomedical Ontologies (OBO) as a de facto standard. Two of the OBO initiatives relevant to CAIS are the Common Anatomy Reference Ontology (CARO) and PATO: An Ontology of Phenotypic Qualities. CARO’s purpose is to provide standards and templates for anatomical ontologies for different species in the interest of interoperability [33]. Ontologies are under development for a wide range of medically important organisms that differ drastically in anatomy, including mouse, fly, tick and mosquito [34], zebrafish [35], and amphibians [36], among others. PATO supports the annotation of phenotypes over a variety of different applications, and is independent of any exchange format or database schema [37]. It permits the composition of single fundamental phenotype units from the ontology into larger units descriptive of phenotypes on a larger scale.
While CARO and PATO are developing standards for principled modeling of future ontologies, it is also the case that ontology development is running ahead of those standards, and many ontologies based on differing or contradictory underlying models are already in use. Many of those ontologies have an established user community, and need to be maintained as legacy applications for that user base, even though they are not in compliance with the OBO standards. An example is GALEN, and Mork, Pottinger, and Bernstein have documented the intensiveness and error rate of aligning GALEN and the FMA [38,39].
In response to the increasing importance of ontology alignment caused by the number of differing medical ontologies being developed, Stuckenschmidt et al. extended the semantics of the Web Ontology Language (OWL) [40]. Their extension of OWL, C-OWL, permits semantic alignment of incompatible ontologies, as well as reasoning about the mappings between those ontologies [40].
By utilizing holes and bridge rules, C-OWL permits two ontologies to be mapped to each other, even if those ontologies represent two mutually contradictory models; they define a mapping between two ontologies as a set of bridge rules between the ontologies [40]. The bridge rules that they define are more-general, more-specific, equivalent, disjoint, and overlapping. These are very similar to the operations we have implemented in CAIS: shared, not-shared, union, is-different?, and is-homologous? As a result, the underlying CAIS conceptual model of types of anatomical transformation across species will translate into C-OWL relatively straightforwardly when we move from the current frame-based representation to a DL-based one.
2.2. The Foundational Model of Anatomy (FMA)
The FMA is a symbolic model of the physical organization of the human body. More specifically, it is an ontology that furnishes a comprehensive set of entities and relationships that describe the human body at all levels of structural organization. At the highest level of abstraction, it consists of the following components: (1) the Anatomical Taxonomy (AT); (2) the Anatomical Structural Abstraction (ASA); (3) the Anatomical Transformation Abstraction (ATA); and (4) the Foundational Model Metaknowledge (Mk)[41,42].
The AT component is a type hierarchy of entities that describes the body at levels of organization from organism down through organ and cell to macromolecule, based on the is-a relationship [42]. Extending it to the mouse involves ascertaining the important entities and terms involved. The AT’s emphasis on entities, rather than terminology, serves us well when deciding what structures to correlate. The ASA component serves to describe the shape, connections, boundaries, location, and orientation of the structures under study, as well as describing units of organization in terms of their component parts. This is where many of the medically important differences in the structures we are studying will be found.
The ATA spells out the “relationships that describe the morphological transformation of anatomical entities during prenatal and postnatal development” [42]. It has not been fully developed and will not be used in our comparisons of species. Mk includes the rules, principles, and axioms underlying the anatomical knowledge it represents. Metaknowledge is used only implicitly in our work.
The FMA was originally developed to represent human anatomy. However, the inclusion in the FMA of high-level abstract classes, such as Organ component and Systemic arterial tree, enables the extension of the FMA to non-human species and the resulting ability to compare corresponding structures across species. Additionally, the FMA’s emphasis on entities rather than on terms permits resolution of inconsistent terminology that has hindered other comparative anatomy systems. Terminology problems such as “ventral” and “anterior” being synonyms in humans but not in other vertebrates, or “anterior prostate” meaning an organ region in humans and a discrete organ in rodents, are handled by associating the various terms with the entities they refer to in slots for preferred, alternate, and deprecated names.
In developing hierarchies for the mouse prostate and mammary gland, we extended the existing human FMA to create mouse organ templates; we then used those templates to map structures at levels of organization from the organ down to the cell, in order to determine where the similarities and differences lie. Additionally, because the mouse anatomical symbolic model is based on the FMA, our comparisons will have to deal with differences between the structures themselves at various levels of organization, but will not need to deal with model or meta-model conflicts.
There are several different interfaces to the FMA. Since it is implemented with the Protégé knowledge-representation system, FMA developers use Protégé’s own interface. In order to make the FMA more accessible to end users, two additional interfaces are available. The Foundational Model Explorer (FME) [43] allows users to view one object class at a time. When a class is viewed, all of its attributes and relationships to other entities are shown. Users can select these other entities for more information, but only one at a time is visible. In contrast, the Emily query interface [44,45] focuses on supporting queries over the relationships among anatomical entities. It allows users to search for entities that are in a given spatial relationship to a selected one, or to find the relationships between two given entities. We will build on previous work on Emily as a basis for our query engine.
2.3. Graph matching
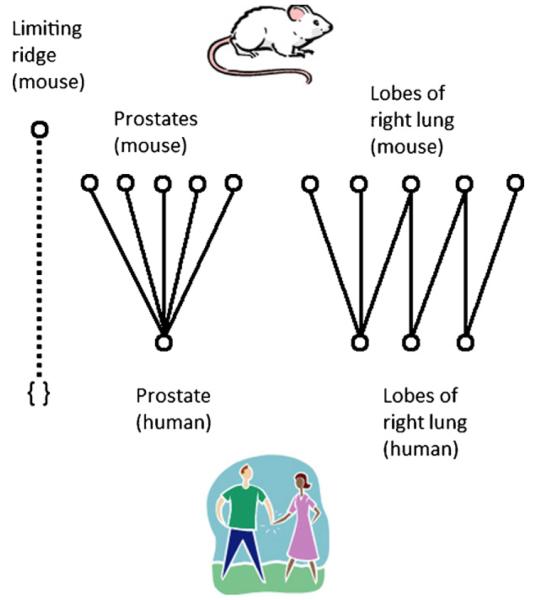
In this section we will introduce the graph-matching frame-work that is used in our work and begin a sequence of examples from anatomy that explain our comparative methodology, which is defined in Section 3. While a running example using the same anatomical structures for our illustrations would be ideal, such an example was not possible, since none of the MMHCC structures under study displayed the full range of similarities and differences. To aid the reader in following the change in anatomical structures from one figure to another, we have developed the following guidelines for orientation (as shown in Fig. 1): rodent structures are always at the top of the figure, and human structures are always at the bottom. If anatomical structures and ontological entities appear in the same figure, then anatomical structures will always be at the left of the figure, and ontological entities will always be at the right of the figure. The ways in which structures can be similar or different are consistently represented by the same style of connecting line from figure to figure. Directionality of relationships is not indicated explicitly in the figures, because the arrows are reserved as a convention for indicating isomorphisms, and most of the isomorphisms considered here will be among nodes; however, the directionality of relations in figures will be explained in the accompanying text wherever it is an important consideration.
Fig. 1.
Key: mouse structures are located toward the top of the figure; human structures toward the bottom. When ontological entities appear in a figure, they appear to the right; anatomical structures appear to the left. The various kinds of relationships, both mapping and ontological, are each represented by a unique style of line.
There is a large body of literature on the application of graphs and graph theory to the description of structural relationships, and especially to their relevance in the representation of medical knowledge [7,46]. Graphs are useful mathematical structures, because the nodes of the graph can be used to represent the anatomical structures under study, while the edges of the graph can be used to represent the relationships among those anatomical structures–a technique fundamental to computer science, which has carried over to the knowledge-representation specialty [47,48]. In that way, we can formally capture what is similar and what is different in comparable structures and relationships, by constructing a graph for each anatomical structure and comparing (matching) the graphs. The comparison of the graphs is effected by comparing each element of the graph to the corresponding element in the other graph, and the graph comparison consists of the set of comparisons of those elements.
Our comparisons involve matching the labeled edges of the graphs (including their directionality), as well as the nodes. This was a deliberate modeling choice, made in order to enable the comparison of spatial and other relationships across species, in addition to comparing the anatomical entities themselves. By permitting this comparison between relationships, CAIS permits the modeling of ontogenetic, evolutionary, physiological, and pathological transformations. For example, certain species of flatfish, such as flounder, are bilaterally symmetrical as hatchlings, and experience migration of crucial organs, such as eyes and renal system components, as they mature [49]. Comparing only the nodes would be insufficient to represent these developmental transformations, because both organs would exist as discrete entities in the initial state and in the transformed state, implying a false isomorphism between those states. To represent such changes as the ontogenetic transformation from symmetric left and right eyes in the flounder hatchling to both eyes on one side in the mature fish, or the evolutionary transformation from two more-or-less bilaterally symmetrical kidneys in basal vertebrates to the widely separated head and trunk kidneys in flatfish, on the other hand, requires formal comparison of spatial relationships between anatomical structures. For this reason, CAIS was designed with the ability to model comparisons among labeled edges (relationships), as well as among nodes (anatomical structures).
Let GA =(A, EA) be a graph with node set A and edge set EA, and let GB =(B, EB) be a second graph. A graph isomorphism is a one-to-one, onto mapping f : A ↦ B such that (a, a’) ∈ GA iff (f(a), f(a’)) ∈ GB. This means that if there is an edge between nodes a and a’ in GA, there must be an edge between the corresponding nodes f(a) and f(a’) in GB, and vice versa. This is called a relational constraint.
For example, let graph A be a tree representation of the human heart (H), and graph B be a tree representation of the mouse heart (M), as depicted in Fig. 2. (For simplicity of illustration, we limit the graph to Cardiac chambers.) The root of each tree is Heart, and each one has four leaf nodes, connected to Heart by two inverse (complementary) relationships: (1) has-part (from Heart, pointing to the chambers), as well as (2) part-of (from the chambers, pointing to Heart): Left atrium, Left ventricle, Right atrium, and Right ventricle.
Fig. 2.
Mapping the human heart (H) to the mouse heart (M).
In mapping the nodes of graph A to the nodes of graph B, mouse Heart matches human Heart, Right atrium matches Right atrium, and so forth. Similarly, the four has-part edges match, as do the four part-of edges. The mapping is therefore one-to-one and onto, and the relational constraints are satisfied, which constitutes a graph isomorphism. If a graph is isomorphic to a subgraph of another graph, the relationship between the graphs is that of a subgraph isomorphism.
In addition to isomorphism, which denotes an exact match between the structures under comparison, the concept of homomorphism, or relationship-preserving partial mapping, is useful in analyzing similar structures. Shapiro and Haralick [50] formally define a relational homomorphism, in order to create a construct that will map the nodes of one graph to those of a second graph, in a way that preserves the interrelationships among the nodes. These comparisons open up the concept of relational distance, or how different or similar graphs are to one another [51]. The relational distance is computed based on a least-error mapping from the nodes of one graph to those of the other, where errors represent failed relationships.
Sanfeliu and Fu [52] worked on a similar problem in the context of pattern recognition. They categorized the different methods of computing a distance measure between attributed graphs, and proposed a distance measure based on cost functions. Given two graphs, a source graph and a reference graph, the cost functions were used to compute the cost of a mapping from the nodes of the source graph to those of the reference graph. Their mapping cost is a summation of the number of node insertions, node deletions, edge insertions, and edge deletions that must be performed to transform the source graph into the reference graph. The minimal mapping cost over all possible mappings (cf. Shapiro and Haralick’s relational distance [51]) is the distance between the graphs.
The formalisms we have outlined are for simple graphs, but the frame-based representation of the FMA in Protégé is much more complex than a simple graph since (1) it has attributed nodes (e.g., has-mass; has-inherent-3D-shape), (2) it has subproperties, such as subslot relation (e.g., has-regional-part is a subslot of has-part) and reified or attributed relations (e.g., attributed-part), and (3) it has multiple relationships (e.g., is-a, has-part, continuous-with, adjacent-to). The edges of the complex graph structure of the FMA represent this rich mixture of structures and relationships. We have found that similarities and differences between two graphs can occur at all levels, as well as across levels, and that, as expected, there are more similarities than differences.
2.4. Ontology matching
Ontology matching is a domain closely related to our application, with a large body of very active work. The difference between CAIS and ontology matching applications is that CAIS does not do the matching of entities between two ontologies itself–it carries out the comparison of two species after the mappings have been generated in some other fashion. CAIS, and the underlying SDM, are the method for comparison, and while they rely on the methods for mapping as a prerequisite for generating input, CAIS and SDM are a separate and subsequent set of operations.
Similar graph-matching algorithms form the basis for ontology matching systems such as Anchor-PROMPT, which enhances semi-automated ontology merging applications by analyzing not only local context (directly related classes and slots between two ontologies), but brings in non-local context by seeking and evaluating possible candidate classes and slots that may also constitute similarities that should be mapped [53]. Just as CAIS does, Anchor-PROMPT operates on an ontology representation as a directed labeled graph, where the classes are represented by nodes, and the slots are represented by edges. Using a set of “anchors” (pairs of related terms from the ontologies specified as related), Anchor-PROMPT extends these relations to analyze the non-local context by traversing the paths between those anchors, returning potential candidates for similarity, and computing cumulative similarity scores for the terms involved [53] (as contrasted to a description of the similarities and differences of the ontologies themselves, as addressed by CAIS).
The approach taken by Anchor-PROMPT holds a great deal of promise for future extension of CAIS, as our system needs a repository of mappings upon which to operate. To generate enough mappings across enough model species for enough anatomical structures, the manual process which was used for the dissertation version of CAIS will quickly become prohibitive, and the Anchor-PROMPT approach holds the potential for efficiently creating a body of mappings that future versions of CAIS will be able to draw upon for comparison.
The Ontology Alignment Evaluation Initiative (OAEI) has been established as a coordinated international initiative to develop consensus on evaluation of methods for schema matching/ontology integration [54]. This motivation has led to the use in recent years of anatomy ontology alignment as one of the tracks in their annual evaluation competition, due not only to the size and complexity of the ontologies involved [55], but also to the pragmatic importance of the problem in applied biomedical informatics. In 2005, the evaluation compared the FMA and OpenGalen, in order to find alignment between classes in each human anatomy ontology; the 2008 competition crossed species by matching Adult Mouse Anatomy and the NCI Anatomy Thesaurus [56]. The systems evaluated show progress over the years in coverage and in ability to deal with complexity–for example, in the ability to compare corresponding concepts that have different compositional names (terms) in different ontologies. The problematic issues they encountered at various steps in the process (inability of systems to completely cover ontologies, lack of a gold standard against which to compare generated mappings and the difficulties encountered in attempting to generate a gold standard subset, intractability of manual curation of mappings, inadequacy of precision and recall as measurement of the quality of the mappings, for example [55,56]) are indicative of the complexity of the problem, and of the need for enhancing methods of validating the semantics of these approaches when applied to cross-species model organism anatomy ontologies–a need which CAIS is a first step toward addressing, in that it is a rigorous description of the similarities and differences among the ontologies in question.
Euzenat and Shvaiko have written a book on ontology matching research efforts [57]. As a survey book, it provides the user with a guide to the terrain, explaining the underlying problem and reviewing various approaches, including evaluating their performance. As CAIS is developed further, we anticipate that it will encounter some of the research questions and challenges that Euzenat and Shvaiko delineate, and will build upon the ongoing efforts in ontology matching systems.
2.5. Model management
Pottinger, Bernstein, and Halevy [58,59] have conducted research in the area of model management to formulate an approach to mapping and merging two different models–for example, the inventory merger of a bookstore with that of a video store. Some of the issues and challenges they have dealt with are directly relevant to developing and querying our model. They have proposed a model-matching-and-merging approach to deal with the problems of combining two or more different schemas in a database environment. Their schemas are represented as graph structures, as are ours. They allow a node in one graph to map to a node in the other graph if they are identical or “similar” entities. Using a very simple definition of similarity, they have developed a matching algorithm to find a mapping from one graph to another. The resulting match is represented as a graph structure itself, a very nice idea which we have implemented in our work.
One of the most important aspects of the work of Pottinger et al. is that the mapping between two models is itself a model–i.e., itis a first-class object, and thus can undergo the same operations as the original models. They outline a set of model management operators, of which the following will be relevant to our SDM: (1) match, (2) apply, (3) compose, and (4) difference. Due to semantic differences between the domains, their operators were not entirely appropriate for our purposes–for example, the fact that two homologous anatomical structures are very different from each other across species does not justify trying to find a better match with a different structure, as their operators would permit. But the underlying logic of their operators suggested the usefulness of specific types of comparison in CAIS for answering queries about what types of anatomical transformations can occur between species–and we accordingly incorporated certain aspects of their logic as an underlying basis for our types of queries, anchored by the relationships similar-to, different-from, shared, not-shared, and union.
Having introduced the domains from which CAIS draws, and the relevant literature informing it, we now describe CAIS’ design and implementation.
3. Comparative anatomy and the structural difference method
The structural difference method (SDM) is a formalism for representing similarities and differences between anatomical structures across two different species. The SDM uses graph isomorphism to illustrate anatomical correspondence, and any deviation from isomorphism to represent a difference in the anatomical entities compared. It allows comparisons on levels from the gross anatomical to the cellular for each species under comparison, and provides the user with the mappings between anatomical entities at each level.
Isomorphism, or graph identity, indicates that there is no difference at a given level of organization; in other words, the mappings between the entities across species are one-to-one and onto. Examples include the Heart chambers (shown in Fig. 2), the Left and Right lung (in mammals), and the mouse and human stomachs at the Organ level. If two structures are isomorphic at some level of abstraction and resolution, they are identical at that level. But if they are not isomorphic, how do we gauge the difference between two corresponding structures?
Based on our preliminary studies and the relational distance work of Shapiro and Haralick [50,51], we propose the following types of differences for our approach: node (structure) differences and edge (relationship) differences. Node mappings may be one-to-one and onto (isomorphism), one-to-one but not onto (sub-graph isomorphism), one-to-nothing (null mapping), one-to-many, many-to-one, or many-to-many. Furthermore, the edges provide relational constraints that may or may not be satisfied (edge similarities and differences). We illustrate each type of symbolic difference with examples, treating the node differences first, and then proceeding to edge differences.
Node set differences are differences between the number of entities in the source species and the corresponding entities in the target species–in other words, a structure that exists in one species but does not exist at all in the other species, or it does exist, but the correspondences are distributed among a different number of entities than in the source species. Examples of such mappings include null mappings, which may be one-to-zero (one Limiting ridge (mouse) to none in the human) or many-to-zero (two Areola[e] of breast (human) to none in the Mammary gland (mouse)). Node set differences are illustrated in Fig. 3.
Fig. 3.
Node set differences for various structures in the human and the mouse.
Additionally, there are mappings that may be one-to-n (one human prostate Organ to five mouse Organ[s]), or n-to-m (three Lobe[s] of right lung (human) to five Lobe[s] of right lung (mouse); two Mammary gland[s] (human) to twelve Mammary gland[s] (mouse). The 1:5 mapping between the human prostate and the mouse prostate organs is illustrated in Fig. 4.
Fig. 4.

The 1:5 correspondence between the human and mouse Prostates at the Organ level.
Node attribute differences are differences in the existence of an attribute between two corresponding structures in the source and target species–in other words, the structure exists in each species, but it occupies a different place in the AT, and thus, the slots required for a sound and complete description of the structure differ across species. For example, has-member (which is a specialization of the partonomic relationship constrained in the FMA to Anatomical sets) is an attribute of the node Set of mouse prostates. In this partonomic scheme, Anatomical set is made up of member Organs. In the human, the prostate is a single organ. The class Organ, however, lacks the attribute has-member, and therefore a node attribute difference exists between the Prostates of the two species. This category of differences is necessary, because it is the only explicit way of acknowledging the difference in roles of the different structures in the AT. In accordance with Stevens’ principle that the parameters of a measurement system be exhaustive and mutually exclusive [60], these attributes are necessary to fully describe the structure and its anatomical role. To correspond to another kind of structure in the AT is to lose those specific attributes of its role in the other species, as well as to gain other attributes, and this category of differences accounts for that shift in anatomical role across species.
Node attribute value differences are differences in values of corresponding attributes shared between corresponding nodes of two species–in other words, the structure exists in both species, and (to some extent) shares an anatomical role, but there is some difference in the values of its attributes from one species to the other. For example, an isomorphism exists between the mouse (or rat) and human Stomachs at the levels of whole Organ and Organ part: the mapping is one-to-one and onto for {Fundus of stomach, Body of stomach, Pyloric antrum}. The isomorphism propagates to the next level of organization, namely, the Stomach wall, the parts of which are: {Mucosa (GM), Submucosa (SM), Muscularis (M) and Serosa (S)}. The difference between the mouse and human Stomachs begins to emerge in the attribute values for the node Mucosa. Unlike the Body of the stomach (human) (HS), which is lined throughout by the Glandular mucosa (GM), the Mucosa of the Body of the stomach (mouse) (MS) is divided into two structurally different regions: Glandular mucosa (GM) and Non-glandular mucosa (NGM). GM and NGM are demarcated from one another by the Limiting ridge (LR), which has no corresponding node in the human [61], as shown in Fig. 5.
Fig. 5.
Node set and node attribute value differences between the human and rodent stomachs.
Edge set differences are differences in the existence of relationships (edges) between structures across species. For example, the Dorsolateral prostates of the mouse are adjacent-to the Coagulating glands, which do not exist as organs in the human. Another example is the Inguinal mammary glands of the mouse, which are adjacent-to the Inguinal ligament (mouse), whereas the human Mammary glands are adjacent only to the Pectoralis major muscle (human). Because they are located in different places in the body in different species, the spatial relationships (such as continuous-with or adjacent-to) among the anatomical entities are changed, and this change is reflected in the relationship differences across species. Edge attribute value differences are differences in the attributes of existing relationships between structures across species. In the same way that nodes can have attributes, edges can as well, and the differences between those attributes can also be expressed symbolically.
There is an asymmetry between the number of node differences and the number of edge differences, due to the lack of edge attribute differences, which would correspond to node attribute differences. This category of edge difference does not exist, because there is no hierarchy of spatial relationships to correspond to the structural hierarchy in the AT.
4. System design
CAIS accepts queries posed by the user about similarities and differences in human, rat, and mouse anatomy. The implementation of this version of the comparative anatomy system is a single database of mappings, from which the query engine accesses and returns a result set. Automatic and dynamic generation of mappings from separate databases by species is a possible future goal of this research, but is specifically outside the scope of this stage of the project. The anatomical mapping data structure and the syntax and semantics of the system’s query language are particularly significant, and will be discussed in more detail below.
4.1. Mappings
Mappings are the data structure at the heart of the proposed information system. As developed in [62], there are two main kinds of mapping classes: Node mappings and Edge mappings, corresponding to the components of the directed graph described by the FMA. The structures which are mapped across species are selected on the basis of homology (evolutionary relatedness); homoplasy (similarity of appearance) and analogy (similarity of function) are not considered in creating mappings. Node mappings are further divided into Node set mappings, Node attribute mappings, and Node attribute value mappings, and Edge mappings are further divided into Edge set mappings and Edge attribute value mappings as specified by the SDM.
The underlying Mapping data structure (shown in Fig. 6) contains pointers in both directions between species: i.e., the human can be either the source or the target species, as can the mouse or rat. Both directions are necessary for a complete answer to queries on similarities and differences between species, as, from the user’s point of view, the answer returned to the query “what is the difference between the human and mouse (or rat) prostates?” should be the same as the answer returned to the query “what is the difference between the mouse (or rat) and human prostates?” This data structure provides that consistency of response, yet at the same time allows a more refined query to return a more granular answer, depending on the level of detail the user wishes to specify. Although the usual query will be bidirectional, there will be users who want information in one direction only. For example, a user may want to know what Prostatic zone in the human is homologous to the mouse Dorsal prostate. This structure is able to accommodate those queries as well.
Fig. 6.
Abstraction of the data structure representing a cross-species comparison between the human and mouse prostates.
The examples for each type of Mapping are taken from [3]. As a class, Mappings are first-class objects (cf. Pottinger and Bernstein [59]), and can thus undergo the same operations as the models from which they are derived. Mappings are thus objects comprised of two species-specific Anatomical structures and the mapping relationship between them.
Mappings are implemented in Protégé in the following manner: the Protégé template slots for Mapping are the two Species being compared, and the two corresponding Anatomical structures. Much of the time the structures will have the same name across species (Left lung (mouse) and Left lung (human)), but not always (cf. Oviduct (dogfish shark) and Fallopian tube (human)). Species names are required to always be single; Anatomical structures can be one or more in a particular Species. Cardinality specifies whether the correspondence is 1:null, null:1, 1:1, 1:many, many:1, many:many, many:null, or null:many.
4.2. Syntax and semantics of the query language
For the purpose of defining CAIS, it is useful to draw a distinction between different kinds of queries, based on how many species models the system handles at a time. These classifications will specify what types of queries our system handles, and what is outside its scope. We define the classification of a query as follows: single-species queries hold for species models taken one at a time. For example, in the human, the Heart is inside the Thoracic cavity, so the query “what is the relationship between Heart and Thoracic cavity [implied: in the human]?” is a single-species query.
Note that a single-species query can be simple or compound; the classification of the query refers not to the complexity of the query, but to the number of species models participating in the query. Single-species queries are the basis of queries in the FMA using Emily [44,45], and involve existence, location, connectivity, and similar features of anatomical structures. Single-species queries are not implemented in our current CAIS system.
Two-species queries hold for species models taken two at a time, and are the basis of what is unique about our CAIS system. They involve comparisons between anatomical structures across two different species and are the main difference between the CAIS system and Emily. For example, the query “how is the human prostate different from the mouse prostate?” is a two-species query. An answer to that query at the Organ level might be: The human prostate is-a discrete organ; the mouse prostate is-a Anatomical set, called Set of prostates (mouse), consisting of 5 member organs (the ventral prostate, left and right dorsolateral prostates, and left and right coagulating glands). Two-species queries involve similarity, difference, homology, identity, and synonymy of anatomical structures in two different species, as described below. Higher-degree queries (as well as queries taking into account sex and stage of development [63]) represent future work, and are explicitly omitted from this specification, but would be easily extensible from the current design. While the concepts of homology, identity, and synonymy overlap to some degree in natural language, the syntax below suffices to deal with them at the level of the users’ needs.
The following BNF rules define a textual abstraction of allow-able two-species queries, and demonstrate the system’s ability to support compound queries.
<query>::=<entity1><relationship><entity2>
<entity1>::=<species1><anat.ent1> ∣ unknown ∣ <result-set>
<entity2>::=<species2><anat.ent2> ∣ unknown ∣ <result-set>
<species1>::=<name-of-species>
<species2>::=<name-of-species>
<anat.ent1>::=<name-of-anatomical-entity>
<anat.ent2>::=<name-of-anatomical-entity>
Both species1 and species2 can be either human or mouse or rat; anat.ent1 and anat.ent2 can be any of the anatomical structures specified earlier, or any of their parts. The fact that the result set from a previous query can be used as an entity in subsequent queries permits CAIS to support complex and detailed compound queries.
We use this syntax as the basis for queries and responses about anatomical similarities and differences between the human, the mouse, and the rat. This notation represents an abstraction of the basis for the queries and responses; there is a low-level syntax that is used by the system for accessing and returning information, as well as a higher level GUI for the users of the system.
Queries are of two major types: set queries and Boolean queries. Boolean queries return T (True) or F (False) when the user queries whether structures in two different species map to each other. Set queries return result sets, such as the set of shared mappings between two species for a structure at a given level of granularity. The semantics of the operators are as follows.
4.2.1. Set queries
The set query operators are similar-to, different-from, shared, not-shared, and union.
similar-to: returns an anatomical isomorphism (one-to-one and onto correspondence) between the two homologous structures across species at the level of granularity (e.g., Organ, Organ part, Cell) of the query if there is one, and returns False otherwise. For example, the Left and Right atria and Left and Right ventricles of the Heart are similar between the mouse and the human.
different-from: returns a non-null correspondence other than anatomical isomorphism (e.g., a one-to-many relationship) between two homologous structures across species at the level of granularity of the query if there is one, and False if there is no mapping in the database. For example, the Lobe[s] of the mouse and human Right lung[s] are different because they are in a 4:3 relationship.
shared: returns all the parts of the structure which occur in both species to the level of granularity specified. For example, the human and mouse Brain[s] both contain an Amygdala, so Amygdala would be one of the structures returned on a shared query on human and mouse Brain.
not-shared: returns all the parts of the structure which occur in one species or the other, but not both, to the level of granularity specified; this is the set complement of the structures returned by shared. For example, the human Brain includes Gyri and Sulci that mouse Brains do not, so the not-shared relation between human and mouse Brains would contain those Gyri and Sulci (among other structures).
union: returns all the parts of the structure that occur either in one species or the other, or in both, to the level of granularity specified: in other words, the set union of the structures returned by the CAIS relationships shared and not-shared.
4.2.2. Boolean queries
The Boolean query operators are is-homologous? and is-different?.
is-homologous? returns True if the two entities selected for the query are homologous, and False if they are not.
is-different? is the opposite of is-homologous?–it returns False if the two entries selected for the query are homologous, and True if they are not.
These Boolean and set query operators suffice to deal with the questions of similarity and difference that a user would ask the system about the comparisons between mouse (or rat) and human anatomy, and this design serves to provide the structure (syntactic and semantic) for those operators.
4.3. CAIS user interface and sample queries
To make the CAIS query functionality available to users, we have designed and implemented a GUI. The CAIS interface is written in Java, and uses the Java API to access the Protégé-2000 database, in which rat, mouse, and human anatomical structures comprise a single hierarchy [64,65]. The CAIS interface provides the following functionalities:
choose the pair of species to compare from all species in the database;
select an anatomical entity from a hierarchy or search for one that the user has entered and give him/her a choice if the entry is ambiguous;
inform the user if selected entities cannot be directly compared and indicate reasonable alternatives if they exist;
select the query operator from a list of choices;
show the user query in a string form as the user constructs it from the GUI;
compare the selected structures at multiple levels of the parts hierarchy as selected by the user (default is one level);
keep track of results from prior queries so the user can return to them; and
show the output in multiple forms including text, tree, graphics, and references.
Fig. 7 shows a screen shot of the full user interface. The user has selected the species “Human” on the left and “Mouse” on the right. She has typed “prostate” in the search area on the left, and the system has found the human prostate in the hierarchy and displayed it. She has also typed “prostate” into the search area on the right. The system has responded with the message “Select from search results,” and displayed four possibilities from which the user has selected “Set of prostates (mouse)”. She has then selected the query operator similar to and clicked on the Execute Query button. The query has been executed, and the results displayed in text mode, since the text tab is the default display tab.
Fig. 7.
CAIS user interface.
As the text display mode (Fig. 8) is very verbose, the user may wish next to look at the results in tree (Fig. 9) or graphics display modes (Fig. 10). Tree results are returned as a structured hierarchy, down as many levels of the tree as were specified in the selected recursion level. In the graphics results a representative graphic is included at each level of the hierarchy. Fig. 11 shows the contents of the References tab, subsequent to the query repeated in the first line of the text–Unknown similar-to Left dorsolateral prostate (rat). The References tab shows the provenance of the information in the peer-reviewed literature, or from domain experts, in narrative form.
Fig. 8.
Text display mode.
Fig. 9.
Tree display mode.
Fig. 10.
Graphics display mode.
Fig. 11.
References display mode.
5. Results
We do not determine the content of the knowledge base. Rather, we model expert consensus [3], and that fact determines how we evaluate the application in regard to the correctness of content. Results, therefore, are correct if they match those provided by the domain expert or reference source. That means that they have to “survive” (1) the process of normalization, according to our syntax and semantics, and (2) entry into Protégé in such a way that the result set based on that information corresponds to what the resource originally said in natural language.
The testing process for the application consisted of developing and carrying out a suite of test cases through the GUI, based on selected scenarios and associated queries. The test cases were all associated with an underlying query, and consisted of the query and the expected results, to be verified against the results obtained when the query was actually run. Table 1 contains a set of representative test cases. In all, 157 test queries were submitted to the CAIS system, and all of them were correctly answered.
Table 1.
Representative test queries.
| Query | Expected response | Obtained expected response? |
|---|---|---|
| Left dorsolateral prostate (rat) similar-to Unknown (human) | Dorsal lobe of prostate (human) | Yes |
| Ventral prostate (rat) is-homologous? Anterior lobe of prostate (human) | F | Yes |
| Unknown (mouse) similar-to Upper lobe of left lung (human) | {} | Yes |
| Right peri-anal mammary gland (mouse) similar-to Unknown (human) | TBD—not null (human) | Yes |
| Mammary gland (human) is-homologous? Mammary gland (mouse) | F | Yes |
| Unknown (human) similar-to Mammary gland (mouse) | Lactiferous duct tree (human) | Yes |
In addition to testing the content of the knowledge base, we also evaluated the interface. Five biomedical and health informatics students tried the interface. Each student was given approximately 5 min of preliminary instruction on how to perform a query, including a demonstration, and then asked to perform a different query using the system. They were then asked to evaluate the interface in terms of clarity and ease of use. This testing determined that the application works well, and is fairly intuitive to use, but users want to see more clarification of the meaning of the different types of possible queries. Some of the interface issues will naturally be resolved as we refine our conceptual model to deal with partial and complex homologies in the content.
6. Discussion
The CAIS system and its associated methods are expected to be useful to biologists and translational medicine researchers. Possible applications range from supporting theoretical work in clarifying and modeling ontogenetic, physiological, pathological, and evolutionary transformations, to concrete techniques to improve the analysis of genotype–phenotype relationships among various animal models.
6.1. Significance
From a biological perspective, the significance of this work lies in the development of a formal, sound, and rigorous technique for modeling anatomical similarities and differences across any pair of species. The importance of anatomy as an essential underpinning of medical knowledge in almost any context from the bench to the clinic has been remarked upon by many observers. The interpretation of almost any kind of medical data, and the inferences drawn from those interpretations, make use of anatomy as an implicit or explicit reference point for diagnosis, treatment, and communication [42].
As the first example of extending the FMA to non-human species, and by permitting the direct comparison of any two species on their own terms, rather than in reference to an anatomical standard species, CAIS also removes the implicit biological assumptions based on an anthropocentric model–a highly significant shift in perspective, considering how much of an outlier species humans are in terms of comparative anatomy. In contrast to the various ontologies currently being developed one at a time under the CARO umbrella (e.g., mouse, fly, amphibian [36]), CAIS enables another aspect of modeling–comparison among those single models. It thus moves beyond static models, and introduces the ability to model dynamic change, whether developmental, pathological, or evolutionary, and it permits those comparisons to be carried out at many different levels of relationship.
The significance of this ability to model transformation describing universal principles of dynamic change in multicellular animals to the field of evolutionary developmental biology is underscored by Myers, who asserts that “the important focus should be on developmental logic, rather than developmental details” [66]. Mabee’s opinion piece calling for phenotype ontologies to connect genomics and evolution [67] agrees with the importance of this focus on the bigger picture, and identifies the inadequacy of current data repositories and computational approaches as one of the major hindrances on the way to this goal. In its ability to provide mappings between the separate ontologies being developed by the CARO collaboration, CAIS provides the opportunity to represent the developmental logic called for by Myers. Additionally, it can serve as a first step in representing the logic of other types of medically significant anatomical transformations (physiological, pathological, and evolutionary), a valuable component of the powerful approach to modeling biological problems of the scope of phenotype–genotype correlation and other applications advocated by Mabee and many others.
Additionally, the introduction of the ability to compare enables the possibility of identifying and resolving conflicts among discrete models. Two anatomical models of different species may each be internally consistent, but conflict with each other, as in the example of the different meanings of “anterior prostate” in humans and rodents. Drawing the mapping between those single-species models identifies the points of conflict, and the reliance of CAIS on entities rather than terms, drawn from the underlying FMA model, provides a means of resolving those conflicts, promoting semantic interoperability among the datasets in different ontologies without rewriting or otherwise changing the underlying data in the models themselves.
6.2. Limitations
The practical benefit of this tool remains limited at the moment, since the knowledge it helps process and query – the anatomical ontologies and especially the mappings – is not yet readily available. However, the basis of the mappings has the potential for semi-automation, so a projected enhancement for a future version of CAIS is the ability – given two models – to speed up the population of the mappings by inferring and proposing potential mappings for the user to approve or reject. This ability will be facilitated by conversion to OBO and support for OWL-DL, which will bring CAIS in line with the emerging preferences of the biology community, and will solve the limitations of the frames structure, which was an artifact of CAIS’ reliance on the existing FMA for templates.
6.3. Structural similarity vs. other forms of similarity
The choice of basing mappings on homology rather than on other forms of similarity was made deliberately, and has significant implications for the use of CAIS within the biological community. A proximate goal is the furthering of genotype–phenotype correlations, especially in the context of health and disease. Homology is the only type of anatomical similarity that concerns itself directly with genotypes across species over evolutionary time. Unlike the other types of similarity (homoplasy and analogy), homology provides a quantitative and objective basis for comparison, based on cladistic analysis, with the corresponding higher confidence in the entities involved. This choice of underlying similarity for modeling carries implications for modeling the differences and similarities of practical importance that biologists care about. One possibility is that, while the mouse prostates offer a valid model for prostate cancer in humans at a high level, it may be more important that they are globally equivalent (at least functionally and for modeling prostate cancer) than to expose the minute differences in their structures. While such an observation is appropriate for the formulation of a hypothesis, it is premature to assume that it is necessarily true, and thus constitutes a major objection to structural modeling.
The ultimate goal of CAIS is to support the theoretical underpinnings of biology and medicine. It will do so by rigorously and formally modeling the aspects of developmental, physiological, pathological, and evolutionary transformation that we do understand, and by shining a spotlight on those areas – indicated either by the lack of mappings or by conflicts among existing mappings – that remain to be explained.
7. Summary and future work
In this paper, we describe a comparative anatomy information system for querying on similarities and differences across species, the knowledge base it operates upon, the method it uses for determining the answer to the queries, and the user interface it employs to present the results. The relevant informatics contributions of our work include (1) the development and application of the structural difference method, a formalism for symbolically representing anatomical similarities and differences across species; (2) the design of the structure of a mapping between the anatomical models of two different species and its application to information about specific structures in humans, mice, and rats; and (3) the design of the internal syntax and semantics of the query language. These contributions provide the foundation for the development of a working system that allows users to submit queries about the similarities and differences between mouse, rat, and human anatomy; delivers result sets that describe those similarities and differences in symbolic terms; and serves as a prototype for the extension of the knowledge base to any number of species. Additionally, we made an initial foray into the validation of the application and its content by means of user questionnaires, software testing, and other feedback.
Based on user feedback, we plan to develop interface and feature enhancements for CAIS. One of the first priorities in future work will be to determine appropriate and more rigorous methods of validation for our approach, including increased evaluation by comparative anatomy domain experts. To that end, we will expand the mappings in the content of the knowledge base to include more of the anatomical structures involved in the MMHCC site cancer working groups. Migrating from the current frame-based incarnation to a DL-based CAIS is also a priority. While using the already existing frames version of the FMA was the most practical choice for a project of the scope of a dissertation, continuing to use frames will hinder future development of CAIS, making the migration a high priority.
On a more theoretical basis, we plan to extend the foundations of the application through the development of models, metamodels, and an anatomical algebra for dealing with them. Stuckenschmidt’s work in developing C-OWL [40] and Bernstein’s semantics for model management operators [68] provide a solid foundation for this expansion of CAIS. This will permit CAIS to provide the basis for a truly integrative anatomical ontology across species.
While the current scope of CAIS is standard anatomy, the methods will apply to mutant phenotypes as well. The wide range of phenotypes involved in comparative medicine means that CAIS will be confronted with a wide variation in phenotypes, rather than one idealized synthetic canonical example. As referred to above, the FMA has already been extended from its traditional canonical representations to deal with instantiated anatomy [7–9], and, based on those preliminary results, we expect that CAIS will correspondingly be able to represent the variations in anatomical features in and among various types of mutants of a given species, including the range of differences among anatomical features displayed by mutant phenotypes of the same species. Specifying, carrying out, and validating those representations will be another high priority in future work.
References
- [1].Jones CE, Baumann U, Brown AL. Automated methods of predicting the function of biological sequences using GO and BLAST. BMC Bioinformatics. 2005 November;6:272. doi: 10.1186/1471-2105-6-272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Biering SF. Evidence-based medicine in treatment and rehabilitation of spinal cord injured. Spinal Cord. 2005 October;43(10):587–92. doi: 10.1038/sj.sc.3101815. [DOI] [PubMed] [Google Scholar]
- [3].Travillian RS, Rosse C, Shapiro LG. An approach to the anatomical correlation of species through the Foundational Model of Anatomy. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2003. pp. 669–73. [PMC free article] [PubMed] [Google Scholar]
- [4].Eckman BA, Kosky AS, Laroco LA., Jr Extending traditional query-based integration approaches for functional characterization of post-genomic data. Bioinformatics. 2001 July;17(7):587–601. doi: 10.1093/bioinformatics/17.7.587. [DOI] [PubMed] [Google Scholar]
- [5].Pagon RA, Tarczy-Hornoch P, Baskin PK, Edwards JE, Covington ML, Espeseth M, et al. GeneTests-GeneClinics: genetic testing information for a growing audience. Hum Mutat. 2002 May;19(5):501–9. doi: 10.1002/humu.10069. [DOI] [PubMed] [Google Scholar]
- [6].Schulz S, Hahn U. Part-whole representation and reasoning in formal biomedical ontologies. Artif Intell Med. 2005 July;34(3):179–200. doi: 10.1016/j.artmed.2004.11.005. [DOI] [PubMed] [Google Scholar]
- [7].Cook DL, Mejino JL, Rosse C. Evolution of a Foundational Model of Physiology: symbolic representation for functional bioinformatics. Stud Health Technol Inform. 2004;107(Pt 1):336–40. [PubMed] [Google Scholar]
- [8].Rosse C, Kumar A, Mejino JL, Jr, Cook DL, Detwiler LT, Smith B. A strategy for improving and integrating biomedical ontologies. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2005. pp. 639–43. [PMC free article] [PubMed] [Google Scholar]
- [9].Smith B, Kumar A, Ceusters W, Rosse C. On carcinomas and other pathological entities. Comp Funct Genomics. 2005;6(7–8):379–87. doi: 10.1002/cfg.497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Noy NF, Crubezy M, Fergerson RW, Knublauch H, Tu SW, Vendetti J, et al. Protégé-2000: an open-source ontology-development and knowledge-acquisition environment. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2003. p. 953. [PMC free article] [PubMed] [Google Scholar]
- [11].Brown SD, Hancock JM, Gates H. Understanding mammalian genetic systems: the challenge of phenotyping in the mouse. PLoS Genet. 2006 August;2(8):e118. doi: 10.1371/journal.pgen.0020118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Bult CJ. From information to understanding: the role of model organism databases in comparative and functional genomics. Anim Genet. 2006 August;37(Suppl. 1):28–40. doi: 10.1111/j.1365-2052.2006.01475.x. [DOI] [PubMed] [Google Scholar]
- [13].Bao L, Peirce JL, Zhou M, Li H, Goldowitz D, Williams RW, et al. An integrative genomics strategy for systematic characterization of genetic loci modulating phenotypes. Hum Mol Genet. 2007 June;16(11):1381–90. doi: 10.1093/hmg/ddm089. [DOI] [PubMed] [Google Scholar]
- [14].Prokisch H, Andreoli C, Ahting U, Heiss K, Ruepp A, Scharfe C, et al. MitoP2: the mitochondrial proteome database–now including mouse data. Nucleic Acids Res. 2006 January;34(Database issue):D705–11. doi: 10.1093/nar/gkj127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Agrafioti I, Stumpf MP. SNPSTR: a database of compound microsatellite-SNP markers. Nucleic Acids Res. 2007 January;35(Database issue):D71–5. doi: 10.1093/nar/gkl806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Friedberg EC, Meira LB. Database of mouse strains carrying targeted mutations in genes affecting biological responses to DNA damage Version 7. DNA Repair (Amst) 2006 February;5(2):189–209. doi: 10.1016/j.dnarep.2005.09.009. [DOI] [PubMed] [Google Scholar]
- [17].Bogue MA, Grubb SC, Maddatu TP, Bult CJ. Mouse Phenome Database (MPD) Nucleic Acids Res. 2007 January;35(Database issue):D643–9. doi: 10.1093/nar/gkl1049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Paigen K, Eppig JT. A mouse phenome project. Mamm Genome. 2000 September;11(9):715–7. doi: 10.1007/s003350010152. [DOI] [PubMed] [Google Scholar]
- [19].Eppig JT, Blake JA, Bult CJ, Kadin JA, Richardson JE, Mouse Genome Database Group The mouse genome database (MGD): new features facilitating a model system. Nucleic Acids Res. 2007 January;35(Database issue):D630–7. doi: 10.1093/nar/gkl940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Blake JA, Eppig JT, Richardson JE, Davisson MT. The Mouse Genome Database (MGD): a community resource. Status and enhancements. The Mouse Genome Informatics Group. Nucleic Acids Res. 1998 January;26(1):130–7. doi: 10.1093/nar/26.1.130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Blake JA, Richardson JE, Davisson MT, Eppig JT. The Mouse Genome Database (MGD): genetic and genomic information about the laboratory mouse. The Mouse Genome Database Group. Nucleic Acids Res. 1999 January;27(1):95–8. doi: 10.1093/nar/27.1.95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Blake JA, Eppig JT, Bult CJ, Kadin JA, Richardson JE, Mouse Genome Database Group The Mouse Genome Database (MGD): updates and enhancements. Nucleic Acids Res. 2006 January;34(Database issue):D562–7. doi: 10.1093/nar/gkj085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Twigger SN, Shimoyama M, Bromberg S, Kwitek AE, Jacob HJ, RGD Team The Rat Genome Database, update 2007–easing the path from disease to data and back again. Nucleic Acids Res. 2007 January;35(Database issue):D658–62. doi: 10.1093/nar/gkl988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Lussier Y, Borlawsky T, Rappaport D, Liu Y, Friedman C. PhenoGO: assigning phenotypic context to gene ontology annotations with natural language processing. Proceedings of the Pacific Symposium on Biocomputing; Singapore. World Scientific Publishing; 2006. pp. 64–75. [PMC free article] [PubMed] [Google Scholar]
- [25].Hayamizu TF, Mangan M, Corradi JP, Kadin JA, Ringwald M. [accessed 03.08.09];The Adult Mouse Anatomical Dictionary: a tool for annotating and integrating data. Genome Biol. 2005 6(3):R29. doi: 10.1186/gb-2005-6-3-r29. Available from: http://www.informatics. jax.org/searches/AMA form.shtml. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26]. [accessed 03.08.09]; http://www.informatics.jax.org/searches/anatdict_form.shtml.
- [27]. [accessed 03.08.09]; http://genex.hgu.mrc.ac.uk/
- [28].Green EC, Gkoutos GV, Lad HV, Blake A, Weekes J, Hancock JM. EMPReSS: European mouse phenotyping resource for standardized screens. Bioinformatics. 2005 June;21(12):2930–1. doi: 10.1093/bioinformatics/bti441. [DOI] [PubMed] [Google Scholar]
- [29].Gailus-Durner V, Fuchs H, Becker L, Bolle I, Brielmeier M, Calzada-Wack J, et al. Introducing the German Mouse Clinic: open access platform for standardized phenotyping. Nat Methods. 2005 June;2(6):403–4. doi: 10.1038/nmeth0605-403. [DOI] [PubMed] [Google Scholar]
- [30].Schofield PN, Bard JB, Booth C, Boniver J, Covelli V, Delvenne P, et al. Pathbase: a database of mutant mouse pathology. Nucleic Acids Res. 2004 January;32(Database issue):D512–5. doi: 10.1093/nar/gkh124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Kurc T, Janies DA, Johnson AD, Langella S, Oster S, Hastings S, et al. An XML-based system for synthesis of data from disparate databases. J Am Med Inform Assoc. 2006 May–June;13(3):289–301. doi: 10.1197/jamia.M1848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Butte AJ, Kohane IS. Creation and implications of a phenome–genome network. Nat Biotechnol. 2006 January;24(1):55–62. doi: 10.1038/nbt1150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33]. [accessed 03.08.09]; http://www.bioontology.org/wiki/images/0/0d/CAROchapter.pdf.
- [34].Topalis P, Tzavlaki C, Vestaki K, Dialynas E, Sonenshine DE, Butler R, et al. Anatomical ontologies of mosquitoes and ticks, and their web browsers in VectorBase. Insect Mol Biol. 2008 February;17(1):87–9. doi: 10.1111/j.1365-2583.2008.00781.x. [DOI] [PubMed] [Google Scholar]
- [35].Sprague J, Bayraktaroglu L, Bradford Y, Conlin T, Dunn N, Fashena D, et al. The Zebrafish Information Network: the zebrafish model organism database provides expanded support for genotypes and phenotypes. Nucleic Acids Res. 2008 January;36(Database issue):D768–72. doi: 10.1093/nar/gkm956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Maglia AM, Leopold JL, Pugener LA, Gauch S. An anatomical ontology for amphibians. Proceedings of the Pacific Symposium on Biocomputing; Singapore. World Scientific Publishing; 2007. pp. 367–78. [PubMed] [Google Scholar]
- [37]. [accessed 03.08.09]; http://www.bioontology.org/wiki/index.php/PATO:About.
- [38].Mork P, Pottinger RA, Bernstein PA. Challenges in precisely aligning models of human anatomy using generic schema matching. Proceedings of Medinfo2004; Amsterdam. IOS Press; 2004. pp. 401–5. [PubMed] [Google Scholar]
- [39].Mork P, Bernstein PA. Adapting a generic match algorithm to align ontologies of human anatomy. Proceedings of the International Conference on Data Engineering; Washington, DC. IEEE Computer Society; 2004. pp. 787–90. [Google Scholar]
- [40].Stuckenschmidt H, van Harmelen F, Bouquet P, Giunchiglia F, Serafini L. Using C-OWL for the alignment and merging of medical ontologies. Proceedings of KR-MED 2004, First International Workshop on Formal Biomedical Knowledge Representation; Bethesda, MD. American Medical Informatics Association; 2004. pp. 88–101. [Google Scholar]
- [41].Rosse C, Mejino JL., Jr A reference ontology for biomedical informatics: the Foundational Model of Anatomy. J Biomed Inform. 2003 December;36(6):478–500. doi: 10.1016/j.jbi.2003.11.007. [DOI] [PubMed] [Google Scholar]
- [42].Rosse C, Mejino JL, Modayur BR, Jakobovits R, Hinshaw KP, Brinkley JF. Motivation and organizational principles for anatomical knowledge representation: the digital anatomist symbolic knowledge base. J Am Med Inform Assoc. 1998 January–February;5(1):17–40. doi: 10.1136/jamia.1998.0050017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Detwiler LT, Mejino JV, Jr, Rosse C, Brinkley JF. Efficient web-based navigation of the Foundational Model of Anatomy. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2003. p. 829. [PMC free article] [PubMed] [Google Scholar]
- [44].Shapiro LG, Chung E, Detwiler LT, Mejino JL, Jr, Agoncillo AV, Brinkley JF, et al. Processes and problems in the formative evaluation of an interface to the Foundational Model of Anatomy knowledge base. J Am Med Inform Assoc. 2005 January–February;12(1):35–46. doi: 10.1197/jamia.M1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Detwiler LT, Chung E, Li A, Mejino JL, Jr, Agoncillo A, Brinkley J, et al. A relationcentric query engine for the Foundational Model of Anatomy. Stud Health Technol Inform. 2004;107(Pt 1):341–5. [PubMed] [Google Scholar]
- [46].Joubert M, Miton F, Fieschi M, Robert JJ. A conceptual graphs modeling of UMLS components. Proceedings of Medinfo1995; Amsterdam. North Holland; 1995. pp. 90–4. [PubMed] [Google Scholar]
- [47].Campbell KE, Das AK, Musen MA. A logical foundation for representation of clinical data. J Am Med Inform Assoc. 1994 May–June;1(3):218–32. doi: 10.1136/jamia.1994.95236154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Campbell KE, Musen MA. Representation of clinical data using SNOMED III and conceptual graphs. Proceedings of the Annual Symposium on Computer Applications in Medical Care; Washington, DC. IEEE Computer Society; 1992. pp. 354–8. [PMC free article] [PubMed] [Google Scholar]
- [49].Martinez GM, Bolker JA. Embryonic and larval staging of summer flounder (Paralichthys dentatus) J Morphol. 2003 February;255(2):162–76. doi: 10.1002/jmor.10053. [DOI] [PubMed] [Google Scholar]
- [50].Shapiro LG, Haralick RM. Organization of relational models for scene analysis. IEEE Transactions on Pattern Analysis and Machine Intelligence, PAMI-4; Washington, DC. IEEE Computer Society; 1982. pp. 595–602. [DOI] [PubMed] [Google Scholar]
- [51].Shapiro LG, Haralick RM. A metric for comparing relational descriptions. IEEE Transactions on Pattern Analysis and Machine Intelligence, PAMI-7; Washington, DC. IEEE Computer Society; 1985. pp. 90–4. [DOI] [PubMed] [Google Scholar]
- [52].Sanfeliu A, Fu KS. A distance measure between attributed relational graphs for pattern recognition. IEEE Transactions on Systems, Man, and Cybernetics, SMC-13; New York, NY. IEEE; 1983. pp. 353–62. [Google Scholar]
- [53].Noy NF, Musen MA. Anchor-PROMPT: using non-local context for semantic matching. Proceedings of the Workshop on Ontologies and Information Sharing at the Seventeenth International Joint Conference on Artificial Intelligence (IJCAI-2001); San Francisco, CA. Morgan Kaufmann; 2001. pp. 63–70. [Google Scholar]
- [54]. [accessed 03.08.09]; http://oaei.ontologymatching.org/
- [55]. [accessed 03.08.09]; http://oaei.ontologymatching.org/2005/anatomy/
- [56]. [accessed 03.08.09]; http://oaei.ontologymatching.org/2008/anatomy/
- [57].Euzenat J, Shvaiko P. Ontology Matching. Springer-Verlag; Heidelberg: 2007. [Google Scholar]
- [58].Bernstein PA, Levy AY, Pottinger RA. Microsoft Research Technical Report MSR-TR-2000-53. Microsoft; Redmond, WA: 2000. A vision for management of complex models. [Google Scholar]
- [59].Pottinger RA, Bernstein PA. University of Washington Technical Report UW-CSE-03-02-03. University of Washington; Seattle, WA: 2003. Merging models based on given correspondences. [Google Scholar]
- [60].Stevens SS. On the theory of scales of measurement. Science. 1946 June;103(2684):677–80. [PubMed] [Google Scholar]
- [61].Robert A. Proposed terminology for the anatomy of the rat stomach. Gastroenterology. 1971 February;60(2):344–5. [PubMed] [Google Scholar]
- [62].Travillian RS. MS thesis. University of Washington; Seattle, WA: 2004. From homology to ontology: comparing anatomy across species with the structural difference method. [Google Scholar]
- [63].Aitken S. Formalizing concepts of species, sex and developmental stage in anatomical ontologies. Bioinformatics. 2005 June;21(11):2773–9. doi: 10.1093/bioinformatics/bti409. [DOI] [PubMed] [Google Scholar]
- [64].Travillian RS, Gennari JH, Shapiro LG. Of mice and men: design of a comparative anatomy information system. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2005. pp. 734–8. [PMC free article] [PubMed] [Google Scholar]
- [65].Travillian RS, Diatchka K, Judge TK, Wilamowska K, Shapiro LG. A graphical user interface for a comparative anatomy information system: design, implementation and usage scenarios. Proceedings of the Americal Medical Informatics Association Annual Symposium; Bethesda, MD. American Medical Informatics Association; 2006. pp. 774–8. [PMC free article] [PubMed] [Google Scholar]
- [66]. [accessed 03.08.09]; http://scienceblogs.com/pharyngula/2008/02/plant_and_animal_development_c.php.
- [67].Mabee PM, Ashburner M, Cronk Q, Gkoutos GV, Haendel M, Segerdell E, et al. Phenotype ontologies: the bridge between genomics and evolution. Trends Ecol Evol. 2007 July;22(7):345–50. doi: 10.1016/j.tree.2007.03.013. [DOI] [PubMed] [Google Scholar]
- [68].Bernstein PA, Melnik S. Model management 2.0–manipulating richer mappings. Proceedings of the Special Interest Group on the Management of Data; New York, NY. ACM; 2007. pp. 1–12. [Google Scholar]











