Abstract
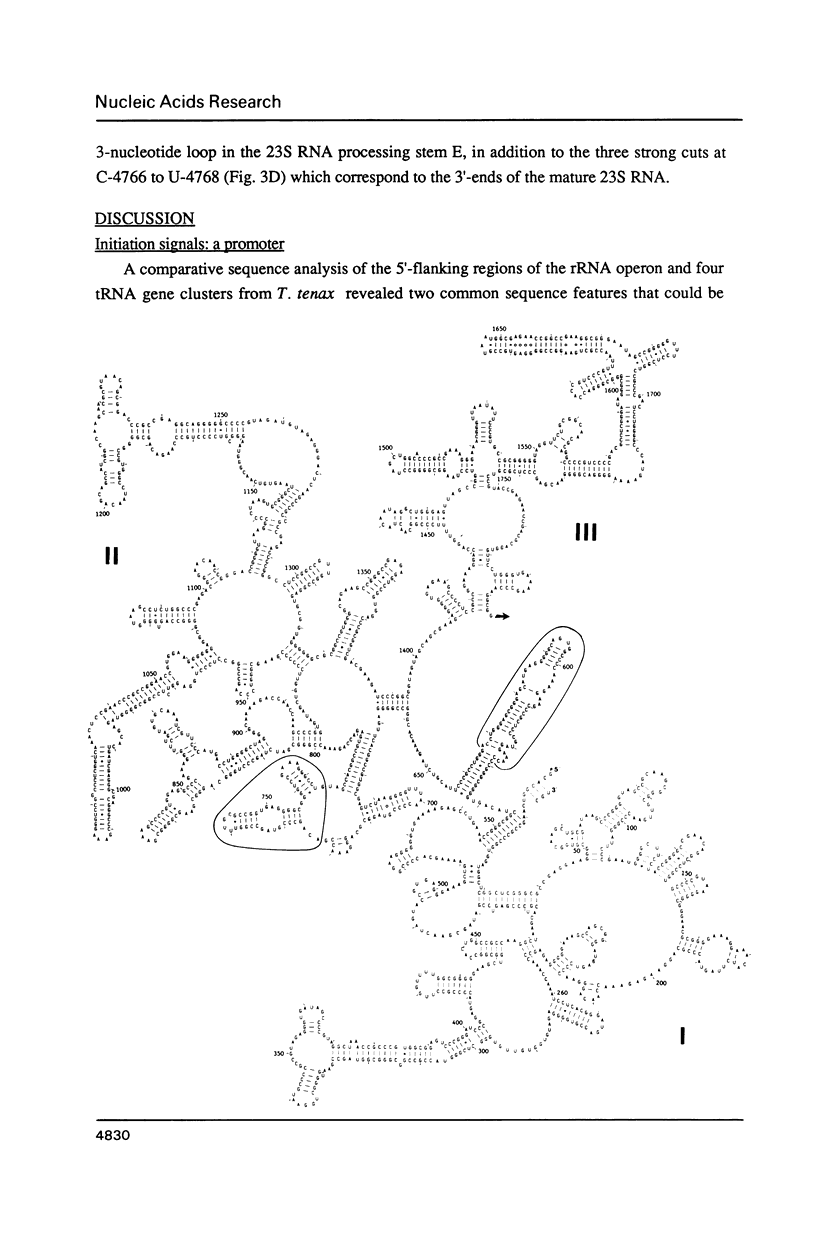
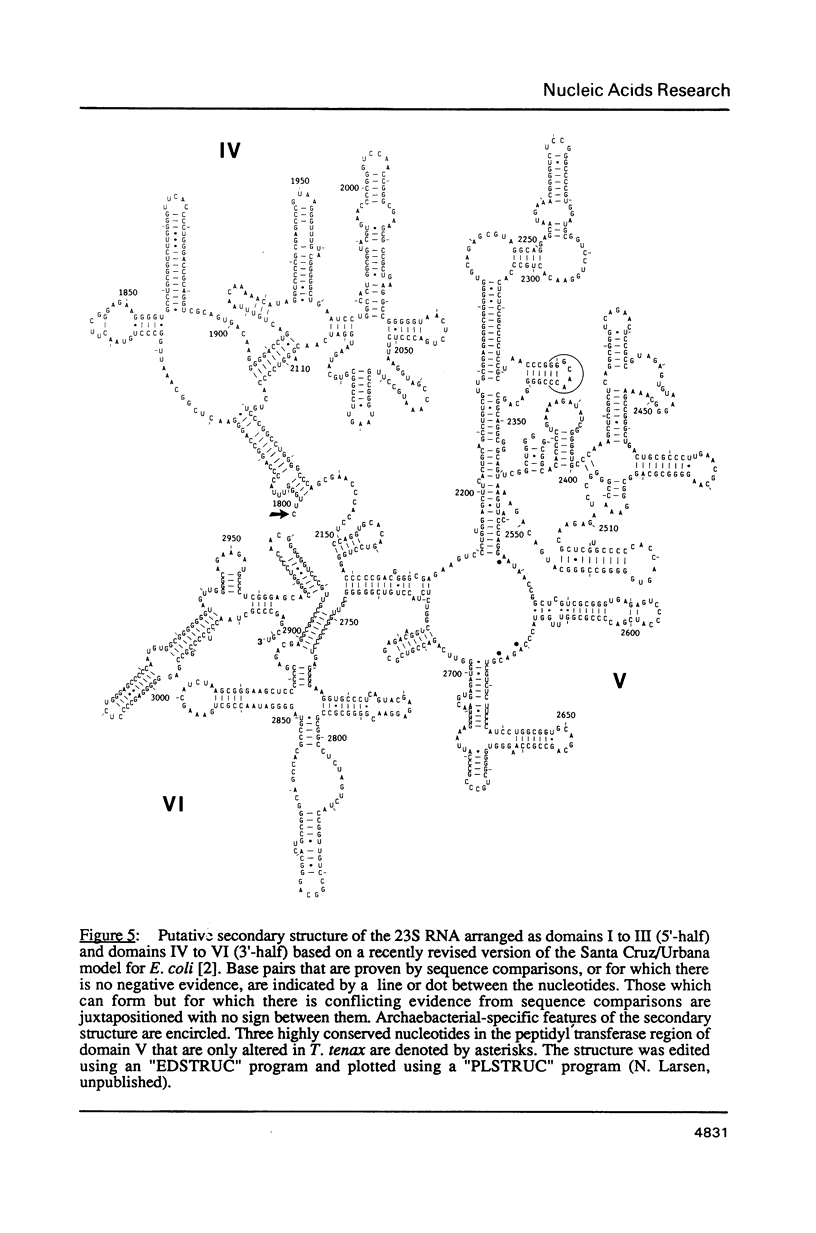
The single ribosomal RNA (rRNA) operon from the extreme thermophile and archaebacterium Thermoproteus tenax was sequenced. Sites of transcriptional initiation and termination were established and the processing sites on the primary transcript were mapped with nuclease S1. The operon contained genes coding for 16S and 23S RNAs but lacked those coding for tRNA and 5S RNA. Transcription initiates 175 bp upstream from the start of the 16S RNA gene (Wich et al., EMBO J. 6, 523-528, 1987) and terminates 49 bp downstream from the 23S RNA gene within a long pyrimidine sequence. An open reading frame downstream from the rRNA operon is transcribed. The sequences bordering both 16S and 23S RNA genes can form putative processing stems in the primary transcript that involve the whole of the 16S-23S RNA spacer. The stems contain irregular features that constitute processing signals and are conserved in other archaebacteria. The 16S RNA stem is cut prior to that of the 23S RNA and RNA maturation follows. An unusual 14 bp helix can form between the extremities of the transcript such that the whole transcript is highly structured and a fork-like structure is formed together with the processing stems. The 23S RNA sequence was aligned with other available 23S-like RNA sequences (Leffers et al., J. Mol. Biol. 195, in press): a putative secondary structure exhibiting archaebacterial-specific features was deduced using comparative sequence analyses. A rooted phylogenetic tree was also derived for the archaebacteria that confirms their division into three major subgroups.
Full text
PDF










Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aiba H., Adhya S., de Crombrugghe B. Evidence for two functional gal promoters in intact Escherichia coli cells. J Biol Chem. 1981 Nov 25;256(22):11905–11910. [PubMed] [Google Scholar]
- Ansorge W., Barker R. System for DNA sequencing with resolution of up to 600 base pairs. J Biochem Biophys Methods. 1984 Mar;9(1):33–47. doi: 10.1016/0165-022x(84)90064-2. [DOI] [PubMed] [Google Scholar]
- Britten R. J. Rates of DNA sequence evolution differ between taxonomic groups. Science. 1986 Mar 21;231(4744):1393–1398. doi: 10.1126/science.3082006. [DOI] [PubMed] [Google Scholar]
- Brosius J., Dull T. J., Sleeter D. D., Noller H. F. Gene organization and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol. 1981 May 15;148(2):107–127. doi: 10.1016/0022-2836(81)90508-8. [DOI] [PubMed] [Google Scholar]
- Chant J., Dennis P. Archaebacteria: transcription and processing of ribosomal RNA sequences in Halobacterium cutirubrum. EMBO J. 1986 May;5(5):1091–1097. doi: 10.1002/j.1460-2075.1986.tb04327.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ettayebi M., Prasad S. M., Morgan E. A. Chloramphenicol-erythromycin resistance mutations in a 23S rRNA gene of Escherichia coli. J Bacteriol. 1985 May;162(2):551–557. doi: 10.1128/jb.162.2.551-557.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Favaloro J., Treisman R., Kamen R. Transcription maps of polyoma virus-specific RNA: analysis by two-dimensional nuclease S1 gel mapping. Methods Enzymol. 1980;65(1):718–749. doi: 10.1016/s0076-6879(80)65070-8. [DOI] [PubMed] [Google Scholar]
- Fox G. E., Stackebrandt E., Hespell R. B., Gibson J., Maniloff J., Dyer T. A., Wolfe R. S., Balch W. E., Tanner R. S., Magrum L. J. The phylogeny of prokaryotes. Science. 1980 Jul 25;209(4455):457–463. doi: 10.1126/science.6771870. [DOI] [PubMed] [Google Scholar]
- Hori H., Osawa S. Evolutionary change in 5S RNA secondary structure and a phylogenic tree of 54 5S RNA species. Proc Natl Acad Sci U S A. 1979 Jan;76(1):381–385. doi: 10.1073/pnas.76.1.381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hui I., Dennis P. P. Characterization of the ribosomal RNA gene clusters in Halobacterium cutirubrum. J Biol Chem. 1985 Jan 25;260(2):899–906. [PubMed] [Google Scholar]
- Lake J. A., Henderson E., Oakes M., Clark M. W. Eocytes: a new ribosome structure indicates a kingdom with a close relationship to eukaryotes. Proc Natl Acad Sci U S A. 1984 Jun;81(12):3786–3790. doi: 10.1073/pnas.81.12.3786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S. C., Squires C. L., Squires C. Antitermination of E. coli rRNA transcription is caused by a control region segment containing lambda nut-like sequences. Cell. 1984 Oct;38(3):851–860. doi: 10.1016/0092-8674(84)90280-0. [DOI] [PubMed] [Google Scholar]
- Mankin A. S., Teterina N. L., Rubtsov P. M., Baratova L. A., Kagramanova V. K. Putative promoter region of rRNA operon from archaebacterium Halobacterium halobium. Nucleic Acids Res. 1984 Aug 24;12(16):6537–6546. doi: 10.1093/nar/12.16.6537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noller H. F. Structure of ribosomal RNA. Annu Rev Biochem. 1984;53:119–162. doi: 10.1146/annurev.bi.53.070184.001003. [DOI] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Wich G., Hummel H., Jarsch M., Bär U., Böck A. Transcription signals for stable RNA genes in Methanococcus. Nucleic Acids Res. 1986 Mar 25;14(6):2459–2479. doi: 10.1093/nar/14.6.2459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wich G., Leinfelder W., Böck A. Genes for stable RNA in the extreme thermophile Thermoproteus tenax: introns and transcription signals. EMBO J. 1987 Feb;6(2):523–528. doi: 10.1002/j.1460-2075.1987.tb04784.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Gupta R., Hahn C. M., Zillig W., Tu J. The phylogenetic relationships of three sulfur dependent archaebacteria. Syst Appl Microbiol. 1984;5:97–105. doi: 10.1016/s0723-2020(84)80054-5. [DOI] [PubMed] [Google Scholar]
- Woese C. R., Olsen G. J. Archaebacterial phylogeny: perspectives on the urkingdoms. Syst Appl Microbiol. 1986;7:161–177. doi: 10.1016/s0723-2020(86)80001-7. [DOI] [PubMed] [Google Scholar]
- Zillig W., Schnabel R., Stetter K. O. Archaebacteria and the origin of the eukaryotic cytoplasm. Curr Top Microbiol Immunol. 1985;114:1–18. doi: 10.1007/978-3-642-70227-3_1. [DOI] [PubMed] [Google Scholar]