Abstract
Meiotic recombination, which promotes proper homologous chromosome segregation at the first meiotic division, normally occurs between allelic sequences on homologues. However, recombination can also take place between non-allelic DNA segments that share high sequence identity. Such non-allelic homologous recombination (NAHR) can markedly alter genome architecture during gametogenesis by generating chromosomal rearrangements. Indeed, NAHR-mediated deletions, duplications, inversions and other alterations have been implicated in numerous human genetic disorders. Studies in yeast have revealed insights into the molecular mechanisms of meiotic NAHR as well as the cellular strategies that limit NAHR.
Gametes are the products of a meiotic programme in which diploid germ cells undergo one round of DNA replication followed by two successive rounds of cell division. During this process, gametes acquire chromosomes comprising new assortments of parental alleles. Subsequent fusion of two gametes during sexual reproduction results in the reconstitution of a diploid genome, yielding offspring that are genetically distinct from their parents.
In most sexually reproducing organisms, homologous recombination lies at the heart of meiosis by promoting proper segregation of HOMOLOGOUS CHROMOSOMES (also referred to as homologues) (reviewed in REF. 1). Prior to the first meiotic division, recombination begins when DNA double-strand breaks (DSBs) are deliberately introduced into each chromosome. The DSBs are then repaired by genetic exchange with allelic sequences, resulting in physical linkages between pairs of homologues. These connections ensure that homologues orient correctly on the meiotic spindle and migrate to opposite spindle poles. Meiotic recombination thus promotes genetic stability and faithful transmission of the genome by limiting the repair of each DSB primarily to DNA sequences at the allelic position on the homologue and ensuring the accurate distribution of chromosomes to gametes.
Errors in meiotic recombination can affect genome stability during gametogenesis. Research spanning several decades has shown that chromosome non-disjunction (that is, missegregation) in meiosis results in constitutive ANEUPLOIDY, which can lead to spontaneous abortion or congenital birth defects (reviewed in REF. 2). More recent work has uncovered a second process that influences genome stability in the germline: aberrant meiotic recombination between non-allelic DNA segments that share high sequence similarity. This process is known as non-allelic homologous recombination (NAHR); the synonymous term `ectopic recombination' is more prevalent in older literature and in yeast studies. NAHR is a more recent, mechanistically evocative term and is used in this Review. Because eukaryotic genomes from yeasts to human harbour repeated blocks of DNA3, 4, a meiotic DSB formed within a repeat has the potential to induce genomic rearrangement through repair with non-allelic sequences. In humans, NAHR-mediated events leading to chromosomal alterations have been implicated in numerous disorders. It is thus important to understand the mechanisms behind NAHR in the germline.
This Review focuses on genome instability induced by meiotic NAHR. We begin with an overview of the mechanism of meiotic recombination and then discuss the types of chromosomal rearrangement and human disorder that are associated with NAHR between duplicated regions of the genome. We next examine insights into NAHR mechanisms obtained from studies in other organisms. Finally, we review cellular strategies that prevent meiotic NAHR and favour recombination between allelic sequences, and consider some of the future challenges in the field.
Meiotic recombination
Meiotic recombination is inherently hazardous because it is initiated by developmentally programmed DSBs at multiple sites in the genome (FIG. 1) (reviewed in REF. 5). A cascade of homologous recombination events, normally templated by DNA at the allelic position on the homologue, results in DSB repair either with reciprocal exchange of chromosome arms that flank the DSB site (a crossover) or without exchange of flanking arms (a noncrossover).
Figure 1. Meiotic recombination pathway.
A double-strand break (DSB) introduced by Spo11 is resected to expose 3′ ssDNA tails. Mediated by Rad51 and Dmc1, one of the 3′ ssDNA tails first invades the homologous chromosome to form a displacement loop (D-loop) and heteroduplex DNA (highlighted), and then initiates DNA synthesis. In the canonical pathway, capture of the second end, DNA synthesis and ligation generate Holliday junctions flanking the DSB site. The Holliday junctions can be resolved by cutting the outside strands (filled arrowheads) or crossed strands (open arrowheads) of each junction (two of the four possible resolution configurations are shown here). In principle, resolution can lead to either a crossover or a noncrossover configuration of the DNA flanking the recombination site. In vivo, however, most resolution products are crossovers. An alternative recombination pathway depicted on the right (known as synthesis-dependent strand annealing (SDSA)) leads primarily to noncrossovers. In both types of pathway, gene conversion can occur by mismatch repair if DNA sequence polymorphisms are incorporated into the heteroduplex DNA that forms as part of the recombination process.
Mechanisms of meiotic recombination
Much of what is known about the molecular events of meiotic recombination comes from studies in the budding yeast Saccharomyces cerevisiae (FIG.1) (reviewed in REFS. 5, 6). Each initiating DSB is formed by a homodimer of the enzyme Spo11, which mediates a topoisomerase-like transesterification reaction. Each Spo11 monomer remains covalently attached to the 5′ terminus of a DNA strand at each DSB and is released following endonucleolytic cleavage of each strand at sites 3′ to the DSB. The two 5′ ends of the DSB are further resected to expose 3′ single-stranded DNA (ssDNA) tails. The strand exchange proteins Rad51 and Dmc1 bind these tails, search for homologous DNA sequences and then catalyse invasion by one of the two 3′ ssDNA tails into the homologous sequence of an intact, double-stranded homologue (that is, not the sister chromatid), thereby generating HETERODUPLEX DNA.This results in separation of the strands of the invaded DNA duplex and the formation of an intermediate known as a displacement loop (D-LOOP). Re-synthesis of DNA strands destroyed by resection is primed by each of the 3′ ssDNA tails, using the strands of the invaded homologue as templates. For some recombination events, branched DNA intermediates containing pairs of HOLLIDAY JUNCTIONS are formed and subsequently resolved, yielding mature recombinant DNA molecules that are primarily in a crossover configuration. Other recombination events are thought to proceed through a mechanism termed synthesis-dependent strand annealing (SDSA) (FIG. 1), in which the invading DNA strand is displaced after DNA synthesis, and then anneals to the complementary ssDNA tail on the other side of the DSB. This mechanism leads primarily to a noncrossover outcome. The choice between crossover and noncrossover outcomes is highly regulated (reviewed in REF. 7).
In both recombination pathways, heteroduplex DNA forms between strands of the two parental alleles, and, importantly, GENE CONVERSION can occur when heterologies that are caused by sequence differences between the two strands are corrected by MISMATCH REPAIR (MMR).Gene conversion can occur with both crossover and noncrossover modes of recombination. The direction of genetic information transfer caused by gene conversion is most often such that the site where the DSB was originally formed is converted to a copy of the sequence of the unbroken homologue . As a consequence of this genetic information transfer, recombination results in sequence homogenization between the participating alleles (reviewed in REFS. 5, 6) (FIG. 1).
Many of the molecular players involved in meiotic recombination in yeast are conserved in larger eukaryotes (reviewed in REFS. 1, 6). For example, disruption of SPO11 homologues in Caenorhabditis elegans8, in Drosophila melanogaster9 and in Mus musculus 10, 11 eliminates meiotic recombination. Furthermore, orthologues of Dmc1 and Rad51, as well as proteins acting later in the crossover and noncrossover pathways, have roles in recombination in larger eukaryotes12, 13. This high degree of conservation means that studies in experimentally tractable organisms provide insight into mechanisms of allelic recombination and NAHR in humans.
Meiotic recombination hotspots
The distribution of meiotic recombination in genomes from yeasts to mammals is not random: most recombination occurs in highly localized 1- to 2-kilobase (kb) genomic domains termed hotspots, which are preferential sites of DSB formation by Spo11 (reviewed in REF. 14). Humans and mice display sex-specific variation of recombination15, 16. Individual human crossover hotspots were first identified at high resolution by sperm typing, a PCR-based approach for recovering crossover DNA molecules directly from sperm DNA14.Subsequently, a genome-wide map of crossover hotspots inferred from population genetic methods was generated, leading to an estimate of 25,000–50,000 crossover hotspots in the human genome17. Genome-wide meiotic DSB and/or recombination maps are also available in budding and fission yeasts18–24. However, despite the hotspot catalogues available, little is known about the factors that govern where DSBs are formed (BOX 1). In perhaps the most significant association in humans to date, a single sequence feature — the degenerate 13-nucleotide motif CCNCCNTNNCCNC — is present in >40% of the meiotic crossover hotspots inferred from population genetic analyses17, 25. The fact that recombination is not distributed randomly across the genome is important for understanding the mechanisms behind NAHR because NAHR events are, by definition, highly dependent on the sequence context in which DSBs form (that is, whether DSBs occur in unique sequences or in repeated DNA).
NAHR in the human genome
The human genome is replete with sequences that are present in a high number of copies, such as transposable elements (for example, Alu elements and long interspersed repetitive elements (LINEs)), or in low number of copies26. Roughly 5% of the human genome comprises low copy repeats (LCRs; also known as segmental duplications), which are blocks of DNA that share ≥90% sequence identity over ≥1 kb (REF. 4). Interest in LCRs has intensified as their roles in human disease, genome architecture and genome evolution have emerged (reviewed in REF. 27).
Genomic disorders: outcomes of NAHR
The high sequence identity over large stretches of DNA renders LCRs susceptible to germline NAHR events that result in deletion, duplication, inversion or ISODICENTRIC CHROMOSOME formation (FIG. 2). Structural alterations that change the constitutional copy number of dosage-sensitive genes, that disrupt genes or that generate gene chimeras result in conditions collectively termed GENOMIC DISORDERS (reviewed in REF. 28). Much of the work on these disorders was pioneered by Lupski and colleagues, who first recognized the role of LCRs in the aetiology of two inherited neurological disorders: CHARCOT-MARIE-TOOTH DISEASE TYPE 1A (CMT1A) and HEREDITARY NEUROPATHY WITH LIABILITY TO PRESSURE PALSIES (HNPP)29, 30. Unequal crossing over between two LCRs on chromosome 17p results in the duplication (causing CMT1A) or deletion (causing HNPP) of genes located in the intervening 1.4-megabase (Mb) region. NAHR-mediated events occurring in the germline are now known to account for over 30 genomic disorders, with phenotypes ranging from mental retardation to blood diseases or male infertility (TABLE 1). NAHR also contributes to copy-number and structural variation in the human genome through LCR-mediated duplications, deletions and inversions that are not pathogenic31–33.
Figure 2. Genome rearrangement by non-allelic homologous recombination.
Crossover recombination between repeated DNA sequences at non-allelic positions can generate a deletion, a duplication, an inversion or an isodicentric chromosome. Depicted here are six chromosomal outcomes of non-allelic homologous recombination (NAHR) between repeats located on the same chromosome: two orientations of repeats relative to one another (direct or inverted) for each of three types of interactions (between homologues, between sister chromatids or in the same chromatid). Homologous chromosomes are shown in blue and red, and sister chromatids are depicted in the same colour (homologous chromosomes are not shown in the schematics depicting inter-sister chromatid or intrachromatid exchanges for simplicity). Low-copy repeats (LCRs) are shown as white and black arrowheads. Figure modified, with permission, from REF. 55.
Table 1.
Recurrent human genomic disorders likely due to meiotic NAHR
| Disorders | NAHR events | Causal genotypes and phenotypes | References |
|---|---|---|---|
| Neurological disorders | |||
| Charcot-Marie-Tooth disease type 1A‡ | 1.4 Mb duplication on 17p |
|
29 |
| Hereditary neuropathy with liability to pressure palsies‡ | 1.4 Mb deletion on 17p |
|
30 |
| Developmental disorders | |||
| 1q21 deletion syndrome‡ | 1.35 deletion on 1q |
|
122 |
| Sotos syndrome‡ | 1.9 Mb deletion on 5q |
|
40 |
| Congenital adrenal hyperplasia§ | 30 kb deletion or gene conversion on 6p |
|
123, 124 |
| Williams-Beuren syndrome‡ | 1.5 Mb deletion on 7q |
|
125 |
| 7q11.23 duplication syndrome‡ | 1.5 Mb duplication on 7q |
|
126 |
| Prader-Willi syndrome‡ | Paternal 4 Mb deletion on 15q |
|
127 |
| Angelman syndrome‡ | Maternal 4 Mb deletion on 15q |
|
127 |
| 15q13.3 deletion syndrome‡ | 680 kb, 1.5 Mb or 3.95 Mb deletion on 15q |
|
128, 129 |
| 15q24 deletion syndrome‡ | 3.9 Mb deletion on 15q |
|
130 |
| Smith-Magenis syndrome‡ | 3.7 Mb deletion on 17p |
|
131 |
| Potocki-Lupski syndrome‡ | 3.7 Mb duplication on 17p |
|
47 |
| 17q21.31 deletion syndrome‡ | 500–650 kb deletion on 17q |
|
132 – 134 |
| Familial isolated growth hormone deficiency, type 1A§ | 6.7 kb deletion on 17q |
|
135 |
| DiGeorge syndrome/velocardiofacial syndrome‡ | 1.5 Mb or 3 Mb deletion on 22q |
|
136 |
| Xp11.22–p11.23 duplication syndrome‡ | Variable-sized duplications on Xp |
|
137 |
| Hunter syndrome* | 20 kb inversion or 30 kb deletion on Xq |
|
138, 139 |
| Metabolic and tissue function disorders | |||
| Bartter syndrome type 3‡ | 11 kb deletion on 1p |
|
140 |
| Gaucher disease§ | 16 kb deletion on 1q |
|
141 |
| Familial juvenile nephronophthisis 1§ | 290 kb deletion on 2q |
|
142 |
| Facioscapulohumeral muscular dystrophy, type 1A‡ | Variable-sized deletions on 4q |
|
143 |
| Glucocorticoid-remediable aldosteronism‡ | 45 kb duplication on 8q |
|
144 |
| Polycystic kidney disease 1‡ | Gene conversion on 16p |
|
145 |
| Neurofibromatosis type 1‡ | 1.5 Mb deletion on 17q |
|
146 |
| 17q12 deletion syndrome‡ | 1.5 Mb deletion on 17q |
|
147 |
| Ichthyosis* | 1.5 Mb deletion on Xp |
|
148 |
| Incontinentia pigmenti* | 10 kb deletion on Xq |
|
149, 150 |
| Red-green colour blindness* | Variable-sized deletions and duplications or gene conversion on Xq |
|
151 |
| Blood diseases | |||
| β-thalassaemia§ | 7 kb deletion on 11p |
|
152 |
| α-thalassaemia□ | 3.7 kb or 4.2 kb deletion on 16p |
|
153 |
| Haemophilia A* | 500 kb or 600 kb inversion on Xq |
|
154 |
| Sex disorders | |||
| Isodicentric Y chromosome formation† | Variable-sized isodicentric Y chromosomes |
|
55 |
| Male infertility or subfertility† | Variable-sized deletions on Yq |
|
56 – 61 |
Mode of inheritance is autosomal dominant,
Mode of inheritance is autosomal recessive,
Mode of inheritance is X-linked
Mode of inheritance is Y-linked.
There are two α globin genes at the α-globin locus. The severity of the thalassaemia is correlated with the number of functional α globin genes. −/ α −/α or −/− α /α: mild anaemia; −/− −/α: severe anaemia; −/− −/−: death in utero or shortly after birth. CHRNA7, neuronal nicotinic acetylcholine receptor; CLCNKB, chloride channel Kb; GBA, glucocerebrosidase; MAPT, microtubule associated protein tau; GH1, growth hormone 1; NF1, neurofibromatosis type 1; PMP22, peripheral myelin protein 22; STS, steroid sulfatase.
The study of genomic disorders has uncovered several parallels between meiotic allelic homologous recombination and NAHR, which suggest that they share common mechanisms and that, by inference, NAHR events occur during prophase of the first meiotic division. First, for some rearrangements caused by NAHR, recombination breakpoints cluster within narrow, 1- to 2-kb hotspots in LCRs, which are reminiscent of allelic recombination hotspots34–40. Interestingly, the allelic recombination hotspot motif CCNCCNTNNCCNC is found in many of these NAHR hotspots25.
Second, allelic crossover hotspots identified by population genetic analysis in the LCRs on chromosome 17p coincide with the hotspots targeted in duplications and deletions that give rise to CMT1A and HNPP, consistent with the hypothesis that allelic homologous recombination and NAHR at these hotspots are both initiated by similarly programmed DSBs41. Hotspots of both allelic and non-allelic recombination also coincide within a set of LCRs on chromosome 17q that undergo recurrent NAHR associated with neurofibromatosis 1, a pleiotropic disorder that includes the development of cutaneous neurofibromas42. However, more general tests of whether allelic recombination and NAHR share crossover hotspots in LCRs is hampered by the difficulty in identifying crossover hotspots in LCRs by population genetic methods because scoring of allelic polymorphisms in one LCR can be confounded by identical sequences in its non-allelic counterpart43.
Third, several studies directly tested the rates of recurrent, reciprocal NAHR-mediated duplications and deletions in DNA isolated from sperm and blood of healthy males (for example, reciprocal duplication and deletion associated with CMT1A and HNPP). One study found that each of the events tested was specific to the germline and that sperm-based frequency estimates of de novo rearrangements encompassed the disease-based estimates. (For example, for the CMT1A duplication, a frequency of 1/23,000–1/79,000 was estimated from sperm analysis and a frequency of 1/23,000–1/41,000 was estimated in the population44.) Further studies showed that LCR-sponsored deletions and duplications at the α-globin gene locus are significantly more common in sperm than in blood45, 46. These findings establish that the NAHR-mediated deletions and duplications under study are detected primarily in cells that have gone through meiosis, which is consistent with the hypothesis that meiotic allelic homologous recombination and NAHR share common mechanisms.
Fourth, for many disorders, the causal NAHR events are predominantly either paternal or maternal. This aspect is congruent with the fact that sex-specific differences in allelic recombination rates at loci throughout the genome are a known feature of allelic meiotic recombination (for example, see REFS. 36, 47) (BOX 1).
Fifth, statistical analysis of the sequence divergence between the LCRs targeted in CMT1A and HNPP uncovered evidence that the repeat sequences have been homogenized by gene conversion. This would be predicted if the repeats were undergoing repeated noncrossover recombination48.
Last, in several HNPP patients, the breakpoint junction in the recombinant LCR remaining after deletion of the intervening sequence displays interspersed patches of sequences from each of the two LCRs. This finding provides evidence of crossover-associated gene conversion, similar to what would occur during allelic recombination35.
Taken together, these results build a strong case that pathological NAHR events in humans are initiated during meiosis by the same type of SPO11-dependent DSBs that initiate normal allelic recombination, although it is possible that at least some NAHR events occur in mitotically dividing cells in the germline. Studies in mice, in which meiotic recombination mutants such as Spo11−/− knockouts are available10, 11, may help resolve this issue.
The Y chromosome: stability and instability by NAHR
Ninety-five percent of the sequence of the human Y chromosome is restricted to males49 (The remaining 5% of its sequence participates in recombination with the X chromosome and is therefore shared with females.). The male-specific region of the Y chromosome (MSY) is unique compared with all other nuclear DNA in that it is haploid and passed on clonally. Until recently, the MSY was thought to not recombine. Because meiotic recombination can purge the genome of deleterious mutations by breaking up linkage groups and segregating otherwise linked loci away from one another, this assumption led some to speculate that, in the absence of recombination, the Y chromosome will disappear in 15 million years50, 51.
At the expense of this `imminent demise' hypothesis, recent sequencing of the human Y chromosome and comparative analyses with primate Y chromosomes have overturned the notion that the MSY does not participate in recombination49, 52, 53. Roughly one third of the MSY is composed of LCRs in direct or inverted orientation49. The most conspicuous of these LCRs are eight palindromes (inverted repeat pairs) that harbour mirror-imaged gene pairs with crucial roles in spermatogenesis49. Ranging in size from 30 kb to 2.9 Mb, the palindromes show >99.9% arm-to-arm sequence identity; such high similarity would normally suggest that they arose on the MSY roughly 200,00 years ago54. However, Y-linked orthologues of these palindromes are found (with equally high arm-to-arm sequence identity) in chimpanzees, indicating that the palindromes predate the human–chimp divergence over 6 millions years ago. Because the arm-to-arm identity of these palindromes in each species is higher than the identity of the palindromes between species (98.6%), it can be inferred that the palindrome arms have been undergoing sequence homogenization since the human-chimp divergence, which would be expected if intrapalindrome NAHR occured repeatedly52. Such concerted evolution, along with evidence in modern human lineages of recent gene conversion in MSY palindromes, led to the hypothesis that the palindromes have been maintained by intrapalindrome arm-to-arm recombination and that such recombination provides a mechanism to preserve the testis-specific genes located there52.
One group55 proposed that this mechanism would come at a cost. They predicted that, if noncrossover resolution of recombination arising from a DSB in a palindrome accounts for intrapalindrome sequence homogenization, then crossover resolution of such a DSB by unequal sister-chromatid exchange should generate an isodicentric Y chromosome (FIG. 2). Indeed, from a group of >2,300 patients, they identified 51 individuals with isodicentric Y chromosomes that had evidently formed by this mechanism. These chromosomes were associated with a wide range of sex-linked reproductive disorders, including failure to produce sperm, anatomic feminization of individuals bearing Y chromosomes and Turner syndrome in women55. Sex reversal is likely caused by the loss of the mitotically unstable isodicentric Y chromosome in the cells of the fetal gonad that determine sex. This model could also account for the features of Turner syndrome observed in some of the individuals.
NAHR in the human Y chromosome is not limited to LCRs in inverted orientation. Several recurrent interstitial deletions ranging in size from 0.8 Mb to 7.7 Mb and associated with reduced spermatogenesis arise by NAHR between LCRs in direct orientation56–61. As predicted by the model of meiotic recombination, reciprocal duplications of these deletions33, 62, evidence of gene conversion63 and hotspots of NAHR56–59 are all observed for Y chromosome-linked LCRs in direct orientation. For at least one pair of LCRs, recurrent reciprocal deletion and duplication events have been shown to be specific to the germline44.
The results of studies of recurrent NAHR events in the human Y chromosome are consistent with the hypothesis that these events are meiotic in origin and are initiated by SPO11-generated DSBs. Furthermore, chromosome-associated complexes of MSH4 (a protein that is required for meiotic recombination in mice and yeast) have been observed on the MSY in cytological studies of spread human spermatocytes64. These findings support the conclusion that meiotic DSBs form on the MSY, although it is not possible from available studies to determine whether DSBs localize to LCRs.
Meiotic NAHR in yeast
The consequences of NAHR in humans emphasize the importance of understanding the molecular mechanisms of this process. Studies in experimentally tractable organisms such as budding yeast have played a key part in addressing this issue. The ease of genetic manipulation of this organism enables facile analysis of genes that are involved in NAHR. Moreover, the ability to recover all four meiotic products (spores) through TETRAD ANALYSIS enables the comprehensive investigation of NAHR-mediated products. Finally, the availability of physical assays to detect directly the DNA intermediates of recombination provides highly detailed views of the molecular steps in this pathway.
Mechanistic parallels to allelic recombination
Meiotic NAHR has been studied by artificially dispersing HETEROALLELES of a gene in various non-allelic positions in the budding yeast genome. Recombination between mutant heteroalleles can restore a wild-type sequence in one of the gene copies, allowing rapid, quantitative analysis of even infrequent recombination events. NAHR between artificially dispersed genes occurs frequently during meiosis (often at 10–100% of allelic frequencies)65–67. Moreover, NAHR in yeast is mechanistically similar to allelic recombination by several criteria. First, NAHR is initiated by Spo11-dependent DSBs68. Second, heteroduplex DNA is formed as an intermediate in NAHR, as assessed by post-meiotic segregation patterns in tetrad analysis69, 70 and by physical assays that directly detect heteroduplexes in genomic DNA extracted from meiotic cells70. Last, the ratio of crossover- to noncrossover-associated gene conversions is similar for allelic recombination and NAHR in at least some instances66.
Nuclear organization of repeat sequences
The frequency of NAHR between artificial repeats is strongly influenced by the spatial organization of the repeated loci in the nucleus65, 67. For example, if repeats are at interstitial locations (≥60 kb away from the closest TELOMERE), on HETEROLOGUOUS CHROMOSOMES (HETEROLOGUES) they recombine approximately 10-fold less efficiently than the same sequences in allelic positions (FIG. 3a). By contrast, repeats dispersed in non-allelic positions on homologues recombine essentially as efficiently as allelic sequences65–67. One interpretation of these findings is that the progressive pairing of homologues during meiosis (discussed below) favours recombination between repeated sequences on homologues by increasing their proximity in the nucleus65, 71. If so, then repeats on heterologues would not benefit from this proximity advantage.
Figure 3. Non-allelic homologous recombination between artificially duplicated repeats in yeast.
a | Non-allelic homologous recombination (NAHR) between interstitial repeats. Artificial repeats (depicted as arrowheads) located interstitially in allelic positions (a) and artificial repeats in non-allelic positions (b) on homologous chromosomes (blue and red) recombine with similar efficiency. By contrast, repeats located on heterologous chromosomes (blue and yellow) recombine ~10-fold less efficiently (c) than allelic sequences. These findings suggest that, compared with repeats on heterologues, repeats on homologues are in a closer proximity, probably through recombination-dependent homologue pairing. b | NAHR between subtelomeric repeats. An artificial repeat located in a subtelomeric region recombines only slightly less efficiently with another subtelomeric repeat (two-six fold less than between allelic repeats on homologues (a)), whether on the opposite chromosome arm (b) of the homologue or on a heterologue (c). The dynamic organization of telomeres during prophase of the first meiotic division (that is, attachment to the nuclear envelope and bouquet formation) might increase the proximity of telomeric regions of different chromosomes and thus influence the frequency of NAHR.
However, the frequency of NAHR is complicated further by the proximity of the artificial repeats to telomeres65, 67. Heteroallelic markers integrated near chromosome ends (<17 kb from each telomere) recombine only two–six-fold less efficiently than allelic sequences, regardless of whether they are on homologues (at opposite chromosome ends) or on heterologues65, 67 (FIG. 3b). During early meiotic prophase, telomeres become physically attached to the inner surface of the nuclear envelope, which is a conserved phenomenon in many organisms, including mammals1, 72. Telomeres then begin rapid, irregular movements and undergo clustering within a small region of the nuclear envelope, in a process known as bouquet formation (reviewed in REFS. 1, 72). Increased proximity driven by this clustering may underlie the increased recombination frequency between non-allelic sequences located near telomeres.
NAHR between natural repeats
The frequency of NAHR between natural repetitive elements is generally lower than NAHR between artificial repeats. The budding yeast genome contains dispersed repetitive elements such as TY ELEMENTS, which are present in ~30–40 copies per haploid genome73. If Ty elements behave similarly to artificial repeats, they could have a marked impact on cell viability. NAHR has been studied between Ty elements engineered to contain different selectable markers and located in allelic or non-allelic positions, as well as between marked Ty elements and endogenous ones74, 75. Allelic Ty elements recombined less frequently than allelic non-Ty sequences, and the frequency of NAHR was dramatically reduced – about 100-fold – compared with allelic recombination73–75. Moreover, recombination products were predominantly noncrossover gene conversions; crossover-associated gene conversions were extremely rare (<1% of recombinants)74.
Similar observations were made in the fission yeast Schizosaccharomyces pombe. tRNA genes, which are another example of dispersed repeat sequences, recombined infrequently and gene conversions were rarely associated with crossovers66. The reasons for the reduced probability of a crossover outcome for NAHR in these two different systems remain to be determined, but possible mechanisms are discussed below.
Insights into meiotic NAHR in mammals
Studies in yeast have shown that meiotic NAHR is mechanistically similar to allelic recombination. The recombination pathways and the players involved are conserved in mammals, reinforcing the view that human germline NAHR is an outcome of aberrant meiotic recombination.
Yeast studies have also shown that the spatial organization of repeats in a nucleus influences NAHR. Non-random nuclear organization of chromosomes has been observed in both somatic and germ cells of mammals, and its importance in the maintenance of genome integrity has begun to be appreciated76. For example, in somatic cells, the frequency of recombinational repair of targeted DSBs varies depending on the location of the homologous sequences that will be used in the recombination reaction: allelic sequences recombine more frequently than nonallelic ones76. Similarly, in mouse germ cells gene conversion frequency between transgenic lacZ heteroalleles varies over a seven-fold range for lacZ transgenes integrated at different positions, indicating that chromosomal location may be a contributing factor to NAHR77. These findings suggest that nuclear organization of repeats may affect the frequency of NAHR in mammals.
Yeast studies have also shown that many naturally occurring repeats show low levels of meiotic NAHR and that crossovers are particularly suppressed. There are as yet no genetic data that directly address this question in mammals. Alu elements are one of the most abundant repetitive elements in the mammalian genome26. Genetic disorders mediated by recombination between non-allelic Alu elements have been identified (reviewed in REF. 78), although it remains unknown whether such recombination occurred during meiosis. In both human and mouse somatic cells, a targeted DSB in a LINE inserted into the genome could be repaired by gene conversion using endogenous copies of the repeat79, 80. Given the prevalence of these multicopy elements in the human genome, understanding their meiotic behaviours is an important challenge.
Restraining NAHR
Because of the substantial impact of NAHR on genome stability, it is not surprising that cells have evolved multiple strategies to suppress or control NAHR. There are several general mechanisms through which NAHR can be discouraged: suppressing Spo11-dependent DSB formation in or near DNA repeats; inhibiting the use of non-allelic homologous templates for recombination and/or favouring the use of allelic templates; and channelling recombination intermediates into pathways that minimize the likelihood of deleterious outcomes. Examples of each type of mechanism have been elucidated and are described below.
Preventing breaks in repeats
The most straightforward way to prevent NAHR is not to form DSBs in susceptible DNA sequences in the first place, and this strategy does indeed seem to operate in many contexts. In budding yeast, the ribosomal DNA (rDNA) contains ~140 tandem repeats of a 9-kb region that encodes ribosomal RNA genes and constitutes almost 10% of the genome81. Despite the large physical distance occupied by this region, interallelic recombination — at least crossover events — occurs 100-fold less frequently than in other regions81, and this low recombination level is largely due to suppression of meiotic DSB formation82. Suppression of DSBs and recombination in the rDNA depends strongly on SIR282, 83, which encodes a histone deacetylase that promotes the formation of a closed, compact chromatin structure in the rDNA and other regions84, 85. DSB formation in budding yeast is favoured in regions with an open (that is, nucleosome free) chromatin structure (BOX 1). Therefore, Sir2 may suppress DSBs in the rDNA in part through the formation of a nucleosomal conformation that is not permissive for Spo11 activity. Other features probably contribute as well. For example, Sir2 mediates the exclusion of Hop1, a chromosome structure protein that promotes DSB formation, from the rDNA86, 87.
Budding yeast chromosome ends contain ~300 base pairs of telomeric repeats and 10–30-kb subtelomeric regions that are enriched in several different repetitive elements88, 89. Subtelomeric regions are evolutionarily dynamic with respect to the composition of these repetitive elements and sequence divergence between them90. For example, the Y′ element, which is adjacent to telomeric repeats on some chromosomes, exhibits only 1% sequence divergence between copies (other subtelomeric repeats show 10–20% divergence on average)88, 91. Mitotic NAHR between Y′ elements has been observed91, suggesting that it contributes to sequence homogenization of Y′ elements88, 90. It remains unknown to what extent Y′ elements undergo meiotic NAHR and whether meiotic telomere clustering (as discussed above) influences the frequency. However, the distal 20 kb of most chromosomes exhibit significantly lower meiotic DSB levels20, 21 and undergo few crossovers23, 89. Similarly to what is observed in the rDNA, DSBs in subtelomeric regions are increased in the absence of Sir2, suggesting that there are active mechanisms to suppress DSBs in this part of the genome82, 92.
As noted above, meiotic NAHR is suppressed between Ty elements. It is possible that, by chance, the regions near the analyzed Ty elements do not undergo frequent DSB formation. However, a more likely explanation is that an active mechanism suppresses DSB formation near Ty elements, because insertion of a Ty element close to a recombination hotspot reduced the frequency of both DSB formation and recombination in this otherwise recombination-active region93. Moreover, insertion of the Ty element converted the chromatin at the hotspot from a open configuration (nuclease hypersensitive) to a closed one (nuclease resistant)93. Although the molecular mechanisms behind these effects are not fully understood, these findings are consistent with the view that recombination of DNA repeats is controlled, at least in part, by establishment of chromatin structures that are not permissive for Spo11-dependent DSB formation.
DSB suppression is likely to be a conserved strategy for minimizing meiotic NAHR for some natural repetitive elements, as DSBs are also rare in the large blocks of repetitive DNA in pericentromeric regions in S. pombe24. Moreover, cytological analyses have shown that recombination-associated protein complexes are rare or absent from the highly repetitive pericentromeric heterochromatin in mice94 and humans64. Although there are no genetic data in mammals to verify that DSB formation is actively inhibited in these regions, it seems likely that DSB suppression in repetitive sequences is a general phenomenon.
Constraining recombination partner choice
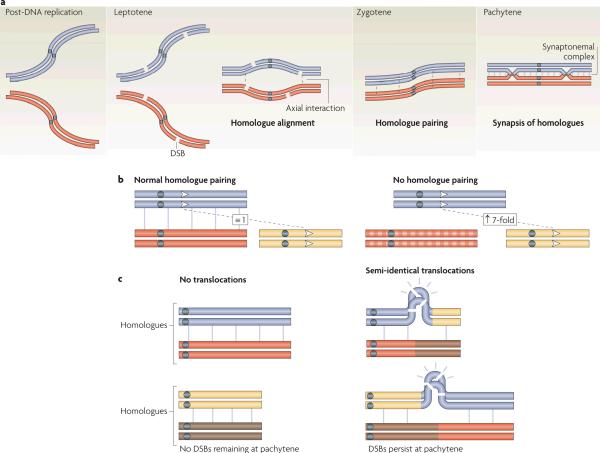
Once a DSB is formed in a repetitive sequence, the choice of homologous template for repair becomes an important factor in determining the likelihood that NAHR will occur. In such circumstances, it seems that the presence of additional recombination events at other locations on the same pair of homologues may help to enforce `good behaviour' of a DSB formed in or near repetitive DNA by promoting the use of an allelic template and/or restricting the use of non-allelic sequences. During early meiotic prophase, homologues find their partners and pair along their entire lengths (FIG. 4a). In many organisms, including fungi (budding yeast and Sordaria), the plant Arabidopsis thaliana and mice, stable homologue pairing depends on DSB formation and early processes of recombination (reviewed in REFS. 1, 72). As an early step in pairing, homologues become aligned at multiple sites. In organisms in which the chromosomes can be easily visualized and studied, alignment is accompanied by the appearance of roughly 400-nm-long bridges between chromosome axes. These bridges are thought to be sites where recombination reactions are in progress (reviewed in REF. 95). As prophase continues, these interactions become increasingly stable, and homologues pair along their entire lengths. In many organisms, homologue pairing culminates in the formation of the SYNAPTONEMAL COMPLEX, a proteinaceous structure that holds homologues together in close juxtaposition (see REFS 96, 97 for details of synaptonemal complex formation and functions of its components). In many fungi, in plants and in mice, neither the initial alignment of homologues nor the subsequent stable pairing interactions are observed in mutants that are defective in DSB formation and strand invasion of DSB ends (for example, spo11Δ and dmc1Δ mutants in yeast)72, 98, emphasizing the importance of early recombination activities in homologue pairing. Note that C. elegans and D. melanogaster use DSB-independent mechanisms for homologue pairing (see REFS. 1, 72).
Figure 4. The influence of homologue pairing on recombination.
a | Double-strand break (DSB)-dependent homologue pairing in budding yeast. After pre-meiotic DNA replication, homologous chromosomes are dispersed in the nucleus. DSBs are formed early in meiotic prophase, providing substrates for recombination proteins to carry out homology search and strand invasion, and resulting in the alignment of homologous chromosome axes at multiple sites (axial interactions)1. As recombination continues, homologue interactions are progressively stabilized, culminating in the formation of the synaptonemal complex96. The development of progressively more stable pairing interactions may inhibit non-allelic homologous recombination (NAHR) by juxtaposing allelic sequences in close proximity. b | The influence of homologue pairing on restraining NAHR. Under normal homologue pairing conditions, NAHR between artificial repeats (indicated by arrowheads) located on one of the paired homologues and on a heterologue occurs at a low level. When pairing between the homologues is disrupted because of sequence divergence between them, the frequency of NAHR between the artificial repeats on heterologues increases ~seven-fold. c | Persistence of recombinational DSBs in unpaired regions of the genome. Mouse chromosomes with semi-identical reciprocal translocations (shown schematically in right pamel) undergo homologous pairing and synapsis along most of their lengths and DSBs are repaired in synapsed regions (left panel), However, DSBs in sequences that are not shared between homologues persist into pachytene despite the presence of identical sequences on heterologues.
In principle, the formation of progressively more stable pairing interactions between homologues could inhibit NAHR through two complementary effects: pairing could favour allelic recombination by placing the allelic copy in close proximity, and pairing may place constraints on the ability of broken DNA segments to find and engage non-allelic sequences that are themselves constrained elsewhere in the nucleus. Elegant experiments in yeast tested this scenario using artificial repeats located on two different chromosomes71 (FIG. 4b). When the homologue of a chromosome carrying a recombination reporter was present (but did not itself carry the recombination reporter), NAHR occurred at a low level. However, when the ability of the homologues to engage in normal pairing was disrupted by substituting their pairing partners with chromosomes from a different yeast strain (≥15% sequence divergence), the frequency of NAHR increased ~seven-fold (FIG. 4b). These findings indicate that homologous pairing in meiosis can contribute substantially to the suppression of NAHR.
There is also evidence that similar effects occur in mammals. An interesting example is provided by mice that carry two different reciprocal translocations between chromosomes 1 and 13 that have slightly different translocation breakpoints99. During meiosis in mice heterozygous for these semi-identical translocations, the two short translocation chromosomes and the two long translocation chromosomes each undergo a separate pairing reaction that culminates in the formation of synaptonemal complexes along most of their lengths. However, at the positions of the translocation breakpoints, the synapsing chromosomes differ from one another and the non-homologous portions form loops that bulge out of the synaptonemal complexes (FIG. 4c). The loop on the larger chromosome pair is homologous to the loop on the smaller chromosome pair. Nevertheless, markers for the presence of unrepaired DSBs were often seen to persist99. A plausible interpretation of this finding is that the pairing and synapsis of homologues imposes a physical constraint that impedes the ability of broken DNA sequences to find homologous templates elsewhere in the genome99.
Controlling the outcome of recombination between divergent sequences
If a DSB forms in a repetitive sequence and a non-allelic homologous partner is then used as template for recombination repair, opportunities still remain to avoid deleterious consequences from NAHR. Specifically, this can be achieved by channelling the recombination events into pathways that give primarily noncrossover outcomes, which do not result in gross chromosomal rearrangements.
One way this may be facilitated is through action of MMR proteins (for example, Pms1 and Msh2 in budding yeast). MMR proteins are conserved among bacteria and eukaryotes and have important roles in mismatch correction during DNA replication and recombination (reviewed in REFS 100, 101). MMR proteins also impede the strand exchange reaction during homologous recombination if a high level of mismatches is present on the heteroduplex DNA (known as heteroduplex rejection)102, which probably promotes noncrossover recombination pathways such as SDSA103–106. Natural repeats (such as retrotransposons) are often more divergent than allelic sequences, thus recombination between non-allelic sequences is expected to generate heteroduplex DNA with more mismatches on average. Although direct evidence is thus far lacking to implicate MMR proteins in crossover suppression during NAHR between natural repeats in yeast and mammals, this is a plausible strategy for stabilizing the genome.
Another means to antagonize NAHR is through the action of any of several helicases that exert anti-recombination activity by disrupting Rad51 or Dmc1 protein-DNA complexes, by dissociating strand exchange intermediates and/or by taking apart Holliday junction recombination intermediates in a manner that disfavours crossover formation. Examples of helicases that can carry one or more of these functions include the RecQ family helicases Sgs1 in yeast and BLM in humans, and the unrelated Srs2 helicase in yeast (reviewed in REFS. 101, 107). Each of these anti-recombinational activities could potentially be involved in preventing deleterious outcomes of meiotic NAHR, although this has yet to be tested.
Perspectives
Recent work has clearly established that homologous recombination between non-allelic repeated sequences in the human genome influences sequence diversity at both the nucleotide and structural levels. It can result in deletions, duplications, inversions and other rearrangements that can lead to human disorders or contribute to architectural polymorphism in the human genome. Paradoxically, NAHR seems to promote both stability and instability of large repeats on the Y chromosome.
Many questions remain to be examined to understand the molecular mechanisms of NAHR. To what extent are the mechanisms of allelic homologous recombination and NAHR similar? Are the DSBs that initiate NAHR part of the cohort of developmentally programmed DSBs intended for resolution by allelic homologous recombination or do they consist of additional, non-programmed DSBs that must nonetheless be repaired? To begin to answer these questions, a high-resolution map of the human meiotic DSB landscape is required. A caveat to the catalogue of 25,000–50,000 human crossover hotspots identified by population genetic analysis is that it fails to capture new hotspots or to mask hotspots that have recently disappeared. Indeed, sperm typing has uncovered emerging crossover hotspots concealed in regions of low historical recombination108, 109, and comparative analysis of the human and chimpanzee genomes has shown that hotspots evolve rapidly110, 111. Therefore historical crossover analysis reveals an incomplete picture of the contemporary recombination map. Maps of meiotic DSB distributions in the human genome will inform us of how DSBs are distributed in repeats such as LCRs and provide further insight into the similarities and differences in the mechanisms of allelic homologous recombination and NAHR.
In addition, studies in experimentally tractable organisms, particularly budding yeast, have begun to flesh out the broad principles behind cellular strategies that have evolved to prevent NAHR, but there is a need to translate experimental findings from yeast to humans. Studies in mice will likely help to bridge this gap but the present inability to study mammalian meiosis in cell culture is a crucial impediment to progress in this area. It will also be interesting to examine, in humans, whether mutants and/or sequence variants in various factors involved in recombination pathways or in the prevention of NAHR prove to be NAHR susceptibility loci. Continuing to examine these mechanisms will provide insights into the events that lead to the genomic disorders and conditions that arise by NAHR in the human genome.
Box 1 | Determinants of meiotic break formation and recombination.
Budding yeast (reviewed in REF. 112)
Meiotic double-strand breaks (DSBs) occur preferentially in intergenic promoter-containing regions rather than in genes.
An open chromatin environment, as assayed by hypersensitivity to nucleases, correlates with increased frequency of recombination, and, at some sites, DSB formation depends on the binding of specific transcription factors. Transcription does not seem to be required for hotspot activity, but there is a weak correlation between the expression level of promoters and their activity as recombination hotspots20.
Although crossovers are suppressed near centromeres, DSBs do form there, indicating that pericentric DSBs may be resolved preferentially by intersister-chromatid recombination20, 23.
DSBs occur infrequently in telomeric regions and retrotransposons.
Trimethylation of lysine 4 of histone H3 is constitutively higher in regions close to DSB sites, and deletion of the Set1 methyltransferase (which generates this methyl mark) dramatically reduces DSB frequency in most hotspots113.
Larger eukaryotes
In humans, crossover rates are lower in genes than in non-coding sequences, based on analysis of a genome-wide catalogue of crossover hotspots inferred from population genetic data17.
The degenerate thirteen-nucleotide motif CCNCCNTNNCCNC accounts for >40% of human hotspots25. This finding was supported by detailed sperm typing studies at one hotspot that contains the motif: a polymorphism that disrupts the consensus sequence was associated with decreased crossover frequencies114.
In the genomes of flies, birds and mammals, there are fewer interspersed repetitive elements in regions of higher recombination115, 116. This may reflect the effects of selection against initiating recombination in such elements because of the potential for NAHR.
A study of recombination across mouse chromosome 1 identified a positive but non-uniform correlation between gene density and crossover rate117.
Several histone modifications are enriched at the known mouse hotspot Psmb9: di- and trimethylation of lysine 4 of histone H3 and acetylation of lysine 9 of histone H3 are thought to be part of the substrate for the initiation of recombination, and H4 hyperacetylation increases after DSB formation by Spo11118.
PRDM9 has recently been identified as an important determinant of meiotic hotspots in humans and mice119, 120. In mouse, PRDM9 has been shown to trimethylate lysine 4 of histone H3 and is expressed specifically in meiotic germ cells121. Both human and mouse PRDM9 have an array of zinc fingers, and human PRDM9 binds specifically to the thirteen-nucleotide consensus motif in vitro. In addition, allelic variation in human PRDM9 correlates with differences in hotspot usage. These findings suggest that the binding of PRDM9 to specific DNA sequences could target recombination to those sites and that allelic variation in PRDM9 could account for variability in hotspot usage between and within species.
Figure 5. Suppression of recombination between divergent sequences.
When a double-strand [see cover letter comment 1] break (DSB) undergoes strand invasion, heteroduplex DNA is formed. When there are few or no mismatched bases, subsequent recombination processes continue. However, when there are many mismatches, mismatch repair (MMR) proteins (such as Pms1 and Msh2) impede strand invasion steps that could lead to crossover formation. Subsequently, heteroduplex DNA might be destabilized (known as heteroduplex rejection) and possibly channelled into the synthesis-dependent strand annealing (SDSA) pathway, which leads to a noncrossover outcome.
Acknowledgements
We thank members of the Keeney lab for comments on the manuscript. Work from the authors' laboratory is supported in part by the Howard Hughes Medical Institute and in part by grants from the US National Institutes of Health.
Glossary
- HOMOLOGOUS CHROMOSOMES (HOMOLOGUES)
In a diploid individual, chromosome pairs that share the same genes, but not necessarily the same alleles, at loci along their lengths
- ANEUPLOIDY
The condition of a cell or organism that is associated with having one or more extra or missing chromosomes, caused by inaccurate chromosome segregation during cell division
- HETERODUPLEX DNA
Hybrid double-stranded DNA that consists of one strand from each homologue; formed by strand invasion of one DSB end into a homologous DNA duplex during recombination
- D-LOOP
A single DNA strand displaced when strand invasion induces the dissociation of a DNA duplex
- HOLLIDAY JUNCTION
Four-stranded, branched DNA structure formed as an intermediate in homologous recombination
- GENE CONVERSION
Transfer of DNA sequence information that “overwrites” the sequence of one allele with the sequence of the other allele; carried out by mismatch repair on heteroduplex DNA
- MISMATCH REPAIR
The repair system that recognizes and corrects mismatches that form during DNA replication and recombination
- ISODICENTRIC CHROMOSOME
A mirror-imaged duplication of part of a chromosome, including the centromere
- GENOMIC DISORDERS
Conditions arising in the germline by submicroscopic, regional chromosomal rearrangements that alter the copy number of dosage-sensitive genes, disrupt genes or generate fusion genes
- CHARCOT-MARIE-TOOTH DISEASE TYPE 1A
Inherited disorder of the nerves that results in weakness and atrophy of muscles of the lower legs and hands
- HEREDITARY NEUROPATHY WITH LIABILITY TO PRESSURE PALSIES
Inherited neuropathy characterized by recurrent episodes of numbness, tingling and/or weakening and atrophy of muscles, most commonly in wrists, elbows and knees
- TETRAD ANALYSIS
Tetrads refer to all four products (spores in yeast) of a single meiosis, which are grouped together in an ascus. Spores in a tetrad be individually isolated by micromanipulation and can be grown in culture for further analysis
- HETEROALLELES
Two different alleles of a gene, each carrying a mutation(s) at a different position within the coding sequence
- TELOMERE
segment at the end of each chromosome arm that consists of repetitive sequences and that prevents the normal chromosome ends from being recognized as a double-strand break
- HETEROLOGOUS CHROMOSOMES (HETEROLOGUES)
Chromosomes that are neither homologues nor sister chromatids
- TY ELEMENT
Retrotransposon in yeast, about 5–6 kb in length, comprising a central domain that is flanked by ~330 bp of long terminal repeats (LTRs) The yeast genome contains full Ty elements as well as solo LTRs and LTR fragments
- SYNAPTONEMAL COMPLEX
A proteinaceous structure that is formed between homologues during meiosis Synaptonemal complexes are composed of lateral elements that are assembled between the entire length of sister chromatids, and transverse filaments that connect lateral elements to the central element. Within the context of synaptonemal complexes, homologues are intimately connected
References
- 1.Gerton JL, Hawley RS. Homologous chromosome interactions in meiosis: diversity amidst conservation. Nat Rev Genet. 2005;6:477–87. doi: 10.1038/nrg1614. [DOI] [PubMed] [Google Scholar]; Review of interactions between homologous chromosomes during meiosis.
- 2.Hassold T, Hall H, Hunt P. The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet. 2007;16(Spec No. 2):R203–8. doi: 10.1093/hmg/ddm243. [DOI] [PubMed] [Google Scholar]
- 3.Goffeau A, et al. Life with 6000 genes. Science. 1996;274:546, 563–7. doi: 10.1126/science.274.5287.546. [DOI] [PubMed] [Google Scholar]
- 4.Bailey JA, et al. Recent segmental duplications in the human genome. Science. 2002;297:1003–7. doi: 10.1126/science.1072047. [DOI] [PubMed] [Google Scholar]; This paper is one of the first to report on the extent of low copy repeats in the human genome.
- 5.Keeney S. In: Recombination and Meiosis: Crossing-Over and Disjunction. Lankenau DH, editor. Springer-Verlag; Heidelberg: 2007. pp. 81–123. [Google Scholar]
- 6.Hunter N. In: Topics in Current Genetics, Molecular Genetics of Recombination. Aguilera A, Rothstein R, editors. Springer-Verlag; Heidelberg: 2006. pp. 381–442. [Google Scholar]
- 7.Bishop DK, Zickler D. Early decision; meiotic crossover interference prior to stable strand exchange and synapsis. Cell. 2004;117:9–15. doi: 10.1016/s0092-8674(04)00297-1. [DOI] [PubMed] [Google Scholar]
- 8.Dernburg AF, et al. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell. 1998;94:387–98. doi: 10.1016/s0092-8674(00)81481-6. [DOI] [PubMed] [Google Scholar]
- 9.McKim KS, et al. Meiotic synapsis in the absence of recombination. Science. 1998;279:876–8. doi: 10.1126/science.279.5352.876. [DOI] [PubMed] [Google Scholar]
- 10.Baudat F, Manova K, Yuen JP, Jasin M, Keeney S. Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell. 2000;6:989–98. doi: 10.1016/s1097-2765(00)00098-8. [DOI] [PubMed] [Google Scholar]
- 11.Romanienko PJ, Camerini-Otero RD. The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell. 2000;6:975–87. doi: 10.1016/s1097-2765(00)00097-6. [DOI] [PubMed] [Google Scholar]
- 12.Yoshida K, et al. The mouse RecA-like gene Dmc1 is required for homologous chromosome synapsis during meiosis. Mol Cell. 1998;1:707–18. doi: 10.1016/s1097-2765(00)80070-2. [DOI] [PubMed] [Google Scholar]
- 13.Pittman DL, et al. Meiotic prophase arrest with failure of chromosome synapsis in mice deficient for Dmc1, a germline-specific RecA homolog. Mol Cell. 1998;1:697–705. doi: 10.1016/s1097-2765(00)80069-6. [DOI] [PubMed] [Google Scholar]
- 14.Kauppi L, Jeffreys AJ, Keeney S. Where the crossovers are: recombination distributions in mammals. Nat Rev Genet. 2004;5:413–24. doi: 10.1038/nrg1346. [DOI] [PubMed] [Google Scholar]; This review examines the distribution of recombination in humans and mice in comparison with results in budding yeast, and discusses its implications of recombination hotspots in genome evolution.
- 15.Kong A, et al. A high-resolution recombination map of the human genome. Nat Genet. 2002;31:241–7. doi: 10.1038/ng917. [DOI] [PubMed] [Google Scholar]
- 16.Lawrie NM, Tease C, Hulten MA. Chiasma frequency, distribution and interference maps of mouse autosomes. Chromosoma. 1995;104:308–14. doi: 10.1007/BF00352262. [DOI] [PubMed] [Google Scholar]
- 17.Myers S, Bottolo L, Freeman C, McVean G, Donnelly P. A fine-scale map of recombination rates and hotspots across the human genome. Science. 2005;310:321–4. doi: 10.1126/science.1117196. [DOI] [PubMed] [Google Scholar]
- 18.Baudat F, Nicolas A. Clustering of meiotic double-strand breaks on yeast chromosome III. Proc Natl Acad Sci U S A. 1997;94:5213–8. doi: 10.1073/pnas.94.10.5213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gerton JL, et al. Inaugural article: global mapping of meiotic recombination hotspots and coldspots in the yeast Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 2000;97:11383–90. doi: 10.1073/pnas.97.21.11383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Blitzblau HG, Bell GW, Rodriguez J, Bell SP, Hochwagen A. Mapping of meiotic single-stranded DNA reveals double-stranded-break hotspots near centromeres and telomeres. Curr Biol. 2007;17:2003–12. doi: 10.1016/j.cub.2007.10.066. [DOI] [PubMed] [Google Scholar]; With reference 21, this article presents a high-resolution, genome-wide map of meiotic DSB hotspots in budding yeast.
- 21.Buhler C, Borde V, Lichten M. Mapping meiotic single-strand DNA reveals a new landscape of DNA double-strand breaks in Saccharomyces cerevisiae. PLoS Biol. 2007;5:e324. doi: 10.1371/journal.pbio.0050324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mancera E, Bourgon R, Brozzi A, Huber W, Steinmetz LM. High-resolution mapping of meiotic crossovers and non-crossovers in yeast. Nature. 2008;454:479–85. doi: 10.1038/nature07135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen SY, et al. Global analysis of the meiotic crossover landscape. Dev Cell. 2008;15:401–15. doi: 10.1016/j.devcel.2008.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cromie GA, et al. A discrete class of intergenic DNA dictates meiotic DNA break hotspots in fission yeast. PLoS Genet. 2007;3:e141. doi: 10.1371/journal.pgen.0030141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Myers S, Freeman C, Auton A, Donnelly P, McVean G. A common sequence motif associated with recombination hot spots and genome instability in humans. Nat Genet. 2008;40:1124–9. doi: 10.1038/ng.213. [DOI] [PubMed] [Google Scholar]; This paper reports the identification of a 13-nucleotide DNA sequence motif that is present in >40% of all human allelic recombination hotspots, as well as in several NAHR hotspots.
- 26.Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 27.Bailey JA, Eichler EE. Primate segmental duplications: crucibles of evolution, diversity and disease. Nat Rev Genet. 2006;7:552–64. doi: 10.1038/nrg1895. [DOI] [PubMed] [Google Scholar]
- 28.Zhang F, Gu W, Hurles ME, Lupski JR. Copy number variation in human health, disease, and evolution. Annu Rev Genomics Hum Genet. 2009;10:451–81. doi: 10.1146/annurev.genom.9.081307.164217. [DOI] [PMC free article] [PubMed] [Google Scholar]; A comprehensive review of copy number variation in the human genome, including mechanisms of formation and the role of variants in disease.
- 29.Pentao L, Wise CA, Chinault AC, Patel PI, Lupski JR. Charcot-Marie-Tooth type 1A duplication appears to arise from recombination at repeat sequences flanking the 1.5 Mb monomer unit. Nat Genet. 1992;2:292–300. doi: 10.1038/ng1292-292. [DOI] [PubMed] [Google Scholar]
- 30.Chance PF, et al. Two autosomal dominant neuropathies result from reciprocal DNA duplication/deletion of a region on chromosome 17. Hum Mol Genet. 1994;3:223–8. doi: 10.1093/hmg/3.2.223. [DOI] [PubMed] [Google Scholar]
- 31.Tuzun E, et al. Fine-scale structural variation of the human genome. Nat Genet. 2005;37:727–32. doi: 10.1038/ng1562. [DOI] [PubMed] [Google Scholar]
- 32.Redon R, et al. Global variation in copy number in the human genome. Nature. 2006;444:444–54. doi: 10.1038/nature05329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Repping S, et al. High mutation rates have driven extensive structural polymorphism among human Y chromosomes. Nat Genet. 2006;38:463–467. doi: 10.1038/ng1754. [DOI] [PubMed] [Google Scholar]
- 34.Reiter LT, et al. A recombination hotspot responsible for two inherited peripheral neuropathies is located near a mariner transposon-like element. Nat Genet. 1996;12:288–97. doi: 10.1038/ng0396-288. [DOI] [PubMed] [Google Scholar]
- 35.Reiter LT, et al. Human meiotic recombination products revealed by sequencing a hotspot for homologous strand exchange in multiple HNPP deletion patients. Am J Hum Genet. 1998;62:1023–33. doi: 10.1086/301827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lopez-Correa C, et al. Recombination hotspot in NF1 microdeletion patients. Hum Mol Genet. 2001;10:1387–92. doi: 10.1093/hmg/10.13.1387. [DOI] [PubMed] [Google Scholar]
- 37.Jenne DE, et al. Molecular characterization and gene content of breakpoint boundaries in patients with neurofibromatosis type 1 with 17q11.2 microdeletions. Am J Hum Genet. 2001;69:516–27. doi: 10.1086/323043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Bayes M, Magano LF, Rivera N, Flores R, Perez Jurado LA. Mutational mechanisms of Williams-Beuren syndrome deletions. Am J Hum Genet. 2003;73:131–51. doi: 10.1086/376565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bi W, et al. Reciprocal crossovers and a positional preference for strand exchange in recombination events resulting in deletion or duplication of chromosome 17p11.2. Am J Hum Genet. 2003;73:1302–15. doi: 10.1086/379979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kurotaki N, Stankiewicz P, Wakui K, Niikawa N, Lupski JR. Sotos syndrome common deletion is mediated by directly oriented subunits within inverted Sos-REP low-copy repeats. Hum Mol Genet. 2005;14:535–42. doi: 10.1093/hmg/ddi050. [DOI] [PubMed] [Google Scholar]
- 41.Lindsay SJ, Khajavi M, Lupski JR, Hurles ME. A chromosomal rearrangement hotspot can be identified from population genetic variation and is coincident with a hotspot for allelic recombination. Am J Hum Genet. 2006;79:890–902. doi: 10.1086/508709. [DOI] [PMC free article] [PubMed] [Google Scholar]; With reference 42, this article demonstrates the coincidence allelic recombination hotspots and NAHR hotspots.
- 42.De Raedt T, et al. Conservation of hotspots for recombination in low-copy repeats associated with the NF1 microdeletion. Nat Genet. 2006;38:1419–23. doi: 10.1038/ng1920. [DOI] [PubMed] [Google Scholar]
- 43.Frazer KA, et al. A second generation human haplotype map of over 3.1 million SNPs. Nature. 2007;449:851–61. doi: 10.1038/nature06258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Turner DJ, et al. Germline rates of de novo meiotic deletions and duplications causing several genomic disorders. Nat Genet. 2008;40:90–5. doi: 10.1038/ng.2007.40. [DOI] [PMC free article] [PubMed] [Google Scholar]; This paper compares the rates of NAHR events in the germline and soma, and demonstrates that the events examined are specific to the germline.
- 45.Lam KW, Jeffreys AJ. Processes of copy-number change in human DNA: the dynamics of {alpha}-globin gene deletion. Proc Natl Acad Sci U S A. 2006;103:8921–7. doi: 10.1073/pnas.0602690103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lam KW, Jeffreys AJ. Processes of de novo duplication of human alpha-globin genes. Proc Natl Acad Sci U S A. 2007;104:10950–5. doi: 10.1073/pnas.0703856104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Potocki L, et al. Molecular mechanism for duplication 17p11.2- the homologous recombination reciprocal of the Smith-Magenis microdeletion. Nat Genet. 2000;24:84–7. doi: 10.1038/71743. [DOI] [PubMed] [Google Scholar]
- 48.Hurles ME. Gene conversion homogenizes the CMT1A paralogous repeats. BMC Genomics. 2001;2:11. doi: 10.1186/1471-2164-2-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Skaletsky H, et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature. 2003;423:825–37. doi: 10.1038/nature01722. [DOI] [PubMed] [Google Scholar]
- 50.Aitken RJ, Marshall Graves JA. The future of sex. Nature. 2002;415:963. doi: 10.1038/415963a. [DOI] [PubMed] [Google Scholar]
- 51.Graves JA, Koina E, Sankovic N. How the gene content of human sex chromosomes evolved. Curr Opin Genet Dev. 2006;16:219–24. doi: 10.1016/j.gde.2006.04.007. [DOI] [PubMed] [Google Scholar]
- 52.Rozen S, et al. Abundant gene conversion between arms of palindromes in human and ape Y chromosomes. Nature. 2003;423:873–6. doi: 10.1038/nature01723. [DOI] [PubMed] [Google Scholar]
- 53.Hughes JF, et al. Conservation of Y-linked genes during human evolution revealed by comparative sequencing in chimpanzee. Nature. 2005;437:100–3. doi: 10.1038/nature04101. [DOI] [PubMed] [Google Scholar]
- 54.Agulnik AI, et al. Evolution of the DAZ gene family suggests that Y-linked DAZ plays little, or a limited, role in spermatogenesis but underlines a recent African origin for human populations. Hum Mol Genet. 1998;7:1371–7. doi: 10.1093/hmg/7.9.1371. [DOI] [PubMed] [Google Scholar]
- 55.Lange J, et al. Isodicentric Y chromosomes and sex disorders as byproducts of homologous recombination that maintains palindromes. Cell. 2009;138:855–69. doi: 10.1016/j.cell.2009.07.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Sun C, et al. Deletion of azoospermia factor a (AZFa) region of human Y chromosome caused by recombination between HERV15 proviruses. Hum Mol Genet. 2000;9:2291–6. doi: 10.1093/oxfordjournals.hmg.a018920. [DOI] [PubMed] [Google Scholar]
- 57.Kamp C, Hirschmann P, Voss H, Huellen K, Vogt PH. Two long homologous retroviral sequence blocks in proximal Yq11 cause AZFa microdeletions as a result of intrachromosomal recombination events. Hum Mol Genet. 2000;9:2563–72. doi: 10.1093/hmg/9.17.2563. [DOI] [PubMed] [Google Scholar]
- 58.Blanco P, et al. Divergent outcomes of intrachromosomal recombination on the human Y chromosome: male infertility and recurrent polymorphism. J Med Genet. 2000;37:752–8. doi: 10.1136/jmg.37.10.752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Repping S, et al. Recombination between palindromes P5 and P1 on the human Y chromosome causes massive deletions and spermatogenic failure. Am J Hum Genet. 2002;71:906–22. doi: 10.1086/342928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kuroda-Kawaguchi T, et al. The AZFc region of the Y chromosome features massive palindromes and uniform recurrent deletions in infertile men. Nat Genet. 2001;29:279–86. doi: 10.1038/ng757. [DOI] [PubMed] [Google Scholar]
- 61.Repping S, et al. Polymorphism for a 1.6-Mb deletion of the human Y chromosome persists through balance between recurrent mutation and haploid selection. Nat Genet. 2003;35:247–51. doi: 10.1038/ng1250. [DOI] [PubMed] [Google Scholar]
- 62.Bosch E, Jobling MA. Duplications of the AZFa region of the human Y chromosome are mediated by homologous recombination between HERVs and are compatible with male fertility. Hum Mol Genet. 2003;12:341–7. doi: 10.1093/hmg/ddg031. [DOI] [PubMed] [Google Scholar]
- 63.Bosch E, Hurles ME, Navarro A, Jobling MA. Dynamics of a human interparalog gene conversion hotspot. Genome Res. 2004;14:835–44. doi: 10.1101/gr.2177404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Oliver-Bonet M, Turek PJ, Sun F, Ko E, Martin RH. Temporal progression of recombination in human males. Mol Hum Reprod. 2005;11:517–22. doi: 10.1093/molehr/gah193. [DOI] [PubMed] [Google Scholar]
- 65.Goldman AS, Lichten M. The efficiency of meiotic recombination between dispersed sequences in Saccharomyces cerevisiae depends upon their chromosomal location. Genetics. 1996;144:43–55. doi: 10.1093/genetics/144.1.43. [DOI] [PMC free article] [PubMed] [Google Scholar]; With reference 65, this paper demonstrates that the spatial organization of repeated sequences within a nucleus influences the frequency of meiotic NAHR in budding yeast.
- 66.Petes TD, Hill CW. Recombination between repeated genes in microorganisms. Annu Rev Genet. 1988;22:147–68. doi: 10.1146/annurev.ge.22.120188.001051. [DOI] [PubMed] [Google Scholar]
- 67.Schlecht HB, Lichten M, Goldman AS. Compartmentalization of the yeast meiotic nucleus revealed by analysis of ectopic recombination. Genetics. 2004;168:1189–203. doi: 10.1534/genetics.104.029157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Steele DF, Morris ME, Jinks-Robertson S. Allelic and ectopic interactions in recombination-defective yeast strains. Genetics. 1991;127:53–60. doi: 10.1093/genetics/127.1.53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Nag DK, Petes TD. Meiotic recombination between dispersed repeated genes is associated with heteroduplex formation. Mol Cell Biol. 1990;10:4420–3. doi: 10.1128/mcb.10.8.4420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Allers T, Lichten M. Differential timing and control of noncrossover and crossover recombination during meiosis. Cell. 2001;106:47–57. doi: 10.1016/s0092-8674(01)00416-0. [DOI] [PubMed] [Google Scholar]
- 71.Goldman AS, Lichten M. Restriction of ectopic recombination by interhomolog interactions during Saccharomyces cerevisiae meiosis. Proc Natl Acad Sci U S A. 2000;97:9537–42. doi: 10.1073/pnas.97.17.9537. [DOI] [PMC free article] [PubMed] [Google Scholar]; This study provides strong evidence that homologue pairing restricts meiotic NAHR between artificial repeats in budding yeast.
- 72.Bhalla N, Dernburg AF. Prelude to a division. Annu Rev Cell Dev Biol. 2008;24:397–424. doi: 10.1146/annurev.cellbio.23.090506.123245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Mieczkowski PA, Lemoine FJ, Petes TD. Recombination between retrotransposons as a source of chromosome rearrangements in the yeast Saccharomyces cerevisiae. DNA Repair (Amst) 2006;5:1010–20. doi: 10.1016/j.dnarep.2006.05.027. [DOI] [PubMed] [Google Scholar]
- 74.Kupiec M, Petes TD. Meiotic recombination between repeated transposable elements in Saccharomyces cerevisiae. Mol Cell Biol. 1988;8:2942–54. doi: 10.1128/mcb.8.7.2942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Kupiec M, Petes TD. Allelic and ectopic recombination between Ty elements in yeast. Genetics. 1988;119:549–59. doi: 10.1093/genetics/119.3.549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Meaburn KJ, Misteli T, Soutoglou E. Spatial genome organization in the formation of chromosomal translocations. Semin Cancer Biol. 2007;17:80–90. doi: 10.1016/j.semcancer.2006.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Murti JR, Bumbulis M, Schimenti JC. Gene conversion between unlinked sequences in the germline of mice. Genetics. 1994;137:837–43. doi: 10.1093/genetics/137.3.837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Batzer MA, Deininger PL. Alu repeats and human genomic diversity. Nat Rev Genet. 2002;3:370–9. doi: 10.1038/nrg798. [DOI] [PubMed] [Google Scholar]
- 79.Tremblay A, Jasin M, Chartrand P. A double-strand break in a chromosomal LINE element can be repaired by gene conversion with various endogenous LINE elements in mouse cells. Mol Cell Biol. 2000;20:54–60. doi: 10.1128/mcb.20.1.54-60.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.D'Anjou H, Chabot C, Chartrand P. Preferential accessibility to specific genomic loci for the repair of double-strand breaks in human cells. Nucleic Acids Res. 2004;32:6136–43. doi: 10.1093/nar/gkh952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Eickbush TH, Eickbush DG. Finely orchestrated movements: evolution of the ribosomal RNA genes. Genetics. 2007;175:477–85. doi: 10.1534/genetics.107.071399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Mieczkowski PA, Dominska M, Buck MJ, Lieb JD, Petes TD. Loss of a histone deacetylase dramatically alters the genomic distribution of Spo11p-catalyzed DNA breaks in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 2007;104:3955–60. doi: 10.1073/pnas.0700412104. [DOI] [PMC free article] [PubMed] [Google Scholar]; Using a microarray-based approach to map DSBs genome-wide in budding yeast, this study demonstrates that meiotic double-strand breaks are suppressed in subtelomeric regions and in the rDNA in a Sir2-dependent manner.
- 83.Gottlieb S, Esposito RE. A new role for a yeast transcriptional silencer gene, SIR2, in regulation of recombination in ribosomal DNA. Cell. 1989;56:771–6. doi: 10.1016/0092-8674(89)90681-8. [DOI] [PubMed] [Google Scholar]
- 84.Blander G, Guarente L. The Sir2 family of protein deacetylases. Annu Rev Biochem. 2004;73:417–35. doi: 10.1146/annurev.biochem.73.011303.073651. [DOI] [PubMed] [Google Scholar]
- 85.Fritze CE, Verschueren K, Strich R, Easton Esposito R. Direct evidence for SIR2 modulation of chromatin structure in yeast rDNA. EMBO J. 1997;16:6495–509. doi: 10.1093/emboj/16.21.6495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.San-Segundo PA, Roeder GS. Pch2 links chromatin silencing to meiotic checkpoint control. Cell. 1999;97:313–24. doi: 10.1016/s0092-8674(00)80741-2. [DOI] [PubMed] [Google Scholar]
- 87.Smith AV, Roeder GS. The yeast Red1 protein localizes to the cores of meiotic chromosomes. J Cell Biol. 1997;136:957–67. doi: 10.1083/jcb.136.5.957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Louis EJ. The chromosome ends of Saccharomyces cerevisiae. Yeast. 1995;11:1553–73. doi: 10.1002/yea.320111604. [DOI] [PubMed] [Google Scholar]
- 89.Barton AB, Pekosz MR, Kurvathi RS, Kaback DB. Meiotic recombination at the ends of chromosomes in Saccharomyces cerevisiae. Genetics. 2008;179:1221–35. doi: 10.1534/genetics.107.083493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Louis EJ, Naumova ES, Lee A, Naumov G, Haber JE. The chromosome end in yeast: its mosaic nature and influence on recombinational dynamics. Genetics. 1994;136:789–802. doi: 10.1093/genetics/136.3.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Louis EJ, Haber JE. Mitotic recombination among subtelomeric Y' repeats in Saccharomyces cerevisiae. Genetics. 1990;124:547–59. doi: 10.1093/genetics/124.3.547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Lichten M. In: Recombination and Meiosis: Models, Means, and Evolution. Lankenau DH, editor. Springer-Verlag; Heidelberg: 2008. pp. 165–193. [Google Scholar]
- 93.Ben-Aroya S, Mieczkowski PA, Petes TD, Kupiec M. The compact chromatin structure of a Ty repeated sequence suppresses recombination hotspot activity in Saccharomyces cerevisiae. Mol Cell. 2004;15:221–31. doi: 10.1016/j.molcel.2004.06.002. [DOI] [PubMed] [Google Scholar]
- 94.Anderson LK, Reeves A, Webb LM, Ashley T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics. 1999;151:1569–79. doi: 10.1093/genetics/151.4.1569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Kleckner N. Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma. 2006;115:175–94. doi: 10.1007/s00412-006-0055-7. [DOI] [PubMed] [Google Scholar]
- 96.Page SL, Hawley RS. The genetics and molecular biology of the synaptonemal complex. Annu Rev Cell Dev Biol. 2004;20:525–58. doi: 10.1146/annurev.cellbio.19.111301.155141. [DOI] [PubMed] [Google Scholar]
- 97.Anuradha S, Muniyappa K. Molecular aspects of meiotic chromosome synapsis and recombination. Prog Nucleic Acid Res Mol Biol. 2005;79:49–132. doi: 10.1016/S0079-6603(04)79002-9. [DOI] [PubMed] [Google Scholar]
- 98.Burgess SM. Homologous chromosome associations and nuclear order in meiotic and mitotically dividing cells of budding yeast. Adv Genet. 2002;46:49–90. doi: 10.1016/s0065-2660(02)46004-x. [DOI] [PubMed] [Google Scholar]
- 99.Mahadevaiah SK, et al. Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet. 2001;27:271–6. doi: 10.1038/85830. [DOI] [PubMed] [Google Scholar]
- 100.Schofield MJ, Hsieh P. DNA mismatch repair: molecular mechanisms and biological function. Annu Rev Microbiol. 2003;57:579–608. doi: 10.1146/annurev.micro.57.030502.090847. [DOI] [PubMed] [Google Scholar]
- 101.Surtees JA, Argueso JL, Alani E. Mismatch repair proteins: key regulators of genetic recombination. Cytogenet Genome Res. 2004;107:146–59. doi: 10.1159/000080593. [DOI] [PubMed] [Google Scholar]; A review that discusses roles of MMR proteins in repairing mismatches during meiotic recombination as well as their roles in suppressing recombination between divergent sequences.
- 102.Worth L, Jr., Clark S, Radman M, Modrich P. Mismatch repair proteins MutS and MutL inhibit RecA-catalyzed strand transfer between diverged DNAs. Proc Natl Acad Sci U S A. 1994;91:3238–41. doi: 10.1073/pnas.91.8.3238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Borts RH, Haber JE. Meiotic recombination in yeast: alteration by multiple heterozygosities. Science. 1987;237:1459–65. doi: 10.1126/science.2820060. [DOI] [PubMed] [Google Scholar]
- 104.Borts RH, et al. Mismatch repair-induced meiotic recombination requires the pms1 gene product. Genetics. 1990;124:573–84. doi: 10.1093/genetics/124.3.573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Hunter N, Chambers SR, Louis EJ, Borts RH. The mismatch repair system contributes to meiotic sterility in an interspecific yeast hybrid. EMBO J. 1996;15:1726–33. [PMC free article] [PubMed] [Google Scholar]
- 106.Elliott B, Jasin M. Repair of double-strand breaks by homologous recombination in mismatch repair-defective mammalian cells. Mol Cell Biol. 2001;21:2671–82. doi: 10.1128/MCB.21.8.2671-2682.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Hunter N. The RecQ DNA helicases: Jacks-of-all-trades or master-tradesmen? Cell Res. 2008;18:328–30. doi: 10.1038/cr.2008.33. [DOI] [PubMed] [Google Scholar]
- 108.Jeffreys AJ, Neumann R, Panayi M, Myers S, Donnelly P. Human recombination hot spots hidden in regions of strong marker association. Nat Genet. 2005;37:601–6. doi: 10.1038/ng1565. [DOI] [PubMed] [Google Scholar]
- 109.Kauppi L, Stumpf MP, Jeffreys AJ. Localized breakdown in linkage disequilibrium does not always predict sperm crossover hot spots in the human MHC class II region. Genomics. 2005;86:13–24. doi: 10.1016/j.ygeno.2005.03.011. [DOI] [PubMed] [Google Scholar]
- 110.Winckler W, et al. Comparison of fine-scale recombination rates in humans and chimpanzees. Science. 2005;308:107–11. doi: 10.1126/science.1105322. [DOI] [PubMed] [Google Scholar]
- 111.Ptak SE, et al. Fine-scale recombination patterns differ between chimpanzees and humans. Nat Genet. 2005;37:429–34. doi: 10.1038/ng1529. [DOI] [PubMed] [Google Scholar]
- 112.Petes TD. Meiotic recombination hot spots and cold spots. Nat Rev Genet. 2001;2:360–9. doi: 10.1038/35072078. [DOI] [PubMed] [Google Scholar]
- 113.Borde V, et al. Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J. 2009;28:99–111. doi: 10.1038/emboj.2008.257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Jeffreys AJ, Neumann R. Reciprocal crossover asymmetry and meiotic drive in a human recombination hot spot. Nat Genet. 2002;31:267–71. doi: 10.1038/ng910. [DOI] [PubMed] [Google Scholar]
- 115.Jensen-Seaman MI, et al. Comparative recombination rates in the rat, mouse, and human genomes. Genome Res. 2004;14:528–38. doi: 10.1101/gr.1970304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Hillier LW, et al. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature. 2004;432:695–716. doi: 10.1038/nature03154. [DOI] [PubMed] [Google Scholar]
- 117.Paigen K, et al. The recombinational anatomy of a mouse chromosome. PLoS Genet. 2008;4:e1000119. doi: 10.1371/journal.pgen.1000119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Buard J, Barthes P, Grey C, de Massy B. Distinct histone modifications define initiation and repair of meiotic recombination in the mouse. EMBO J. 2009;28:2616–24. doi: 10.1038/emboj.2009.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Baudat F, et al. PRDM9 Is a Major Determinant of Meiotic Recombination Hotspots in Humans and Mice. Science. 2009 doi: 10.1126/science.1183439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Myers S, et al. Drive Against Hotspot Motifs in Primates Implicates the PRDM9 Gene in Meiotic Recombination. Science. 2009 doi: 10.1126/science.1182363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Hayashi K, Yoshida K, Matsui Y. A histone H3 methyltransferase controls epigenetic events required for meiotic prophase. Nature. 2005;438:374–8. doi: 10.1038/nature04112. [DOI] [PubMed] [Google Scholar]
- 122.Mefford HC, et al. Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N Engl J Med. 2008;359:1685–99. doi: 10.1056/NEJMoa0805384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.White PC, Vitek A, Dupont B, New MI. Characterization of frequent deletions causing steroid 21-hydroxylase deficiency. Proc Natl Acad Sci U S A. 1988;85:4436–40. doi: 10.1073/pnas.85.12.4436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Tusie-Luna MT, White PC. Gene conversions and unequal crossovers between CYP21 (steroid 21-hydroxylase gene) and CYP21P involve different mechanisms. Proc Natl Acad Sci U S A. 1995;92:10796–800. doi: 10.1073/pnas.92.23.10796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Urban Z, et al. 7q11.23 deletions in Williams syndrome arise as a consequence of unequal meiotic crossover. Am J Hum Genet. 1996;59:958–62. [PMC free article] [PubMed] [Google Scholar]
- 126.Somerville MJ, et al. Severe expressive-language delay related to duplication of the Williams-Beuren locus. N Engl J Med. 2005;353:1694–701. doi: 10.1056/NEJMoa051962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Carrozzo R, et al. Inter- and intrachromosomal rearrangements are both involved in the origin of 15q11–q13 deletions in Prader-Willi syndrome. Am J Hum Genet. 1997;61:228–31. doi: 10.1086/513907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Sharp AJ, et al. A recurrent 15q13.3 microdeletion syndrome associated with mental retardation and seizures. Nat Genet. 2008;40:322–8. doi: 10.1038/ng.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Shinawi M, et al. A small recurrent deletion within 15q13.3 is associated with a range of neurodevelopmental phenotypes. Nat Genet. 2009;41:1269–71. doi: 10.1038/ng.481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Sharp AJ, et al. Characterization of a recurrent 15q24 microdeletion syndrome. Hum Mol Genet. 2007;16:567–72. doi: 10.1093/hmg/ddm016. [DOI] [PubMed] [Google Scholar]
- 131.Chen KS, et al. Homologous recombination of a flanking repeat gene cluster is a mechanism for a common contiguous gene deletion syndrome. Nat Genet. 1997;17:154–63. doi: 10.1038/ng1097-154. [DOI] [PubMed] [Google Scholar]
- 132.Koolen DA, et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet. 2006;38:999–1001. doi: 10.1038/ng1853. [DOI] [PubMed] [Google Scholar]
- 133.Shaw-Smith C, et al. Microdeletion encompassing MAPT at chromosome 17q21.3 is associated with developmental delay and learning disability. Nat Genet. 2006;38:1032–7. doi: 10.1038/ng1858. [DOI] [PubMed] [Google Scholar]
- 134.Sharp AJ, et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet. 2006;38:1038–42. doi: 10.1038/ng1862. [DOI] [PubMed] [Google Scholar]
- 135.Vnencak-Jones CL, Phillips JA, 3rd, Chen EY, Seeburg PH. Molecular basis of human growth hormone gene deletions. Proc Natl Acad Sci U S A. 1988;85:5615–9. doi: 10.1073/pnas.85.15.5615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Edelmann L, Pandita RK, Morrow BE. Low-copy repeats mediate the common 3-Mb deletion in patients with velo-cardio-facial syndrome. Am J Hum Genet. 1999;64:1076–86. doi: 10.1086/302343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Giorda R, et al. Complex segmental duplications mediate a recurrent dup(X)(p11.22–p11.23) associated with mental retardation, speech delay, and EEG anomalies in males and females. Am J Hum Genet. 2009;85:394–400. doi: 10.1016/j.ajhg.2009.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Bondeson ML, et al. Inversion of the IDS gene resulting from recombination with IDS-related sequences is a common cause of the Hunter syndrome. Hum Mol Genet. 1995;4:615–21. doi: 10.1093/hmg/4.4.615. [DOI] [PubMed] [Google Scholar]
- 139.Karsten SL, et al. Two distinct deletions in the IDS gene and the gene W: a novel type of mutation associated with the Hunter syndrome. Genomics. 1997;43:123–9. doi: 10.1006/geno.1997.4811. [DOI] [PubMed] [Google Scholar]
- 140.Simon DB, et al. Mutations in the chloride channel gene, CLCNKB, cause Bartter's syndrome type III. Nat Genet. 1997;17:171–8. doi: 10.1038/ng1097-171. [DOI] [PubMed] [Google Scholar]
- 141.Zimran A, et al. A glucocerebrosidase fusion gene in Gaucher disease. Implications for the molecular anatomy, pathogenesis, and diagnosis of this disorder. J Clin Invest. 1990;85:219–22. doi: 10.1172/JCI114415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Saunier S, et al. Characterization of the NPHP1 locus: mutational mechanism involved in deletions in familial juvenile nephronophthisis. Am J Hum Genet. 2000;66:778–89. doi: 10.1086/302819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Wijmenga C, et al. Chromosome 4q DNA rearrangements associated with facioscapulohumeral muscular dystrophy. Nat Genet. 1992;2:26–30. doi: 10.1038/ng0992-26. [DOI] [PubMed] [Google Scholar]
- 144.Lifton RP, et al. A chimaeric 11 beta-hydroxylase/aldosterone synthase gene causes glucocorticoid-remediable aldosteronism and human hypertension. Nature. 1992;355:262–5. doi: 10.1038/355262a0. [DOI] [PubMed] [Google Scholar]
- 145.Watnick TJ, Gandolph MA, Weber H, Neumann HP, Germino GG. Gene conversion is a likely cause of mutation in PKD1. Hum Mol Genet. 1998;7:1239–43. doi: 10.1093/hmg/7.8.1239. [DOI] [PubMed] [Google Scholar]
- 146.Dorschner MO, Sybert VP, Weaver M, Pletcher BA, Stephens K. NF1 microdeletion breakpoints are clustered at flanking repetitive sequences. Hum Mol Genet. 2000;9:35–46. doi: 10.1093/hmg/9.1.35. [DOI] [PubMed] [Google Scholar]
- 147.Mefford HC, et al. Recurrent reciprocal genomic rearrangements of 17q12 are associated with renal disease, diabetes, and epilepsy. Am J Hum Genet. 2007;81:1057–69. doi: 10.1086/522591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Van Esch H, et al. Deletion of VCX-A due to NAHR plays a major role in the occurrence of mental retardation in patients with X-linked ichthyosis. Hum Mol Genet. 2005;14:1795–803. doi: 10.1093/hmg/ddi186. [DOI] [PubMed] [Google Scholar]
- 149.Smahi A, et al. Genomic rearrangement in NEMO impairs NF-kappaB activation and is a cause of incontinentia pigmenti. The International Incontinentia Pigmenti (IP) Consortium. Nature. 2000;405:466–72. doi: 10.1038/35013114. [DOI] [PubMed] [Google Scholar]
- 150.Aradhya S, et al. A recurrent deletion in the ubiquitously expressed NEMO (IKK-gamma) gene accounts for the vast majority of incontinentia pigmenti mutations. Hum Mol Genet. 2001;10:2171–9. doi: 10.1093/hmg/10.19.2171. [DOI] [PubMed] [Google Scholar]
- 151.Nathans J, Piantanida TP, Eddy RL, Shows TB, Hogness DS. Molecular genetics of inherited variation in human color vision. Science. 1986;232:203–10. doi: 10.1126/science.3485310. [DOI] [PubMed] [Google Scholar]
- 152.Metzenberg AB, Wurzer G, Huisman TH, Smithies O. Homology requirements for unequal crossing over in humans. Genetics. 1991;128:143–61. doi: 10.1093/genetics/128.1.143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Higgs DR, Hill AV, Bowden DK, Weatherall DJ, Clegg JB. Independent recombination events between the duplicated human alpha globin genes; implications for their concerted evolution. Nucleic Acids Res. 1984;12:6965–77. doi: 10.1093/nar/12.18.6965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Lakich D, Kazazian HH, Jr., Antonarakis SE, Gitschier J. Inversions disrupting the factor VIII gene are a common cause of severe haemophilia A. Nat Genet. 1993;5:236–41. doi: 10.1038/ng1193-236. [DOI] [PubMed] [Google Scholar]