Abstract
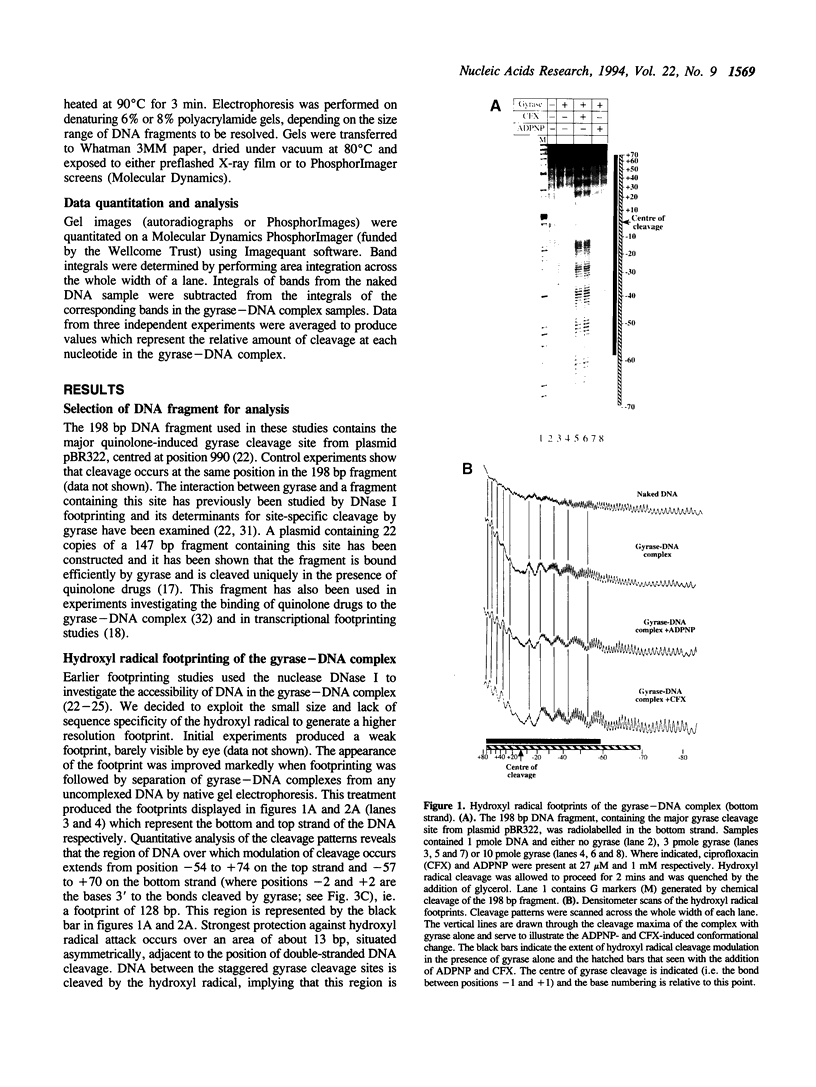
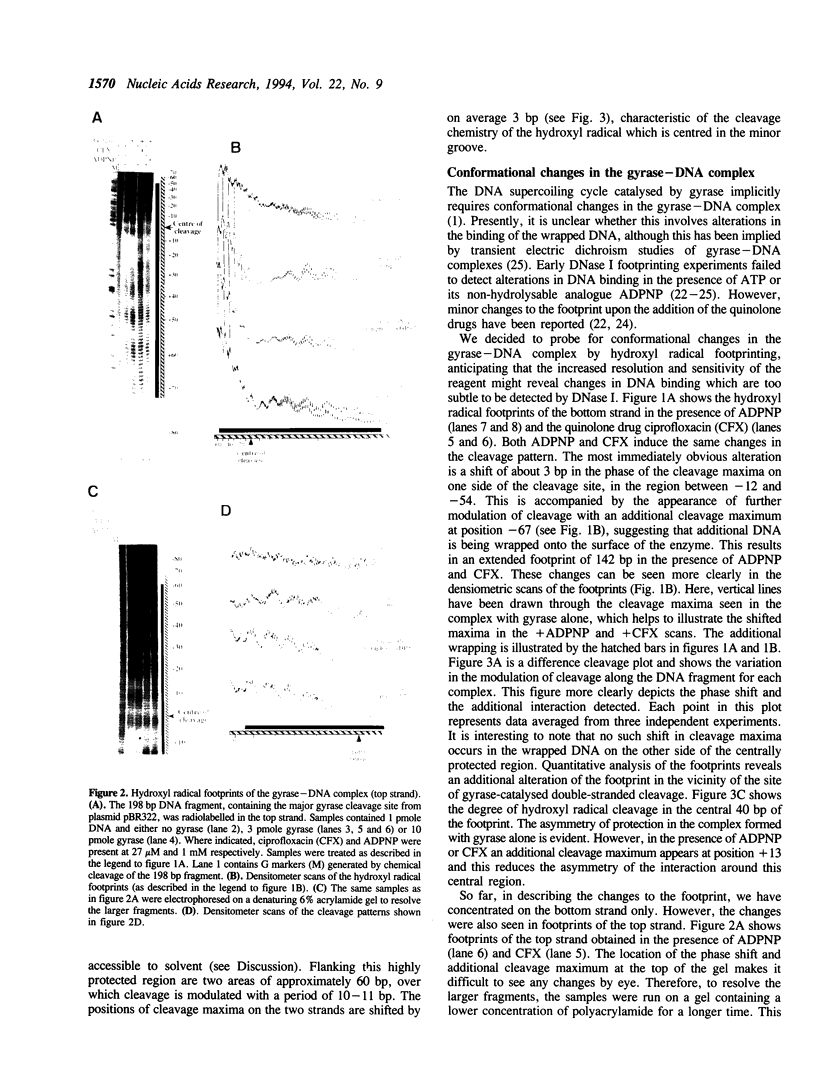
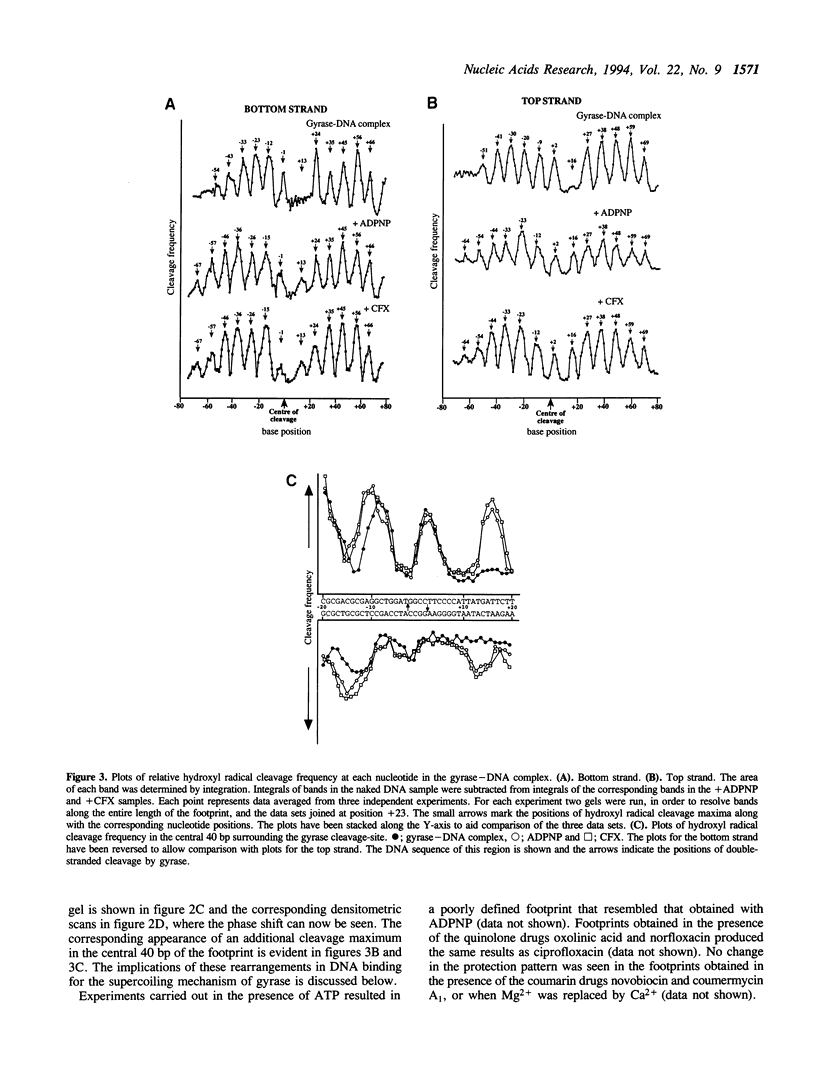
We have used the technique of hydroxyl radical footprinting to probe the complex between DNA gyrase and a 198 bp DNA fragment containing the preferred gyrase cleavage site from plasmid pBR322. We find that gyrase protects 128 bp from the hydroxyl radical with the central 13 bp (adjacent to the gyrase cleavage site) being most strongly protected. Flanking the central region are arms showing periodic protection from the reagent suggesting a helical repeat of 10.6 bp, consistent with the DNA being wrapped upon the enzyme surface. The presence of 5'-adenylyl-beta,gamma-imidodiphosphate or a quinolone drug causes alteration of the protection pattern consistent with a conformational change in the complex involving one arm of the wrapped DNA. The significance of these results for the mechanism of DNA supercoiling by gyrase is discussed.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adachi T., Mizuuchi M., Robinson E. A., Appella E., O'Dea M. H., Gellert M., Mizuuchi K. DNA sequence of the E. coli gyrB gene: application of a new sequencing strategy. Nucleic Acids Res. 1987 Jan 26;15(2):771–784. doi: 10.1093/nar/15.2.771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ali J. A., Jackson A. P., Howells A. J., Maxwell A. The 43-kilodalton N-terminal fragment of the DNA gyrase B protein hydrolyzes ATP and binds coumarin drugs. Biochemistry. 1993 Mar 16;32(10):2717–2724. doi: 10.1021/bi00061a033. [DOI] [PubMed] [Google Scholar]
- Bates A. D., Maxwell A. DNA gyrase can supercoil DNA circles as small as 174 base pairs. EMBO J. 1989 Jun;8(6):1861–1866. doi: 10.1002/j.1460-2075.1989.tb03582.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown P. O., Peebles C. L., Cozzarelli N. R. A topoisomerase from Escherichia coli related to DNA gyrase. Proc Natl Acad Sci U S A. 1979 Dec;76(12):6110–6114. doi: 10.1073/pnas.76.12.6110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dobbs S. T., Cullis P. M., Maxwell A. The cleavage of DNA at phosphorothioate internucleotidic linkages by DNA gyrase. Nucleic Acids Res. 1992 Jul 25;20(14):3567–3573. doi: 10.1093/nar/20.14.3567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drlica K., Coughlin S. Inhibitors of DNA gyrase. Pharmacol Ther. 1989;44(1):107–121. doi: 10.1016/0163-7258(89)90093-4. [DOI] [PubMed] [Google Scholar]
- Fisher L. M., Barot H. A., Cullen M. E. DNA gyrase complex with DNA: determinants for site-specific DNA breakage. EMBO J. 1986 Jun;5(6):1411–1418. doi: 10.1002/j.1460-2075.1986.tb04375.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher L. M., Mizuuchi K., O'Dea M. H., Ohmori H., Gellert M. Site-specific interaction of DNA gyrase with DNA. Proc Natl Acad Sci U S A. 1981 Jul;78(7):4165–4169. doi: 10.1073/pnas.78.7.4165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gellert M., Fisher L. M., O'Dea M. H. DNA gyrase: purification and catalytic properties of a fragment of gyrase B protein. Proc Natl Acad Sci U S A. 1979 Dec;76(12):6289–6293. doi: 10.1073/pnas.76.12.6289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gellert M., Mizuuchi K., O'Dea M. H., Itoh T., Tomizawa J. I. Nalidixic acid resistance: a second genetic character involved in DNA gyrase activity. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4772–4776. doi: 10.1073/pnas.74.11.4772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hallett P., Grimshaw A. J., Wigley D. B., Maxwell A. Cloning of the DNA gyrase genes under tac promoter control: overproduction of the gyrase A and B proteins. Gene. 1990 Sep 1;93(1):139–142. doi: 10.1016/0378-1119(90)90148-k. [DOI] [PubMed] [Google Scholar]
- Hayes J. J., Clark D. J., Wolffe A. P. Histone contributions to the structure of DNA in the nucleosome. Proc Natl Acad Sci U S A. 1991 Aug 1;88(15):6829–6833. doi: 10.1073/pnas.88.15.6829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Tullius T. D., Wolffe A. P. The structure of DNA in a nucleosome. Proc Natl Acad Sci U S A. 1990 Oct;87(19):7405–7409. doi: 10.1073/pnas.87.19.7405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horowitz D. S., Wang J. C. Torsional rigidity of DNA and length dependence of the free energy of DNA supercoiling. J Mol Biol. 1984 Feb 15;173(1):75–91. doi: 10.1016/0022-2836(84)90404-2. [DOI] [PubMed] [Google Scholar]
- Kirchhausen T., Wang J. C., Harrison S. C. DNA gyrase and its complexes with DNA: direct observation by electron microscopy. Cell. 1985 Jul;41(3):933–943. doi: 10.1016/s0092-8674(85)80074-x. [DOI] [PubMed] [Google Scholar]
- Kirkegaard K., Wang J. C. Mapping the topography of DNA wrapped around gyrase by nucleolytic and chemical probing of complexes of unique DNA sequences. Cell. 1981 Mar;23(3):721–729. doi: 10.1016/0092-8674(81)90435-9. [DOI] [PubMed] [Google Scholar]
- Krueger S., Zaccai G., Wlodawer A., Langowski J., O'Dea M., Maxwell A., Gellert M. Neutron and light-scattering studies of DNA gyrase and its complex with DNA. J Mol Biol. 1990 Jan 5;211(1):211–220. doi: 10.1016/0022-2836(90)90021-D. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. DNA-DNA gyrase complex: the wrapping of the DNA duplex outside the enzyme. Cell. 1978 Nov;15(3):979–984. doi: 10.1016/0092-8674(78)90281-7. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. Micrococcus luteus DNA gyrase: active components and a model for its supercoiling of DNA. Proc Natl Acad Sci U S A. 1978 May;75(5):2098–2102. doi: 10.1073/pnas.75.5.2098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxwell A., Gellert M. Mechanistic aspects of DNA topoisomerases. Adv Protein Chem. 1986;38:69–107. doi: 10.1016/s0065-3233(08)60526-4. [DOI] [PubMed] [Google Scholar]
- Maxwell A. The interaction between coumarin drugs and DNA gyrase. Mol Microbiol. 1993 Aug;9(4):681–686. doi: 10.1111/j.1365-2958.1993.tb01728.x. [DOI] [PubMed] [Google Scholar]
- Maxwell A. The molecular basis of quinolone action. J Antimicrob Chemother. 1992 Oct;30(4):409–414. doi: 10.1093/jac/30.4.409. [DOI] [PubMed] [Google Scholar]
- Morrison A., Cozzarelli N. R. Contacts between DNA gyrase and its binding site on DNA: features of symmetry and asymmetry revealed by protection from nucleases. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1416–1420. doi: 10.1073/pnas.78.3.1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peck L. J., Wang J. C. Sequence dependence of the helical repeat of DNA in solution. Nature. 1981 Jul 23;292(5821):375–378. doi: 10.1038/292375a0. [DOI] [PubMed] [Google Scholar]
- Rau D. C., Gellert M., Thoma F., Maxwell A. Structure of the DNA gyrase-DNA complex as revealed by transient electric dichroism. J Mol Biol. 1987 Feb 5;193(3):555–569. doi: 10.1016/0022-2836(87)90266-x. [DOI] [PubMed] [Google Scholar]
- Reece R. J., Maxwell A. DNA gyrase: structure and function. Crit Rev Biochem Mol Biol. 1991;26(3-4):335–375. doi: 10.3109/10409239109114072. [DOI] [PubMed] [Google Scholar]
- Reece R. J., Maxwell A. Probing the limits of the DNA breakage-reunion domain of the Escherichia coli DNA gyrase A protein. J Biol Chem. 1991 Feb 25;266(6):3540–3546. [PubMed] [Google Scholar]
- Reece R. J., Maxwell A. The C-terminal domain of the Escherichia coli DNA gyrase A subunit is a DNA-binding protein. Nucleic Acids Res. 1991 Apr 11;19(7):1399–1405. doi: 10.1093/nar/19.7.1399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reece R. J., Maxwell A. Tryptic fragments of the Escherichia coli DNA gyrase A protein. J Biol Chem. 1989 Nov 25;264(33):19648–19653. [PubMed] [Google Scholar]
- Rhodes D., Klug A. Sequence-dependent helical periodicity of DNA. Nature. 1981 Jul 23;292(5821):378–380. doi: 10.1038/292378a0. [DOI] [PubMed] [Google Scholar]
- Roca J., Wang J. C. The capture of a DNA double helix by an ATP-dependent protein clamp: a key step in DNA transport by type II DNA topoisomerases. Cell. 1992 Nov 27;71(5):833–840. doi: 10.1016/0092-8674(92)90558-t. [DOI] [PubMed] [Google Scholar]
- Rádl S. Structure-activity relationships in DNA gyrase inhibitors. Pharmacol Ther. 1990;48(1):1–17. doi: 10.1016/0163-7258(90)90014-s. [DOI] [PubMed] [Google Scholar]
- Shore D., Baldwin R. L. Energetics of DNA twisting. II. Topoisomer analysis. J Mol Biol. 1983 Nov 15;170(4):983–1007. doi: 10.1016/s0022-2836(83)80199-5. [DOI] [PubMed] [Google Scholar]
- Sugino A., Higgins N. P., Brown P. O., Peebles C. L., Cozzarelli N. R. Energy coupling in DNA gyrase and the mechanism of action of novobiocin. Proc Natl Acad Sci U S A. 1978 Oct;75(10):4838–4842. doi: 10.1073/pnas.75.10.4838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugino A., Peebles C. L., Kreuzer K. N., Cozzarelli N. R. Mechanism of action of nalidixic acid: purification of Escherichia coli nalA gene product and its relationship to DNA gyrase and a novel nicking-closing enzyme. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4767–4771. doi: 10.1073/pnas.74.11.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wigley D. B., Davies G. J., Dodson E. J., Maxwell A., Dodson G. Crystal structure of an N-terminal fragment of the DNA gyrase B protein. Nature. 1991 Jun 20;351(6328):624–629. doi: 10.1038/351624a0. [DOI] [PubMed] [Google Scholar]
- Willmott C. J., Maxwell A. A single point mutation in the DNA gyrase A protein greatly reduces binding of fluoroquinolones to the gyrase-DNA complex. Antimicrob Agents Chemother. 1993 Jan;37(1):126–127. doi: 10.1128/aac.37.1.126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshida H., Nakamura M., Bogaki M., Ito H., Kojima T., Hattori H., Nakamura S. Mechanism of action of quinolones against Escherichia coli DNA gyrase. Antimicrob Agents Chemother. 1993 Apr;37(4):839–845. doi: 10.1128/aac.37.4.839. [DOI] [PMC free article] [PubMed] [Google Scholar]





