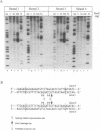
Abstract
The Escherichia coli RuvC protein is an endonuclease that resolves Holliday junctions. In vitro, the protein shows efficient structure-specific binding of Holliday junctions, yet the rate of junction resolution is remarkably low. We have mapped the sites of cleavage on a synthetic junction through which a crossover can branch migrate through 26 bp and find that > or = 90% of the junctions were cleaved at one site. This observation of sequence-specific cleavage suggests that inefficient resolution may be due to DNA binding events which occur away from the cleavage site and are therefore non-productive. Holliday junction resolution by RuvC protein can be stimulated by a number of factors including: (i) the presence of Mn2+ (rather than Mg2+) as the divalent metal cofactor, (ii) alkaline pH (< or = 10), and (iii) elevated temperature. These observations may indicate that other proteins are required for efficient RuvC-mediated resolution.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baker T. A., Mizuuchi M., Mizuuchi K. MuB protein allosterically activates strand transfer by the transposase of phage Mu. Cell. 1991 Jun 14;65(6):1003–1013. doi: 10.1016/0092-8674(91)90552-a. [DOI] [PubMed] [Google Scholar]
- Bennett R. J., Dunderdale H. J., West S. C. Resolution of Holliday junctions by RuvC resolvase: cleavage specificity and DNA distortion. Cell. 1993 Sep 24;74(6):1021–1031. doi: 10.1016/0092-8674(93)90724-5. [DOI] [PubMed] [Google Scholar]
- Benson F. E., West S. C. Substrate specificity of the Escherichia coli RuvC protein. Resolution of three- and four-stranded recombination intermediates. J Biol Chem. 1994 Feb 18;269(7):5195–5201. [PubMed] [Google Scholar]
- Dunderdale H. J., Benson F. E., Parsons C. A., Sharples G. J., Lloyd R. G., West S. C. Formation and resolution of recombination intermediates by E. coli RecA and RuvC proteins. Nature. 1991 Dec 19;354(6354):506–510. doi: 10.1038/354506a0. [DOI] [PubMed] [Google Scholar]
- Dunderdale H. J., Sharples G. J., Lloyd R. G., West S. C. Cloning, overexpression, purification, and characterization of the Escherichia coli RuvC Holliday junction resolvase. J Biol Chem. 1994 Feb 18;269(7):5187–5194. [PubMed] [Google Scholar]
- Kosak H. G., Kemper B. W. Large-scale preparation of T4 endonuclease VII from over-expressing bacteria. Eur J Biochem. 1990 Dec 27;194(3):779–784. doi: 10.1111/j.1432-1033.1990.tb19469.x. [DOI] [PubMed] [Google Scholar]
- Mandal T. N., Mahdi A. A., Sharples G. J., Lloyd R. G. Resolution of Holliday intermediates in recombination and DNA repair: indirect suppression of ruvA, ruvB, and ruvC mutations. J Bacteriol. 1993 Jul;175(14):4325–4334. doi: 10.1128/jb.175.14.4325-4334.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Müller B., Burdett I., West S. C. Unusual stability of recombination intermediates made by Escherichia coli RecA protein. EMBO J. 1992 Jul;11(7):2685–2693. doi: 10.1002/j.1460-2075.1992.tb05334.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller B., Jones C., Kemper B., West S. C. Enzymatic formation and resolution of Holliday junctions in vitro. Cell. 1990 Jan 26;60(2):329–336. doi: 10.1016/0092-8674(90)90747-3. [DOI] [PubMed] [Google Scholar]
- Parsons C. A., Tsaneva I., Lloyd R. G., West S. C. Interaction of Escherichia coli RuvA and RuvB proteins with synthetic Holliday junctions. Proc Natl Acad Sci U S A. 1992 Jun 15;89(12):5452–5456. doi: 10.1073/pnas.89.12.5452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsons C. A., West S. C. Formation of a RuvAB-Holliday junction complex in vitro. J Mol Biol. 1993 Jul 20;232(2):397–405. doi: 10.1006/jmbi.1993.1399. [DOI] [PubMed] [Google Scholar]
- Parsons C. A., West S. C. Specificity of binding to four-way junctions in DNA by bacteriophage T7 endonuclease I. Nucleic Acids Res. 1990 Aug 11;18(15):4377–4384. doi: 10.1093/nar/18.15.4377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Picksley S. M., Parsons C. A., Kemper B., West S. C. Cleavage specificity of bacteriophage T4 endonuclease VII and bacteriophage T7 endonuclease I on synthetic branch migratable Holliday junctions. J Mol Biol. 1990 Apr 20;212(4):723–735. doi: 10.1016/0022-2836(90)90233-C. [DOI] [PubMed] [Google Scholar]
- Sharples G. J., Benson F. E., Illing G. T., Lloyd R. G. Molecular and functional analysis of the ruv region of Escherichia coli K-12 reveals three genes involved in DNA repair and recombination. Mol Gen Genet. 1990 Apr;221(2):219–226. doi: 10.1007/BF00261724. [DOI] [PubMed] [Google Scholar]
- Tsaneva I. R., Müller B., West S. C. ATP-dependent branch migration of Holliday junctions promoted by the RuvA and RuvB proteins of E. coli. Cell. 1992 Jun 26;69(7):1171–1180. doi: 10.1016/0092-8674(92)90638-s. [DOI] [PubMed] [Google Scholar]
- Tsaneva I. R., Müller B., West S. C. RuvA and RuvB proteins of Escherichia coli exhibit DNA helicase activity in vitro. Proc Natl Acad Sci U S A. 1993 Feb 15;90(4):1315–1319. doi: 10.1073/pnas.90.4.1315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vermote C. L., Halford S. E. EcoRV restriction endonuclease: communication between catalytic metal ions and DNA recognition. Biochemistry. 1992 Jul 7;31(26):6082–6089. doi: 10.1021/bi00141a018. [DOI] [PubMed] [Google Scholar]
- West S. C. The processing of recombination intermediates: mechanistic insights from studies of bacterial proteins. Cell. 1994 Jan 14;76(1):9–15. doi: 10.1016/0092-8674(94)90168-6. [DOI] [PubMed] [Google Scholar]