Abstract
The presence of an α-crystallin domain documents the evolutionary relatedness of the ubiquitous family of small heat shock proteins. Sequence and three-dimensional structure provide no evidence for the presence of such a domain in HSPC034, recently proposed as the 11th member of the human HSPB family. Also, phylogenetic analyses detect no relationship between HSPC034 and the human HSPB1–10 sequences. Arguments are provided as to why inclusion in the HSPB family of proteins like HSPC034, which resemble small heat shock proteins in being heat-inducible and having chaperone-like properties and a low monomeric mass, but are evolutionarily unrelated, is misleading and confusing.
Electronic supplementary material
The online version of this article (doi:10.1007/s12192-009-0155-4) contains supplementary material, which is available to authorized users.
Keywords: Chaperone-like, α-crystallin, HSPB11, Phylogeny, Protein family, Protein nomenclature, Small heat shock protein
Kampinga et al. (2009) recently made an applaudable attempt to bring order to the nomenclature of the human heat shock proteins (HSPs). While such an endeavor is badly needed for most HSP families, this is not the case for the α-crystallin-related small heat shock proteins (sHSPs). Ever since the discovery that Drosophila sHSPs share a homologous region of about 90 residues with the α-crystallins (Ingolia and Craig 1982), the “α-crystallin domain” (ACD) has been the unmistakable and unquestioned hallmark of the sHSP family, from bacteria to humans. Thorough database searches have traced ten human members of the sHSP family, designated as HSPB1–10 (Fontaine et al. 2003; Kappé et al. 2003). It was therefore a surprise that Kampinga et al. (2009) present a protein alternatively known as HSPC034, C1orf41, PP25, Hsp16.2, or IFT20 (indeed, illustrating the need for standard nomenclature) as a new member of the human sHSP family under the name HSPB11. The authors provide no reasons to consider HSPC034 as an sHSP, but acknowledge that sHSPs “are characterized by a signature conserved crystallin domain flanked by variable N- and C-termini.” Bellyei et al. (2007b) indeed reported a “reasonably high” sequence similarity of the human C1orf41 protein with some human sHSPs, and dubbed it Hsp16.2.
Hsp16.2 is small, heat-inducible, and displays chaperone-like properties, like many sHSPs (Bellyei et al. 2007a, b); whether it also forms oligomers (Bohn and Winckler 1991; Bellyei et al. 2007b) or rather monomers (Ramelot et al. 2009) remains unclear. But does the reported sequence similarity reflect the presence of a genuine ACD-like sequence? Figure 1 presents an alignment of human HSPB1–10, a plant and an archaeal sHSP, human HSPC034 (or Hsp16.2), and five of its orthologs from other organisms. The alignment highlights a pronounced sequence conservation of HSPC034, from protist to humans. Amongst the sHSPs, overall sequence homology is only observed in the C-terminal half, corresponding with the ACD; in the N-terminal halves and C-terminal tails (not shown in Fig. 1), sequence homology is only present within certain subgroups of sHSPs. Most significantly, sequence “signatures” that are characteristic for the ACD, like DELxV (or similar residues) at positions 120–124, VxI (131–133), LPxxV (165–169), and GxLxV (182–186) are not present in the HSPC034 sequences, and vice versa. This is reflected by the fact that an InterProScan Sequence Search (http://www.ebi.ac.uk) readily predicts an ACD sequence even in the quite deviating HSPB10, functioning as outer dense fiber protein Odf1 in sperm tails, but not in HSPC034. Therefore, at the sequence level, there is no evidence for the presence of an ACD-like sequence in HSPC034.
Fig. 1.
Alignment and secondary structure elements of sHSP and HSPC034 sequences. Human HSPB1–10 are aligned with wheat Tae Hsp16.9 and the archaeal Methanococcus jannasschii Mj Hsp16.5 because of the latters’ known crystal structures. HSPC034 sequences are from human (Hs), mouse (Mm), chicken (Gg), amphioxus (Bf), Trichoplax adhaerens, a placozoan, the lowest metazoans (Tad), and T. vaginalis, a unicellular eukaryote (Tv). No prokaryotic homologs of HSPC034 could be retrieved. For accession numbers, see the legends of Supplementary Fig. S1. This set of sequences is taken without change from the larger alignment in Supplementary Fig. S1, which was used for the construction of the cladogram in Fig. 2, and also includes mouse and chicken HSPB1–10. Alignment was obtained with ClustalW v1.83 at default settings, without manual editing, to avoid any subjective influences. White letters on dark gray background: >80% identity or similarity in the six HSPC034 sequences; black letters on light gray background: >80% identity or similarity in the 12 sHSP sequences; white letters on black background: >80% identity or similarity both in HSPC034 and sHSP sequences. Similarity groups are: D/N, V/L/I/M, F/Y/W, K/R, E/D, and Q/E. To make the alignment more compact, the C-terminal tails of most sHSPs are not shown (slashes) and an internal sequence of 20 residues is deleted from Hs HspB10 (number sign at position 97). The demarcations of the discoidin domain in human HSPC034 (positions 39–171) and the ACD in human HSPB5 (positions 94–191) are indicated. Asterisks at positions 117 and 119 indicate the F and P residues that are diagnostic for metazoan sHSPs (de Jong et al. 1998; Bagnéris et al. 2009). The positions of experimentally determined β-strands and α-helices in human αB-crystallin (Hs HspB5; Bagnéris et al. 2009; Jehle et al. 2009), rat Hsp20 (Rn HspB6; Bagnéris et al. 2009), wheat Tae Hsp16.9 (van Montfort et al. 2001), M. jannasschii Mj Hsp16.5 (Kim et al. 1998), and human HSPC034 (Ramelot et al. 2009) are shown below the alignment; for Hs_HspB5 and rat Rn_HspB6, only the structures of the ACDs are known. The remarkable interruptions in the β2 strands of HspB5, Hsp16.9, and Hsp16.5 would normally have been corrected by manual editing
NMR and X-ray structures of human HSPC034 (Ramelot et al. 2009) and of the ACDs of human αB-crystallin (HSPB5; Jehle et al. 2009; Bagnéris et al. 2009) and rat Hsp20 (HSPB6; Bagnéris et al. 2009) have most recently been published. All three proteins possess a β-sandwich fold, but HSPC034 contains—as predicted by InterProScan—a discoidin (coagulation factor 5/8 type C; PFAM accession PF00754) or galactose-binding-like domain (IPR008979), whereas the ACDs of αB-crystallin and Hsp20 belong to the HSP20/alpha crystallin domain family (PF00011). In contrast to the ACD, and as marked in Fig. 1, the discoidin domain also comprises most of the N-terminal sequence of HSPC034. Positions and lengths of the secondary structure elements indicated in Fig. 1 reflect the different folding of the ACD and discoidin domains. Notably, whereas the demarcations of most β-strands in the different ACDs coincide exactly, this is not the case for the β-strands in HSPC034. Considering that during divergent evolution the 3D structures of proteins are better conserved than their primary structures, we conclude that the 3D structure of HSPC034 presents no evidence for evolutionary relatedness with the sHSPs.
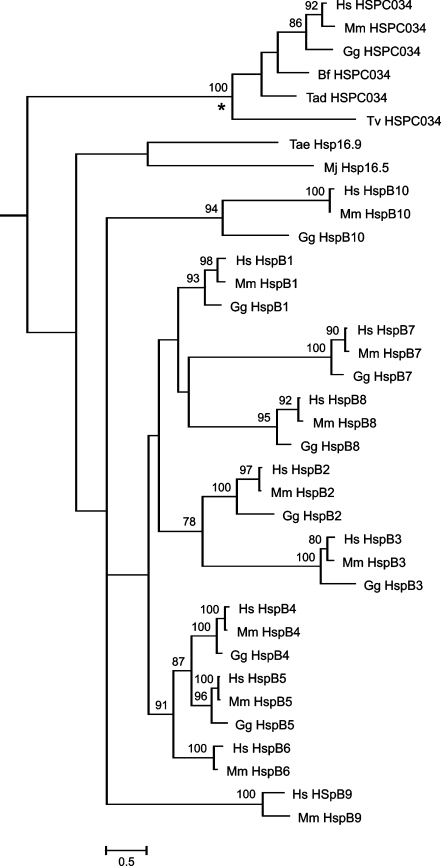
Nevertheless, one still might wonder whether the traces of sequence similarity between HSPC034 and sHSPs, such as the residues marked in black in Fig. 1, might reflect that HSPC034 has dwindled away from an sHSP ancestor, losing its ACD and associated conserved residues, and transformed it somehow in a discoidin domain. We therefore performed a phylogenetic analysis on an extended dataset (Supplementary Fig. S1). As expected from the sequence signatures in Fig. 1, the resulting cladogram (Fig. 2) displays a highly supported HSPC034 clade, and a separate sHSP clade that accurately presents the major relationships amongst vertebrate HSPB1–10, as earlier established independently by Fontaine et al. (2003), Franck et al. (2004), and Elicker and Hutson (2007). It thus shows that the sequence similarities between human HSPC034 and human HSPB1 (Hsp27), HSPB2, HSPB5, and HSPB9, as reported by Bellyei et al. (2007b), do not reflect any specific evolutionary relatedness of human HSPC034 with these HSPB members. What else can we infer about the relationship between HSPC034 and the sHSPs from the topology in Fig. 2? One should realize that a phylogenetic program necessarily joins all proteins in a dataset, even if totally unrelated, in one single “tree”; any unrelated proteins will be placed in the poorly resolved basal part of the tree. Thus, whether we accept the mid-point rooting as shown in Fig. 2 or force the root towards the prokaryote Mj Hsp16.5, we can only conclude that HSPC034 is either unrelated or “diverged” from the sHSPs immeasurably deep in evolutionary time. This should have been long before the divergence of protists and higher eukaryotes because the last common ancestor of Trichomonas vaginalis and Homo sapiens already had a full-fledged HSPC034-like sequence, and thus a discoidin domain (asterisk in Fig. 2). The HSPC034 clade also remains far outside the vertebrate HSPB1–10 clade, which finds its origin deep within the metazoan realm (de Jong et al. 1998; Waters and Vierling 1999; Fontaine et al. 2003).
Fig. 2.
Cladogram resulting from a maximum likelihood analysis of the aligned complete sHSP and HSPC034 sequences as present in Supplementary Fig. S1. These are the same as in Fig. 1, plus the mouse (Mm) and chicken (Gg) orthologs of human (Hs) HSPB1–10. The analysis was performed with PhyML v3.0 at http://www.atgc-montpellier.fr/phyml/ using the WAG substitution model, a fixed proportion of invariable sites, four substitution rate categories, and estimated gamma shape parameter. Nodal support was estimated by bootstrap analysis with 100 replicates. Bootstrap values above 70 are shown. The asterisk indicates the oldest divergence in the HSPC034 clade, between the protist T. vaginalis and the metazoa
It may be clear that HSPC034 has no ACD or ACD-like sequence, and also otherwise shows no evidence of evolutionary relationship with the α-crystallin-related sHSPs. But are heat-inducibility, chaperone-like properties, and small monomer size not good reasons to welcome HSPC034 in the human HSPB family, even in the absence of evolutionary relatedness, and similarly incorporate other unrelated proteins with such properties, like, for example, the 19.6-kD human translationally controlled tumor protein (TCTP) (Gnanasekar et al. 2009)? At present, the α-crystallin-related sHSPs form a well-defined family of evolutionarily related though structurally and functionally diverse proteins, occurring in all domains of life. The sole unifying characteristic for inclusion in the sHSP family is the presence of a recognizable ACD-like sequence. It is, in fact, difficult to imagine how a protein without an ACD-like sequence could otherwise be demonstrated to be evolutionarily related with the α-crystallin-related sHSP family. Unlike many other structural domains, including the discoidin domain (Kiedzierska et al. 2007), the ACD is not known to recombine with other domains in chimaeric proteins. This lack of promiscuity and the easy recognition of the ACD generally leave little doubt as to whether or not a protein belongs to the sHSP family, even when characteristic residues in the ACD sequence have been lost, as in Odf1/HSPB10 and some plant sHSPs (Waters et al. 2008). Also, the presence of conserved regions or motifs in the N-terminal domains of specific sHSP subfamilies, notably in plants (e.g., Waters and Vierling 1999), never hampers their recognition as sHSPs. Including HSPC034 and other unrelated proteins amongst the human HSPBs means that the α-crystallin-related sHSPs no longer constitute a protein family in the established evolutionary sense and would falsely suggest that these novel members possess an ACD-like sequence with its associated properties.
Skipping the requirement for the presence of an ACD-like sequence, and making the sHSP “family” a grouping of unrelated small, heat-inducible, and chaperone-like proteins would have other remarkable consequences. Not only the sperm tail protein Odf1/HSPB10 should be separated from its relatives, but also many if not most of the other present sHSP family members should be excluded, because they are not (proven to be) heat-inducible or have no (proven) in vitro and/or in vivo chaperone-like properties (see, e.g., de Jong et al. 1998; Narberhaus 2002; Heikkila 2004; Haslbeck et al. 2005; Sun and MacRae 2005; Taylor and Benjamin 2005; Nakamoto and Vigh 2007; Siddique et al. 2008; Waters et al. 2008; van de Schootbrugge and Boelens 2009). Curious dilemmas would arise. For example, should duck αB-crystallin (HSPB5) be expelled but not its close mammalian orthologs because the former is not heat-inducible while the latter are (Wistow and Graham 1995)? Above all, stress-inducibility and chaperone-like properties are problematic criteria to apply. And who decides when a protein becomes too large to be a “small” HSP? It is difficult to envisage how the ACD sequence, as evidence of evolutionary relationship, can be given up as sole criterion for membership of the sHSP family without making this “family” an uninformative assemblage of evolutionarily unrelated proteins. Moreover, if sequence homology is no longer the essential criterion for inclusion in the α-crystallin-related sHSP family, how then can members of this “family” ever be identified in newly determined genomic sequences?
Fortunately, the solution is not so difficult. There are indeed proteins, like human HSPC034 (Bellyei et al. 2007b) and TCTP (Gnanasekar et al. 2009), which certainly deserve to be described as “small heat shock proteins”, but lack an ACD-like sequence. As long as they are not implied to be α-crystallin-related sHSPs, there is no confusion. One should make a clear distinction between proteins that resemble the sHSPs in certain functional or structural (i.e., small, oligomeric) respects, but have no ACD-like sequence, and those that are evolutionarily related with the α-crystallin-related sHSPs. Unless strong arguments to the contrary are provided, the formal name HSPB should remain restricted to human proteins, and their vertebrate orthologs, that can be shown to have an ACD-like sequence. For the time being, it is less confusing to leave HSPC034 as the orphan small heat shock protein Hsp16.2 (Bellyei et al. 2007a, b; Pozsgai et al. 2007), with various additional functions and names, as PP25 in placenta (Bohn and Winckler 1991), C1orf41 in the human protein–protein interaction network (Rual et al. 2005), IFT25 in the primary cilium (Follit et al. 2009), or HSPC034 in protein structure analysis (Ramelot et al. 2009), than naming it HSPB11, misleadingly implying that it has an evolutionary relationships with human HSPB1–10. Proposing an adequate formal name for HSPC034 and its aliases is beyond the scope of this communication.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Alignment of sHSP and HSPC034 sequences. Names and accession numbers are: human Hs_HspB1 (P04792), Hs_HspB2 (Q16082), Hs_HspB3 (Q12988), Hs_HspB4(P02489), Hs_HspB5 (P02511), Hs_HspB6 (O14558), Hs_HspB7 (Q9UBY9), Hs_HspB8 (Q9UJY1), Hs_HSpB9 (Q9BQS6), Hs_HspB10 (Q14990); mouse Mm_HspB1 (P14602), Mm_HspB2 (Q99PR8), Mm_HspB3 (Q9QZ57), Mm_HspB4 (P24622), Mm_HspB5 (P23927), Mm_HspB6 (Q5EBG6), Mm_HspB7 (P35385), Mm_HspB8 (Q9JK92), Mm_HspB9 (Q9DAM3), Mm_HspB10 (Q61999); chicken Gg_HspB1 (Q00649), Gg_HspB2 (NP_001001527), Gg_HspB3 (XP_001231558), Gg_HspB4 (P02504), Gg_HspB5 (Q05713), Gg_HspB7 (XP_427836), Gg_HspB8 (XP_415280), Gg_HspB10(XP_418368); wheat Tae_Hsp16.9 (CAA45902); Methanococcus jannasschii Mj_Hsp16.5 (Q57733); human Hs_HSPC034 (Q9Y547), mouse Mm_HSPC034 (Q9D6H2), chicken Gg_HSPC034 (XP_422488), amphioxus Bf_HSPC0234 (XP_002207873), Trichoplax adhaerens Tad_HSPC034 (XP_002114382), and T. vaginalis Tv_HSPC034 (XP_001326124). Alignment is obtained with ClustalW v1.83 at default settings, without manual editing, to avoid any subjective influences (GIF 121 KB)
Abbreviation
- ACD
α-crystallin domain
References
- Bagnéris C, Bateman OA, Naylor CE, Cronin N, Boelens WC, Keep NH, Slingsby C. Crystal structures of α-crystallin domain dimers of αB-crystallin and Hsp20. J Mol Biol. 2009;392:1242–1252. doi: 10.1016/j.jmb.2009.07.069. [DOI] [PubMed] [Google Scholar]
- Bellyei S, Szigeti A, Boronkai A, Pozsgai E, Gomori E, Melegh B, Janaky T, Bognar Z, Hocsak E, Sumegi B, Gallyas F., Jr Inhibition of cell death by a novel 16.2 kD heat shock protein predominantly via Hsp90 mediated lipid rafts stabilization and Akt activation pathway. Apoptosis. 2007;12:97–112. doi: 10.1007/s10495-006-0486-x. [DOI] [PubMed] [Google Scholar]
- Bellyei S, Szigeti A, Pozsgai E, Boronkai A, Gomori E, Hocsak E, Farkas R, Sumegi B, Gallyas F., Jr Preventing apoptotic cell death by a novel small heat shock protein. Eur J Cell Biol. 2007;86:161–171. doi: 10.1016/j.ejcb.2006.12.004. [DOI] [PubMed] [Google Scholar]
- Bohn H, Winckler W. Isolation and characterization of five new soluble placental tissue proteins (PP22, PP23, PP24, PP25, PP26) Arch Gynecol Obstet. 1991;248:111–115. doi: 10.1007/BF02390087. [DOI] [PubMed] [Google Scholar]
- Jong WW, Caspers GJ, Leunissen JAM. Genealogy of the alpha-crystallin–small heat-shock protein superfamily. Int J Biol Macromol. 1998;22:151–162. doi: 10.1016/S0141-8130(98)00013-0. [DOI] [PubMed] [Google Scholar]
- Elicker KS, Hutson LD. Genome-wide analysis and expression profiling of the small heat shock proteins in zebrafish. Gene. 2007;403:60–69. doi: 10.1016/j.gene.2007.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Follit JA, Xu F, Keady BT, Pazour GJ. Characterization of mouse IFT complex B. Cell Motil Cytoskeleton. 2009;66:457–468. doi: 10.1002/cm.20346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fontaine JM, Rest JS, Welsh MJ, Benndorf R. The sperm outer dense fiber protein is the 10th member of the superfamily of mammalian small stress proteins. Cell Stress Chaperones. 2003;8:62–69. doi: 10.1379/1466-1268(2003)8<62:TSODFP>2.0.CO;2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franck E, Madsen O, Rheede T, Ricard G, Huynen MA, Jong WW. Evolutionary diversity of vertebrate small heat shock proteins. J Mol Evol. 2004;59:792–805. doi: 10.1007/s00239-004-0013-z. [DOI] [PubMed] [Google Scholar]
- Gnanasekar M, Dakshinamoorthy G, Ramaswamy K. Translationally controlled tumor protein is a novel heat shock protein with chaperone-like activity. Biochem Biophys Res Commun. 2009;386:333–337. doi: 10.1016/j.bbrc.2009.06.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haslbeck M, Franzmann T, Weinfurtner D, Buchner J. Some like it hot: the structure and function of small heat-shock proteins. Nat Struct Mol Biol. 2005;12:842–846. doi: 10.1038/nsmb993. [DOI] [PubMed] [Google Scholar]
- Heikkila JJ. Regulation and function of small heat shock protein genes during amphibian development. J Cell Biochem. 2004;93:672–680. doi: 10.1002/jcb.20237. [DOI] [PubMed] [Google Scholar]
- Ingolia TD, Craig EA. Four small Drosophila heat shock proteins are related to each other and to mammalian alpha-crystallin. Proc Natl Acad Sci U S A. 1982;79:2360–2364. doi: 10.1073/pnas.79.7.2360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jehle S, Rossum B, Stout JR, Noguchi SM, Falber K, Rehbein K, Oschkinat H, Klevit RE, Rajagopal P. αB-crystallin: a hybrid solid-state/solution-state NMR investigation reveals structural aspects of the heterogeneous oligomer. J Mol Biol. 2009;385:1481–1497. doi: 10.1016/j.jmb.2008.10.097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kampinga HH, Hageman J, Vos MJ, Kubota H, Tanguay RM, Bruford EA, Cheetham ME, Chen B, Hightower LE. Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones. 2009;14:105–111. doi: 10.1007/s12192-008-0068-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kappé G, Franck E, Verschuure P, Boelens WC, Leunissen JA, Jong WW. The human genome encodes 10 α-crystallin-related small heat shock proteins: HspB1–10. Cell Stress Chaperones. 2003;8:53–61. doi: 10.1379/1466-1268(2003)8<53:THGECS>2.0.CO;2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiedzierska A, Smietana K, Czepczynska H, Otlewski J. Structural similarities and functional diversity of eukaryotic discoidin-like domains. Biochim Biophys Acta. 2007;1774:1069–1078. doi: 10.1016/j.bbapap.2007.07.007. [DOI] [PubMed] [Google Scholar]
- Kim KK, Kim R, Kim SH. Crystal structure of a small heat-shock protein. Nature. 1998;394:595–599. doi: 10.1038/29106. [DOI] [PubMed] [Google Scholar]
- Nakamoto H, Vigh L. The small heat shock proteins and their clients. Cell Mol Life Sci. 2007;64:294–306. doi: 10.1007/s00018-006-6321-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narberhaus F. Alpha-crystallin-type heat shock proteins: socializing minichaperones in the context of a multichaperone network. Microbiol Mol Biol Rev. 2002;66:64–93. doi: 10.1128/MMBR.66.1.64-93.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pozsgai E, Gomori E, Szigeti A, Boronkai A, Gallyas F, Jr, Sumegi B, Bellyei S. Correlation between the progressive cytoplasmic expression of a novel small heat shock protein (Hsp16.2) and malignancy in brain tumors. BMC Cancer. 2007;7:233. doi: 10.1186/1471-2407-7-233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramelot TA, Raman S, Kuzin AP, Xiao R, Ma LC, Acton TB, Hunt JF, Montelione GT, Baker D, Kennedy MA. Improving NMR protein structure quality by Rosetta refinement: a molecular replacement study. Proteins. 2009;75:147–167. doi: 10.1002/prot.22229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J, Fraughton C, Llamosas E, Cevik S, Bex C, Lamesch P, Sikorski RS, Vandenhaute J, Zoghbi HY, Smolyar A, Bosak S, Sequerra R, Doucette-Stamm L, Cusick ME, Hill DE, Roth FP, Vidal M. Towards a proteome-scale map of the human protein–protein interaction network. Nature. 2005;437:1173–1178. doi: 10.1038/nature04209. [DOI] [PubMed] [Google Scholar]
- Siddique M, Gernhard S, Koskull-Döring P, Vierling E, Scharf KD. The plant sHSP superfamily: five new members in Arabidopsis thaliana with unexpected properties. Cell Stress Chaperones. 2008;13:183–197. doi: 10.1007/s12192-008-0032-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Y, MacRae TH. Small heat shock proteins: molecular structure and chaperone function. Cell Mol Life Sci. 2005;62:2460–2476. doi: 10.1007/s00018-005-5190-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor RP, Benjamin IJ. Small heat shock proteins: a new classification scheme in mammals. J Mol Cell Cardiol. 2005;38:433–444. doi: 10.1016/j.yjmcc.2004.12.014. [DOI] [PubMed] [Google Scholar]
- Schootbrugge C, Boelens WC. Introduction to small heat shock proteins. In: Arrigo AP, Simon S, editors. Small stress proteins and human diseases. New York: Nova Sciences; 2009. [Google Scholar]
- Montfort RL, Basha E, Friedrich KL, Slingsby C, Vierling E. Crystal structure and assembly of a eukaryotic small heat shock protein. Nat Struct Biol. 2001;8:1025–1030. doi: 10.1038/nsb722. [DOI] [PubMed] [Google Scholar]
- Waters ER, Vierling E Chloroplast small heat shock proteins: evidence for atypical evolution of an organelle-localized protein. Proc Natl Acad Sci U S A. 1999;96:14394–14399. doi: 10.1073/pnas.96.25.14394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waters ER, Aevermann BD, Sanders-Reed Z Comparative analysis of the small heat shock proteins in three angiosperm genomes identifies new subfamilies and reveals diverse evolutionary patterns. Cell Stress Chaperones. 2008;13:127–142. doi: 10.1007/s12192-008-0023-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wistow G, Graham C. The duck gene for alpha B-crystallin shows evolutionary conservation of discrete promoter elements but lacks heat and osmotic shock response. Biochim Biophys Acta. 1995;1263:105–113. doi: 10.1016/0167-4781(95)00087-w. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Below is the link to the electronic supplementary material.
Alignment of sHSP and HSPC034 sequences. Names and accession numbers are: human Hs_HspB1 (P04792), Hs_HspB2 (Q16082), Hs_HspB3 (Q12988), Hs_HspB4(P02489), Hs_HspB5 (P02511), Hs_HspB6 (O14558), Hs_HspB7 (Q9UBY9), Hs_HspB8 (Q9UJY1), Hs_HSpB9 (Q9BQS6), Hs_HspB10 (Q14990); mouse Mm_HspB1 (P14602), Mm_HspB2 (Q99PR8), Mm_HspB3 (Q9QZ57), Mm_HspB4 (P24622), Mm_HspB5 (P23927), Mm_HspB6 (Q5EBG6), Mm_HspB7 (P35385), Mm_HspB8 (Q9JK92), Mm_HspB9 (Q9DAM3), Mm_HspB10 (Q61999); chicken Gg_HspB1 (Q00649), Gg_HspB2 (NP_001001527), Gg_HspB3 (XP_001231558), Gg_HspB4 (P02504), Gg_HspB5 (Q05713), Gg_HspB7 (XP_427836), Gg_HspB8 (XP_415280), Gg_HspB10(XP_418368); wheat Tae_Hsp16.9 (CAA45902); Methanococcus jannasschii Mj_Hsp16.5 (Q57733); human Hs_HSPC034 (Q9Y547), mouse Mm_HSPC034 (Q9D6H2), chicken Gg_HSPC034 (XP_422488), amphioxus Bf_HSPC0234 (XP_002207873), Trichoplax adhaerens Tad_HSPC034 (XP_002114382), and T. vaginalis Tv_HSPC034 (XP_001326124). Alignment is obtained with ClustalW v1.83 at default settings, without manual editing, to avoid any subjective influences (GIF 121 KB)