Abstract
The MspI restriction-modification system, which recognizes the sequence 5'-CCGG-3', has been previously cloned and sequenced (1). We subcloned the methyltransferase gene (M.MspI) downstream of the ptac promoter in the multicopy vector pUC119 and overexpressed it in E. coli. Upon induction with IPTG, M.MspI constitutes more than 10% of cellular protein. A scheme has been devised to purify large amounts of biologically active M.MspI to apparent homogeneity from these overexpressing E. coli cells. Approximately 0.8 mg of pure M.MspI per gram of cells (wet weight) can be obtained. The apparent molecular weight of M.MspI is 49 kD, by SDS gel electrophoresis and 48-54 kD by gel filtration. At low concentrations (less than 0.4 mg/ml), the methyltransferase is a monomer in solution but at higher concentrations (greater than 3.0 mg/ml) it exists predominantly as a dimer. Polyclonal antibodies raised against M.MspI cross-react with the DNA-methyltransferases of several other restriction-modification systems.
Full text
PDF






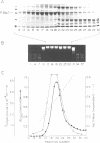
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amann E., Brosius J., Ptashne M. Vectors bearing a hybrid trp-lac promoter useful for regulated expression of cloned genes in Escherichia coli. Gene. 1983 Nov;25(2-3):167–178. doi: 10.1016/0378-1119(83)90222-6. [DOI] [PubMed] [Google Scholar]
- Bergerat A., Guschlbauer W. The double role of methyl donor and allosteric effector of S-adenosyl-methionine for Dam methylase of E. coli. Nucleic Acids Res. 1990 Aug 11;18(15):4369–4375. doi: 10.1093/nar/18.15.4369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Chandrasegaran S., Wu L. P., Valda E., Smith H. O. Overproduction and purification of the M.HhaII methyltransferase from Haemophilus haemolyticus. Gene. 1988 Dec 25;74(1):15–21. doi: 10.1016/0378-1119(88)90240-5. [DOI] [PubMed] [Google Scholar]
- Chen L., MacMillan A. M., Chang W., Ezaz-Nikpay K., Lane W. S., Verdine G. L. Direct identification of the active-site nucleophile in a DNA (cytosine-5)-methyltransferase. Biochemistry. 1991 Nov 19;30(46):11018–11025. doi: 10.1021/bi00110a002. [DOI] [PubMed] [Google Scholar]
- Cheng S. C., Kim R., King K., Kim S. H., Modrich P. Isolation of gram quantities of EcoRI restriction and modification enzymes from an overproducing strain. J Biol Chem. 1984 Sep 25;259(18):11571–11575. [PubMed] [Google Scholar]
- Everett E. A., Falick A. M., Reich N. O. Identification of a critical cysteine in EcoRI DNA methyltransferase by mass spectrometry. J Biol Chem. 1990 Oct 15;265(29):17713–17719. [PubMed] [Google Scholar]
- Friedman S., Som S., Yang L. F. The core element of the EcoRII methylase as defined by protease digestion and deletion analysis. Nucleic Acids Res. 1991 Oct 11;19(19):5403–5408. doi: 10.1093/nar/19.19.5403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldschmidt R. In vivo degradation of nonsense fragments in E. coli. Nature. 1970 Dec 19;228(5277):1151–1154. doi: 10.1038/2281151a0. [DOI] [PubMed] [Google Scholar]
- Holliday R. The inheritance of epigenetic defects. Science. 1987 Oct 9;238(4824):163–170. doi: 10.1126/science.3310230. [DOI] [PubMed] [Google Scholar]
- Howe M. M. Prophage deletion mapping of bacteriophage Mu-1. Virology. 1973 Jul;54(1):93–101. doi: 10.1016/0042-6822(73)90118-9. [DOI] [PubMed] [Google Scholar]
- Kaszubska W., Aiken C., O'Connor C. D., Gumport R. I. Purification, cloning and sequence analysis of RsrI DNA methyltransferase: lack of homology between two enzymes, RsrI and EcoRI, that methylate the same nucleotide in identical recognition sequences. Nucleic Acids Res. 1989 Dec 25;17(24):10403–10425. doi: 10.1093/nar/17.24.10403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaul S., Pfeifer G. P., Drahovsky D. Preparation of monoclonal antibodies against DNA-cytosine-5-methyltransferase from human placenta. Eur J Cell Biol. 1984 Jul;34(2):330–335. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lauster R., Trautner T. A., Noyer-Weidner M. Cytosine-specific type II DNA methyltransferases. A conserved enzyme core with variable target-recognizing domains. J Mol Biol. 1989 Mar 20;206(2):305–312. doi: 10.1016/0022-2836(89)90480-4. [DOI] [PubMed] [Google Scholar]
- Lewis J., Bird A. DNA methylation and chromatin structure. FEBS Lett. 1991 Jul 22;285(2):155–159. doi: 10.1016/0014-5793(91)80795-5. [DOI] [PubMed] [Google Scholar]
- Lin P. M., Lee C. H., Roberts R. J. Cloning and characterization of the genes encoding the MspI restriction modification system. Nucleic Acids Res. 1989 Apr 25;17(8):3001–3011. doi: 10.1093/nar/17.8.3001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modrich P. DNA mismatch correction. Annu Rev Biochem. 1987;56:435–466. doi: 10.1146/annurev.bi.56.070187.002251. [DOI] [PubMed] [Google Scholar]
- Pfeifer G. P., Kohlmaier L., Tomassetti A., Schleicher R., Follmann H., Pfohl-Leszkowicz A., Dirheimer G., Drahovsky D. Polypeptide composition and an immunological analysis of DNA methyltransferases from different species. Arch Biochem Biophys. 1989 Jan;268(1):388–392. doi: 10.1016/0003-9861(89)90599-7. [DOI] [PubMed] [Google Scholar]
- Pósfai J., Bhagwat A. S., Pósfai G., Roberts R. J. Predictive motifs derived from cytosine methyltransferases. Nucleic Acids Res. 1989 Apr 11;17(7):2421–2435. doi: 10.1093/nar/17.7.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raleigh E. A., Wilson G. Escherichia coli K-12 restricts DNA containing 5-methylcytosine. Proc Natl Acad Sci U S A. 1986 Dec;83(23):9070–9074. doi: 10.1073/pnas.83.23.9070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao D. N., Page M. G., Bickle T. A. Cloning, over-expression and the catalytic properties of the EcoP15 modification methylase from Escherichia coli. J Mol Biol. 1989 Oct 20;209(4):599–606. doi: 10.1016/0022-2836(89)90597-4. [DOI] [PubMed] [Google Scholar]
- Reich N. O., Maegley K. A., Shoemaker D. D., Everett E. Structural and functional analysis of EcoRI DNA methyltransferase by proteolysis. Biochemistry. 1991 Mar 19;30(11):2940–2946. doi: 10.1021/bi00225a030. [DOI] [PubMed] [Google Scholar]
- Roberts R. J., Macelis D. Restriction enzymes and their isoschizomers. Nucleic Acids Res. 1991 Apr 25;19 (Suppl):2077–2109. doi: 10.1093/nar/19.suppl.2077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubin R. A., Modrich P. EcoRI methylase. Physical and catalytic properties of the homogeneous enzyme. J Biol Chem. 1977 Oct 25;252(20):7265–7272. [PubMed] [Google Scholar]
- Schumacher G., Sizmann D., Haug H., Buckel P., Böck A. Penicillin acylase from E. coli: unique gene-protein relation. Nucleic Acids Res. 1986 Jul 25;14(14):5713–5727. doi: 10.1093/nar/14.14.5713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegel L. M., Monty K. J. Determination of molecular weights and frictional ratios of proteins in impure systems by use of gel filtration and density gradient centrifugation. Application to crude preparations of sulfite and hydroxylamine reductases. Biochim Biophys Acta. 1966 Feb 7;112(2):346–362. doi: 10.1016/0926-6585(66)90333-5. [DOI] [PubMed] [Google Scholar]
- Som S., Friedman S. Identification of a highly conserved domain in the EcoRII methyltransferase which can be photolabeled with S-adenosyl-L-[methyl-3H]methionine. Evidence for UV-induced transmethylation of cysteine 186. J Biol Chem. 1991 Feb 15;266(5):2937–2945. [PubMed] [Google Scholar]
- Thompson J. P., Simkevich C. P., Holness M. A., Kang A. H., Raghow R. In vitro methylation of the promoter and enhancer of Pro alpha 1(I) collagen gene leads to its transcriptional inactivation. J Biol Chem. 1991 Feb 5;266(4):2549–2556. [PubMed] [Google Scholar]
- Vanyushin B. F. Replicative DNA methylation in animals and higher plants. Curr Top Microbiol Immunol. 1984;108:99–114. doi: 10.1007/978-3-642-69370-0_7. [DOI] [PubMed] [Google Scholar]
- Walder R. Y., Langtimm C. J., Chatterjee R., Walder J. A. Cloning of the MspI modification enzyme. The site of modification and its effects on cleavage by MspI and HpaII. J Biol Chem. 1983 Jan 25;258(2):1235–1241. [PubMed] [Google Scholar]
- Williams D. C., Van Frank R. M., Muth W. L., Burnett J. P. Cytoplasmic inclusion bodies in Escherichia coli producing biosynthetic human insulin proteins. Science. 1982 Feb 5;215(4533):687–689. doi: 10.1126/science.7036343. [DOI] [PubMed] [Google Scholar]
- Wu J. C., Santi D. V. High level expression and purification of HhaI methyltransferase. Nucleic Acids Res. 1988 Jan 25;16(2):703–717. doi: 10.1093/nar/16.2.703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J. C., Santi D. V. Kinetic and catalytic mechanism of HhaI methyltransferase. J Biol Chem. 1987 Apr 5;262(10):4778–4786. [PubMed] [Google Scholar]
- de Boer H. A., Comstock L. J., Vasser M. The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc Natl Acad Sci U S A. 1983 Jan;80(1):21–25. doi: 10.1073/pnas.80.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de la Campa A. G., Kale P., Springhorn S. S., Lacks S. A. Proteins encoded by the DpnII restriction gene cassette. Two methylases and an endonuclease. J Mol Biol. 1987 Aug 5;196(3):457–469. doi: 10.1016/0022-2836(87)90024-6. [DOI] [PubMed] [Google Scholar]