Abstract
Human bocavirus 2 (HBoV2) is a parvovirus that has been recently identified in stool samples from children. Any association between the virus and clinical disease is unclear. A rapid, reliable diagnostic method is necessary to address this issue. In this study, we developed a sensitive and specific HBoV2 quantitative real-time PCR assay that targets the HBoV2 NP-1 gene, based on the TaqMan method. The assay could reproducibly detect 10 copies of a recombinant DNA plasmid containing a partial region of the HBoV2 genome, with a dynamic range of 8 log units (101 to 108 copies). A clinical evaluation detected HBoV2 in 85 (24.6%) of 345 children with gastroenteritis, with viral loads ranging from 1.67 × 102 to 4.27 × 109 copies per ml of stool specimen.
INTRODUCTION
The family Parvoviridae consists of the subfamilies Parvovirinae and Densovirinae. The subfamily Parvovirinae is classified into five genera: Parvovirus, Erythrovirus, Dependovirus, Amdovirus, and Bocavirus. Parvovirus B19, in the genus Erythrovirus, is the only parvovirus known to cause human disease (11). Human bocavirus (HBoV), in the genus Bocavirus, was discovered recently and is detected frequently in nasopharyngeal aspirate, fecal, urine, and blood samples from children with respiratory or gastrointestinal tract infections (1, 2, 15, 18, 20). Serology studies demonstrated that most children and elderly people have been exposed to HBoV (4, 9, 12, 14). Although current data suggest that HBoV is the second parvovirus pathogenic in humans, it is very difficult to prove this without animal and cell models of HBoV.
In 2009, Kapoor et al. discovered human bocavirus 2 (HBoV2) in stool specimens from Pakistani children with acute flaccid paralysis (13). Subsequently, Arthur et al. reported that HBoV2 was the third most-prevalent virus in children with acute gastroenteritis (3). They found an association between HBoV2 infection and gastroenteritis using multivariate conditional logistic regression analysis. However, the exact clinical significance and epidemiological characteristics of HBoV2 remain to be determined, as discussed here. While it is frequently detected in fecal samples, HBoV2 is rarely detected in respiratory samples (7, 10).
To date, HBoV2 has been detected only by using nested PCR assays, which are laborious and prone to false-positive results. A sensitive and specific real-time PCR assay is necessary to address the epidemiology and disease association of HBoV2. In the present study, we developed a real-time PCR assay targeting the HBoV2 NP-1 gene and used it to detect HBoV2 in stool specimens from acute gastroenteritis patients in Taiyuan, China.
MATERIALS AND METHODS
Clinical specimens.
Stool specimens were collected from 345 children with acute gastroenteritis who were hospitalized in Taiyuan Children Hospital, Taiyuan, Shanxi, China, between November 2007 and October 2008. Informed consent was obtained from the parents of all of the children who provided specimens. The study protocol was approved by the hospital ethics committee. The stool specimens were transported to the China Center for Disease Control (CDC) on dry ice and stored at −20°C until further analysis. Viral DNA was extracted into 50 μl of elution buffer manually from 140 μl of a 10% fecal suspension in phosphate-buffered saline using the QIAamp viral RNA minikit (Qiagen, Hilden, Germany), according to the manufacturer's protocol.
Primers and probe.
A conserved region of the HBoV2 NP-1 gene distinct from that of HBoV was identified from alignments of nucleotide sequences available from GenBank (for HBoV, these were DQ000495, EF203921, EF203922, EF450721, EF450723, EF450727, and NC_007455; for HBoV2 these were FJ170279, FJ170278, FJ170280, GU138990, GU138991, and GU138992) using CLUSTAL W (6). Primer and probe sets were designed for the conserved region using the Primer Express software (version 2.0.0; Applied Biosystems, Foster City, CA). The best primer and probe set selected by the software that showed no major nonspecific homologies in a BLAST analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) was synthesized by Takara Biotechnology (Dalian). The TaqMan probe was labeled at the 5′ end with the reporter molecule 6-carboxyfluorescein and at the 3′ end with eclipse dark quencher (Table 1).
Table 1.
HBoV2 primers and probe used in the real-time PCR assay for gene target NP-1
| Primer or probea | Sequence (5′–3′) | Amplicon length (bp) | Positionb |
|---|---|---|---|
| Primer, forward | TCAGACCAAGCGACGAAGAC | 145 | 2531–2550 |
| Primer, reverse | CTCTAGCAAGYCTAGTAGAATGCC | 145 | 2675–2652 |
| Probe | AACCCACACCATCCAGGAGCATCTG | 145 | 2646–2622 |
The probe was 5′ end labeled with 6-carboxyfluorescein and 3′ end labeled with eclipse dark quencher.
Nucleotide numbering is based on HBoV2 isolate strain PK-5510 (accession no. NC_012042).
Plasmid standard construction.
A 587-bp DNA subgenomic fragment of HBoV2 was amplified from a positive stool specimen using primers bracketing the real-time PCR target region in the NP-1 gene (forward 5′-ATGAGCTCCGAATCTATG-3′ and reverse 5′-CCTCTGATTCCTGTGAAG-3′). The product was cloned into plasmid vector pGEM-T Easy vector (Promega, Madison, WI) and sequenced for verification (GenBank accession no. GU138990). The pGEM-T NP-1 HBoV2 plasmid was purified by using a QIAprep miniprep kit (Qiagen) and quantified by using an Eppendorf 6131 nuclear detector (Eppendorf). To generate standard curves for quantitative determinations and to assess the amplification efficiency, replicate serial 10-fold dilutions of the pGEM-T NP-1 HBoV2 plasmid were prepared in 10 mM Tris-EDTA buffer containing 100 μg of herring sperm DNA (Promega, Madison, WI)/ml and stored at −20°C until use. To minimize the potential for contamination, the HBoV2 plasmid DNA was prepared in a separate laboratory.
HBoV2 real-time PCR assay.
The real-time PCR assays were performed using TaqMan Gene Expression Master Mix (Applied Biosystems); each 25-μl reaction mixture contained 0.5 μM concentrations of forward and reverse primers, a 0.1 μM concentration of probe, and 2.5 μl of nucleic acid extract. The Rotor-Gene 3000 real-time thermal cycling system (Corbett Research, Mortlake, New South Wales, Australia) was used for DNA amplification. The thermocycling conditions consisted of 10 min at 95°C for activating the AmpliTaq Gold, UP enzyme, and 45 cycles of 15 s at 95°C and 1 min at 60°C. Each run included serial 10-fold dilutions of the pGEM-T NP-1 HBoV2 plasmid as a positive control and for the standard curve. A duplicate no-template control was extracted and amplified for every run.
Conventional HBoV2 PCR.
The real-time HBoV2 PCR assay was compared to a nested-primer PCR using conditions described by Kapoor et al. (13). The first-round PCR primers were HBoV2-sf1 (AACAGATGGGCAAGCAGAAC) and HBoV2-sr1 (AGGACAAAGGTCTCCAAGAGG); the second-round PCR primers were HBoV2-sf2 (TGCTTCAACAGGCAAAACAA) and HBoV2-sr2 (TCCAAGAGGAAATGAGTTTGG), which amplified a 455-nucelotide (nt) region within NS1. The PCR products were analyzed by agarose gel electrophoresis, and the positive products were sequenced directly.
RESULTS
Real-time PCR amplification efficiency and limit of detection.
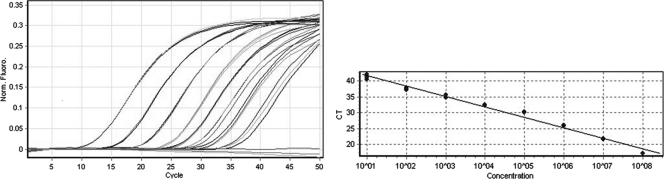
An optimized PCR assay targeting a conserved region of the NP-1 gene was evaluated for amplification efficiency and the limit of target detection. PCR amplification of serial 10-fold dilutions of the pGEM-T NP-1 HBoV2 plasmid achieved >95% efficiency, with linear amplification over an 8-log dynamic range, from 101 to 108 copies per reaction (Fig. 1 and Table 2). To assess the limit of detection of the assay, 1, 5, and 10 copies of the pGEM-T NP-1 HBoV2 plasmid per reaction were tested in 12 replicates. At 10 copies 100% of the replicates were positive, at 5 copies eight (67%) replicates were positive, and at 1 copy three (25%) replicates were positive.
Fig. 1.
NP-1 real-time PCR assay showing representative results obtained with serial 10-fold dilutions (101 to 108 copies per reaction) of the pGEM-T NP-1 HBoV2 plasmid, run in triplicate. The left panel shows baseline subtractive curve fit views of the data with relative fluorescence units (RFU) plotted against cycle number. The right panel shows a standard curve analysis of the DNA amplification plots with cycle threshold (CT) values plotted against the logarithm of the input copy number. The dynamic range of the NP-1 PCR assay spanned 8 log units with a slope of −3.3 and an R2 value of 1.0.
Table 2.
Efficiency of the HBoV2 real-time PCR assay
| Estimated pGEM-T NP-1 HBoV2 plasmid DNA copy no. | Mean CT ± SDa |
|---|---|
| 101 | 41.32 ± 0.80 |
| 102 | 37.62 ± 0.37 |
| 103 | 35.06 ± 0.07 |
| 104 | 32.47 ± 0.07 |
| 105 | 30.13 ± 0.11 |
| 106 | 26.03 ± 0.10 |
| 107 | 21.69 ± 0.06 |
| 108 | 17.18 ± 0.04 |
The values shown are the means of triplicate samples. The slope, −3.31, was determined from the formula y = y intercept − slope log10. The slope was calculated from 101 to 108 copies per reaction. The percent efficiency was 100.5. Efficiency = 10(−1/slope) − 1.
Real-time PCR reproducibility.
To assess the intra- and interassay reproducibility, 10-fold serial dilutions of the pGEM-T NP-1 HBoV2 plasmid from 101 to 108 copies per reaction were tested in triplicate on three different days. Over the linear range of the assays, the coefficient of variation of the mean cycle threshold (CT) values within runs ranged from 0.21 to 1.92% and from run to run ranged from 0.36 to 1.72% (Table 3).
Table 3.
Reproducibility of the HBoV2 real-time PCR assay
| Reproducibility type | CV (%) of the no. of pGEM-T NP-1 HBoV2 plasmid DNA for the following copies/reaction:a |
|||||||
|---|---|---|---|---|---|---|---|---|
| 101 | 102 | 103 | 104 | 105 | 106 | 107 | 108 | |
| Intra-assayb | 1.92 | 0.99 | 1.29 | 0.21 | 0.35 | 0.37 | 0.28 | 0.23 |
| Interassayc | 1.72 | 0.77 | 0.37 | 0.83 | 0.36 | 0.37 | 0.66 | 1.37 |
Tenfold serial dilutions of HBoV2 plasmid DNA are indicated. Dilution series were thawed on three different days, and assays were performed in triplicate for each dilution. CV, coefficient of variation.
Determined from three replicates within each assay.
Determined from three independent assays performed on different days.
Real-time PCR specificity.
The specificity of the real-time PCR assay was assessed by testing nucleic acid extracts of stool specimens containing human and microbiological flora DNA, as well as some enteric viruses, including rotavirus, norovirus, sapovirus, astrovirus, enteric adenovirus, and HBoV. We analyzed three or four positive samples for every enteric virus listed above. No specific amplification was detected for any of these samples.
Real-time and conventional nested PCR results from patients with gastroenteritis.
As determined by real-time PCR, 85 (24.6%) of the 345 stool specimens were positive. The median viral load of the 85 positive samples was 1.02 × 104 copies per ml of starting stool material (range, 1.67 × 102 to 4.27 × 109 copies/ml, Fig. 2). With conventional nested PCR, 57 (16.5% [57/345]) were determined to be positive by gel electrophoresis (with sequencing, 52 were confirmed as HBoV2, 4 were confirmed as HBoV, and 1 was confirmed as HBoV3; GenBank accession no. HM853485 to HM853512, HM853514, HM853515, HM853517 to HM853525, HQ012667 to HQ012677, and HQ141389 to HQ141395). The prevalence with the real-time PCR assay was higher than with the nested PCR assay (24.6% versus 16.5%, χ2 = 6.952; P = 0.008). The sensitivity of the conventional nested PCR was measured using dilutions of an NS1 plasmid subclone. Ten, five, and one plasmid copies were detected in 12/12, 6/12, and 2/12 PCR analyses, respectively.
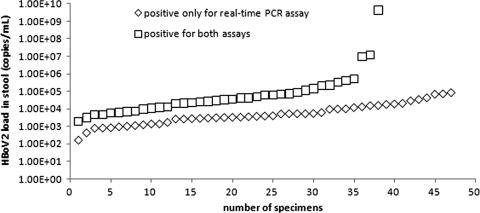
Fig. 2.
Viral loads of HBoV2 in stools according to the real-time PCR assay. The median viral load of 38 positive specimens with both assays was 3.37 × 104 copies/ml, ranging from 1.93 × 103 to 4.27 × 109copies per ml. The median viral load of 47 positive specimens only by real-time PCR was 4.01 × 103 copies/ml, ranging from 1.67 × 102 to 8.76 × 104 copies per ml.
Comparison of the two assays showed that 38 specimens were positive with both assays (median viral load, 3.37 × 104 copies per ml; range, 1.93 × 103 to 4.27 × 109 copies per ml; Fig. 2). Of the 38 specimens, 36 were HBoV2 and 2 were HBoV according to the NS1 gene sequences. For the two HBoV specimens, we sequenced their NP-1 partial genes by using two sets of primers (data not shown) and found that they contained both HBoV2 and HBoV NP-1 sequences (GenBank accession no. HQ398867 to HQ398870). In addition, 47 real-time PCR positive specimens were negative for conventional nested PCR (median viral load, 4.01 × 103 copies per ml; range, 1.67 × 102 to 8.76 × 104 copies per ml, Fig. 2). Nineteen conventional nested PCR-positive specimens were negative for real-time PCR. Of the 19 specimens, 16 were HBoV2, 1 was HBoV3, and 2 were HBoV according to the NS1 gene sequences. For the 16 HBoV2-positive specimens, we sequenced successfully NP-1 partial regions of four specimens and found that two were HBoV2, one was HBoV, and one was HBoV3 in this region (GenBank accession no. HQ398863 to HQ398866). In addition, the NP-1 sequences of the two HBoV2 specimens had no variation at their sites corresponding to the primers and probe. We also repeated the real-time PCR assay three times for the 16 HBoV2-positive specimens and found that 8 of them turned positive once, and the viral loads of the eight specimens were lower than 10 copies per reaction, ranging from 2.01 × 100 to 6.99 × 100 copies.
Epidemiology and clinical symptoms of HBoV2 infection.
The 345 children with gastroenteritis ranged in age between 1 and 50 months (median age, 8 months). The ratio of boys to girls was 1.70:1 (217/128). According to the real-time PCR assay, the 85 HBoV2-positive cases ranged between 1 and 41 months (median age, 10 months), with no significant difference across age groups. The ratio of boys to girls was 1.36:1 (49/36). The positive cases were scattered throughout the year. There was a higher positive rate (34% [25/74]) during the period from February to April (Fig. 3). For the 85 HBoV2-positive cases, common clinical symptoms, in addition to diarrhea, included vomiting (42% [36/85]), fever (41% [35/85]), and respiratory symptoms (31% [26/85]).
Fig. 3.
Seasonal distribution of the children hospitalized with acute gastroenteritis, November 2007 to October 2008, at Taiyuan Children Hospital. The numbers of cases (bars, left y axis) and the HBoV2-positive rate (line, right y axis) varied in different months (x axis). “total” means the number of all of the cases. “HBoV2” means the number of HBoV2-positive cases.
DISCUSSION
Since the first detection of HBoV2, studies of HBoV2 infection have used conventional nested PCR assays with agarose gel electrophoresis (3, 13). To improve the diagnostic methods for HBoV2 detection, we established a real-time PCR assay in the present study. The assay was sensitive, specific, and reliable for HBoV2 DNA amplification, with a reproducible detection limit of 10 target genome copies per reaction and linear amplification over a wide dynamic range, suitable for quantitative applications.
Comparing the real-time PCR assay and the nested PCR assay of HBoV2, there was a significant difference between the detection rates (24.6% versus 16.5%, χ2 = 6.952; P = 0.008) in the clinical evaluation. However, the limits of detection of both assays determined by probit analysis were very similar. This indicates that the TaqMan assay is at least as sensitive as the nested PCR assay. Moreover, the TaqMan assay does give a quantitative measure of viral load, which could potentially correlate with clinical significance. A second advantage of the new assay is that tubes containing the amplified TaqMan product do not ever need to be physically opened in the laboratory, so that the potential for cross-contamination is theoretically much lower, making it more useful in most clinical settings.
In the specificity evaluation with positive samples, our real-time assay did not cross-amplify rotavirus, norovirus, sapovirus, astrovirus, enteric adenovirus, or HBoV. In the clinical evaluation, two HBoV-positive specimens confirmed by NS1 sequencing were detected with the real-time assay. However, further sequence analysis showed that NP-1 sequences of HBoV2 and HBoV coexisted in these two samples, which strengthens the specificity of the real-time assay. Using the nested PCR assay for HBoV2, an HBoV3 (NS1 region) specimen was also detected, while it was negative using real-time PCR. This suggests that our real-time assay appeared to distinguish HBoV3 from HBoV2. However, 16 HBoV2 specimens confirmed by NS1 sequencing after conventional nested PCR were negative for real-time PCR in our clinical evaluation. Further analysis found that this resulted from the low viral loads (lower than the detection limit) and potential recombination between NP-1 and NS1 in the viral genome, which had been noted in other studies (3, 13). Furthermore, as Kapoor et al. pointed out, possible divergence of the NP-1 gene of HBoV2 may also explain this.
Note that HBoV2 had been detected using sensitive nested-PCR but not with single-round PCR in previous studies (3, 13), suggesting low viral loads in the specimens. In the present study, we found that the median viral load of the positive samples was 1.02 × 104copies per ml of stool (range, 1.67 × 102 to 4.27 × 109 copies per ml), which was far below the viral loads in other enteric-virus-associated gastroenteritis. For example, the median viral load of norovirus was at least 8.4 × 105 copies/g or ml of stool specimen in previous studies (5, 16). Based on the hypothesis that high viral loads are an indication of pathogenic relevance, the relatively low virus loads of HBoV2 in stool specimens do not seem to support its clinical significance. The lower viral loads even suggest that there may be a cutoff point and specimens with a viral load below this cutoff point should be classified as “negative,” according to Phillips et al. (17). Further study is necessary to examine the significance of HBoV2 at low viral loads.
In our study, the HBoV2-positive rate was high from February to April. In comparison, Arthur et al. detected HBoV2 predominantly between midsummer and early spring, with no obvious peak (3). Furthermore, most HBoV2-positive patients had a fever and respiratory symptoms. Our results indicate that, like HBoV, HBoV2 is distributed worldwide (3, 8, 10, 13, 19) and may be associated with enteric or respiratory diseases. More investigations are needed to reveal the exact epidemiology of HBoV2 infection and its role in human disease.
The availability of a real-time PCR assay with the pGEM-T NP-1 HBoV2 plasmid control developed in the present study will facilitate further studies to address these issues. In conclusion, we developed a real-time PCR assay that is more sensitive and specific than the reported conventional nested PCR. With this assay, we confirmed the high prevalence of HBoV2 with low viral loads in hospitalized children with gastroenteritis.
ACKNOWLEDGMENTS
This work was partly supported by the China Mega-Project for Infectious Disease (2008ZX10004-008 and 2009ZX10004-101).
Footnotes
Published ahead of print on 16 February 2011.
REFERENCES
- 1. Allander T., et al. 2007. Human bocavirus and acute wheezing in children. Clin. Infect. Dis. 44:904–910 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Allander T., et al. 2005. Cloning of a human parvovirus by molecular screening of respiratory tract samples. Proc. Natl. Acad. Sci. U. S. A. 102:12891–12896 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Arthur J. L., Higgins G. D., Davidson G. P., Givney R. C., Ratcliff R. M. 2009. A novel bocavirus associated with acute gastroenteritis in Australian children. PLoS Pathog. 5:e1000391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Cecchini S., et al. 2009. Evidence of prior exposure to human bocavirus: a retrospective serological study of 404 adult sera in the United States. Clin. Vaccine Immunol. 16:597–604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Chan M. C. W., et al. 2006. Fecal viral load and Norovirus-associated gastroenteritis. Emerg. Infect. Dis. 12:1278–1280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Chenna R., et al. 2003. Multiple sequence alignment with the CLUSTAL series of programs. Nucleic Acids Res. 31:3497–3500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Chieochansin T., Kapoor A., Delwart E., Poovorawan Y., Simmonds P. 2009. Absence of detectable replication of human bocavirus species 2 in respiratory tract. Emerg. Infect. Dis. http://www.cdc.gov/EID/content/15/9/1503.htm doi:10.3201/eid1509.090394 [DOI] [PMC free article] [PubMed]
- 8. Chow B. D., Ou Z., Esper F. P. 2009. Newly recognized bocaviruses (HBoV, HBoV2) in children and adults with gastrointestinal illness in the United States. J. Clin. Virol. doi:10.1016/j.jcv.2009.11.030 [DOI] [PubMed] [Google Scholar]
- 9. Endo R., et al. 2007. Seroepidemiology of human bocavirus in Hokkaido prefecture, Japan. J. Clin. Microbiol. 45:3218–3223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Han T.-H., Chung J.-Y., Hwang E.-S. October 2009. Human bocavirus 2 in children, South Korea. Emerg. Infect. Dis. http://www.cdc.gov/EID/content/15/10/1698.htm doi:10.3201/eid1510.090337 [DOI] [PMC free article] [PubMed]
- 11. Heegaard E. D., Brown K. E. 2002. Human parvovirus B19. Clin. Microbiol. Rev. 15:485–505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Kahn J. S., et al. 2008. Seroepidemiology of human bocavirus defined using recombinant virus-like particles. J. Infect. Dis. 198:41–50 [DOI] [PubMed] [Google Scholar]
- 13. Kapoor A., et al. 2009. A newly identified bocavirus species in human stool. J. Infect. Dis. 199:196–200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Lin F., et al. 2008. ELISAs using human bocavirus VP2 virus-like particles for detection of antibodies against HBoV. J. Virol. Methods 149:110–117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Neske F., et al. 2007. Real-time PCR for diagnosis of human bocavirus infections and phylogenetic analysis. J. Clin. Microbiol. 45:2116–2122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Pang X. L., Preiksaitis J. K., Lee B. 2005. Multiplex real-time RT-PCR for the detection and quantitation of norovirus genogroups I and II in patients with acute gastroenteritis. J. Clin. Virol. 33:168–171 [DOI] [PubMed] [Google Scholar]
- 17. Phillips G., et al. 2009. Diagnosing norovirus-associated infectious intestinal disease using viral load. BMC Infect. Dis. 9:63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Pozo F., et al. 2007. High incidence of human bocavirus infection in children in Spain. J. Clin. Virol. 40:224–228 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Shan T. L., et al. 2009. The first detection of human bocavirus 2 infections in China. J. Clin. Virol. doi:10.1016/j.jcv.2009.07.012 [DOI] [PubMed] [Google Scholar]
- 20. Vicente D., Cilla G., Montes M., Perez-Yarza E. G., Perez-Trallero E. 2007. Human bocavirus, a respiratory and enteric virus. Emerg. Infect. Dis. 13:636–637 [DOI] [PMC free article] [PubMed] [Google Scholar]