Abstract
Protein-induced DNA bending is of importance in the formation of complex nucleoprotein assemblies such as those involved in the initiation of DNA replication or transcription initiation. We have compared the DNA bending characteristics of the Escherichia coli cyclic AMP receptor protein (CRP or CAP), an archetypal DNA bending protein, to those of TFIID, the eukaryotic TATA-element binding transcription factor. By altering the helical phasing between a CRP binding site and the E. coli melR promoter we have mapped a DNA sequence-directed bend in the downstream region of the promoter. This intrinsic DNA bend may be important in the regulation of the melR promoter by CRP in vivo. Gel retardation assays and DNAse I footprinting show that human TFIID binds to the melR promoter - 10 region. Taking advantage of this fact, and using the CRP-induced DNA bend as a standard, we have employed phase sensitive detection to show that the DNA bend angle induced by TFIID is far less than that induced by CRP. Further evidence to support this conclusion comes from a comparison of the relative mobilities of CRP-DNA and TFIID-DNA complexes. These results place limits on the role of any DNA bending induced by TFIID alone in the initiation of transcription.
Full text
PDF





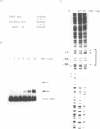
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bell A. I., Gaston K. L., Cole J. A., Busby S. J. Cloning of binding sequences for the Escherichia coli transcription activators, FNR and CRP: location of bases involved in discrimination between FNR and CRP. Nucleic Acids Res. 1989 May 25;17(10):3865–3874. doi: 10.1093/nar/17.10.3865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossi L., Smith D. M. Conformational change in the DNA associated with an unusual promoter mutation in a tRNA operon of Salmonella. Cell. 1984 Dec;39(3 Pt 2):643–652. doi: 10.1016/0092-8674(84)90471-9. [DOI] [PubMed] [Google Scholar]
- Bracco L., Kotlarz D., Kolb A., Diekmann S., Buc H. Synthetic curved DNA sequences can act as transcriptional activators in Escherichia coli. EMBO J. 1989 Dec 20;8(13):4289–4296. doi: 10.1002/j.1460-2075.1989.tb08615.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breathnach R., Chambon P. Organization and expression of eucaryotic split genes coding for proteins. Annu Rev Biochem. 1981;50:349–383. doi: 10.1146/annurev.bi.50.070181.002025. [DOI] [PubMed] [Google Scholar]
- Buratowski S., Hahn S., Guarente L., Sharp P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell. 1989 Feb 24;56(4):549–561. doi: 10.1016/0092-8674(89)90578-3. [DOI] [PubMed] [Google Scholar]
- Busby S., Kotlarz D., Buc H. Deletion mutagenesis of the Escherichia coli galactose operon promoter region. J Mol Biol. 1983 Jun 25;167(2):259–274. doi: 10.1016/s0022-2836(83)80335-0. [DOI] [PubMed] [Google Scholar]
- Busby S., Spassky A., Chan B. RNA polymerase makes important contacts upstream from base pair -49 at the Escherichia coli galactose operon P1 promoter. Gene. 1987;53(2-3):145–152. doi: 10.1016/0378-1119(87)90002-3. [DOI] [PubMed] [Google Scholar]
- Davison B. L., Egly J. M., Mulvihill E. R., Chambon P. Formation of stable preinitiation complexes between eukaryotic class B transcription factors and promoter sequences. Nature. 1983 Feb 24;301(5902):680–686. doi: 10.1038/301680a0. [DOI] [PubMed] [Google Scholar]
- Dynlacht B. D., Hoey T., Tjian R. Isolation of coactivators associated with the TATA-binding protein that mediate transcriptional activation. Cell. 1991 Aug 9;66(3):563–576. doi: 10.1016/0092-8674(81)90019-2. [DOI] [PubMed] [Google Scholar]
- Ebright R. H., Cossart P., Gicquel-Sanzey B., Beckwith J. Mutations that alter the DNA sequence specificity of the catabolite gene activator protein of E. coli. Nature. 1984 Sep 20;311(5983):232–235. doi: 10.1038/311232a0. [DOI] [PubMed] [Google Scholar]
- Fried M., Crothers D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 1981 Dec 11;9(23):6505–6525. doi: 10.1093/nar/9.23.6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner M. M., Revzin A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981 Jul 10;9(13):3047–3060. doi: 10.1093/nar/9.13.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gartenberg M. R., Crothers D. M. Synthetic DNA bending sequences increase the rate of in vitro transcription initiation at the Escherichia coli lac promoter. J Mol Biol. 1991 May 20;219(2):217–230. doi: 10.1016/0022-2836(91)90563-l. [DOI] [PubMed] [Google Scholar]
- Gaston K., Bell A., Kolb A., Buc H., Busby S. Stringent spacing requirements for transcription activation by CRP. Cell. 1990 Aug 24;62(4):733–743. doi: 10.1016/0092-8674(90)90118-x. [DOI] [PubMed] [Google Scholar]
- Gaston K., Chan B., Kolb A., Fox J., Busby S. Alterations in the binding site of the cyclic AMP receptor protein at the Escherichia coli galactose operon regulatory region. Biochem J. 1988 Aug 1;253(3):809–818. doi: 10.1042/bj2530809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaston K., Kolb A., Busby S. Binding of the Escherichia coli cyclic AMP receptor protein to DNA fragments containing consensus nucleotide sequences. Biochem J. 1989 Jul 15;261(2):649–653. doi: 10.1042/bj2610649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghosaini L. R., Brown A. M., Sturtevant J. M. Scanning calorimetric study of the thermal unfolding of catabolite activator protein from Escherichia coli in the absence and presence of cyclic mononucleotides. Biochemistry. 1988 Jul 12;27(14):5257–5261. doi: 10.1021/bi00414a046. [DOI] [PubMed] [Google Scholar]
- Goodman S. D., Nash H. A. Functional replacement of a protein-induced bend in a DNA recombination site. Nature. 1989 Sep 21;341(6239):251–254. doi: 10.1038/341251a0. [DOI] [PubMed] [Google Scholar]
- Greenblatt J. Roles of TFIID in transcriptional initiation by RNA polymerase II. Cell. 1991 Sep 20;66(6):1067–1070. doi: 10.1016/0092-8674(91)90027-v. [DOI] [PubMed] [Google Scholar]
- Gronenborn A. M., Nermut M. V., Eason P., Clore G. M. Visualization of cAMP receptor protein-induced DNA kinking by electron microscopy. J Mol Biol. 1984 Nov 15;179(4):751–757. doi: 10.1016/0022-2836(84)90166-9. [DOI] [PubMed] [Google Scholar]
- Hagerman P. J. Sequence dependence of the curvature of DNA: a test of the phasing hypothesis. Biochemistry. 1985 Dec 3;24(25):7033–7037. doi: 10.1021/bi00346a001. [DOI] [PubMed] [Google Scholar]
- Hahn S., Buratowski S., Sharp P. A., Guarente L. Yeast TATA-binding protein TFIID binds to TATA elements with both consensus and nonconsensus DNA sequences. Proc Natl Acad Sci U S A. 1989 Aug;86(15):5718–5722. doi: 10.1073/pnas.86.15.5718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heumann H., Ricchetti M., Werel W. DNA-dependent RNA polymerase of Escherichia coli induces bending or an increased flexibility of DNA by specific complex formation. EMBO J. 1988 Dec 20;7(13):4379–4381. doi: 10.1002/j.1460-2075.1988.tb03336.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horikoshi M., Bertuccioli C., Takada R., Wang J., Yamamoto T., Roeder R. G. Transcription factor TFIID induces DNA bending upon binding to the TATA element. Proc Natl Acad Sci U S A. 1992 Feb 1;89(3):1060–1064. doi: 10.1073/pnas.89.3.1060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horikoshi M., Yamamoto T., Ohkuma Y., Weil P. A., Roeder R. G. Analysis of structure-function relationships of yeast TATA box binding factor TFIID. Cell. 1990 Jun 29;61(7):1171–1178. doi: 10.1016/0092-8674(90)90681-4. [DOI] [PubMed] [Google Scholar]
- Jayaraman P. S., Peakman T. C., Busby S. J., Quincey R. V., Cole J. A. Location and sequence of the promoter of the gene for the NADH-dependent nitrite reductase of Escherichia coli and its regulation by oxygen, the Fnr protein and nitrite. J Mol Biol. 1987 Aug 20;196(4):781–788. doi: 10.1016/0022-2836(87)90404-9. [DOI] [PubMed] [Google Scholar]
- Kolb A., Spassky A., Chapon C., Blazy B., Buc H. On the different binding affinities of CRP at the lac, gal and malT promoter regions. Nucleic Acids Res. 1983 Nov 25;11(22):7833–7852. doi: 10.1093/nar/11.22.7833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotlarz D., Fritsch A., Buc H. Variations of intramolecular ligation rates allow the detection of protein-induced bends in DNA. EMBO J. 1986 Apr;5(4):799–803. doi: 10.1002/j.1460-2075.1986.tb04284.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koudelka G. B. Bending of synthetic bacteriophage 434 operators by bacteriophage 434 proteins. Nucleic Acids Res. 1991 Aug 11;19(15):4115–4119. doi: 10.1093/nar/19.15.4115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu-Johnson H. N., Gartenberg M. R., Crothers D. M. The DNA binding domain and bending angle of E. coli CAP protein. Cell. 1986 Dec 26;47(6):995–1005. doi: 10.1016/0092-8674(86)90814-7. [DOI] [PubMed] [Google Scholar]
- Marini J. C., Levene S. D., Crothers D. M., Englund P. T. Bent helical structure in kinetoplast DNA. Proc Natl Acad Sci U S A. 1982 Dec;79(24):7664–7668. doi: 10.1073/pnas.79.24.7664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsui T., Segall J., Weil P. A., Roeder R. G. Multiple factors required for accurate initiation of transcription by purified RNA polymerase II. J Biol Chem. 1980 Dec 25;255(24):11992–11996. [PubMed] [Google Scholar]
- Menendez M., Kolb A., Buc H. A new target for CRP action at the malT promoter. EMBO J. 1987 Dec 20;6(13):4227–4234. doi: 10.1002/j.1460-2075.1987.tb02771.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakajima N., Horikoshi M., Roeder R. G. Factors involved in specific transcription by mammalian RNA polymerase II: purification, genetic specificity, and TATA box-promoter interactions of TFIID. Mol Cell Biol. 1988 Oct;8(10):4028–4040. doi: 10.1128/mcb.8.10.4028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porschke D., Hillen W., Takahashi M. The change of DNA structure by specific binding of the cAMP receptor protein from rotation diffusion and dichroism measurements. EMBO J. 1984 Dec 1;3(12):2873–2878. doi: 10.1002/j.1460-2075.1984.tb02223.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rojo F., Zaballos A., Salas M. Bend induced by the phage phi 29 transcriptional activator in the viral late promoter is required for activation. J Mol Biol. 1990 Feb 20;211(4):713–725. doi: 10.1016/0022-2836(90)90072-t. [DOI] [PubMed] [Google Scholar]
- Samuels M., Fire A., Sharp P. A. Separation and characterization of factors mediating accurate transcription by RNA polymerase II. J Biol Chem. 1982 Dec 10;257(23):14419–14427. [PubMed] [Google Scholar]
- Schmidt M. C., Kao C. C., Pei R., Berk A. J. Yeast TATA-box transcription factor gene. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7785–7789. doi: 10.1073/pnas.86.20.7785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz S. C., Shields G. C., Steitz T. A. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science. 1991 Aug 30;253(5023):1001–1007. doi: 10.1126/science.1653449. [DOI] [PubMed] [Google Scholar]
- Spassky A., Busby S., Buc H. On the action of the cyclic AMP-cyclic AMP receptor protein complex at the Escherichia coli lactose and galactose promoter regions. EMBO J. 1984 Jan;3(1):43–50. doi: 10.1002/j.1460-2075.1984.tb01759.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Søgaard-Andersen L., Mironov A. S., Pedersen H., Sukhodelets V. V., Valentin-Hansen P. Single amino acid substitutions in the cAMP receptor protein specifically abolish regulation by the CytR repressor in Escherichia coli. Proc Natl Acad Sci U S A. 1991 Jun 1;88(11):4921–4925. doi: 10.1073/pnas.88.11.4921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Travers A. A. DNA conformation and protein binding. Annu Rev Biochem. 1989;58:427–452. doi: 10.1146/annurev.bi.58.070189.002235. [DOI] [PubMed] [Google Scholar]
- Ulanovsky L. E., Trifonov E. N. Estimation of wedge components in curved DNA. Nature. 1987 Apr 16;326(6114):720–722. doi: 10.1038/326720a0. [DOI] [PubMed] [Google Scholar]
- Webster C., Gaston K., Busby S. Transcription from the Escherichia coli melR promoter is dependent on the cyclic AMP receptor protein. Gene. 1988 Sep 7;68(2):297–305. doi: 10.1016/0378-1119(88)90032-7. [DOI] [PubMed] [Google Scholar]
- Wobbe C. R., Struhl K. Yeast and human TATA-binding proteins have nearly identical DNA sequence requirements for transcription in vitro. Mol Cell Biol. 1990 Aug;10(8):3859–3867. doi: 10.1128/mcb.10.8.3859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]
- Zinkel S. S., Crothers D. M. DNA bend direction by phase sensitive detection. Nature. 1987 Jul 9;328(6126):178–181. doi: 10.1038/328178a0. [DOI] [PubMed] [Google Scholar]
- de Crombrugghe B., Busby S., Buc H. Cyclic AMP receptor protein: role in transcription activation. Science. 1984 May 25;224(4651):831–838. doi: 10.1126/science.6372090. [DOI] [PubMed] [Google Scholar]