Abstract
Vibrio vulnificus is a natural inhabitant of estuarine waters. The three known biotypes include (i) most human pathogens, (ii) primarily eel pathogens, and (iii) pathogens associated with fish and with human wound infections in Israel. Despite the frequently lethal consequences of V. vulnificus infections, the growth rates of the various biotypes and their response to environmental changes are not well characterized. We compared the specific growth rates (μ) of a representative of each biotype by culture and quantitative PCR (qPCR) analysis in a defined medium under varied pH, temperature, and salinity. Growth rates based on culturable concentrations were always higher than those based on qPCR estimates; however, both enumeration methods yielded comparable results on the influence of environmental factors on growth rates. Temperature (25°C, 30°C, 37°C), pH (7.0, 8.0), and salinity (5 to 40‰) all had significant effects on the μ of each biotype. Temperature had the greatest effect on the μ of biotype 1 (CMCP6), whereas salinity had the greatest effect on the μ of biotypes 2 (ATCC 33147) and 3 (302/99). The biotypes' growth rates varied significantly; biotype 1 grew most rapidly, while biotype 3 grew most slowly. The highest growth rates were achieved at 37°C, pH 7.0, and salinities of 15 to 30‰ (μ = 4.0, 2.9, and 2.4 generations h−1 for biotypes 1, 2, and 3, respectively). Other strains of the biotypes yielded comparable results, suggesting that the physiological responses of the biotypes are differentially affected by parameters that are highly variable both in estuarine environments and between the free-living and pathogen states of V. vulnificus.
INTRODUCTION
Vibrio vulnificus is an autochthonous estuarine inhabitant that is widely distributed in warm, saline waters; however, it is also an opportunistic human pathogen that causes wound infections, gastroenteritis, and rapidly fulminating fatal septicemia (13, 20, 21, 33). Confirmed cases are commonly associated with the consumption of contaminated raw seafood, specifically oysters. Moreover, contact with seawater, oysters, or fish contaminated with V. vulnificus can result in necrotizing fasciitis and limb amputation, particularly if the individual is immunocompromised (13, 27). The species has been linked to over 95% of deaths related to consumption of seafood in the United States (17). Medical conditions such as hemochromatosis, renal failure, HIV, chronic liver disease, and immunosuppression act as predisposing factors for infection (6, 14, 20, 33). Vibrio vulnificus strains are categorized into three biotypes determined by biochemical testing methods (4, 6). Biotype 1 strains are most frequently responsible for human infections, whereas biotype 2 strains are associated with disease in eels (3). Biotype 3 was identified in a series of outbreaks in Israel that were associated with the handling of tilapia (6, 21).
Vibrio vulnificus is found in tropical and temperate estuaries, where it faces frequently changing aspects of its environment, such as temperature, salinity, pH, nutrient levels, and pollutants (19, 28, 31). The highest concentrations of V. vulnificus in environmental waters are observed during warm months (16, 28, 30, 36), and the bacteria may enter a viable but not culturable state in water temperatures below 15°C (25). Various works have provided a fragmented picture of the effect of salinity on V. vulnificus (11, 15, 18, 19, 28, 30, 31, 36). The species can tolerate a wide range of salinities, as evidenced by detection in brackish river water at 2.6‰ (11) and culturing from oysters harvested from waters ranging from 2 to 34‰ salinity (28). A survival study conducted in microcosms found that the optimal salinity for biotype 2 eel isolates (strains E22, E86, and E105) was 3 to 5‰ at 25°C and 15 to 38‰ at 12°C; therefore, optimal salinity was influenced by temperature under the experimental conditions (18). Others found that the abundance of V. vulnificus in seawater was lower when salinity rose above 25‰ (15). Another study, which used culture-independent PCR methods, found a salinity optimum of 5 to 10‰ in coastal waters of New Jersey (31).
Whereas distribution and survival studies that explore physical parameters (such as temperature and salinity) of the bacterium's niche have been performed (11, 15, 18, 19, 25, 28, 30, 31, 36), little information about their effects on the species' specific growth rate (μ) exists. In particular, little is known about the comparative growth characteristics of the various biotypes. One study showed that V. vulnificus grew most rapidly at elevated temperatures (∼37°C) and moderate salinity (10 to 20‰) (16). Others report that optimal growth conditions are 22°C, 10 to 40‰ salinity, and pH above 5.2 (35). However, these studies analyzed growth of environmental isolates of unknown biotype or genotype. To our knowledge, no studies have systematically examined the effects of biotype and variation in physical-chemical parameters on V. vulnificus growth rate.
The use of quantitative PCR (qPCR) analysis allows rapid detection of targeted sequences without the requirement of culturability or enrichment procedures (7, 10, 12, 27, 31); therefore, qPCR analysis can quantify cells that might be omitted by culture-dependent methods. Bacterial concentrations determined by qPCR are comparable to those measured by culturing during the exponential phase of growth (32). However, under less optimal conditions, such as during the stationary phase of growth, when nutrients are depleted and metabolites accumulate (22), culturable concentrations decrease, while qPCR-based concentrations remain unchanged (32).
In this study, the growth of one representative of each of the three V. vulnificus biotypes was observed by three methods: (i) increase in optical density (OD), (ii) culturable plate counts, and (iii) total cell concentrations of a single-copy hemolysin gene (vvhA) estimated by qPCR analysis. Logistic curves were used for growth rate calculations under a variety of physical-chemical conditions, including ranges of temperature, pH, and salinity.
MATERIALS AND METHODS
Bacterial strains.
One representative of each biotype was included in the detailed study: V. vulnificus CMCP6 (biotype 1, septicemia isolate, South Korea), V. vulnificus ATCC 33147 (biotype 2, diseased eel isolate, Japan), and V. vulnificus 302/99 (biotype 3, wound infection isolate, Israel). Biotype 2 and 3 strains were kindly provided by Anita C. Wright (University of Florida, Gainesville, FL). Supplementary testing was performed with V. vulnificus 33814 (biotype 1), V. vulnificus 33149 (biotype 2), and V. vulnificus 313/98 (biotype 3) to support the finding of different growth rates among the biotypes.
Culture conditions.
Bacteria were grown overnight at room temperature (∼25°C) in 30 ml CAYEG (34) broth (3% Casamino Acids [Difco Laboratories Inc., Detroit, MI], 0.3% yeast extract, 0.2% glucose, 0.05% KH2PO4, pH 8.4) with agitation at 155 rpm. Cells from overnight cultures were concentrated by centrifugation for 5 min at 10,000 rpm. The supernatant was removed, and cells were washed once with 1× phosphate-buffered saline (PBS) (pH 7.0). The pellet was then resuspended in 50 ml of defined (modified M9) minimal medium (23.2 mM Na2HPO4, 11.02 mM KH2PO4, 9.34 mM NH4Cl, 1.0 mM MgSO4) augmented with 0.06% Casamino Acids, 0.006% yeast extract (Difco Laboratories Inc., Detroit, MI), and 4.5 mM glucose. This medium has a base salinity of 5‰. Salinity was adjusted by adding NaCl, and stipulated values include the base salinity of the medium. Each microcosm was inoculated with washed cells so that the initial cell concentration was approximately 107 CFU ml−1. Cultures were agitated at 155 rpm. Dilution series were made in 0.85% NaCl, and plating was performed on Trypticase soy agar (Difco Laboratories Inc., Detroit, MI) supplemented with NaCl (5.0 g liter−1).
Measurements of growth.
Microcosm cultures (50 ml) were grown in triplicate, and the experiment was repeated once to provide a total of six replicates for each salinity-pH combination: 5, 10, 15, 20, 25, 30, 35, and 40‰ salinity at pH 7.0 and pH 8.0. Growth at all salinity-pH combinations was observed at temperatures of 25, 30 and 37°C. The pH of each microcosm was adjusted to initial values during optical density readings (approximately 20-min intervals) with 300 mM NaOH. The OD was measured at 600 nm (OD600) using a NanoDrop 2000 spectrophotometer (Thermo Scientific Inc., Wilmington, DE). Plating was performed at least three times during the exponential phase of growth to assess the relationship between OD and plate counts. Plates were incubated for 24 h at 37°C.
Conditions of near-optimal salinity (15 to 25‰) and temperature (37°), as determined from the rate of increase in OD and corresponding plate counts, were chosen for additional analysis by qPCR at both pH levels. Boiling lysis was used to extract DNA (7). One milliliter of culture was centrifuged for 10 min at 10,000 rpm. The pellet was washed with 1× PBS, and the centrifugation was repeated. The supernatant was discarded, and the pellet was resuspended in 1 ml of 1× PBS and then boiled at 100°C for 10 min to lyse the cells. The product of boiling lysis served as a template for qPCR performed with an ABI 7500 real-time PCR system (Applied Biosystems Inc., Carlsbad, CA). BRYT Green fluorescent dye (Promega Corporation, Madison, WI) was used to visualize the increase in the vvhA hemolysin gene amplicon by using previously published primers and procedure (7). The size of the expected 100-bp amplicon was confirmed by agarose gel electrophoresis. The average melting temperature (±standard deviation) of the amplicon was 81.23 ± 0.91°C, the average efficiency of the qPCR was 94.8%, and R2 of the standard curve was 0.996. The relationships between qPCR estimate of cell concentration, OD, and culture-dependent concentrations were determined at four time points during the exponential growth.
Curve fitting and statistical analyses.
Bacterial growth curves can be described by sigmoid functions, such as the logistic model (29, 38). The logistic equation has been previously modified from general mathematical to biological parameters that include specific growth rate (μ) (38). Relationships between OD, concentrations determined by plate counts (CFU ml−1), and total cell concentrations determined by qPCR analysis (cells ml−1) were calculated on log10-transformed data sets. Pairwise relationships between OD versus CFU ml−1 and cells ml−1 were determined by fitting a linear regression curve to each data set (Table 1). The corresponding data points were plotted versus time for each replicate, and self-start logistic curves were fitted using the nonlinear regression function of R 2.10.1 software (R Development Core Team, 2008; available at http://cran.r-project.org/). Logistic curves and their parameters were established following previously described procedures:
where y is cell concentration, A is the asymptote of the curve, λ is lag time (in hours), and μ is maximum growth rate (38).
Table 1.
Mean growth rates of six replicates based on CFU ml−1 versus time and total cells ml−1 versus time
| Salinity (ppt) | Mean growth rate (generations h−1) |
|||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Biotype 1 (CMCP6) |
Biotype 2 (ATCC 33147) |
Biotype 3 (302/99) |
||||||||||||||||
| pH 7 |
pH 8 |
pH 7 |
pH 8 |
pH 7 |
pH 8 |
|||||||||||||
| 25°C | 30°C | 37°C | 25°C | 30°C | 37°C | 25°C | 30°C | 37°C | 25°C | 30°C | 37°C | 25°C | 30°C | 37°C | 25°C | 30°C | 37°C | |
| CFU ml−1 vs time | ||||||||||||||||||
| 5 | 1.80 | 2.15 | 2.03 | 1.30 | 1.79 | 1.95 | 0.38 | 0.50 | 0.73 | 0.35 | 0.55 | 1.03 | 0.41 | 0.70 | 0.72 | 0.30 | 0.39 | 1.15 |
| 10 | 2.03 | 2.57 | 3.24 | 1.35 | 2.69 | 3.31 | 1.45 | 2.25 | 1.23 | 1.42 | 1.55 | 1.95 | 0.88 | 1.48 | 0.97 | 0.90 | 1.13 | 1.42 |
| 15 | 2.16 | 2.94 | 3.31 | 1.39 | 2.76 | 3.65 | 1.85 | 2.25 | 2.28 | 1.02 | 1.76 | 1.91 | 1.99 | 1.90 | 2.15 | 0.88 | 1.92 | 2.14 |
| 20 | 1.77 | 2.87 | 3.56 | 1.46 | 2.09 | 2.84 | 1.38 | 2.01 | 1.97 | 1.09 | 1.74 | 2.60 | 0.98 | 1.24 | 1.97 | 0.89 | 1.36 | 1.89 |
| 25 | 1.83 | 2.78 | 3.97 | 1.97 | 2.20 | 2.42 | 1.33 | 1.88 | 2.17 | 1.34 | 1.63 | 2.27 | 1.15 | 1.40 | 2.38 | 1.28 | 1.42 | 2.15 |
| 30 | 1.73 | 3.41 | 3.19 | 1.84 | 2.44 | 2.64 | 0.90 | 1.61 | 2.91 | 1.18 | 1.74 | 1.86 | 1.12 | 1.58 | 2.04 | 1.39 | 1.36 | 1.95 |
| 35 | 1.43 | 2.66 | 2.48 | 1.77 | 2.54 | 3.08 | 0.79 | 1.31 | 1.88 | 0.90 | 1.29 | 1.97 | 1.20 | 1.76 | 2.01 | 1.31 | 1.32 | 1.52 |
| 40 | 1.22 | 2.45 | 2.76 | 1.71 | 2.00 | 1.85 | 0.78 | 1.25 | 1.45 | 0.78 | 0.79 | 1.24 | 1.11 | 1.72 | 1.90 | 1.34 | 1.38 | 1.77 |
| Total cells ml−1 vs time (qPCR analysis) | ||||||||||||||||||
| 5 | 1.20 | 1.48 | 1.38 | 1.25 | 1.20 | 1.30 | 0.33 | 0.80 | 0.73 | 0.34 | 0.54 | 0.94 | 0.32 | 0.59 | 0.64 | 0.25 | 0.35 | 1.00 |
| 10 | 1.33 | 1.59 | 1.30 | 1.22 | 1.67 | 2.49 | 1.38 | 2.12 | 1.18 | 1.06 | 1.46 | 1.86 | 0.79 | 1.21 | 0.80 | 0.78 | 0.93 | 1.09 |
| 15 | 1.49 | 1.85 | 2.37 | 1.05 | 1.84 | 2.58 | 1.76 | 2.08 | 1.97 | 0.98 | 1.64 | 1.79 | 1.49 | 1.44 | 1.52 | 0.88 | 1.46 | 1.74 |
| 20 | 1.12 | 1.86 | 2.51 | 1.05 | 1.43 | 1.96 | 1.30 | 1.90 | 1.90 | 1.06 | 1.62 | 2.47 | 0.82 | 1.48 | 1.53 | 0.78 | 1.03 | 1.55 |
| 25 | 1.26 | 1.78 | 2.48 | 1.33 | 1.51 | 2.04 | 1.27 | 1.79 | 2.06 | 1.22 | 1.63 | 2.12 | 1.00 | 1.42 | 1.92 | 0.84 | 1.16 | 1.55 |
| 30 | 1.22 | 2.84 | 2.26 | 0.98 | 1.84 | 1.90 | 0.85 | 1.50 | 2.72 | 1.07 | 1.67 | 1.74 | 0.79 | 1.23 | 1.54 | 0.93 | 1.10 | 1.47 |
| 35 | 1.38 | 1.19 | 1.71 | 1.12 | 1.65 | 1.93 | 0.75 | 1.26 | 1.76 | 1.06 | 1.22 | 1.82 | 1.10 | 1.21 | 1.60 | 0.91 | 1.04 | 1.31 |
| 40 | 1.11 | 1.84 | 1.79 | 1.11 | 0.93 | 1.48 | 0.79 | 1.20 | 1.39 | 0.71 | 0.76 | 1.15 | 1.07 | 1.20 | 1.56 | 0.98 | 1.00 | 1.30 |
Growth rates, identified as maximum rates of change of the logistic model, μ (tangent lines at the point of inflection), were compared. Three-way analysis of variance (ANOVA) (Minitab Statistical software; State College, PA) was carried out to identify the statistical significance between all tested physical variables (temperature, pH, and salinity), as well as any interactions among them. The growth rates based on CFU ml−1 concentrations were further analyzed by one-way ANOVA with post hoc testing (Tukey-Kramer) to better identify statistical significance of differences among salinity levels within each temperature-pH group. Results were considered significant at the α-level of 0.05 (see Fig. 2).
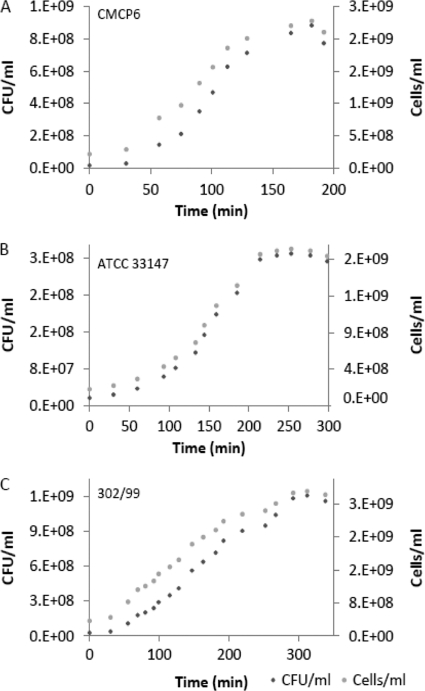
Fig. 2.
Comparison of growth curves obtained by culturing versus qPCR at 30°C, pH 7.0, and 20‰ salinity. (A) Biotype 1; (B) biotype 2; (C) biotype 3. Note the different scales of the y axes.
RESULTS
Growth rate calculations.
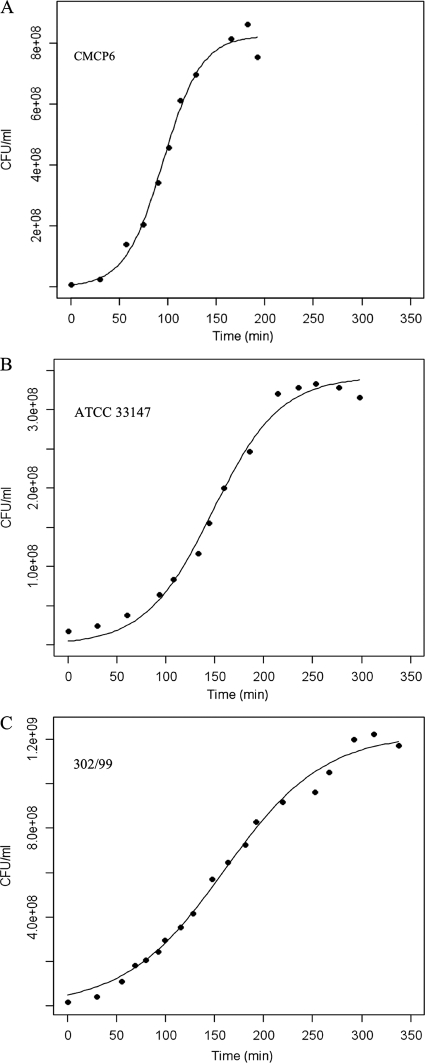
Logistic curves were fitted to culture and qPCR data generated during the exponential phase of growth to calculate μ, the specific growth rate for each condition. Figure 1 shows a comparison of the growth curve of the three biotypes based on culture data (CFU ml−1) at 30°C, pH 7.0, and 20‰ salinity. Growth rates under these conditions were 3.05, 1.75, and 1.20 generations h−1 for biotypes 1, 2, and 3, respectively. The fitting of data generated from other conditions and qPCR measurements to the logistic model was comparable to the strong fit shown in Fig. 1. Growth rates based on culture and qPCR data for all conditions are included in Table 1.
Fig. 1.
Growth curves based on culturable concentrations (CFU ml−1) for each V. vulnificus biotype at 30°C, pH 7.0, and 20‰ salinity fitted with the logistic model. (A) Biotype 1; (B) biotype 2; (C) biotype 3. The growth rates are 3.05, 1.75, and 1.20 generations h−1, respectively. Dots are actual data points, while the line represents the fitted logistic curve. Note the different scales of the y axes.
Measurements of V. vulnificus concentrations.
Growth of the three V. vulnificus biotypes was measured by three methods: (i) OD600 and (ii) culturable plate counts (CFU ml−1) for all conditions and (iii) total cell concentrations (cells ml−1) by qPCR analysis of the vvhA gene for conditions of 37°C, pH 7.0, and 15, 20, and 25‰ salinities. These conditions were all within the optimal range for each biotype, and no significant difference in growth rates among these salinity levels was observed. Pairwise relationships between OD, CFU ml−1, and cells ml−1 were determined by fitting a linear regression curve to each combination of data points during exponential growth (Table 2). The greatest correlations between concentrations determined by the three methods were always observed for biotype 1 (R2 = 0.67 to 0.90), followed by biotype 3 (R2 = 0.52 to 0.85) and biotype 2 (R2 = 0.40 to 0.80) (Table 2).
Table 2.
Relationships between estimates of cell concentrations obtained by various methods: optical density versus culture or optical density versus qPCR analysisa
| Biotype | Relationship |
||
|---|---|---|---|
| Log10 OD600 vs log10 CFU ml−1 | Log10 OD600 vs log10 cells ml−1 | Log10 CFU ml−1 vs log10 cells ml−1 | |
| 1 | y = 1.70x + 10.70 (R2 = 0.90) | y = 1.01x + 10.40 (R2 = 0.76) | y = 0.5279x + 4.6093 (R2 = 0.67) |
| 2 | y = 1.24x + 9.77 (R2 = 0.80) | y = 1.16x + 10.48 (R2 = 0.63) | y = 0.598x + 4.0589 (R2 = 0.40) |
| 3 | y = 1.70x + 10.92 (R2 = 0.85) | y = 1.13x + 10.64 (R2 = 0.70) | y = 0.5163x + 4.6614 (R2 = 0.52) |
The culture conditions were 37°C and pH 7.0, and data were combined for salinities of 15, 20, and 25‰.
Vibrio vulnificus concentrations determined by plate count (CFU ml−1) were always lower than corresponding average estimates of total cell numbers by qPCR analysis (cells ml−1) for all biotypes at all treatment and level combinations. An example of this relationship for each biotype is shown in Fig. 2, where culture conditions were 30°C, pH 7.0, and 20‰ salinity. The largest difference between the plate count and qPCR measurements was always observed during the lag phase. As the growth progressed, the magnitude of this difference declined (Fig. 2). The greatest discrepancy between the plate count and qPCR measurements was observed for biotype 1. A similar pattern was observed for biotype 3, while the least difference was observed for biotype 2.
Comparison of growth rates among biotypes.
The growth rates of the biotypes were compared using calculations based on qPCR and plate count data. The differences among the growth rates of all three biotypes were significant under all environmental conditions and for both measurement methods. When the specific growth rates measured at each environmental condition were compared, biotype 1 grew on average 1.7 and 1.9 times faster than biotypes 2 and 3, and biotype 2 grew on average 1.2 times faster than biotype 3. However, under the conditions nearest the optimum (37°C, pH 7.0, 15 to 30‰ salinity), the growth rates of biotype 3 were not significantly different from those of biotype 2. The growth rates based on plate counts were on average greater than those based on qPCR analysis by a factor of 1.5 for biotype 1, 1.1 for biotype 2, and 1.3 for biotype 3.
The effects of temperature, salinity, and pH on growth rates.
Three-way ANOVA of growth rates calculated from plate count data (CFU ml−1) was performed for all possible treatment and level combinations of environmental factors (Table 3). All strains grew best at 37°C, pH 7.0, and within the salinity range of 15 to 30‰. The factor that most affected the growth rate of biotype 1 was temperature, which was responsible for 43.1% of total variation among the growth rates. Salinity and pH resulted in 12.2 and 4.3% of total variation, respectively. The interactions of the variables, with the exception of pH and temperature, were significant, contributing to 19.6% of total variation. The factor that most affected growth of biotype 2 was salinity, followed by temperature and pH, which represented 34.0, 24.9, and 1.5% of total variation, respectively. Interactions of all parameters were significant and contributed 26.1% of the total variation. Biotype 3 produced results that were most similar to those of biotype 2, as the order of the parameters' effect on μ was salinity, temperature, and then pH. Interactions of all variables, except those between pH and temperature, were significant and accounted for 15.6% of total variation among the growth rates. Overall, the effects of the measured parameters and interactions among them accounted for 77.0, 84.2, and 79.2% of total variation in growth rate for biotypes 1, 2, and 3, respectively. The remainder of the variation was not accounted for (stochastic).
Table 3.
Contribution of individual variables and interaction among them (*) to the growth rates based on culture-dependent methods and qPCR for each V. vulnificus biotypea
| Biotype | Contribution to growth rate (%) |
|||||||
|---|---|---|---|---|---|---|---|---|
| Individual variable |
Interaction between variables |
All physical variables and interactions | ||||||
| pH | Temp | Salinity | pH*temp | pH*salinity | Temp*salinity | pH*temp*salinity | ||
| Culture-dependent methods | ||||||||
| CMCP6 biotype 1 | 4.3 | 43.1 | 12.2 | 0.5† | 3.7 | 6.6 | 8.7 | 77 |
| ATCC 33147 biotype 2 | 1.5 | 24.9 | 34 | 1 | 2.8 | 11.6 | 10.6 | 84.2 |
| 302/99 biotype 3 | 0.6 | 26.8 | 38.9 | 0.6† | 2 | 6.2 | 7.4 | 79.2 |
| qPCR analysis | ||||||||
| CMCP6 biotype 1 | 1 | 40.4 | 12.2 | 0.7† | 5.7 | 10.2 | 9.4 | 77.3 |
| ATCC 33147 biotype 2 | 0.3 | 23.4 | 42.6 | 0.1† | 7.7 | 11.4 | 11 | 87.1 |
| 302/99 biotype 3 | 5.1 | 26.5 | 42.1 | 1.2 | 1.7 | 3.9 | 7.8 | 85.4 |
The proportion of the variation in growth rate contributed by each variable is given. All factors significantly affected growth rates, except those marked with a dagger (†), which denotes that results are not significantly different.
Analyses of the growth rates based on qPCR analysis yielded comparable results (Table 3); i.e., the dominant factors affecting the growth rate of each biotype remained the same, although the attributed percentage varied slightly from that of culture-dependent methods.
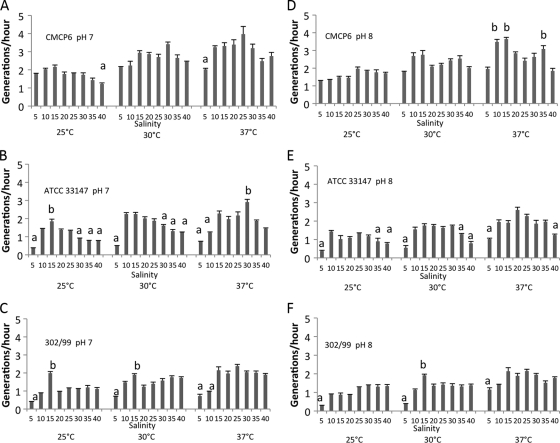
To further explore the effects of salinity, growth rates based on plate counts were compared (one-way ANOVA with a post hoc [Tukey-Kramer] test). The effects of salinity were analyzed separately for cultures grown at pH 7.0 and pH 8.0 and for each temperature (Fig. 3). Biotype 1 maintained comparable growth rates across the majority of the salinity range at pH 7.0 (Fig. 3A and D). The only significant differences (lower growth rates) were found at 5‰ salinity for cultures grown at 37°C and at 40‰ salinity for cultures grown at 25°C. At pH 8.0 and 37°C, the biotype 1 strain grew significantly faster at salinities of 10, 15 and 35‰ than at other salinities under the same temperature-pH conditions.
Fig. 3.
Growth rates of the three V. vulnificus biotypes determined by plate count (CFU ml−1) at salinities from 5 to 40‰ at three temperatures (25, 30, 37°C) and at pH 7.0 (A to C) and 8.0 (D to F). (A, D) Biotype 1; (B, E) biotype 2; (C, F) biotype 3. Values that share a letter designation (including no letter) within each temperature-pH group are not significantly different. Error bars represent standard errors.
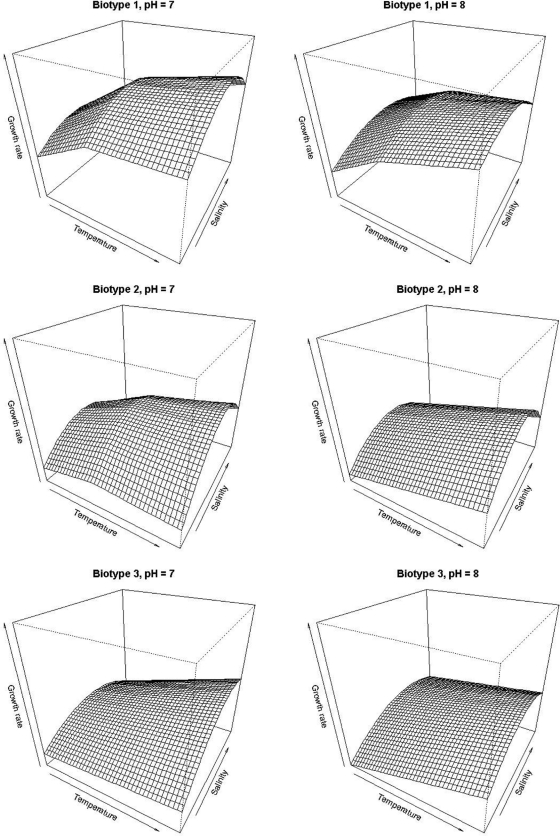
Variations in salinity had a greater effect than temperature and pH on the μ of biotype 2 (Fig. 3B and E). Cultures grown in media with salinities at the low and high ends of the range had consistently the lowest growth rates at both pH levels. At pH 7.0, a significant increase in μ was observed for 15‰ salinity at 25°C and at 30‰ at 37°C. Biotype 3 (Fig. 3C and F) grew significantly more slowly at the low end of the salinity range. Biotype 3 cultures grown at pH 7.0 at 15‰ and 25°C and 30°C had significantly higher growth rates than other salinities. Growth rates for each biotype were modeled as a function of salinity and temperature at pH 7.0 and 8.0 (Fig. 4). Growth rates were determined from plate count (CFU ml−1) concentrations. This figure is helpful for visually comparing the overall response of the biotypes' growth rates to the variables; e.g., note the great effect of low salinity on the growth rate of biotype 3 and insignificant effect of higher salinity.
Fig. 4.
Growth responses modeled as a function of temperature and salinity. Mean growth rates were standardized for the entire data set, where mean was set to zero and standard deviation was set to one, as supported by IAC (Intelligent Adaptive Curiosity; R Development Core Team, 2008; available at http://cran.r-project.org/). Temperature, salinity, and interactions were squared for the model. Temperature range was 25 to 37°C, salinity range was 5 to 40‰.
Supplementary measurements of growth rates were performed with V. vulnificus strains ATCC 33814 (biotype 1), ATCC 33149 (biotype 2), and 313/98 (biotype 3) (8) under selected conditions (37°C, pH 7.0, 15 to 25‰ salinity) (data not shown). The mean growth rates were compared to those observed for CMCP6, ATCC 33147, and 302/99 under the identical conditions. The same trends in growth rates were observed among these examples of the three biotypes, i.e., biotype 1 had a greater growth rate than biotype 2, which had a growth rate greater than or equal to that of biotype 3. Furthermore, no significant differences were found between the growth rates of strains within a biotype; e.g., the growth rate of biotype 1 CMCP6 was not significantly different than that of ATTC 33814.
DISCUSSION
In previous studies, the effects of environmental conditions on the growth rates of V. vulnificus were described for strains of unknown biotype (16, 35). We have determined that the growth rate of V. vulnificus in a nonselective, defined broth differed by biotype and that it was strongly affected by temperature and salinity and to a lesser extent by pH. These effects were observed whether the enumeration method was by culture (plate count) or qPCR analysis.
During our study, the fastest average generation time observed by culture methods was 15 min for biotype 1 (37°C, pH 7.0, 15 to 30‰ salinity). The generation time estimated by qPCR measurements under the same conditions was 24 min. The genome of V. vulnificus, as well as that of other members of the family Vibrionaceae, consists of two chromosomes. It has been suggested that the presence of two chromosomes could contribute to the high growth rate of V. vulnificus and other Vibrionaceae species (37). For example, V. parahaemolyticus and V. alginolyticus can double in 12 to 14 min. The most rapid generation time reported for Vibrio natriegens is 9.8 min (1).
Quantitative PCR analysis is a relatively novel method for measuring bacterial growth compared to culturable plate counts or optical density; we therefore compared growth rate measurements obtained using these methods. Enumeration by culture typically underestimates total cell concentrations. Optical density is a measurement of light scattering that reflects relative bacterial concentrations and must be correlated with another method, such as culturable counts, to determine cell concentrations (16, 23, 35). Total cell concentrations can be determined by qPCR; however, these measurements may also include free DNA and that of nonreplicating cells.
Concentrations based on qPCR measurements were always higher than concentrations based on plate counts for each biotype. Findings by others support the existence of discrepancies between culture-dependent and culture-independent measurements, with concentrations determined by culture-dependent methods being smaller in each case (12, 32). In our study, the greatest differences were always observed during the lag phase, and these discrepancies decreased as the cultures entered exponential phase. This phenomenon may be explained by the formation of viable but not culturable (VBNC) cells (26) in the inoculum (overnight) culture that were carried over into the fresh cultures. In that case, one may well be measuring the growth rate of two populations as the culture enters early exponential phase: one that is actively dividing and a minority population that has not yet begun to divide. Resuscitation (regaining culturability) of the VBNC cells during exponential phase may have closed the gap in concentrations determined by the two methods. The qPCR measurements would estimate a lower growth rate, because they would account for the initially unculturable cells, while the plate counts would not detect them. The existence of V. vulnificus VBNC cells and the health concerns they impose on shellfish consumers and recreational water users have been well documented (24); therefore, it is informative to compare the culture-dependent and culture-independent measurements of V. vulnificus levels.
Growth rates, determined as maximal rates of change of the fitted logistic curves (tangent lines at the point of inflection), were always larger when based on plate counts than when based on qPCR analysis. Whereas this is perhaps counterintuitive based on the fact that values of cell concentrations have the opposite relationship (estimates by qPCR are greater than those by culture methods), others have also observed differences between growth rates determined from culture-dependent and culture-independent estimates (32). Since the initial populations (lag phase and early exponential phase) likely contained relatively high proportions of nonculturable targets, the exponential phase of the respective logistic curves produced more shallow slopes. Consequently, the slopes of tangent lines at points of inflection of each logistic curve, which define the growth rate (38), were smaller. We therefore conclude that concentrations determined by plate counts yield higher estimates of specific growth rates of V. vulnificus than qPCR analysis, which must be taken into account if one is comparing data generated by the two methods.
Vibrio vulnificus strains are native to estuaries, which experience a high degree of physical-chemical variability, and are also human and fish pathogens (19, 30, 33). Therefore, the species must possess the capability to grow in a wide variety of different environmental conditions. In our study, all three physical variables (temperature, pH, and salinity) exerted significant effects on the growth of V. vulnificus. The growth of each biotype was affected primarily by temperature and salinity. The effect of pH was also significant but less prominent. All biotypes grew fastest at 37°C and pH 7.0 in this study. Biotype 1 was less affected by salt concentrations than biotypes 2 and 3. This suggests that biotype 1, a common estuarine bacterium which is responsible for most human infections (14), is better adapted for growth under diverse environmental conditions. The lesser tolerance to low or high salinity levels by biotypes 2 and 3 may contribute to their restricted host distribution in eels (4) and tilapia (5), respectively, as well as narrow geographical boundaries (14).
The interactions among individual variables, except between pH and temperature for biotypes 1 and 3, were significant contributors to total variability among the growth rates. Therefore, we conclude that pH, temperature, and salinity are interdependent variables with regard to the growth of V. vulnificus. Interestingly, analyses performed on growth rates based on plate counts (CFU ml−1), as well as qPCR analysis (cells ml−1), produced similar results, making this method suitable for evaluation of the effects of physical parameters on V. vulnificus growth. Moreover, the growth rates of additional strains (ATCC 33814 biotype 1, ATCC 33149 biotype 2, 2095/00 biotype 3), determined for selected, near-optimal conditions, supported the detailed growth rate studies on the other examples of the three biotypes. These data strongly suggest that growth rates of V. vulnificus differ by biotype.
Human infections are most frequently caused by biotype 1 strains, which are also most commonly isolated from environmental waters (2). Vibrio vulnificus infections are often characterized by rapidly fulminating septicemia and cellulitis or sepsis of a wound infection (13, 20). Biotype 1 V. vulnificus can be classified into E and C genotypes by randomly amplified polymorphic DNA (RAPD) (36). Environmental and clinical isolates commonly (but not exclusively) belong to genotypes E and C, respectively. During this study, strains representing biotype 1, which were both genotype C and of clinical origin, grew significantly faster than biotype 2 (both strains were confirmed eel pathogens) and biotype 3 under all experimental conditions. The fast onset of exponential phase and rapid growth may well contribute to the relative virulence of biotype 1 strains toward humans. This finding also may offer an explanation for why biotype 1 strains account for the majority of human pathogens, as the human body represents an assembly of conditions beneficial for their growth: stable temperature of 37°C, NaCl concentrations in blood of ∼9.0‰ (9), and blood pH just above 7.0.
In summary, temperature, pH, and salinity are interdependent variables that significantly affect the growth of all three V. vulnificus biotypes. Furthermore, the growth of V. vulnificus differs by biotype. Moreover, biotype 1 is more tolerant to low and high salinities and grows significantly faster than biotypes 2 and 3. These qualities likely contribute to its ubiquitous distribution, as well as to relatively frequent pathogenicity in humans. The qPCR method for quantification used in this study was well correlated with plate count measurements, particularly during mid- and late exponential phase, and served as a good indicator of the effects of environmental factors on V. vulnificus.
ACKNOWLEDGMENTS
We thank Theresa Tran for help with data generation, Kathleen Scott for help with medium development, Gordon Fox (USF) for help with statistical analysis, and Anita Wright (University of Florida) for providing V. vulnificus strains and for helpful discussions.
Footnotes
Published ahead of print on 22 April 2011.
REFERENCES
- 1. Aiyar S. E., Gaal T., Gourse R. L. 2002. rRNA promoter activity in the fast-growing bacterium Vibrio natriegens. J. Bacteriol. 184:1349–1358 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Amaro C., Biosca E. G., Fouz B., Garay E. 1992. Electrophoretic analysis of heterogeneous lipopolysaccharides from various strains of Vibrio vulnificus biotype-1 and biotype-2 by silver staining and immunoblotting. Curr. Microbiol. 25:99–104 [DOI] [PubMed] [Google Scholar]
- 3. Amaro C., et al. 1999. Isolation of Vibrio vulnificus serovar E from aquatic habitats in Taiwan. Appl. Environ. Microbiol. 65:1352–1355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Biosca E. G., Amaro C., Larsen J. L., Pedersen K. 1997. Phenotypic and genotypic characterization of Vibrio vulnificus: proposal for the substitution of the subspecific taxon biotype for serovar. Appl. Environ. Microbiol. 63:1460–1466 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Bisharat N., et al. 1999. Clinical, epidemiological, and microbiological features of Vibrio vulnificus biogroup 3 causing outbreaks of wound infection and bacteraemia in Israel. Lancet 354:1421–1424 [DOI] [PubMed] [Google Scholar]
- 6. Bisharat N., et al. 2005. Hybrid Vibrio vulnificus. Emerg. Infect. Dis. 11:30–35 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Campbell M. S., Wright A. C. 2003. Real-time PCR analysis of Vibrio vulnificus from oysters. Appl. Environ. Microbiol. 69:7137–7144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Chatzidaki-Livanis M., Hubbard M. A., Gordon K., Harwood V. J., Wright A. C. 2006. Genetic distinctions among clinical and environmental strains of Vibrio vulnificus. Appl. Environ. Microbiol. 72:6136–6141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Daniels R. 2004. Fluid, electrolyte, and acid-base balance, p. 971 In Nursing fundamentals: caring and clinical decision making. Thomson Learning, Inc., Clifton Park, NY [Google Scholar]
- 10. Fey A., et al. 2004. Establishment of a real-time PCR-based approach for accurate quantification of bacterial RNA targets in water, using Salmonella as a model organism. Appl. Environ. Microbiol. 70:3618–3623 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Fukushima H., Seki R. 2004. Ecology of Vibrio vulnificus and Vibrio parahaemolyticus in brackish environments of the Sada River in Shimane Prefecture, Japan. FEMS Microb. Ecol. 48:221–229 [DOI] [PubMed] [Google Scholar]
- 12. Gordon K. V., Vickery M. C., DePaola A., Staley C., Harwood V. J. 2008. Real-time PCR assays for quantification and differentiation of Vibrio vulnificus strains in oysters and water. Appl. Environ. Microbiol. 74:1704–1709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Gulig P. A., Bourdage K. L., Starks A. M. 2005. Molecular pathogenesis of Vibrio vulnificus. J. Microbiol. 43:118–131 [PubMed] [Google Scholar]
- 14. Jones M. K., Oliver J. D. 2009. Vibrio vulnificus: disease and pathogenesis. Infect. Immun. 77:1723–1733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Kaspar C. W., Tamplin M. L. 1993. Effects of temperature and salinity on the survival of Vibrio vulnificus in seawater and shellfish. Appl. Environ. Microbiol. 59:2425–2429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kelly M. T. 1982. Effect of temperature and salinity on Vibrio (Beneckea) vulnificus occurrence in a Gulf Coast environment. Appl. Environ. Microbiol. 44:820–824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Linkous D. A., Oliver J. D. 1999. Pathogenesis of Vibrio vulnificus. FEMS Microbiol. Lett. 174:207–214 [DOI] [PubMed] [Google Scholar]
- 18. Marco-Noales E., Biosca E. G., Amaro C. 1999. Effects of salinity and temperature on long-term survival of the eel pathogen Vibrio vulnificus biotype 2 (serovar E). Appl. Environ. Microbiol. 65:1117–1126 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Motes M. L., et al. 1998. Influence of water temperature and salinity on Vibrio vulnificus in northern Gulf and Atlantic Coast oysters (Crassostrea virginica). Appl. Environ. Microbiol. 64:1459–1465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Muldrew K. L., Miller R. R., Kressin M., Tang Y. W., Stratton C. 2007. Necrotizing fasciitis from Vibrio vulnificus in a patient with undiagnosed hepatitis and cirrhosis. J. Clin. Microbiol. 45:1058–1062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Nudelman A., Edelson G., Linden A., Raz R. 1997. Infection by Vibrio vulnificus after a prick from from the spine of a tilapia. Harefuah 133:444–445,502. [PubMed] [Google Scholar]
- 22. Nystrom T. 2004. Stationary-phase physiology. Ann. Rev. Microbiol. 58:161–181 [DOI] [PubMed] [Google Scholar]
- 23. Oh W. S., Im Y. S., Yeon K. Y., Yoon Y. J., Kim J. W. 2007. Phosphate and carbon source regulation of alkaline phosphatase and phospholipase in Vibrio vulnificus. J. Microbiol. 45:311–317 [PubMed] [Google Scholar]
- 24. Oliver J. D. 2010. Recent findings on the viable but nonculturable state in pathogenic bacteria. FEMS Microbiol. Rev. 34:415–425 [DOI] [PubMed] [Google Scholar]
- 25. Oliver J. D., Hite F., McDougald D., Andon N. L., Simpson L. M. 1995. Entry into, and resuscitation from, the viable but nonculturable state by Vibrio vulnificus in an estuarine environment. Appl. Environ. Microbiol. 61:2624–2630 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Oliver J. D., Nilsson L., Kjelleberg S. 1991. Formation of nonculturable Vibrio vulnificus cells and its relationship to the starvation state. Appl. Environ. Microbiol. 57:2640–2644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Panicker G., Bej A. K. 2005. Real-time PCR detection of Vibrio vulnificus in oysters: comparison of oligonucleotide primers and probes targeting vvhA. Appl. Environ. Microbiol. 71:5702–5709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Parvathi A., Kumar H. S., Karunasagar I., Karunasagar I. 2004. Detection and enumeration of Vibrio vulnificus in oysters from two estuaries along the southwest coast of India, using molecular methods. Appl. Environ. Microbiol. 70:6909–6913 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Perni S., Andrew P. W., Shama G. 2005. Estimating the maximum growth rate from microbial growth curves: definition is everything. Food Microbiol. 22:491–495 [Google Scholar]
- 30. Pfeffer C. S., Hite M. F., Oliver J. D. 2003. Ecology of Vibrio vulnificus in estuarine waters of eastern North Carolina. Appl. Environ. Microbiol. 69:3526–3531 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Randa M. A., Polz M. F., Lim E. 2004. Effects of temperature and salinity on Vibrio vulnificus population dynamics as assessed by quantitative PCR. Appl. Environ. Microbiol. 70:5469–5476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Reichert-Schwillinsky F., Pin C., Dzieciol M., Wagner M., Hein I. 2009. Stress- and growth rate-related differences between plate count and real-time PCR data during growth of Listeria monocytogenes. Appl. Environ. Microbiol. 75:2132–2138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Strom M. S., Paranjpye R. N. 2000. Epidemiology and pathogenesis of Vibrio vulnificus. Microbes Infect. 2:177–188 [DOI] [PubMed] [Google Scholar]
- 34. Tamplin M. L., Colwell R. R. 1986. Effects of microcosm salinity and organic substrate concentration on production of Vibrio cholerae enterotoxin. Appl. Environ. Microbiol. 52:297–301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Wang Y. L., Gu J. D. 2005. Influence of temperature, salinity and pH on the growth of environmental Aeromonas and Vibrio species isolated from Mai Po and the Inner Deep Bay Nature Reserve Ramsar Site of Hong Kong. J. Basic Microbiol. 45:83–93 [DOI] [PubMed] [Google Scholar]
- 36. Williams L. A., LaRock P. A. 1985. Temporal occurrence of Vibrio species and Aeromonas hydrophila in estuarine sediments. Appl. Environ. Microbiol. 50:1490–1495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Yamaichi Y., Iida T., Park K. S., Yamamoto K., Honda T. 1999. Physical and genetic map of the genome of Vibrio parahaemolyticus: presence of two chromosomes in Vibrio species. Mol. Microbiol. 31:1513–1521 [DOI] [PubMed] [Google Scholar]
- 38. Zwietering M. H., Jongenburger I., Rombouts F. M., Vantriet K. 1990. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 56:1875–1881 [DOI] [PMC free article] [PubMed] [Google Scholar]