Abstract
Modulation of intracellular calcium ([Ca2+]i) by erythropoietin (Epo) is an important signaling pathway controlling erythroid proliferation and differentiation. Transient receptor potential (TRP) channels TRPC3 and homologous TRPC6 are expressed on normal human erythroid precursors, but Epo stimulates an increase in [Ca2+]i through TRPC3 but not TRPC6. Here, the role of specific domains in the different responsiveness of TRPC3 and TRPC6 to erythropoietin was explored. TRPC3 and TRPC6 TRP domains differ in seven amino acids. Substitution of five amino acids (DDKPS) in the TRPC3 TRP domain with those of TRPC6 (EERVN) abolished the Epo-stimulated increase in [Ca2+]i. Substitution of EERVN in TRPC6 TRP domain with DDKPS in TRPC3 did not confer Epo responsiveness. However, substitution of TRPC6 TRP with DDKPS from TRPC3 TRP, as well as swapping the TRPC6 distal C terminus (C2) with that of TRPC3, resulted in a chimeric TRPC6 channel with Epo responsiveness similar to TRPC3. Substitution of TRPC6 with TRPC3 TRP and the putative TRPC3 C-terminal AMP-activated protein kinase (AMPK) binding site straddling TRPC3 C1/C2 also resulted in TRPC6 activation. In contrast, substitution of the TRPC3 C-terminal leucine zipper motif or TRPC3 phosphorylation sites Ser-681, Ser-708, or Ser-764 with TRPC6 sequence did not affect TRPC3 Epo responsiveness. TRPC3, but not TRPC6, and TRPC6 chimeras expressing TRPC3 C2 showed significantly increased plasma membrane insertion following Epo stimulation and substantial cytoskeletal association. The TRPC3 TRP domain, distal C terminus (C2), and AMPK binding site are critical elements that confer Epo responsiveness. In particular, the TRPC3 C2 and AMPK site are essential for association of TRPC3 with the cytoskeleton and increased channel translocation to the cell surface in response to Epo stimulation.
Keywords: AMP-activated kinase (AMPK), Calcium Channels, Erythropoeisis, Protein Motifs, TRP Channels, TRP Domain
Introduction
The importance of erythropoietin (Epo)2 in red blood cell production is demonstrated by the severe anemia and subsequent death at 11–15 days of gestation following homozygous deletion of the Epo or Epo receptor (Epo-R) genes in mice (1, 2). Many signaling pathways have been elucidated through which erythropoietin mediates its effects on erythroid proliferation and differentiation, but much remains unknown (3–7). For example, erythropoietin regulation of the intracellular calcium concentration ([Ca2+]i) has been shown to be important in the control of erythroid proliferation and differentiation (8–16). However, only recently have specific ion channels, members of the canonical transient receptor potential (TRPC) family, been identified in erythroid cells through which erythropoietin modulates calcium influx (17–21). TRPC3 is expressed on normal human erythroid progenitors and precursors and is regulated by erythropoietin (17). In contrast, the highly homologous TRPC6, also expressed on human erythroid cells, is not gated by erythropoietin and can inhibit calcium influx through TRPC3 (21).
The TRP superfamily of ion channels is a recently discovered group of calcium-permeable cation channels expressed in nonexcitable cells. These channels mediate a broad range of physiological processes (22–30). TRP channels function as homotetramers or heterotetramers, with the pore formed by loops between the fifth and sixth transmembrane domains. Interaction of heteromeric monomers in the heterotetramer or homomeric channel isoforms in the homotetramer is a modality of channel regulation (31–34). There is a growing number of diseases in which TRP channel involvement is recognized (35).
The TRPC subfamily, including TRPC3 and TRPC6, has important biological functions (36–43). TRPC3 activation involves a number of signaling pathways. TRPC3, like many other TRPC channels, associates with phospholipase Cγ (PLCγ) and the inositol 1,4,5-trisphosphate receptor, which modulate its activation (17, 43). TRPC3 can also be activated by phospholipase D (40). TRPC3 cell surface translocation is stimulated by a number of agonists, including erythropoietin and vasopressin (21, 36, 44). TRPC3 is of physiological importance in a number of tissues, including brain (42), hematopoietic cells (17, 41), and heart (45, 46). TRPC3 and TRPC6 have significant sequence homology. Both TRPC3 and TRPC6 can be activated by VEGF (47, 48), promote cardiac hypertrophy (39, 49), and contribute to the progression of certain malignancies (50–53). However, their sequence is sufficiently different that a number of activation pathways and agonist responses differ. For example, TRPC3 but not TRPC6 is activated by erythropoietin to increase [Ca2+]i. In addition, TRPC3 expression increases during normal human progenitor differentiation, but TRPC6 expression decreases. These observations suggest that the ratio of TRPC3/TRPC6 expression and activity differences are physiologically important (21).
Because TRPC3 and TRPC6 are involved in a number of diverse physiological processes, the goal of this work was to identify domains of functional importance. Individual domains that differ between TRPC3 and TRPC6 were examined to determine their role in the Epo-stimulated rise in [Ca2+]i. The TRP domain of TRPC3 was required for channel response to Epo, and substitution of five amino acids with those of TRPC6 abolished channel function. In contrast, the TRP domain of TRPC3 alone was not sufficient to restore Epo responsiveness to TRPC6. However, substitution of TRPC6 with both TRPC3 TRP and either the distal C terminus of TRPC3 (C2) or TRPC3 amino acids 741–748, the putative AMP-activated protein kinase (AMPK) binding site that straddles C1 and C2, resulted in a significant increase in [Ca2+]i in response to Epo. In contrast, TRPC3 leucine zipper motifs and specific phosphorylation sites were substituted with corresponding sequence from TRPC6 without substantial loss of activity. TRPC3, TRPC6, and all chimeras had substantial membrane localization. However, an increase in channel membrane insertion was observed after Epo stimulation of cells expressing Epo-R and TRPC3 or TRPC6 chimeras containing the distal C terminus of TRPC3 or the AMPK site but not wild type TRPC6. Cell fractionation also revealed that only TRPC3, TRPC6 chimeras expressing the distal TRPC3 C terminus, or the AMPK site showed substantial cytoskeletal association. This suggests that both TRPC3 C2 and the AMPK site may have a functional role in membrane insertion in response to Epo. We conclude that the TRP domain, the distal C terminus, and the AMPK site of TRPC3 are essential for channel responsiveness to Epo. We hypothesize that the Epo-induced [Ca2+]i increase is at least partly mediated by translocation of TRPC3 channels to the cell membrane via the cytoskeletal network.
EXPERIMENTAL PROCEDURES
Tissues and Cell Lines
Human embryonic kidney (HEK) 293T cells were cultured in Dulbecco's modified Eagle's medium (DMEM) with 10% fetal calf serum. In some experiments, cells were treated with 40 units/ml erythropoietin (Amgen, Thousand Oak, CA). UT-7/Epo cells, an erythropoietin-dependent cell line derived from the human leukemia cell line UT-7, were cultured in Iscove's modified Dulbecco's medium with 10% fetal calf serum and 0.5 unit/ml Epo.
Transfection of Human TRPC3, Human TRPC6, and Epo-R into HEK 293T Cells
Human TRPC3 and TRPC6 (both gifts of Dr. Lutz Birnbaumer) were subcloned into pQBI50 (QBiogene, Carlsbad, CA; BFP-TRPC3), pcDNA 3.1/V5-His TOPO (Invitrogen), or pCMV-Tag (Stratagene, La Jolla, CA). Chimeric channels were constructed in which domains in the C terminus of TRPC3 and TRPC6 were exchanged (Fig. 1). The construction of TRPC3-C6C1 and TRPC3-C6C2 chimeras was described previously (21). TRPC3-C6TRP, TRPC3-C6LZ, TRPC3 S681A, TRPC3 S708N, TRPC3 S764H, TRPC3-C6 802–809, TRPC3-C6 802–807, TRPC3-C6 804–809, TRPC6-C3TRP, TRPC6-C3TRP-C3C2, TRPC6-C3LZ, and TRPC6-C3TRP-C3 741–748 substitution mutants and chimeras were generated as described below and cloned into vectors as noted. HEK 293T cells at 50–70% confluence were transfected with these vectors and/or pTracer-CMV expressing Epo-R using Lipofectamine 2000 (Invitrogen) in accordance with the manufacturers' recommended protocols. Cells were routinely transfected with 1 μg of each plasmid in 7.5 ml of Opti-MEM in a 100-mm dish. HEK 293T cells were routinely studied 24–48 h after transfection.
FIGURE 1.
Schema of TRPC3/TRPC6 chimera. A, domains and motifs in the TRPC3 C terminus. B, representation of the proximal (C1) and distal (C2) C termini of TRPC3 and TRPC6. C, schematic models of TRPC3 chimeras: TRPC3-C6C1, TRPC3-C6C2, and TRPC3-C6TRP. D, schematic models of TRPC6 chimeras: TRPC6-C3C1, TRPC6-C3C2, TRPC6-C3TRP, and TRPC6-C3TRP-C3C2. For B–D, the origin of the TRP domain is shown by the color of the oval. The amino acid (AA) number at the beginning and at the end of the C1 and C2 domains contributed by TRPC3 (top) or TRPC6 (bottom) is indicated. FLAG-tagged chimeras expressed FLAG at the N terminus, and V5-tagged chimeras expressed V5 at the C terminus of the channel.
Generation of Chimeras
TRPC6-C3C1 and TRPC6-C3C2
TRPC6-C3C1 included TRPC6 amino acids 1–727 and 808–931 and TRPC3 amino acids 671–746 (C3C1) (Fig. 1D). TRPC6-C3C2 included TRPC6 amino acids 1–807, and TRPC3 included amino acids 747–848 (C3C2) (Fig. 1D). The 5′ (C1) and 3′ (C2) parts of C termini were exchanged between TRPC3 and TRPC6 using a megaprimer-mediated domain swapping PCR technique. For FLAG-TRPC6-C3C1, a 255-bp megaprimer was amplified using the FLAG-TRPC6-C3C chimera as a template and the following primers: forward primer, 5′- TTAATTGCCATGATCAATAGCTCATATCAAG-3′; reverse primer, 5′- AATTCCAAGTTTCTTCTTTGAGTTACCCATTC-3′. Normal type indicates the sequence of primers derived from TRPC6, and boldface type indicates the sequence derived from TRPC3. For FLAG-TRPC6-C3C2, 733-bp megaprimer was amplified using FLAG-TRPC3-C6C1 as a template and the following primers: forward primer, 5′-AACAGTTCATTCCAGGAAATTG-3′; reverse primer, 5′-TAATACGACTCACTATAGGGCG-3′ (binding to the T7 promoter in the pCMV vector of the FLAG construct). All chimeras were subcloned into vectors as noted, and clones were verified by sequencing.
TRPC3-C6TRP, TRPC6-C3TRP, and TRPC6-C3TRP-C3C2 Chimeras
Substitution of TRP domains was performed with PCR-based site-directed mutagenesis using the QuikChange multisite-directed mutagenesis kit (Stratagene) according to the manufacturer's protocol. To construct V5-TRPC3-C6TRP, V5-TRPC3 was used as a template with the PAGE-purified single reversed primer 5′- TGGACTAGGAACTAGATTGAAAGGTACAGGTAATGTTCTTCCCTCCTCAAAATAGGATAACCAAAGTTTTGAACGAG-3′ (the six nucleotides marked in boldface type introduced five amino acid changes in the C3TRP domain; Figs. 1C and 2A). To construct FLAG-TRPC6-C3TRP, FLAG-TRPC6 was used as a template with the PAGE-purified single forward primer 5′- ACTCTGGTTTTCCTACTTTGATGATGGCAAAACACTTCCTCCACCCTTCAGTCTGGTGCCGA-3′ (the six nucleotides marked in boldface type introduced five amino acids changes in the C3TRP domain; Figs. 1D and 2A). Generation of the TRPC6-C3TRP-C3C2 chimera was as described for the TRPC6-C3TRP chimera with the exception that TRPC6-C3C2 template was used instead of the TRPC6 template.
FIGURE 2.
TRP domains, leucine zipper motifs, and AMPK binding site in TRPC3 and TRPC6. A, amino acid compositions of TRP domains and leucine zipper motifs of TRPC3 and TRPC6 are shown. Letters in boldface type indicate exchanged amino acids in the chimeras. For TRP domain exchange, the boldface sequences in the TRP domain of TRPC6 were exchanged with those of TRPC3 to create TRPC3-C6TRP. The boldface amino acids of TRPC3 TRP were exchanged with those of TRPC6 to create TRPC6-C3TRP. For leucine zipper exchange, the boldface sequences in the leucine zipper of TRPC6 were exchanged with those of TRPC3 to create TRPC3-C6LZ. The boldface amino acids of TRPC3 were exchanged with those of TRPC6 leucine zipper to create TRPC6-C3LZ. B, exchange of TRPC3 741–748 and TRPC6 802–809 is shown. Amino acids contributed by TRPC3 (top) or TRPC6 (bottom) are indicated, and localization to the C1 or C2 part of the C terminus is shown.
TRPC3-C6LZ and TRPC6-C3LZ Chimeras
Leucine zipper motif exchange chimeras were prepared by PCR-directed mutagenesis. V5-TRPC3 was used as a template for TRPC3 chimera, and FLAG-TRPC6 was used as a template for TRPC6 chimeras. The following primers were used: for TRPC3-C6LZ, forward primer (5′-CCTAGTCCAAAATCATTATTTTATCTCCTCTTGAAGATTGTTAACTTTCCCAAATGC-3′) and reverse primer (5′-GCATTTGGGAAAGTTAACAATCTTCAAGAGGAGATAAAATAATGATTTTGGACTAGG-3′) (substituted nucleotides introduced the substitution of six amino acids marked in boldface type in Fig. 2A); for TRPC6-C3LZ, forward primer (5′-GGTGCCGAGTCCAAAGTCCTTTGTTTATTTCATAATGCGGCTTAAAAAATGG-3′) and reverse primer (5′-CCATTTTTTAAGCCGCATTATGAAATAAACAAAGGACTTTGGACTCGGCACC-3′) (substituted nucleotides introduced the substitution of the six amino acids marked in boldface type in Fig. 2A).
TRPC3 Phosphorylation Mutants
S681A, S708N, and S764H were prepared with PCR-based site-directed mutagenesis using the QuikChange multisite-directed mutagenesis kit (Stratagene) and PAGE-purified primers according to the manufacturer's protocol. The V5-TRPC3 was used as a template. The following single, reversed primers were as follows: GCAAACTTCCATTCTACATCAGCGTCATCCTCAATTTCTTGATA (two nucleotide changes substituting alanine for serine 681 are marked in boldface type); GATTTTGGACTAGGAACTAGATTGAAAGGTGGAGGTAATGTTT (one nucleotide change substituting asparagine for serine 708 is marked in boldface type); TGATTGAGAATGCTGTTAAAATGGTGTGATTCAAAAACTCTTGA (two nucleotide changes substituting histidine for serine 764 are marked in boldface type).
TRPC3-C6 802–809 Mutants
The TRPC3-C6 802–809 mutant was prepared using two-step PCR. In the first step, two separated but overlapping fragments were amplified using TRPC3 as a template; a fragment containing mutation 802–809 was amplified with forward primer 5′-AACAAGATTAACGAAGAGAAGAAGTTAAACCTCTTCACTCAGTCT-3′ and reverse primer 5′-CATTCACATCTCAGCATGCTG-3′, and another fragment was amplified using forward primer 5′-TCCATGGAGGGAAGCCCATCC-3′ and reverse primer 5′-CTTCTTCTCTTCGTTAATCTTGTCCATTTCTATATCCTTCTGAAG-3′. The sequence in boldface type represents the 802–809 substitution (Fig. 2B). In the second step, both products were mixed. Forward primer for the second step fragment and reverse primer for the first fragment were used to amplify the final product. TRPC3-C6 802–807 and the TRPC3-C6 804–809 chimeras were prepared with the QuikChange multisite-directed mutagenesis kit (Stratagene) using TRPC3-C6 802–809 as a template. TRPC3-C6 802–807 used the primer 5′-AGTGAAGAGGTTTAACCTGGACTCTTCGTTAATCTT-3′, and TRPC3-C6 804–809 used the primer 5′-CTTCTCTTCGTTAATCATTCCCATTTCTATATCCTT-3′.
TRPC6-C3 741–748 Chimera
This chimera was prepared using the QuikChange multisite-directed mutagenesis kit (Stratagene), FLAG-TRPC6 as a template, and the following primer: 5′-GATGCAGAGATGGGCATGGGAAATTCAAAATCGAGACTTGGAATTTTA-3′ (Fig. 2B).
Measurement of [Ca2+]i with Digital Video Imaging
HEK 293T cells were transfected with 1 μg/1.5 ml empty pQBI50 vector; pQBI50 vector expressing wild type TRPC3, TRPC6, and/or chimeric TRPC3/TRPC6 channels; and pTracer-CMV expressing Epo-R in a 35-mm dish. pTracer-CMV contains a CMV promoter, which drives expression of Epo-R and an SV40 promoter, which drives expression of GFP. The pQBI50 vector uses a CMV promoter to drive expression of BFP fused to the indicated channel. Successful transfection of individual HEK 293T cells with pQBI50 vectors was verified by detection of BFP (excitation, 380 nm; emission, 435 nm) and transfection of pTracer-CMV by detection of GFP (excitation, 478 nm; emission, 535 nm) with our fluorescence microscopy-coupled digital video imaging system (8, 16, 18). To study changes in [Ca2+]i in transfected cells, we used the fluorescent indicator Fura Red (excitation, 440 and 490 nm; emission, 600 nm long pass), a dual wavelength excitation probe (54, 55). At 48 h post-transfection, HEK 293T cells were loaded with 5 μm Fura Red-AM (Molecular Probes, Inc., Eugene, OR) for 20–25 min at 37 °C. The extracellular buffer routinely contained 0.68 mm CaCl2. HEK 293T cells were then treated with 0–40 units/ml recombinant Epo or vehicle (PBS). [Ca2+]i was measured in individual cells at base line and at 1–5-min intervals for 20 min by determining the fluorescence intensity ratio R (F440/F490).
Immunoblotting and Immunoprecipitation
For Western blotting, whole cell lysates (100 μg/lane) or immunoprecipitates were separated on 8 or 10% polyacrylamide gels, followed by transfer to Hybond-C Extra membranes (Amersham Biosciences). Western blotting was performed as described previously (19). Blots were incubated with anti-TRPC3 (1:400; Alomone Laboratories, Jerusalem, Israel), anti-TRPC3-C targeted to the human C-terminal sequence RRRRLQDIEMGMGNSKSRLN (1:500 to 1:1000, Bethyl Laboratories, Inc., Montgomery, TX) (33), anti-TRPC3-N targeted to the murine N-terminal sequence LNGDLESAEPLERHGHKASL (1:1000; Bethyl Laboratories, Inc.) (56), anti-TRPC6 (1:250; Alomone Laboratories), anti-V5-HRP (1:10,000; Invitrogen), anti-FLAG (1:1000; Sigma), anti-PLCγ (SC-81, 1:1000; Santa Cruz Biotechnology, Inc., Santa Cruz, CA), anti-Epo-R (SC-697; 1:1000; Santa Cruz Biotechnology, Inc.), anti-actin (1:10,000; Sigma), anti-tubulin (1:10,000; Sigma), anti-GAPDH (1:5000; Cell Signaling Technology, Inc., Boston, MA), anti-Na+K+-ATPase (1:1000; Cell Signaling Technology, Inc.), anti-lamin (1:1000; Cell Signaling Technology, Inc.), anti-vimentin (1:1000; Cell Signaling Technology), or streptavidin-HRP (Pierce) antibodies. Blots were washed and incubated with the appropriate horseradish peroxidase (HRP)-conjugated antibodies (1:2000). Enhanced chemiluminescence (ECL) was used for the detection of signal.
To examine the association of TRPC3 with Epo-R or PLCγ, HEK 293T cells were transfected with TRPC3 or TRPC3-C6TRP in pcDNA3.1/V5-His TOPO or in pCMV-tag and in human Epo-R (in pcDNA3) or rat PLCγ1 (in pcDNA3). Cells were washed in ice-cold Hanks' balanced salt solution and lysed in buffer (50 mm Tris, pH 7.5, 150 mm NaCl, 1 mm EDTA, 1% Triton X-100) supplemented with Complete protease inhibitor mixture (Roche Applied Science) and phosphatase inhibitor 2 (Sigma). Protein lysates (1 mg/0.5 ml) were incubated with protein A-Sepharose (50 μl of 50%) for 1 h at 4 °C. Precleared lysates were collected and incubated with preimmune rabbit serum (4 μg/500 μl) or anti-V5 (Invitrogen), anti-Epo-R, or anti-PLCγ1 antibodies at 4 μg/500 μl and Protein G-Sepharose (50 μg of 50% in 500 μl) or anti-FLAG-agarose (40 μg of 50% in 500 μl; Sigma) for 2 h at 4 °C. Immunoprecipitates were washed three times. FLAG immunoprecipitates were eluted using FLAG peptide (Sigma). Sample buffer (2×) was added to the pellets or eluates. The samples were heated at 60 °C for 30 min. Western blotting was performed as described above, and blots were probed with anti-V5-HRP, or anti-FLAG, anti-Epo-R, anti-PLCγ1, or anti-actin antibodies, followed by the appropriate HRP-conjugated secondary antibodies and ECL.
Cell Surface Localization and Cell Fractionation
To determine the cell surface localization of TRPC3, TRPC6, and chimeras, cell surface biotinylation was performed, followed by immunoprecipitation and Western blotting as described previously (21). To investigate the differences in protein localization and protein-protein interaction in different cell compartments, cell fractionation was performed on either transfected HEK 293T cells or UT-7/Epo cells using the Qiagen cell fractionation kit according to the manufacturer's protocol. Proteins obtained from different cell fractions were concentrated using Pall Life Sciences (Ann Arbor, MI) Nanosep 10K Omega centrifugal devices, and Western blotting was performed by loading equivalent amounts of protein in each lane for each fraction. To confirm results, HEK 293T cells were also fractionated with 0.5% Triton X-100 extraction modified from a previously published method with 1% Triton (57).
RESULTS
TRPC3 C Terminus Is Required for Epo Activation
We previously demonstrated that the TRPC3 C terminus contains domains critical for TRPC3 channel activation by Epo (17, 21). When the proximal TRPC3 C terminus, C3C1, was substituted with the sequences of C6C1 (TRPC3-C6C1; Fig. 1C), the Epo-stimulated increase in [Ca2+]i, expressed as F440/F490, observed in cells expressing Epo-R and wild type TRPC3 was abolished (supplemental Fig. 1A). Substitution of the distal TRPC3 C terminus, C3C2, with the sequences of C6C2, TRPC3-C6C2, modestly but significantly reduced the Epo-stimulated increase in F440/F490 compared with TRPC3 (supplemental Fig. 1B). By contrast, substitution of neither the C1 nor C2 domains of TRPC6 with TRPC3 sequences (TRPC6-C3C1 and TRPC6-C3C2; Fig. 1D) reconstituted channel activation by Epo (supplemental Fig. 1). These data suggest that both the proximal (C3C1) and distal (C3C2) C-terminal segments of TRPC3 contain elements that are necessary but not sufficient for channel activation by Epo.
The TRPC3 TRP Domain Is Essential for Epo Activation
The role of specific domains in the C1 region in TRPC3 activation by Epo was next examined, initially focusing on the TRP domain and leucine zipper motifs (Fig. 2A). Substitution of five amino acids in the TRP domain of TRPC3 (DDKPS) with those of TRPC6 (EERVN) (TRPC3-C6TRP) significantly reduced the response of TRPC3 to Epo stimulation (Fig. 3). However, substitution of the TRP domain of TRPC6 with that of TRPC3 (TRPC6-C3TRP) did not confer Epo responsiveness to the chimeric TRPC6 channel (Fig. 3). These data demonstrate that the C3TRP domain is essential but not sufficient for channel activation by Epo.
FIGURE 3.
Role of TRP domains in regulation of TRPC3 and TRPC6 by Epo-R. HEK 293T cells were transfected with BFP-TRPC3, BFP-TRPC6, BFP-TRPC3-C6TRP, or BFP-TRPC6-C3TRP chimeras and Epo-R. Fura Red-loaded cells were treated with 40 units/ml Epo. To quantitate [Ca2+]i, F440/F490 was measured at base line and by monitoring over 20 min after Epo stimulation. Shown is the percentage increase in F440/F490 above base line (mean ± S.E. (error bars) percentage increase) = peak F440/F490 divided by base line F440/F490 × 100% − 100% (base line). The numbers of individual cells studied were as follows: BFP-TRPC3 (PBS 17, Epo 21), BFP-TRPC6 (PBS 17, Epo 22), BFP-TRPC3-C6TRP (PBS 16, Epo 22), or BFP-TRPC6-C3TRP (PBS 18, Epo 21). The Epo-stimulated increase in cells expressing TRPC6 and Epo-R is not statistically different from cells expressing Epo-R alone and is thought to be secondary to Epo-R activation of low levels of endogenous channels (17). *, significantly greater percentage increase in F440/F490 compared with Epo-stimulated cells expressing wild type TRPC6 (p < 0.001). **, significantly less percentage increase in F440/F490 compared with Epo-stimulated cells expressing wild type TRPC3 (p < 0.001).
Substitution of the leucine zipper motif of TRPC3 with that of TRPC6 (TRPC3-C6LZ; Fig. 2A) did not alter the Epo-stimulated rise in [Ca2+]i through TRPC3 (supplemental Fig. 2). The reciprocal substitution (TRPC6-C3LZ) minimally increased the response of TRPC6. These data demonstrate that the TRPC3 leucine zipper motif has a minimal role in Epo-R-induced [Ca2+]i increase.
C3TRP and C3C2 Domains Are Sufficient to Reconstitute TRPC6 Response to Epo Activation
When cells were transfected with TRPC6-C3TRP-C3C2 (Fig. 1D), the Epo-stimulated [Ca2+]i increase was significantly higher than in cells transfected with either wild type TRPC6, TRPC6-C3TRP, or TRPC6-C3C2 chimeras and approached the response of wild type TRPC3 (Fig. 4). These data suggest that TRPC3-C2 contains domains that interact with C3TRP to form an Epo-responsive channel.
FIGURE 4.
Role of the distal C terminus of TRPC3 and TRPC6 in regulation by Epo-R. HEK 293T cells were transfected with BFP-TRPC3, BFP-TRPC6, BFP-TRPC6-C3TRP, BFP-TRPC6-C3C2, BFP-TRPC6-C3TRP-C3C2, BFP-TRPC3-C6 802–809, or BFP-TRPC6-C3TRP-C3 741–748 chimeras and Epo-R. Fura Red-loaded cells were treated with 40 units/ml Epo. To quantitate [Ca2+]i, F440/F490 was measured at base line and by monitoring over 20 min after Epo stimulation. Shown is the percentage increase in F440/F490 above base line (mean ± S.E. (error bars) percentage increase) = peak F440/F490 divided by base line F440/F490 × 100% − 100% (base line). The numbers of individual cells studied were as follows: BFP-TRPC3 (PBS 56, Epo 102), BFP-TRPC6 (PBS 57, Epo 103), BFP-TRPC6-C3TRP (PBS 20, Epo 34), BFP-TRPC6-C3C2 (PBS 20, Epo 35), BFP-TRPC6-C3TRP-C3C2 (PBS 30, Epo 64), BFP-TRPC3-C6 802–809 (PBS 26, Epo 49), or BFP-TRPC6-C3TRP-C3 741–748 (PBS 12, Epo 30). *, significantly greater percentage increase in F440/F490 compared with Epo-stimulated cells expressing wild type TRPC6 (p < 0.001). **, significantly less percentage increase in F440/F490 compared with Epo-stimulated cells expressing wild type TRPC3 (p < 0.001).
TRPC3 Amino Acids 741–748 Are Involved in Epo Activation
Three potential sites of serine phosphorylation of TRPC3 were identified and mutated; serine 681 (predicted casein kinase site; World Wide Web motif search) was mutated to alanine (S681A), serine 708 (predicted GSK site) was mutated to asparagine (S708N), and serine 764 (predicted PDK1 site) was mutated to histidine (S764H). Cells expressing these TRPC3 single substitution mutants had a response to Epo activation similar to that of wild type TRPC3 (data not shown).
The predicted binding site for AMPK in TRPC3 lies in a region spanning residues 740–754, with serine 747 as a potential phosphorylation site. When eight amino acids (residues 741–748) in TRPC3 were substituted with the corresponding amino acids in TRPC6 (TRPC3-C6 802–809; Fig. 2B), there was significant reduction in the rise in [Ca2+]i after Epo activation (Fig. 4). Constructs consisting of TRPC3-C6 802–807 (52 ± 1% increase in F440/F490 above base line) and TRPC3-C6 804–809 (47 ± 1% increase) also showed significantly less rise in F440/F490 after Epo stimulation compared with TRPC3 (56 ± 0.5% increase). A chimera consisting of TRPC6-C3TRP-C3 741–748 containing both C3TRP and C3 741–748 resulted in Epo-induced increases in F440/F490 that were similar to those of TRPC6-C3TRP-C3C2. This important observation indicates that both the TRPC3 TRP domain and AMPK binding site, which spans both TRPC3 C1 and C2 domains, are involved in conferring Epo-responsiveness.
The TRP Domain Does Not Regulate TRPC3 Channel Association with Epo-R
The role of the TRP domain in interaction of TRPC3 or TRPC6 with Epo-R was examined next. Reciprocal immunoprecipitation with anti-V5 (six experiments), anti-FLAG (seven experiments), or anti-Epo-R was performed. There was no statistically significant difference in the association of Epo-R with V5-TRPC3 compared with V5-TRPC3-C6TRP (Fig. 5A). Results were similar when we compared the association of FLAG-tagged TRPC3 with Epo-R with that of FLAG-TRPC3-C6TRP (Fig. 6A). We previously observed that FLAG-TRPC3 interacted significantly better with Epo-R than did FLAG-TRPC6 (21). These data show that although the TRPC3 TRP domain is of key importance in channel gating, it is not a critical domain in terms of protein-protein interaction between TRPC3 and Epo-R.
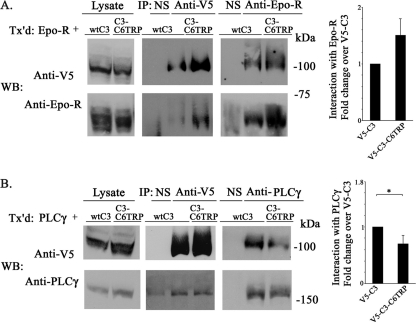
FIGURE 5.
Interaction of V5-TRPC3 and V5-TRPC3-C6TRP with Epo-R and PLCγ. A, HEK 293T cells were transfected (Tx'd) with Epo-R and V5-TRPC3 or V5-TRPC3-C6TRP. Lysates were immunoprecipitated (IP) with anti-V5, anti-Epo-R antibodies, or normal rabbit serum (NS). Western blots (WB) of lysates and immunoprecipitates were probed with anti-V5-HRP or anti-Epo-R and appropriate secondary antibodies. Western blotting of lysates demonstrated equivalent input from each construct. A representative result of six experiments is shown. Band intensities in A were quantitated with densitometry, and the mean ± S.E. (error bars) ratio of Epo-R/channel, normalized to wild type TRPC3, from six experiments is shown. There was no significant difference in interaction of Epo-R with TRPC3 compared with TRPC3-C6TRP. B, HEK 293T cells were transfected with PLCγ and V5-TRPC3 or V5-TRPC3-C6TRP. Lysates were immunoprecipitated with anti-V5, anti-PLCγ antibodies, or normal rabbit serum (NS). Western blots of lysates and immunoprecipitates were probed with anti-V5-HRP or anti-PLCγ and appropriate secondary antibodies. A representative result of four experiments is shown. Band intensities in B were quantitated with densitometry, and mean ± S.E. ratio of PLCγ/channel, normalized to TRPC3, from four experiments is shown. *, significant difference in the ratio compared with V5-TRPC3 (p < 0.05).
FIGURE 6.
Interaction of FLAG-TRPC3 and FLAG-TRPC3-C6TRP with Epo-R and PLCγ. A, HEK 293T cells were transfected (Tx'd) with Epo-R and FLAG-TRPC3 or FLAG-TRPC3-C6TRP. Lysates were immunoprecipitated (IP) with anti-FLAG-agarose. Western blots (WB) of lysates and immunoprecipitates were probed with anti-FLAG or anti-Epo-R and appropriate secondary antibodies. A representative result of seven experiments is shown. B, HEK 293T cells were transfected with PLCγ and FLAG-TRPC3 or FLAG-TRPC3-C6TRP. Lysates were immunoprecipitated with anti-FLAG-agarose. Western blots of lysates and immunoprecipitates were probed with anti-FLAG or anti-PLCγ and appropriate secondary antibodies. A representative result of three experiments is shown. C and D, intensity of bands was quantitated with densitometry, and the mean ± S.E. (error bars) ratio of Epo-R/channel (C) or PLCγ/channel (D), normalized to FLAG-TRPC3, for seven or three experiments, respectively, is shown. *, significant difference in the ratio compared with TRPC3 (p < 0.05).
Substitution of the TRP Domain Significantly Decreased TRPC3 Channel Interaction with PLCγ
Previously, PLCγ was also found to interact strongly with TRPC3 but much less with TRPC6 (21). The reduced interaction was hypothesized to play a role in the failure of TRPC6 to respond to Epo. As shown in Figs. 5 and 6, V5-TRPC3-C6TRP and FLAG-TRPC3-C6TRP, which respond poorly to Epo, showed statistically less association with PLCγ than V5-TRPC3 (Fig. 5B) or FLAG-TRPC3 (Fig. 6B), suggesting a role for the TRP domain in PLCγ interaction with TRPC3.
TRPC3C2 Domain and AMPK Binding Site Are Involved in Epo-induced TRPC3 Channel Translocation
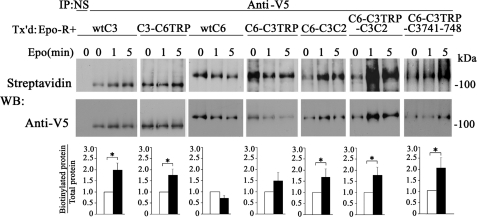
Using biotinylation, we first confirmed that channel chimeras (TRPC3, TRPC3-C6TRP, TRPC6, TRPC6-C3TRP, TRPC6-C3C2, TRPC6-C3TRP-C3C2, and TRPC6-C3TRP-C3 741–748) were present at the cell surface of transfected HEK 293T cells (Fig. 7, Epo time 0). We next examined changes in cell surface expression after stimulation with Epo. Cell surface localization of TRPC3, TRPC3-C6TRP, TRPC6-C3C2, TRPC6-C3TRP-C3C2, and TRPC6-C3TRP-C3 741–748 increased significantly after Epo stimulation, whereas that of TRPC6 and TRPC6-C3TRP did not (Fig. 7). The enhanced membrane expression of TRPC3-C6TRP in response to Epo suggests that the C3TRP domain is not critical in Epo-stimulated cell surface mobilization. In contrast, membrane insertion of chimeric TRPC6-C3C2 and TRPC6-C3 741–748 channels increased in response to Epo, suggesting that the distal C-terminal C3C2 domain and, specifically, amino acids spanning the C1/C2 AMPK site have a role in Epo-stimulated cell surface localization.
FIGURE 7.
Plasma membrane insertion of TRPC3/TRPC6 chimeras detected with cell surface biotinylation. HEK 293T cells transfected (Tx'd) with Epo-R and V5-TRPC3, V5-TRPC3-C6TRP, V5-TRPC6, V5-TRPC6-C3TRP, V5-TRPC6-C3C2, V5-TRPC6-C3TRP-C3C2, or V5-TRPC6-C3TRP-C3 741–748 were stimulated with 0–40 units/ml Epo for 0–5 min. Biotinylation of cell surface proteins was performed, and V5-tagged proteins were immunoprecipitated (IP) from lysates with anti-V5 antibody. Western blots (WB) of immunoprecipitates were probed with streptavidin-HRP to detect biotinylated protein and then stripped and reprobed with anti-V5-HRP to detect total protein. Representative results of Western blots from three experiments are shown. Biotinylated and total protein bands were quantitated with densitometry, and the ratio was normalized to time 0. The mean ± S.E. (error bars) values of the biotinylated/total protein ratios from three experiments after 5 min of stimulation are shown. *, significant difference in the ratio compared with time 0 (p < 0.05).
TRPC3C2 Domain and AMPK Binding Site Are Critical in TRPC3 Channel Association with the Cytoskeleton
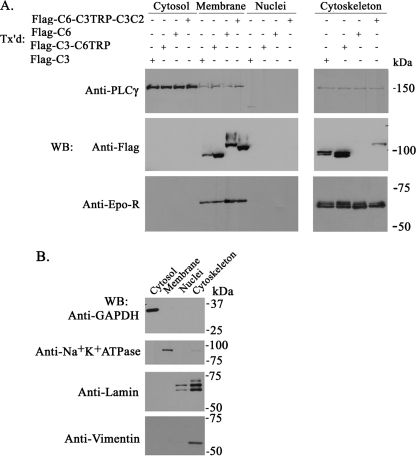
Epo-induced translocation of TRPC3 channels may involve the cytoskeletal network. To test this hypothesis, we performed subcellular fractionation to determine subcellular localization of TRPC3, TRPC6, and chimeric channels. As expected, TRPC3, TRPC6, and channel chimeras were present in the membrane but not in the cytosolic or nuclear compartments (Fig. 8A). TRPC3 and TRPC3-C6TRP but not TRPC6 were also abundant in the cytoskeleton fraction. Association of TRPC6-C3TRP-C3C2 with the cytoskeleton was significantly greater than that of TRPC6. This finding suggested that domains in the TRPC3 C3C2 terminus are important in regulating cytoskeletal localization. Endogenous PLCγ was found primarily in the cytosol, with minor membrane expression. Epo-R was found in the cell membrane but also showed significant cytoskeletal association. The quality of subcellular fractionation was confirmed by probing each fraction with antibodies specific for proteins in that compartment, including anti-GAPDH (cytosol), anti-Na+K+-ATPase (membrane), anti-lamin (nucleus, cytoskeleton), and anti-vimentin (cytoskeleton) antibodies (Fig. 8B).
FIGURE 8.
Subcellular localization of TRPC3, TRPC6, TRPC3/TRPC6 chimeras, PLCγ, and Epo-R. A, proteins from HEK 293T cells transfected with FLAG-TRPC3, FLAG-TRPC3-C6TRP, FLAG-TRPC6, FLAG-TRPC6-C3TRP-C3C2, and Epo-R were fractionated, purified, and analyzed by Western blotting (WB). Transfected FLAG-tagged constructs were detected by probing with anti-FLAG antibody. Transfected Epo-R was detected with anti-Epo-R antibody, and endogenous PLCγ was detected with anti-PLCγ antibody. Results from four fractionation experiments using FLAG-tagged constructs and two experiments using V5-tagged constructs were similar, and representative results with FLAG constructs are shown. B, representative results showing quality of fractionation by probing Western blots with anti-GAPDH, anti-Na+K+-ATPase, anti-lamin, and anti-vimentin (markers for cytosol, membrane, nuclear, and cytoskeletal fractions, respectively).
Chimeric channels were then used to study the roles of TRP and C3C2 domains in cytoskeletal association. Associations of FLAG-TRPC3 and FLAG-TRPC3-C6TRP with the cytoskeletal fraction were similar (Fig. 9A, n = 9). Associations of FLAG-TRPC6 and FLAG-TRPC6-C3TRP were also not statistically different (Fig. 9A, n = 7), demonstrating that the TRPC3 TRP domain does not play a key role. FLAG-TRPC6-C3C2 and FLAG-TRPC6-C3TRP-C3C2 had similar cytoskeletal association (n = 3), which was significantly greater than that of FLAG-TRPC6 (p < 0.01, n = 3) or FLAG-TRPC6-C3TRP (p < 0.02, n = 3; p < 0.05, n = 7 respectively). This demonstrates a role for the C3C2 domain, although FLAG-TRPC3 had significantly greater cytoskeletal association than either FLAG-TRPC6-C3C2 (p < 0.004, n = 3) or FLAG-TRPC6-C3TRP-C3C2 (p < 0.0001, n = 8). Cytoskeletal association of FLAG-TRPC6-C3TRP C3 741–748 was similar to that of FLAG-TRPC6-C3C2 and FLAG-TRPC6-C3TRP-C3C2. To confirm these results, lysates from cells transfected with different TRPC3/TRPC6 chimeras were also fractionated into different subcellular compartments with Triton X-100. Three experiments confirmed results observed with the Qiagen cell fractionation kit (Fig. 9B). This suggests that the AMPK binding motif including the Ser-747 phosphorylation site may be a key domain in facilitating C3C2 cytoskeletal association.
FIGURE 9.
Membrane and cytoskeletal association of TRPC3, TRPC6, and TRPC3/TRPC6 chimeras. Proteins from transfected HEK 293T cells (A–C) or UT-7/Epo cells (C) were fractionated, purified, and analyzed by Western blotting (WB). Transfected FLAG-tagged constructs were detected by probing with anti-FLAG antibody. Quality of fractionation was confirmed by probing Western blots with anti-Na+K+-ATPase (membrane), anti-vimentin (cytoskeletal fraction), and anti-GAPDH (cytosol marker) antibodies. Equivalent input from lysates was confirmed by probing with anti-actin. A, fractionation with the Qiagen cell fractionation kit. Membrane and cytoskeletal association of TRPC3/6 channels and chimeras. Three to nine experiments (depending on the FLAG-tagged construct) were performed, and representative results are shown. B, fractionation using the 0.5% Triton X-100 extraction method. Representative results of three experiments are shown. C, subcellular fractionation of endogenous TRPC3 and TRPC6 in UT-7/Epo cells with the Qiagen fractionation kit. For all preparations, 100 μg/lane was loaded except for the cytoskeletal fraction of transfected cells, where 50 μg was loaded per lane. Endogenous TRPC3 was detected using anti-TRPC3-C antibody. Endogenous TRPC6 was detected using Alomone anti-TRPC6 antibody. Transfected HEK 293T cells were used as controls. Representative results of four experiments are shown. lys, whole cell lysate; M, membrane fraction; Csk, cytoskeletal fraction. *, bands that did not disappear with peptide blocking, indicating that they are nonspecific.
To confirm that endogenous TRPC3 also associated with the cytoskeleton, subcellular association of endogenous TRPC3 and TRPC6 was examined in Epo-responsive UT-7 cells (Fig. 9C). TRPC3 was found in the membrane fraction but was strongly expressed in the cytoskeleton. By contrast, TRPC6 was detected primarily in the membrane fraction. Increased cytoskeletal association of endogenous TRPC3 compared with TRPC6 in Epo-responsive cells is consistent with our hypothesis that the cytoskeletal network is intimately involved in Epo-induced translocation of TRPC channels to the cell surface.
DISCUSSION
TRPC3 and TRPC6 are highly homologous TRPC channels, which are activated by some of the same growth factors (47, 48) and share a number of functions (49–52). However, TRPC3 is activated by Epo, whereas TRPC6 is not (21). Here, the domains that differ between these two channels were examined to identify sites involved in specificity of channel activation and cell surface localization. These studies identified domains, including the TRP domain, and sequences in the distal C terminus, including an AMPK binding site, that are critical for the selective activation of TRPC3 by Epo and involved in channel subcellular localization.
Previously, the proximal part of the TRPC3 carboxyl domain (C3C1) was found to be critical for activation by Epo (21). Here, C3C1 was confirmed to be important for TRPC3 gating by Epo. Substitution of C3C1 with C6C1 sequence eliminated the Epo-stimulated rise in [Ca2+]i. However, C3C1 was not sufficient, because little increase in [Ca2+]i was observed after Epo stimulation of the TRPC6-C3C1 chimera. Domains in C3C1 that are required for the response to Epo were examined. The first important finding of this work is that the TRP domain is critical for activation of the TRPC3 channel by Epo. Substitution of five amino acids in the TRP domain of TRPC3 with those of TRPC6 significantly reduced the Epo-stimulated [Ca2+]i increase, whereas reciprocal substitution of leucine zipper motifs had minimal effect on the Epo-stimulated rise in [Ca2+]i through TRPC3 or TRPC6. Substitution of the TRP domain of TRPC6 with that of TRPC3 did not result in an increase in [Ca2+]i after Epo stimulation, demonstrating that the C3TRP domain is essential but not sufficient for activation by Epo. The important role of the TRP domain in activation of other TRP channels has previously been demonstrated (58–60). The TRP domain plays an important role in TRPM8 channel activation by affecting sensitivity to agonists menthol and icilin (60). Mutations in the TRP domain were hypothesized to mediate their effect through either disruption of ligand binding to the channel or effects on the ability to translate ligand binding into conformational changes that cause channel opening. TRP domains have been found to interact with modulators of channel function. For example, the TRPC1 TRP domain binds Homer, an adapter protein that regulates its activation (61). Sequences in the TRP domains of TRPM8, TRPM5, and TRPV5 are sites of phosphatidylinositol 4,5-bisphosphate interaction involved in sensitivity to and regulation by phosphatidylinositol 4,5-bisphosphate (62). Sequences in the TRPV1 TRP domain are involved in voltage-, capsaicin-, and heat-dependent channel activation (63). Our data show that although the TRPC3 TRP domain is not critical for protein-protein interaction between TRPC3 and Epo-R, it is essential for mediating Epo-induced Ca2+ influx through TRPC3. Furthermore, immunoprecipitation studies suggested that the TRP domain may be involved in PLCγ binding, but sites of TRPC3 binding to PLCγ have been controversial, and this was not pursued here (17, 64, 65).
The second important finding of this work is the identification of domains in the distal TRPC3 C terminus that are involved in TRPC3 activation by Epo. The TRPC6-C3TRP-C3C2 chimera responds to Epo with an increase in [Ca2+]i, which approached that of wild type TRPC3. The critical roles of both the TRPC3 TRP domains and the C terminus in TRPC3 activation by Epo are consistent with the central localization of these two regions in a recently proposed three-dimensional model of TRPC3 (66). One site of potential importance located in the TRPC3 distal C terminus is the conserved CRIB (calmodulin/IP3R binding) domain (17). Affinities of the TRPC3 and TRPC6 CRIB domains differ 3-fold and could have an impact on channel function (67). Another site of potential importance is the AMPK binding site at 740–754 and the phosphorylation site at TRPC3 Ser-747. Disruption of either the putative AMPK binding site at TRPC3 741–746 (TRPC3-C6 802–807) or 741–748 (TRPC3-C6 802–809), substitution of serine 747 (TRPC3 C6C2), or both (TRPC3-C6 802–809) significantly reduced TRPC3 activity. Replacement of this site in TRPC6 with TRPC3 sequence, in conjunction with substituting the C3 TRP domain (TRPC6-C3TRP-C3 741–748), conferred Epo responsiveness to the chimeric TRPC6 channel (Fig. 4). Recently, AMPK was implicated in regulation of oxidative stress and life span in erythrocytes (68). These data suggest that AMPK and TRPC3 could be part of the same signaling pathway involved in erythrocyte survival.
The mechanisms by which Epo induced [Ca2+]i increase partly involve increased membrane insertion of Epo responsive TRPC3 channels, as indicated by surface biotinylation experiments before and 5 min after the addition of Epo (Fig. 7). Of particular relevance is that chimeric TRPC6 channels (TRPC6-C3TRP-C3C2 and TRPC6-C3TRP-C3 741–748) that exhibited Epo responsiveness in terms of [Ca2+]i increase also increased membrane insertion. This observation is important in two respects. The first is that increased Ca2+ influx after Epo stimulation is at least partly mediated by increased density of TRPC3 channels on the cell membrane. The second is that the same channel domains (C3C2 and AMPK binding site) are responsible for Epo-induced [Ca2+]i rise, channel translocation, and cytoskeletal association. Selected TRP channels have previously been shown to associate with the cytoskeleton, either directly or via scaffold proteins (69, 70), and this association has been reported to regulate channel accumulation in the plasma membrane (69, 71). For example, direct interaction between TRPC4 and the spectrin cytoskeleton is required for EGF-induced TRPC4 membrane insertion and involved in TRPC4 interaction (71). TRPC3 has been shown to assemble in a multimolecular complex in caveolae, and localization of the complex and subsequent movement to the plasma membrane are influenced by the status of the cytoskeleton (72).
In summary, domains of TRPC3 that are different from those present in TRPC6 are functionally important in its response to Epo. These include the TRPC3 TRP domain, which is required for channel gating in response to Epo and involved in PLCγ interaction, and an AMPK binding and phosphorylation site in TRPC3C2, which together with TRPC3 C3TRP is sufficient for TRPC6 channel activation by Epo. Both TRPC3C2 and the AMPK site are involved in increased channel insertion into the cell surface in response to Epo and association of the channel with cytoskeletal elements. Finally, substitution of TRPC6 sequence with as few as five amino acids in the TRP domain (DDKPS) and eight amino acids in the C terminus (GMGNSKSR) of TRPC3 can result in a functional channel.
Supplementary Material
Acknowledgment
We appreciate the assistance of Tina Brissette in preparation of the manuscript.
This work was supported, in whole or in part, by National Institutes of Health Grants R01 DK46778 (to B. A. M.) and R01 HL58672 and R01 HL74854 (to J. Y. C.). This work was also supported by the Four Diamonds Fund of the Pennsylvania State University College of Medicine.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. 1 and 2.
- Epo
- erythropoietin
- Epo-R
- erythropoietin receptor
- TRP
- transient receptor potential
- TRPC
- canonical transient receptor potential
- PLC
- phospholipase C.
REFERENCES
- 1. Lin C. S., Lim S. K., D'Agati V., Costantini F. (1996) Genes Dev. 10, 154–164 [DOI] [PubMed] [Google Scholar]
- 2. Kieran M. W., Perkins A. C., Orkin S. H., Zon L. I. (1996) Proc. Natl. Acad. Sci. U.S.A. 93, 9126–9131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Orkin S. H., Zon L. I. (2008) Cell 132, 631–644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Mulloy J. C., Cancelas J. A., Filippi M. D., Kalfa T. A., Guo F., Zheng Y. (2010) Blood 115, 936–947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Graf T., Enver T. (2009) Nature 462, 587–594 [DOI] [PubMed] [Google Scholar]
- 6. Sims B., Clarke M., Njah W., Hopkins E. S., Sontheimer H. (2010) Brain Res. 1321, 88–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Richmond T. D., Chohan M., Barber D. L. (2005) Trends Cell Biol. 15, 146–155 [DOI] [PubMed] [Google Scholar]
- 8. Miller B. A., Barber D. L., Bell L. L., Beattie B. K., Zhang M. Y., Neel B. G., Yoakim M., Rothblum L. I., Cheung J. Y. (1999) J. Biol. Chem. 274, 20465–20472 [DOI] [PubMed] [Google Scholar]
- 9. Miller B. A., Scaduto R. C., Jr., Tillotson D. L., Botti J. J., Cheung J. Y. (1988) J. Clin. Invest. 82, 309–315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Miller B. A., Cheung J. Y., Tillotson D. L., Hope S. M., Scaduto R. C., Jr. (1989) Blood 73, 1188–1194 [PubMed] [Google Scholar]
- 11. Misiti J., Spivak J. L. (1979) J. Clin. Invest. 64, 1573–1579 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Gillo B., Ma Y. S., Marks A. R. (1993) Blood 81, 783–792 [PubMed] [Google Scholar]
- 13. Hensold J. O., Dubyak G., Housman D. E. (1991) Blood 77, 1362–1370 [PubMed] [Google Scholar]
- 14. Levenson R., Housman D., Cantley L. (1980) Proc. Natl. Acad. Sci. U.S.A. 77, 5948–5952 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Cheung J. Y., Elensky M. B., Brauneis U., Scaduto R. C., Jr., Bell L. L., Tillotson D. L., Miller B. A. (1992) J. Clin. Invest. 90, 1850–1856 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Cheung J. Y., Zhang X. Q., Bokvist K., Tillotson D. L., Miller B. A. (1997) Blood 89, 92–100 [PubMed] [Google Scholar]
- 17. Tong Q., Hirschler-Laszkiewicz I., Zhang W., Conrad K., Neagley D. W., Barber D. L., Cheung J. Y., Miller B. A. (2008) J. Biol. Chem. 283, 10385–10395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Chu X., Cheung J. Y., Barber D. L., Birnbaumer L., Rothblum L. I., Conrad K., Abrasonis V., Chan Y. M., Stahl R., Carey D. J., Miller B. A. (2002) J. Biol. Chem. 277, 34375–34382 [DOI] [PubMed] [Google Scholar]
- 19. Chu X., Tong Q., Cheung J. Y., Wozney J., Conrad K., Mazack V., Zhang W., Stahl R., Barber D. L., Miller B. A. (2004) J. Biol. Chem. 279, 10514–10522 [DOI] [PubMed] [Google Scholar]
- 20. Tong Q., Chu X., Cheung J. Y., Conrad K., Stahl R., Barber D. L., Mignery G., Miller B. A. (2004) Am. J. Physiol. Cell Physiol. 287, C1667–C1678 [DOI] [PubMed] [Google Scholar]
- 21. Hirschler-Laszkiewicz I., Tong Q., Conrad K., Zhang W., Flint W. W., Barber A. J., Barber D. L., Cheung J. Y., Miller B. A. (2009) J. Biol. Chem. 284, 4567–4581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Clapham D. E. (2003) Nature 426, 517–524 [DOI] [PubMed] [Google Scholar]
- 23. Sharif-Naeini R., Folgering J. H., Bichet D., Duprat F., Delmas P., Patel A., Honoré E. (2010) J. Mol. Cell Cardiol. 48, 83–89 [DOI] [PubMed] [Google Scholar]
- 24. Ribelayga C. (2010) Curr. Biol. 20, R278–280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Miller B. A. (2006) J. Membr. Biol. 209, 31–41 [DOI] [PubMed] [Google Scholar]
- 26. Dong X. P., Wang X., Xu H. (2010) J. Neurochem. 113, 313–328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Di A., Malik A. B. (2010) Curr. Opin. Pharmacol. 10, 127–132 [DOI] [PubMed] [Google Scholar]
- 28. Birnbaumer L. (2009) Annu. Rev. Pharmacol. Toxicol. 49, 395–426 [DOI] [PubMed] [Google Scholar]
- 29. Montell C. (2005) Sci. STKE 2005, re3. [DOI] [PubMed] [Google Scholar]
- 30. Venkatachalam K., Montell C. (2007) Annu. Rev. Biochem. 76, 387–417 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Zhang W., Chu X., Tong Q., Cheung J. Y., Conrad K., Masker K., Miller B. A. (2003) J. Biol. Chem. 278, 16222–16229 [DOI] [PubMed] [Google Scholar]
- 32. Hofmann T., Schaefer M., Schultz G., Gudermann T. (2002) Proc. Natl. Acad. Sci. U.S.A. 99, 7461–7466 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Goel M., Sinkins W. G., Schilling W. P. (2002) J. Biol. Chem. 277, 48303–48310 [DOI] [PubMed] [Google Scholar]
- 34. Chu X., Tong Q., Wozney J., Zhang W., Cheung J. Y., Conrad K., Mazack V., Stahl R., Barber D. L., Miller B. A. (2005) Cell Calcium 37, 173–182 [DOI] [PubMed] [Google Scholar]
- 35. Nilius B., Owsianik G. (2010) Pflugers Arch. 460, 437–450 [DOI] [PubMed] [Google Scholar]
- 36. Goel M., Zuo C. D., Schilling W. P. (2010) Am. J. Physiol. Renal. Physiol. 298, F988–F996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Sun Y. H., Li Y. Q., Feng S. L., Li B. X., Pan Z. W., Xu C. Q., Li T. T., Yang B. F. (2010) Biochem. Biophys. Res. Commun. 394, 955–961 [DOI] [PubMed] [Google Scholar]
- 38. Tu P., Gibon J., Bouron A. (2010) J. Neurochem. 112, 204–213 [DOI] [PubMed] [Google Scholar]
- 39. Wu X., Eder P., Chang B., Molkentin J. D. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 7000–7005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Glitsch M. D. (2010) FASEB J. 24, 318–325 [DOI] [PubMed] [Google Scholar]
- 41. Numaga T., Nishida M., Kiyonaka S., Kato K., Katano M., Mori E., Kurosaki T., Inoue R., Hikida M., Putney J. W., Jr., Mori Y. (2010) J. Cell Sci. 123, 927–938 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Trebak M. (2010) Pflugers Arch. 459, 369–375 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Woodard G. E., López J. J., Jardín I., Salido G. M., Rosado J. A. (2010) J. Biol. Chem. 285, 8045–8053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Smyth J. T., Lemonnier L., Vazquez G., Bird G. S., Putney J. W., Jr. (2006) J. Biol. Chem. 281, 11712–11720 [DOI] [PubMed] [Google Scholar]
- 45. Guinamard R., Bois P. (2007) Biochim. Biophys. Acta 1772, 885–894 [DOI] [PubMed] [Google Scholar]
- 46. Bush E. W., Hood D. B., Papst P. J., Chapo J. A., Minobe W., Bristow M. R., Olson E. N., McKinsey T. A. (2006) J. Biol. Chem. 281, 33487–33496 [DOI] [PubMed] [Google Scholar]
- 47. Poteser M., Graziani A., Eder P., Yates A., Mächler H., Romanin C., Groschner K. (2008) FEBS Lett. 582, 2696–2702 [DOI] [PubMed] [Google Scholar]
- 48. Ge R., Tai Y., Sun Y., Zhou K., Yang S., Cheng T., Zou Q., Shen F., Wang Y. (2009) Cancer Lett. 283, 43–51 [DOI] [PubMed] [Google Scholar]
- 49. Onohara N., Nishida M., Inoue R., Kobayashi H., Sumimoto H., Sato Y., Mori Y., Nagao T., Kurose H. (2006) EMBO J. 25, 5305–5316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Chigurupati S., Venkataraman R., Barrera D., Naganathan A., Madan M., Paul L., Pattisapu J. V., Kyriazis G. A., Sugaya K., Bushnev S., Lathia J. D., Rich J. N., Chan S. L. (2010) Cancer Res. 70, 418–427 [DOI] [PubMed] [Google Scholar]
- 51. Yang S. L., Cao Q., Zhou K. C., Feng Y. J., Wang Y. Z. (2009) Oncogene 28, 1320–1328 [DOI] [PubMed] [Google Scholar]
- 52. Bomben V. C., Sontheimer H. W. (2008) Cell Prolif. 41, 98–121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. El Boustany C., Bidaux G., Enfissi A., Delcourt P., Prevarskaya N., Capiod T. (2008) Hepatology 47, 2068–2077 [DOI] [PubMed] [Google Scholar]
- 54. Kurebayashi N., Harkins A. B., Baylor S. M. (1993) Biophys. J. 64, 1934–1960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Wu Y., Clusin W. T. (1997) Am. J. Physiol. 273, H2161–H2169 [DOI] [PubMed] [Google Scholar]
- 56. Goel M., Sinkins W. G., Zuo C. D., Estacion M., Schilling W. P. (2006) Am. J. Physiol. Renal Physiol. 290, F1241–F1252 [DOI] [PubMed] [Google Scholar]
- 57. Wills J., Credle J., Haggerty T., Lee J. H., Oaks A. W., Sidhu A. (2011) PLoS One 6, e17953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Varnum M. D., Black K. D., Zagotta W. N. (1995) Neuron 15, 619–625 [DOI] [PubMed] [Google Scholar]
- 59. Tibbs G. R., Goulding E. H., Siegelbaum S. A. (1997) Nature 386, 612–615 [DOI] [PubMed] [Google Scholar]
- 60. Bandell M., Dubin A. E., Petrus M. J., Orth A., Mathur J., Hwang S. W., Patapoutian A. (2006) Nat. Neurosci. 9, 493–500 [DOI] [PubMed] [Google Scholar]
- 61. Yuan J. P., Kiselyov K., Shin D. M., Chen J., Shcheynikov N., Kang S. H., Dehoff M. H., Schwarz M. K., Seeburg P. H., Muallem S., Worley P. F. (2003) Cell 114, 777–789 [DOI] [PubMed] [Google Scholar]
- 62. Rohács T., Lopes C. M., Michailidis I., Logothetis D. E. (2005) Nat. Neurosci. 8, 626–634 [DOI] [PubMed] [Google Scholar]
- 63. Valente P., García-Sanz N., Gomis A., Fernández-Carvajal A., Fernández-Ballester G., Viana F., Belmonte C., Ferrer-Montiel A. (2008) FASEB J. 22, 3298–3309 [DOI] [PubMed] [Google Scholar]
- 64. van Rossum D. B., Patterson R. L., Sharma S., Barrow R. K., Kornberg M., Gill D. L., Snyder S. H. (2005) Nature 434, 99–104 [DOI] [PubMed] [Google Scholar]
- 65. Wen W., Yan J., Zhang M. (2006) J. Biol. Chem. 281, 12060–12068 [DOI] [PubMed] [Google Scholar]
- 66. Ko K. D., Bhardwaj G., Hong Y., Chang G. S., Kiselyov K., van Rossum D. B., Patterson R. L. (2009) Commun. Integr. Biol. 2, 133–137 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Tang J., Lin Y., Zhang Z., Tikunova S., Birnbaumer L., Zhu M. X. (2001) J. Biol. Chem. 276, 21303–21310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Wang S., Dale G. L., Song P., Viollet B., Zou M. H. (2010) J. Biol. Chem. 285, 19976–19985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Clark K., Middelbeek J., van Leeuwen F. N. (2008) Eur. J. Cell Biol. 87, 631–640 [DOI] [PubMed] [Google Scholar]
- 70. Goswami C., Hucho T. (2008) FEBS J. 275, 4684–4699 [DOI] [PubMed] [Google Scholar]
- 71. Odell A. F., Van Helden D. F., Scott J. L. (2008) J. Biol. Chem. 283, 4395–4407 [DOI] [PubMed] [Google Scholar]
- 72. Lockwich T., Singh B. B., Liu X., Ambudkar I. S. (2001) J. Biol. Chem. 276, 42401–42408 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.