Abstract
Through forward genetic screening in the mouse, a recessive mutation (couch potato, cpto) has been discovered that dramatically reduces plasma cholesterol levels across all lipoprotein classes. The cpto mutation altered a highly conserved residue in the Src homology domain 3 (SH3) domain of the Mia2 protein. Full-length hepatic Mia2 structurally and functionally resembled the related Mia3 protein. Mia2 localized to endoplasmic reticulum (ER) exit sites, suggesting a role in guiding proteins from the ER to the Golgi. Similarly to the Mia3 protein, Mia2’s cytosolic C terminus interacted directly with COPII proteins Sec23 and Sec24, whereas its lumenal SH3 domain may facilitate interactions with secretory cargo. Fractionation of plasma revealed that Mia2cpto/cpto mice had lower circulating VLDL, LDL, HDL, and triglycerides. Mia2 is thus a novel, hepatic, ER-to-Golgi trafficking protein that regulates cholesterol metabolism.
Keywords: endoplasmic reticulum, lipoprotein, secretion, Mia3, Tango1, Ctage5
In an effort to reveal novel pathways of cholesterol homeostasis in vivo, N-ethyl-N-nitrosourea- (ENU) mutagenized, C57BL/6 (B6) mice were screened, and affected families showing altered levels of plasma cholesterol were selected. One family derived from this screen, couch potato (cpto) mutant mice, had dramatically low plasma levels of both total cholesterol and HDL-cholesterol (HDL-C). The cpto phenotype was associated with a Phe-to-Ser substitution at a highly conserved position within the Src homology domain 3 (SH3)-encoding domain of the Mia2 gene.
MIA2 was originally described as a human gene that encodes a protein with an N-terminal SH3 domain homologous to those of two other genes, melanoma inhibitory activity (MIA) and otoraplin (OTOR), both of which encode secreted, largely SH3-only proteins (1). MIA2 also shows homology to a fourth MIA family member, MIA3/TANGO1 (2–4), which contains additional functional domains beyond the SH3 motif, and which may contribute to cardiovascular disease risk in humans (5, 6). MIA2’s expression is liver specific in humans (1, 7), and changes in hepatocyte MIA2 expression correlate with multiple hepatic disease states (7, 8).
An additional Mia family member was found in a genome-wide knockdown screen in Drosophila tissue culture. Drosophila Tango1 and one of its mammalian orthologs, Mia3/Tango1, both encode proteins that retain similarity to other Mia family members in their N-terminal SH3 motifs, but are not secreted. Instead, these genes produce longer, transmembrane proteins that regulate the secretory process (4, 9). Mia2 sequences were also noted to extend beyond the SH3 domain, but the reported cDNAs were truncated prior to the conserved coiled-coil and proline-rich domains of Tango1 or Mia3 (1). We show that mouse hepatic Mia2 protein has extensive structural and functional homology to MIA3 and Drosophila Tango1, through downstream exons previously characterized as a separate gene, Ctage5/Mgea6 (10).
We also demonstrate that, like Mia3/Tango1 (4) the Mia2 protein localizes to endoplasmic reticulum (ER) exit sites. Furthermore, the apparent topological organization of Mia2 appears to place the cpto mutation-bearing SH3 domain within the ER lumen. Although Mia2 may act as a regulator of sorting in the secretory pathway, the phenotypic effects of its mutation are seen as a systemic reduction in plasma levels of cholesterol and triglycerides in VLDL, LDL, and HDL in Mia2 mutant mice.
MATERIALS AND METHODS
Animals
All mice were housed, fed, bred, and maintained under standard conditions. All procedures were conducted in conformity with the Public Health Service Policy on the Humane Care and Use of Laboratory Animals and were approved by the Scripps Research Institute's Institutional Animal Care and Use Committee. The mouse N-ethyl-N-nitrosourea screen was performed as described (11, 12), using C57BL/6J animals for the initial mutagenesis. For mapping studies, progeny of ENU-mutagenized founders were bred to the 129S1/SvIMJ background.
Histology
Animals were euthanized for tissues as adults. Livers were fixed in formalin at 4°C, then processed for paraffin embedding by the Scripps Histology Core Facility, and sectioned, mounted, and stained with hematoxylin and eosin.
Mapping and identifying the gene responsible for the cpto phenotype
The affected (cpto) family was identified in the G3 generation, and backcrossed to B6 to demonstrate heritability. Affected (cpto) mice in the B6 background were bred to 129S1/SvIMJ mice, and the resulting (heterozygous) progeny were intercrossed to generate the F2 generation. Genomes of affected and unaffected F2 mice were screened for correlation of the low-cholesterol phenotype with inheritance of B6 strain-specific , single-nucleotide polymorphisms (SNPs), as described (13). Further recombination refined the interval to chromosome 12, at 58.2–60.3 Mbases. Exons from all genes within the interval were amplified and sequenced until the mutation in Mia2 was identified.
cDNA construction
Mouse Mia2 was amplified in three pieces (a 5′ fragment from bp 1–984, a 2,334 bp segment spanning the internal EcoRI and SbfI sites, and a 1,275 bp fragment from the SbfI site to the 3′ end of the open reading frame) that were ligated together using overlapping EcoRI and SbfI sites. For wild-type (WT) and cpto mutant versions of the first two fragments, RNA was prepared from WT and Mia2cpto livers using Trizol (Sigma), and cDNA was prepared using SuperScript III reverse transcriptase (Invitrogen) before amplification by PCR. For the third fragment (SbfI site to 3′ end) of mouse Mia2, a mouse Ctage5 cDNA clone (ID BC064355) was obtained from the Origene full-length cDNA collection, and used as template for PCR. Mouse Mia2x1–4 and Mia2x1–6 cDNAs were generated using the full-length Mia2 cDNA as template in PCR, using primers containing the alternative 3′ coding sequences. For human MIA2, RNA was prepared from HepG2 cells. cDNA was reverse transcribed using SuperScript III (Invitrogen), and used as template for PCR. All PCR-generated clones were verified by double-stranded sequencing.
Western blots
Polyclonal rabbit anti-human MIA2 antibodies (Antigenix America; Huntington Station, NY), polyclonal rabbit anti-human CTAGE5 antibodies (Atlas Antibodies/Sigma), polyclonal rabbit anti-human apoA-I antibodies (Calbiochem; San Diego, CA), polyclonal goat anti-human apoB antibodies (Calbiochem), and monoclonal mouse anti-GM130 and anti-Sec31A antibodies (both from BD Biosciences Pharmingen) were purchased and used according to the manufacturers’ instructions. Proteins were routinely separated by electrophoresis on 3–8% NuPAGE Tris-Acetate or 4–12% NuPAGE Bis-Tris precast gels (Invitrogen) following the manufacturer's instructions, and transferred to polyvinylidene difluoride membranes. The proteins were detected by Western blot, using the above antibodies, or mouse monoclonal anti-FLAG M2 (Sigma), followed by HRP-coupled secondary antibodies and ECL Plus Western blot detection reagent (Amersham).
Immunocytochemistry
Cells (HuH-7, NIH-3T3) were grown overnight in 12-well culture dishes on glass coverslips, rinsed in 1× PBS, then fixed in 10% formalin for 15 min at room temperature. Cells were then permeabilized in either dilution buffer (1× TBS, 0.05% Tween-20, 0.5 M NaCl, 3% BSA, 0.3% Triton X-100) or detergent-free dilution buffer plus digitonin (1× TBS, 0.5 M NaCl, 3% BSA plus 50 μg/ml digitonin) for 30 min at room temperature. Primary antibodies were diluted 1:200 in the same buffers, and incubated on the cells for 1 h at room temperature. Following two washes in 1× PBS, Alexa 488- or Alexa 546-conjugated secondary antibodies were added to the cells for 30 min at room temperature. After two washes in 1× PBS, coverslips were inverted on a drop of ProLong Gold with DAPI (Invitrogen) on a glass microscope slide for mounting, and staining was examined via fluorescent microscopy.
Yeast two-hybrid assay
The yeast reporter strain L40, which contains an integrated LacZ reporter under the control of the LexA operon (14), was transformed with plasmids expressing an LexA-Mia21106-1397 fusion protein (bearing the TRP1 selectable marker) and one of several VP16 fusion proteins (bearing the LEU2 selectable marker). Yeast, grown on solid media lacking tryptophan and leucine to maintain the plasmids, was transferred to nitrocellulose and lysed by freeze/thaw in liquid nitrogen. The yeast-containing nitrocellulose was then incubated in X-gal substrate, and the appearance of blue precipitate was recorded as described (15).
Body composition and plasma metabolite analysis
Glucose, triglyceride, total and HDL-C, and nonesterified FA levels in plasma from individual, 4 hour-fasted mice were measured on a clinical blood chemistry analyzer (AU400e; Olympus). Insulin levels in duplicate samples were determined by enzyme-linked immunosorbent assay (Crystalchem). Body composition analysis was performed using the EchoMRI-100 whole-body composition analyzer (EchoMRI).
Plasma fractionation and cholesterol and #8232triglyceride assay
Fresh mouse plasma was pooled by Mia2 genotype (four to five animals per pool), and frozen until use. Thawed plasma pools were fractionated by fast-protein liquid chromatography (FPLC), over Superdex 200 10/300 GL and Superdex 200 HR10/30 columns in series. Forty 0.5 ml fractions were collected per genotype pool. Total cholesterol in FPLC fractions was quantitated using a ThermoDMA 2350-400H kit, following the manufacturer's instructions. Triglycerides were measured with a Raichem 84098 kit, following the manufacturer's instructions. Apolipoproteins in the fractions were estimated after visualization in SDS-PAGE gels and protein scanning.
Liver lipid extraction and quantitation
Hepatic lipids were extracted following an adaptation of published methods (16). Briefly, pieces of mouse liver were weighed, then homogenized in 10 vols of 1× PBS. Homogenate (120 μl) was then extracted with 0.8 ml of a 2:1 (v:v) mixture of chloroform- methanol, with vortexing to mix thoroughly. After adding 100 μl more of 1× PBS and remixing, samples were spun at 4,200 g at 4°C for 10 min. Two hundred microliters of the organic (lower) phase was air-dried in a fresh tube then resuspended in 100 μl of 1% Triton X-100 in ethanol. This was then air dried and resuspended in 500 μl 1× PBS, for the final lipid extract. Fifty microliter aliquots of this extract were used to measure either cholesterol, using the Amplex Red cholesterol assay kit (#A12216, Life Technologies; Carlsbad, CA) or triglycerides (Triglyceride Assay Kit #10010303, Cayman Chemicals; Ann Arbor, MI).
RNA quantitation
RNA was isolated from various tissues with Trizol (Invitrogen) according to the manufacturer's instructions. Primer/probe combinations for quantitative reverse-transcriptase PCR (qRT-PCR) were purchased from Applied Biosystems (ABI), catalog number Mm00616368_m1 for mouse Mia2 and Mm00524692_m1 for mouse Ctage5. qRT-PCR was done on an ABI 7900HT sequence detection system. RNA was quantified by the comparative CT method using m36b4 as an internal standard. For each primer/probe combination (Mia2 or Ctage5), values for different tissues were normalized to values for liver.
RESULTS
The couch potato mutation alters the SH3 domain of Mia2
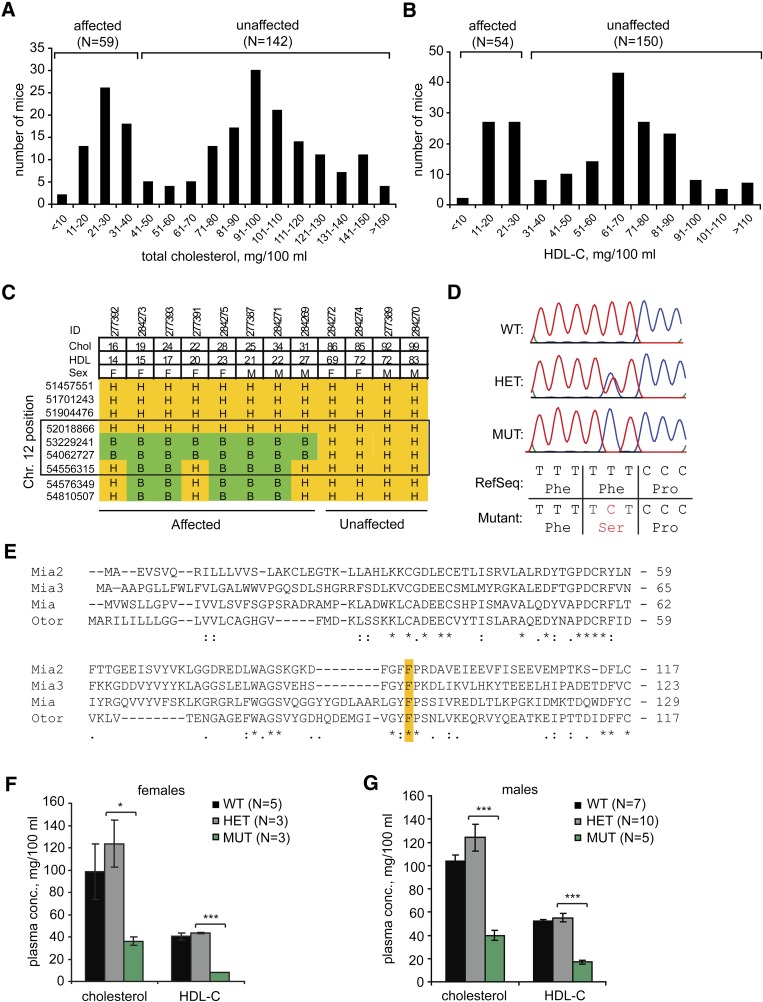
The couch potato (cpto) mutation was initially identified in a screen of ENU-mutagenized B6 mice, designed to reveal recessive mutations that confer significant changes in plasma cholesterol. One family derived from a single ENU-mutant founder displayed a distinctly bimodal distribution in cholesterol levels, with roughly one-fourth of the animals having dramatically lower cholesterol levels (“affected,” Fig. 1A), consistent with inheritance of a single recessive allele. Similar proportions of animals also had low plasma HDL-C (Fig. 1B). This phenotype was dubbed couch potato, and the family was selected for further analysis: affected mice were crossed to the 129S1/SvImJ (129) background for mapping, or backcrossed to B6 for physiological studies. The cpto phenotype was initially mapped to a 13.7 Mb interval on mouse chromosome 12 in 99 B6.129 hybrid F2 progeny, using a genome-wide array of SNPs as molecular markers. Continued backcrosses to 129 and mapping with a denser set of SNP markers revealed a tight correlation between the cpto low-cholesterol/low HDL-C phenotype and inheritance of an interval from 52.0 to 54.6 Mb (mouse genome build NCBIM33) from the B6 (mutant) background (Fig. 1C).
Fig. 1.
Mapping, identification and phenotype of the cpto mutation. Distribution of total plasma cholesterol (A), and HDL-C levels in couch potato (cpto) mice (B). Two peaks were visible in each distribution, with “affected” animals defined as those covered by the lower cholesterol or lower HDL peaks. C: In animals bred with 129S1/SvImJ, phenotypically affected (cpto) mice were homozygous for C57BL6/J-specific SNP markers between 52 and 54.6 Mb of mouse chromosome 12 (genome build NCBIM33). “H”/orange represents heterozygous (one copy B6, one copy 129) and “B”/green represents homozygous B6 at each SNP. Animals were sorted left-to-right by ascending HDL levels. D: Electropherograms and corresponding nucleotide and amino acid sequence of exon 3 of Mia2 from wild-type (WT) mice or mice heterozygous (HET) or homozygous mutant (MUT) for the Mia2F91S mutation. E: Alignment of the N-termini of all four mouse Mia2-homologous proteins. Highlighting shows that F91 of Mia2 is conserved in the other three paralogs (Mia, Otor, Mia3). F, G: The cpto low-cholesterol/low-HDL phenotype followed inheritance of the Mia2F91S mutation in female (F) and male (G) cpto mice, after five generations of backcrossing into the B6 background. Data are expressed as mean ±SEM. Student's t-test, * P < 0.05, *** P < 0.001.
Sequencing of candidate genes within this interval (see supplementary Fig. I) revealed a T-to-C transition in the third exon of the Mia2 gene, which replaces the WT phenylalanine-91 with a serine in the Mia2 protein (F91S, Fig. 1D). This substitution occurs within Mia2’s N-terminal SH3 domain. Comparison of Mia2’s protein sequence with its three other mouse paralogs (Mia, Otor, and Mia3) showed that this position is otherwise invariant (Fig. 1E). Furthermore, in both the solution structure (17) and crystal structure (18) of MIA's atypical SH3 domain, the analogous residue is located at or adjacent to the surface, in a region shown in other SH3 structures to comprise the ligand-binding interface, indicating that an F91S substitution could be functionally significant. After backcrossing for five generations into the B6 background, this Mia2F91S allele segregated precisely with both the low total cholesterol and low HDL-C cpto phenotypes in a total of 32 mice, both female and male (Fig. 1F, G). Genotypically, in over 11 litters from HET by HET crosses, mice were obtained at roughly the expected 1:2:1 Mendelian ratio (25 MUT: 32 HET: 20 WT). We therefore concluded that the Mia2F91S mutation probably played a causative role in the cpto phenotype, and will henceforth refer to the Mia2F91S mutant allele as Mia2cpto.
Plasma triglycerides, which like cholesterol are transported within lipoproteins, also were substantially lower in Mia2cpto mice (see supplementary Fig. IIA), although plasma NEFAs were not significantly different (see supplementary Fig. IIB). Other metabolic parameters showed little correlation to the Mia2cpto genotype, including glucose (see supplementary Fig. IIC) and insulin levels (see supplementary Fig. IID). Similarly, total body weight, fat mass, and lean mass were not discernibly different between HET and MUT animals, even after high-fat feeding (see supplementary Fig. IIE). Although circulating levels of cholesterol and triglycerides were lower in Mia2cpto mutant mice, there was no visible evidence of hepatic steatosis or of altered adipose tissue morphology (see supplementary Fig. IIF). No major differences in size, lifespan, or fertility between Mia2cpto mutants and their WT counterparts were obvious (data not shown), leaving lowered cholesterol and triglyceride levels as the main observed mutant phenotypes.
Mia2 shows extended homology to Mia3 and #8232Drosophila Tango1
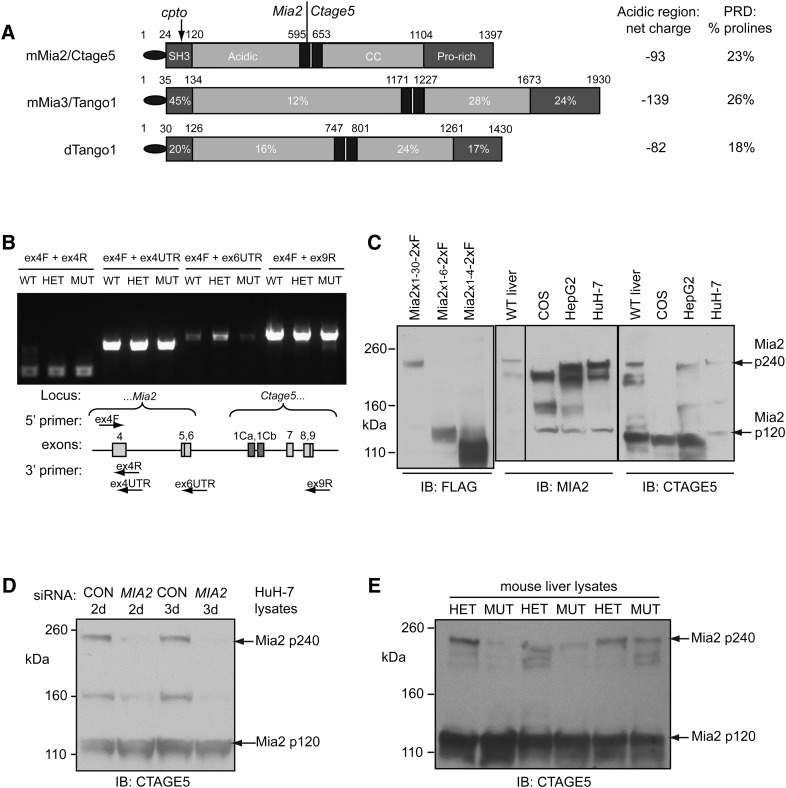
The original report of Mia2 cloned from mice described a gene with four coding exons (1). In examining the sequence of Mia2, we observed that the “gene” immediately distal to Mia2, Ctage5/Mgea6, encoded both coiled-coil and proline-rich motifs highly homologous to the carboxyl termini of both Mia3 and Drosophila Tango1 (Fig. 2A). We therefore sought Mia2/Ctage5 transcripts analogous to those of Mia3 and dTango1. RT-PCR demonstrated the existence of Mia2 transcripts that included both Mia2 and Ctage5 sequences (Fig. 2B; see supplementary Figs. IIIA, IIIB). Combination of these RT-PCR results with published data and expressed sequence tag (EST) sequences led to a model in which Mia2 and Ctage5 are a single gene with three major classes of transcripts, containing solely Mia2-specific exons, solely Ctage5-specific exons (1, 19, 20), or exons from both Mia2 and Ctage5 (see supplementary Fig. IIIC).
Fig. 2.
Structure of Mia2 transcripts and Mia2 protein isoforms. A: Full-length Mia2 protein includes Ctage5-encoded sequences, and structurally resembles Mia3 and Drosophila Tango1. Black oval represents signal sequences (cleaved in Mia2), and black bars indicate putative transmembrane motifs. CC, putative coiled coil domain; PRD, proline-rich domain. Numbers in corresponding domains of mMia3 and dTango1 reflect percent amino acid identity to Mia2 within each region. Although primary sequence conservation is low, amino acid composition in acidic and proline-rich regions is highly similar. B: RT-PCR-based detection of Mia2 splicing events terminating in Mia2 exon 4 (ex4UTR products) and exon 6 (ex6UTR products) and continuing through to Ctage5-specific exons (ex9R products) in total RNA from livers of Mia2+/+ (WT), Mia2cpto/+ (HET) and Mia2cpto/cpto (MUT) mice. Ethidium bromide-stained gel shows RT-PCR products; cartoon shows relative positions of PCR primers. C: Anti-MIA2 and anti-CTAGE5 both detected a ∼240 kDa protein that comigrated with the protein produced by full-length Mia2x1–30 expression vector. D: HuH-7 cells transfected with control (CON) or pooled MIA2-specific siRNAs were harvested 2 or 3 days posttransfection. Western blotting with anti-CTAGE5 showed reduced Mia2 p240 isoform, whereas the Mia2 p120 isoform (which should not be regulated by the MIA2-specific siRNAs) was unaffected. E: In whole-cell lysates from mouse livers, full-length (mutant SH3-containing) Mia2 p240 was present at lower levels in MUT animals relative to HET animals, whereas Mia2 p120 levels did not vary.
Microarray-based analysis of Mia2 expression in mice, using a probe set directed against the Mia2 SH3-encoding exons (viewable at http://biogps.org), detected Mia2 expression robustly in liver, with additional signals in kidney and small intestine (21). This agreed with published reports of liver-specific Mia2 expression in mouse and human tissues (1, 7, 8). In contrast, analysis using a probe set targeting the last two exons of Ctage5 showed expression in mice to be widespread. Similarly, our own qRT-PCR analysis using primer/probe combinations specific for Mia2 or Ctage5 showed that Ctage5-containing transcripts were widely expressed, whereas Mia2-containing transcripts were restricted to liver, intestine, kidney, and heart (see supplementary Fig. IIID).
Because the cpto mutation affects Mia2’s SH3 domain, this limited “Mia2” expression profile should reflect the tissues in which Mia2 function affects cholesterol levels. Notably, the major site of Mia2 expression is the liver, the organ from which nascent HDL and VLDL are secreted.
Mia2 produces 240 and 120 kDa protein isoforms
Detection of multiple alternative Mia2 transcript classes made it important to investigate Mia2 protein isoforms in vivo. The largest endogenous protein detected with both anti-MIA2 and anti-CTAGE5 was ∼240 kDa, and was present in both mouse liver and human hepatoma HuH-7 and HepG2 cell lines (Fig. 2C). This protein also comigrated with a recombinant protein (Mia2x1–30, detected with anti-FLAG) based on the longest transcript class, containing all six Mia2 exons and all but the first Ctage5 exons (Fig. 2C, Mia2 p240). We refer to this recombinant cDNA as full-length Mia2; the sequence of the protein it encodes is shown in supplementary Fig. IVA. Other mouse and human anti-MIA2-reactive protein species were evident as well, although their identity is unclear, inasmuch as the majority of the additional species appear to be much larger than the proteins containing Mia2 exons 1–4 or exons 1–6 (Fig. 2C).
Additional anti-CTAGE5-reactive proteins were detectable, particularly at or below 120 kDa, in most tissues and cell lines examined (Fig. 2C and supplementary Fig. IVB). It should be noted that in human cells, but not in mice, anti-CTAGE5 may cross-react with a family of expressed pseudogenes, including the previously described CTAGE1, CTAGE2, and CTAGE4 (20, 22), all of which should encode proteins of similar size to MIA2 p120 (see supplementary Fig. VA). These proteins are highly homologous to human MIA2/CTAGE5 in the region used as an immunogen for anti-CTAGE5 (see supplementary Fig. VB), but should not react with anti-MIA2. Full-length Mia2 p240 in mice was detectable in liver, and may also be present in lung, but was not detectable in muscle, brain, adipose tissue, or testis (see supplementary Fig. IVB). No CTAGE5-reactive or MIA2-reactive proteins of any size were detectable in mouse plasma (see supplementary Fig. IVB, and data not shown), arguing against a secreted isoform of Mia2.
Because the anti-MIA2-reactive proteins, all of which are larger than 120 kDa, should contain the SH3 domain bearing the cpto mutation, the abundant p240 MIA2 isoform was examined in further detail. A pool of siRNAs specific for MIA2-coding exons was used to knock down MIA2 expression in HuH-7 cells. An anti-CTAGE5 Western blot showed that the MIA2 siRNAs substantially diminished the abundance of the p240, full-length MIA2 isoform, while producing no detectable change in the abundance of the 120 kDa species (Fig. 2D). This confirmed that the p240 isoform was encoded by both MIA2 and CTAGE5 exons, inasmuch as it is recognized by a CTAGE5-specific antibody, but it was also sensitive to the quantities of Mia2 transcripts. It should also be noted that in Mia2cpto mice, compared with heterozygotes, homozygous mutant livers often contained lower amounts of the p240 Mia2 isoform, whereas the abundance of the p120 Mia2 isoform was not altered by the cpto mutation (Fig. 2E). Thus, the cpto mutation may alter the stability of SH3 domain-containing isoforms, in addition to effects of the primary amino acid change on protein function.
Based on this evidence, we propose that Mia2 and Ctage5 should be considered a single gene, with multiple alternative 5′ exons, and that this gene be referred to as Mia2, to acknowledge its homology to Mia and Mia3. Support for this arrangement is seen in the predicted structures of Mia2/Ctage5 genes in other species, such as chimp (XP_001147938), orangutan (NP_001128150), marmoset (XP_002753890), platypus (XP_001513694), and finch (XP_002199906), all of which include isoforms encoded by Mia2-specific exons spliced onto downstream Ctage5 exons. Like Mia2, the Mia3 gene appears to give rise to transcripts analogous to the Ctage5-only transcripts shown in supplementary Fig. IIIC (e.g., EST sequence CR990232). Because both Mia2 and Mia3 are capable of generating both SH3-containing and SH3-lacking protein isoforms, there may be a functional significance to this organization.
MIA2 is an ER exit site protein
The structural homology between Mia2 p240 and Drosophila Tango1 and vertebrate Mia3/Tango1 suggested that Mia2 may also have a similar function. Mia3/Tango1 was identified as an ER exit site protein that interacts with the COPII coat proteins Sec23 and Sec24 through its C-terminal proline-rich domain, and with select Golgi-bound cargo, such as collagen VII, through its N-terminus (4). Similarly, dTango1 is ER exit site-localized (23), and is required for secretion of some proteins in a Drosophila tissue culture system (9). If Mia2 functionally resembles these proteins, Mia2 should first have a similar subcellular location.
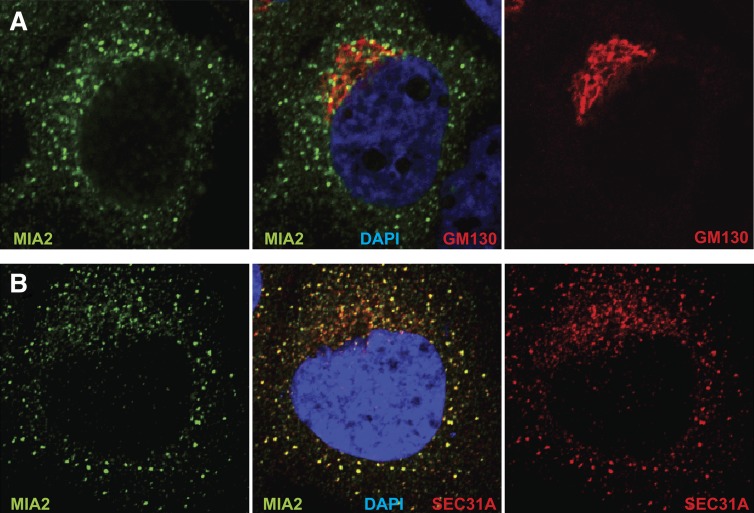
Human HuH-7 cells were used, because they express primarily a full-length MIA2 protein similar to the major species in mouse liver (Fig. 2CndashE). Immunostaining of these cells with MIA2-specific polyclonal antibodies (Fig. 3)identified two discrete subcellular locations: bright, asymmetric staining immediately adjacent to the nucleus, and a dispersed, punctate cytosolic signal. The juxtanuclear MIA2-positive signal was adjacent to but distinct from the early Golgi marker, GM130 (Fig. 3A). Anti-CTAGE5 staining of HuH-7 cells also produced perinuclear and punctate cytosolic signals, but in contrast to anti-MIA2, the anti-CTAGE5 perinuclear signal largely colocalized with GM130 (see supplementary Fig. VIA), although it was unclear whether this staining reflected specific binding of MIA2 p240, MIA2 p120, or of other human CTAGE-family pseudogene proteins (see supplementary Fig. V).
Fig. 3.
MIA2 localizes to ER exit sites in HuH-7 cells. A: Confocal imaging of HuH-7 cells costained with anti-MIA2 (green) and anti-GM130 (red) revealed that the perinuclear anti-MIA2 staining was adjacent to but not overlapping with GM130 signal. B: In this confocal image, costaining with anti-MIA2 (green) and anti-Sec31A (red) showed extensive colocalization of MIA2 with the COPII protein, Sec31. Nuclei were stained with 4’,6-diamidino-2-phenylindole (DAPI, blue).
The majority of the anti-MIA2 signal, however, was in a punctate staining pattern in the cytosol that resembles that of ER exit sites. Indeed, MIA2 staining was coincident with the signal for the COPII subunit, Sec31A (Fig. 3B). Substantially less overlapping signal was seen with anti-CTAGE5 (see supplementary Fig. VIB). MIA2’s subcellular localization thus closely resembled that of MIA3/TANGO1, which remains in the ER, and does not accompany COPII vesicles to the Golgi (4). This argues that lower plasma cholesterol levels resulting from mutations in Mia2 involve some aspect of cholesterol homeostasis that is dependent on ER-to-Golgi trafficking.
Mia2 protein topology and COPII binding resemble that of Mia3/Tango1
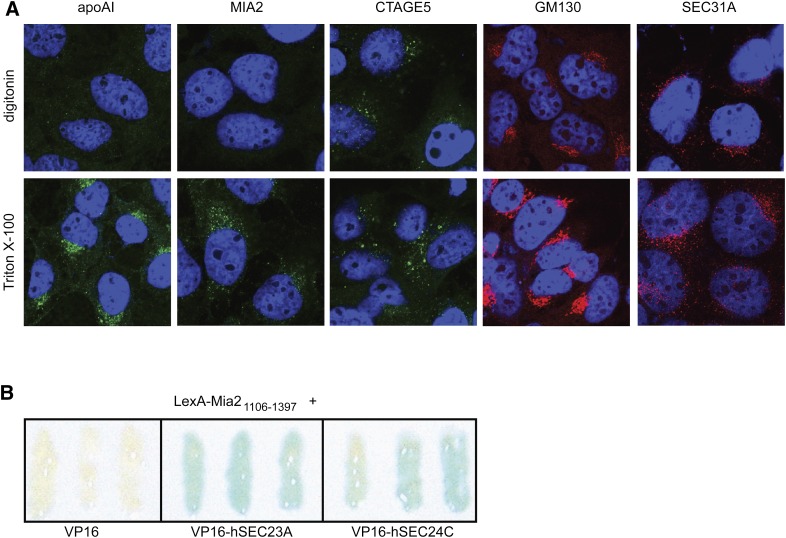
Because Mia2’s SH3 domain carries the cpto mutation, its position relative to the ER is an important component of its function. Differential permeabilization (24) of HuH-7 cells was used to identify regions of MIA2 that were exposed to the cytosol (Fig. 4A). Digitonin preferentially permeabilizes the cellular membrane, but not the internal (Golgi, ER) membranes (25). To allow complete antibody access to all internal compartments, a parallel set of cells was incubated in Triton X-100. As shown in Fig. 4A, Golgi- and/or ER-localized apoA-I proteins were detectable solely with Triton X-100 permeabilization, and not with digitonin, indicating that these proteins were entirely enclosed within organelle membranes. In contrast, Golgi protein GM130 and COPII protein Sec31A were detected under both conditions, indicating that these epitopes resided in the cytosol (Fig. 4A). Anti-MIA2 recognized epitopes including Mia2’s N-terminal SH3 domain and the adjacent acidic region (see supplementary Fig. VIIB) and produced a signal with Triton X-100 permeabilization, but not with digitonin (Fig. 4A). In contrast, anti-CTAGE5, which recognized epitopes in the C-terminal coiled-coil domain (see supplementary Fig. IVB), was positive under both permeabilization conditions (Fig. 4A). Thus the N-terminal half of MIA2, including the SH3 and acidic region, was lumenally localized, whereas the C-terminal coiled-coil and proline-rich portions of MIA2 probably resided in the cytosol. This orientation is identical to the topology recently demonstrated for MIA3/TANGO1 (4).
Fig. 4.
MIA2 topology as revealed by differential permeabilization and staining of MIA2 and CTAGE5 epitopes. A: HuH-7 cells were grown on coverslips, fixed in 4% formaldehyde/1× PBS for 10 min at room temperature, then permeabilized for 15 min at room temperature in either TBST (containing 0.1% Tween-20) plus 0.3% Triton X-100 (bottom row), or TBS (without Tween-20) plus 50 μg/ml digitonin (top row). Both anti-GM130 and anti-Sec31 functioned comparably under either permeabilization condition, indicating a cytosolically exposed epitope. In contrast, apoA-I was detectable in the ER and Golgi only with Triton X-100 permeabilization, indicating that it was lumenally enclosed. Anti-MIA2, which recognizes the N-terminal half of MIA2, matched the lumenal pattern, whereas anti-CTAGE5, which recognizes the C-terminal portion of MIA2, stained a cytosolic epitope. B: Yeast two-hybrid results (β-galactoside assay) showing interaction between mouse Mia2’s C-terminal proline-rich domain (Mia21106–1397) and the human COPII proteins SEC23A and SEC24C. Colonies containing both an LexA- Mia21106–1397 bait vector and either empty VP16 prey vector (as a negative control) or vectors encoding VP16-SEC23A or -SEC24C fusion proteins were streaked and grown together, lifted on a single filter and processed. Blue color indicates protein-protein interaction.
Because anti-CTAGE5 may also have recognized expressed pseudogenes in human cells (see supplementary Fig. V), HuH-7 cells were transfected with expression vectors producing C-terminally FLAG epitope-tagged Mia2 proteins of various lengths (see supplementary Fig. VIIA). In agreement with the anti-MIA2 and anti-CTAGE5 staining, the C terminus of full-length Mia2x1–30 (both WT and cpto mutant versions) was detectable after permeabilization with both digitonin and Triton X-100. A shorter version, Mia2x1–6, which encoded a protein with a truncated/ablated transmembrane domain, appeared to have a lumenally located carboxy terminus (see supplementary Fig. VIIA). It should be noted, however, that all overexpressed MIA2 proteins, including the shorter versions shown in supplementary Fig. VII, appear to mislocalize throughout the ER, and not to ER exit sites.
Because immunostaining indicated Mia2 localized to ER exit sites, Mia2’s ability to interact directly with components of the COPII vesicular coat, as observed for MIA3/TANGO1 (4), was also examined. In a yeast two-hybrid assay, the proline-rich C-terminal region of Mia2 (LexA-Mia21106–1397) interacted specifically with both Sec23A and with Sec24C, components of the COPII inner vesicular coat (Fig. 4B). This suggests that, like MIA3/TANGO1, Mia2 functions in assembly of COPII vesicles, and that its lumenally localized SH3 domain, altered by the cpto mutation, might interact with outgoing cargo.
couch potato mutant mice have low plasma cholesterol and triglycerides, and increased hepatic triglycerides
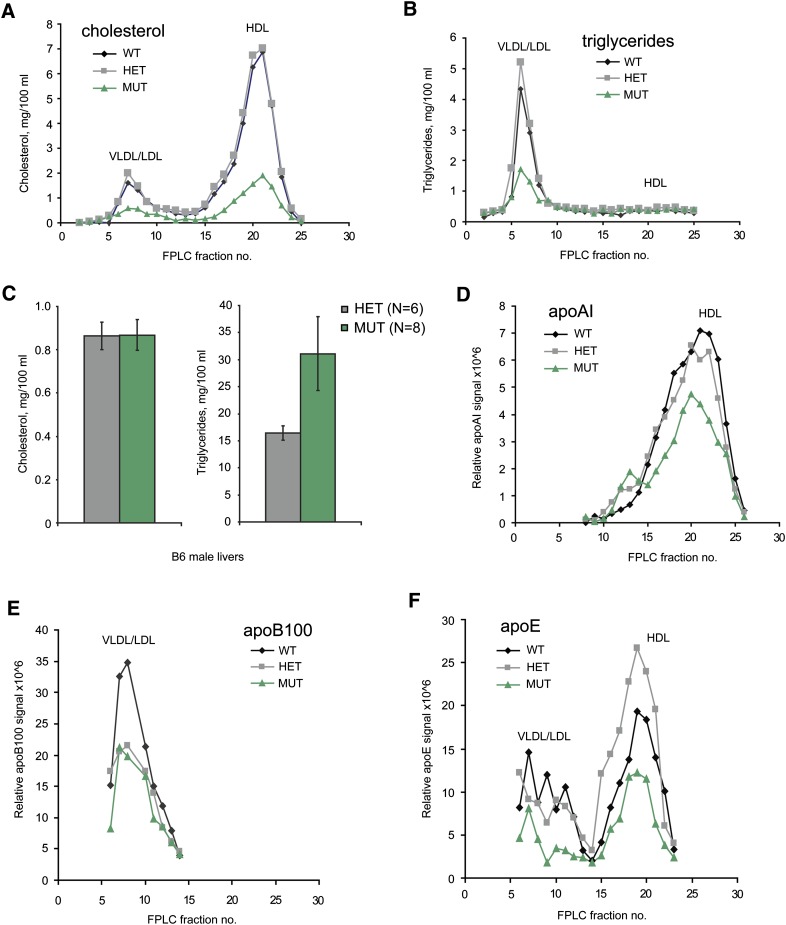
Whole plasma had reduced total cholesterol and HDL-C in Mia2cpto/cpto mice (Fig. 1F). To more directly assess the different lipoproteins and their lipid content, size-fractionated plasma was examined. FPLC plasma fractionation revealed that Mia2 mutant mice had lower cholesterol levels in all lipoprotein fractions, not just in HDL (Fig. 5A). As expected, cholesterol in the HDL fractions (samples 18–22, Fig. 5A) was severely reduced among Mia2cpto/cpto animals (MUT), to approximately 25–30% of the levels observed in WT, whereas cholesterol levels in Mia2cpto/+ animals (HET) were indistinguishable from those in WT. Furthermore, cholesterol levels in fractions from the VLDL/LDL peak (samples 5–9, Fig. 5A) were also decreased to 25–30% of WT levels in MUT animals. Triglyceride, which is largely carried in VLDL particles, was also affected by the cpto mutation, with a significant reduction among homozygous mutants in the VLDL/LDL fractions (samples 5–9, Fig. 5B). The cpto mutation thus affected the lipid content of multiple plasma lipoproteins. It should be noted that the degree of reduction in triglycerides in the fractionated plasma is similar to that of cholesterol in VLDL/LDL and HDL, although the reduction is larger than that observed in whole plasma (see supplementary Fig. IIA). This may be because free glycerol is present in whole plasma, but not the fractionated samples, although this has not been directly tested.
Fig. 5.
Reduced cholesterol and triglycerides in Mia2cpto mice. A: Total cholesterol in FPLC fractions of pooled plasma from Mia2+/+ (WT, n = 4), Mia2cpto/+ (HET, n = 6), and Mia2cpto/cpto (MUT, n = 6) mice. VLDL/LDL and HDL peaks are marked. Cholesterol levels in the MUT pool were lower in fractions 6–24, including VLDL, LDL, and HDL. B: Triglyceride levels in the same FPLC fractions as in part A were again lower in MUT animals. C: Direct measurement of cholesterol (left panel) and triglycerides (right panel) in liver samples from HET and MUT mice revealed no difference in cholesterol levels, but did show a trend toward increased triglyceride levels in MUT livers. Data are expressed as mean ± SEM. D–F: Apolipoproteins in FPLC fractions from part A. D: ApoA-I was modestly lower in HDL fractions from pooled MUT plasma, but not to the same extent as the cholesterol in A. E: Amounts of apoB-100 in MUT VLDL/LDL fractions were not discernibly different from those in HET fractions. F: Although the levels of apoE in MUT fractions are low, they are also highly variable between HET and WT fractions, suggesting that the differences are not significant.
Because Mia2 resembles secretory pathway-regulating proteins such as Mia3 and Tango1, hepatic cholesterol and triglyceride levels were directly measured to determine whether lowered plasma lipids coincides with increased retention of lipids within the liver (Fig. 5C). Interestingly, no significant difference was evident between HET and MUT hepatic cholesterol levels (Fig. 5C, left panel). In contrast, however, there was a distinct trend toward increased triglyceride levels in the liver (Fig. 5C, right panel), although this fell short of statistical significance (P = 0.07). Some inherent regulation of hepatic triglyceride levels in Mia2cpto mice must still be operative, however, because mutant mice did not progress to an overtly steatotic state histologically (see supplementary Fig. IIF).
cpto mutation does not appear to directly alter #8232apolipoprotein secretion
Both Mia3/Tango1 and fly Tango1 appear to regulate secretory processes, and depletion of either protein results in demonstrable defects in the secretion of specific proteins (4, 9). It is possible, then, that defective Mia2 could lower plasma cholesterol levels through reduced secretion of apolipoproteins. ApoA-I is the major protein in newly formed HDL, and apoB-100 is the primary protein in VLDL and LDL (26). Despite apparent increased hepatic triglyceride levels and decreased plasma levels of cholesterol and triglycerides, however, no consistent, Mia2cpto-dependent alteration in circulating apolipoproteins was observed. Analyzing whole, unfractionated plasma from individual mice revealed wide variation in the levels of apoA-I from mouse to mouse, but low levels of apoA-I in whole plasma did not consistently correlate with the Mia2cpto genotype (see supplementary Fig. VIIIA). Levels of plasma apoE, apoB-100, and apoB-48 were similarly variable (see supplementary Fig. VIIIA).
To control for inter-mouse variability, and to determine whether differences in apolipoprotein levels were associated with specific lipoproteins, we identified the proteins in individual fractions of the fractionated plasma pools (Fig. 5A, B) by Western blotting (Fig. 5DndashF). Whereas apoA-I was modestly reduced in fractions of the MUT pool corresponding to HDL, relative to the HET and WT pools (60% of WT levels), this reduction was less dramatic than the overall reduction seen in mutant HDL-C (Fig. 5A, D). Similarly, cholesterol was dramatically reduced in VLDL/LDL fractions of the MUT pool relative to both HET and WT pools (Fig. 5A), yet no obvious reduction in apoB-100 levels was evident in MUT pool fractions relative to HET pool fractions (Fig. 5E). Both MUT and HET pools contained less apoB-100 than did WT pools, but because HET animals were phenotypically unaffected by the Mia2cpto mutation, this difference was not related to the Mia2cpto mutation. Levels of other apolipoproteins, such as apoE (found on HDL and VLDL/LDL) and apoB-48 (chylomicron and LDL in mouse) also did not correlate with the Mia2cpto genotype (Fig. 5F and data not shown). Furthermore, no differences were observed in the levels of apoA-I, apoB-100, or apoE secreted into the media of HuH-7 cells either overexpressing WT versus cpto mutant full-length Mia2 cDNAs, or of HuH-7 cells transfected with MIA2 siRNA pools. Furthermore, these proteins did not detectably accumulate within the cells (see supplementary Fig. IXA, B, and data not shown).
Arguing against a general defect in secretion, albumin, a liver-secreted blood protein, was not significantly different between mutant and WT as measured by Coomassie staining of fractionated plasma shown in Fig. 5 (data not shown). Another liver-secreted plasma protein, Rbp4, was not reduced in mutant plasma, as measured by ELISA (see supplementary Fig. VIIIB). We determined whether the Mia2 protein or its isolated SH3 domain interacted with known apolipoproteins, and if these interactions were affected by the cpto mutation. In coimmunoprecipitation experiments using mouse liver membrane extracts from Mia2cpto/+ or Mia2cpto/cpto animals, no direct binding of endogenous Mia2 to apoA-I, apoB-100, apoB-48, or apoE was observed (see supplementary Fig. VIIIC). Similar results were observed in HuH-7 cells (data not shown). Together, although there is a slight reduction in some apolipoproteins in Mia2cpto/cpto plasma, the low cholesterol levels in Mia2cpto/cpto mice did not result from either a general or an apolipoprotein-specific defect in Mia2-dependent secretion from the liver.
DISCUSSION
Couch potato, a novel ENU-induced mutation that reduced circulating VLDL-C, LDL-C, HDL-C, and triglyceride levels, has been identified in mice. The mutation resides in a poorly characterized gene not previously related to cholesterol metabolism, Mia2. The Mia2cpto mutation alters a highly conserved residue in Mia2’s N-terminal SH3 domain (F91S). Mia2 is a largely liver-specific protein, encoded by exons previously annotated separately as the Mia2 and Ctage5/Mgea6 genes. The transcript that encodes full-length Mia2 (p240) spans both sets of exons, and like the related MIA3/TANGO1 protein (4), localized to ER exit sites, with a lumenally localized SH3 connected to a cytosolic carboxy-terminal proline-rich domain that interacted with the COP II proteins Sec23 and Sec24.
Mia2 is the first non-SH3-only member of this family of genes to have a mutant phenotype in whole animals. The Mia2cpto point mutation modestly reduced the overall steady-state level of full-length Mia2 protein in the liver, although the exact mechanism by which this resulted in lower levels of VLDL-C/LDL-C, HDL-C, and triglycerides in the plasma remains an open question. In vitro reduction of the levels of either fly Tango1 or human MIA3 results in demonstrable, cargo-specific, defects in protein secretion (4, 9). In Mia2cpto/cpto mice, there were variably reduced levels of some liver-secreted apolipoproteins, such as apoA-I and apoE (Fig. 5), yet the degree of reduction in apoA-I protein was inconsistent and smaller than the overall reduction in HDL-C. In addition, we observed a reduction in VLDL-C, but not of the major protein component of VLDL, apoB-100. It remains possible that cargo-specific secretion defects are the mechanism responsible for the lowering of cholesterol levels in Mia2cpto/cpto mice, and that we have not found the cargo that is altered. Attempts to detect protein-protein interactions with the Mia2 N-terminal portion have been hindered by mislocalization of the overexpressed protein (throughout the ER; see supplementary Fig. VIIA), and inability of the human-specific antibodies (anti-MIA2 and anti-CTAGE5) to immunoprecipitate native mouse or human protein. A pilot immunoprecipitation-mass spectrometry approach using an overexpressed, epitope-tagged Mia2 N-terminal fragment failed to detect interaction with any proteins known to be associated with cholesterol metabolism (data not shown).
The Mia2cpto mutation apparently did not directly alter general or cholesterol-specific hepatic protein secretion, although these mice did display a trend toward increased triglyceride levels, but not cholesterol levels, in their livers. Two other possibilities for how Mia2 affected plasma cholesterol levels are an effect at the level of cholesterol and/or lipoprotein synthesis/assembly, or an effect on lipoproteins after they are secreted, via enzymatic activity, distal uptake, or catabolism. Although we cannot rule out an effect on lipoprotein assembly or postsecreted lipoproteins, we have evidence that fails to support an effect on lipid synthesis. Cholesterol synthesis is regulated by the SREBP family of transcription factors, whose own activity is regulated by ER-to-Golgi trafficking (27). If Mia2 is also involved in SREBP regulation, the Mia2cpto mutation could decrease SREBP processing, and thus downregulate SREBP target genes in the liver. The end result of this cascade in the mouse would be reduced circulating cholesterol and triglyceride levels. Analysis of mRNA expression in mouse liver, under conditions in which SREBP-1c (fasting and refeeding; see supplementary Fig. XA) or SREBP2 (lovastatin plus ezetimibe treatment; see supplementary Fig. XB–D) are induced, have failed to show a difference in SREBP target gene expression between Mia2cpto/+ and Mia2cpto/cpto mice. Notably, direct measurement of liver cholesterol and triglyceride levels in these same mice (fasted, fasted-refed, lovastatin plus ezetimibe-treated; see supplementary Fig. XIA–C) showed the same trend of increased triglycerides but unchanged cholesterol levels observed in chow-fed mice (Fig. 5). Regardless of the mechanism, Mia2 represents a novel gene regulating hepatic cholesterol metabolism.
Mia2, Mia3/Tango1, and human disease
Whereas the Mia2cpto mutation produces a dramatic cholesterol phenotype in mice, it is not necessarily the case that MIA2 performs a similar role in humans. Some aspects of cholesterol metabolism differ significantly between rodents and humans, including, for example, the relative levels of LDL and HDL cholesterol, which are nearly inverse between mice and humans. Nonetheless, several aspects of Mia2 biology suggest that human MIA2 could function analogously to its murine counterpart. First, as in mice, MIA2 expression in humans is liver specific (1), and we detected an MIA2 protein in human hepatocyte-like HuH-7 and HepG2 cells that was similar to the major Mia2 species in mouse liver. Still, no studies to date have linked MIA2/CTAGE5 variants to alterations in human cholesterol levels. Multiple genome-wide association studies have, however, identified an association between variants within the human MIA3 locus and cardiovascular disease, although this association does not appear to be mediated by plasma lipids (5, 6, 28). On the basis of our results in mice, we would expect a mutation in human MIA2 to similarly be associated with cholesterol levels and/or cardiovascular disease; although no such human variants have yet been identified or reported.
It was recently shown that a carboxy-terminal portion of MIA2 (CTAGE5, or MIA2 p120) forms heterooligomers with MIA3/TANGO1 in human tissue culture cells, via the two proteins’ cytosolic coiled-coil regions (29). Although we have no direct evidence, we would expect full-length, SH3-containing MIA2 p240 to also dimerize with MIA3/TANGO1, inasmuch as the cytosolic portions of MIA2 p240 and MIA2 p120 are identical. Depletion of CTAGE5 is sufficient to cause cellular retention of the MIA3 cargo, COL7A1 (29), which binds directly to MIA3′s SH3 domain within the ER lumen (4). This argues that the integrity of the heterodimer is important for MIA3 activity, and thus we would expect that changes in the levels or function of either MIA2 or MIA3 should alter heterodimer function.
It is also possible that given that MIA2 and MIA3 are both highly similar in their domain structure, bind to each other, and share some protein partners at their carboxy-terminal ends, MIA3 genetic variants might directly contribute to cardiovascular disease via a mechanism similar to the (as yet unknown) one altered by the Mia2cpto mutation in mice. Although modulating protein-protein interactions that occur within the ER lumen may present a formidable hurdle for drug development, MIA2, MIA3, or both genes could potentially represent novel targets in the ongoing struggle against human cardiovascular disease.
Mia2/Mia3/dTango1: regulators of secretion, metabolism, or both?
Drosophila Tango1 was originally identified as a regulator of secretion and/or Golgi organization because dTango1 knockdown in Drosophila S2 cells resulted in an 80% reduction in secretion of HRP, along with retention in the ER of a green fluorescent protein-tagged version of the Golgi protein Mannosidase II (9). Subsequent work showed that dTango1 regulates secretion of a secretable luciferase, and localized the protein to S2 cell ER exit sites (23). Similarly, knockdown of mammalian MIA3/TANGO1 results in defective secretion of collagen VII (encoded by COL7A1) and HRP, although notably not of collagen I, alkaline phosphatase, or VSV-G; nor is there a detectable reduction in gross protein secretion (9). Interestingly, similar results have also recently been reported for knockdown of a CTAGE5-only transcript in HeLa cells (29). All of these results are consistent with the interpretation that dTango1, Mia3, and related proteins such as Mia2 regulate ER-to-Golgi trafficking of a discrete set of proteins.
Yet knockdown of genes critical either for lipid biogenesis (ACC, Drosophila FASN, SCAP) or for cholesterol/steroid synthesis (HMG-CoA reductase, HMG-CoA synthase, and SCAP again), also produced reductions in HRP secretion similar to or greater than those observed for dTango1, along with similar ER retention of Mannosidase II-GFP (9). These genes were excluded from further analysis as “transport and Golgi organization” genes, because they were known regulators of lipid metabolism. Whereas MIA3 binds the COL7A1 protein in an SH3-dependent fashion, and either MIA3 or CTAGE5 depletion results in impaired COL7A1 secretion (4, 29), the conclusion that MIA3 acts as a specific adaptor for large cargo at ER exit sites is at odds with the requirement for both dTango1 and mammalian MIA3 for secretion of the 305 amino acid HRP protein (4, 9). An alternative hypothesis could be that, as with the Drosophila metabolic genes above, the observed secretion defects when CTAGE5, MIA3, or dTango1 are mutated or depleted may be indirect, and instead result from alteration of lipid and/or cholesterol metabolism (30).
Note added in proof
During review of this manuscript, a relevant paper was published by Wilson et al. describing the phenotype of Mia3 knockout mice, which die neonatally due to a defect in collagen secretion. Importantly, this work supports our claim that alternative transcripts arise from the Mia3 gene similar to those that arise from the Mia2 gene. The complete reference is: Wilson D. G., K. Phamluong, L. Li, M. Sun, T. C. Cao, P. S. Liu, Z. Modrusan, W. N. Sandoval, L. Rangell, R. A. Carano, A. S. Peterson, and M. J. Solloway. 2011. Global defects in collagen secretion in a Mia3/TANGO1 knockout mouse. J. Cell Biol. 193: 935–951.
Supplementary Material
Acknowledgments
The authors thank Sandy Bohan, Lacey Kischassey, Amelia Brown, and Claire Jantsch for their expert technical assistance; Brian Steffy and Tim Wiltshire for their work in mapping and identifying the cpto mutant interval; Sassan Azarian's group for performing the RBP4 ELISA assays; and Bill Kiosses for performing the confocal imaging. The authors are also grateful to Larry Gerace for critical review of the manuscript.
Footnotes
Abbreviations:
- B6
- C57BL/6
- ENU
- N-ethyl-N-nitrosourea
- ER
- endoplasmic reticulum
- EST
- expressed sequence tag
- HDL-C
- HDL-cholesterol
- SH3
- Src homology domain 3
- SNP
- single-nucleotide polymorphism
- WT
- wild-type
This work was supported by grants UL1 RR025774 and R01 DK079925 (N.G.).
The online version of this article (available at http://www.jlr.org) contains supplementary data in the form of eleven figures.
REFERENCES
- 1.Bosserhoff A. K., Moser M., Scholmerich J., Buettner R., Hellerbrand C. 2003. Specific expression and regulation of the new melanoma inhibitory activity-related gene MIA2 in hepatocytes. J. Biol. Chem. 278: 15225–15231. [DOI] [PubMed] [Google Scholar]
- 2.Bosserhoff A. K., Moser M., Buettner R. 2004. Characterization and expression pattern of the novel MIA homolog TANGO. Gene Expr. Patterns. 4: 473–479. [DOI] [PubMed] [Google Scholar]
- 3.Arndt S., Bosserhoff A. K. 2007. Reduced expression of TANGO in colon and hepatocellular carcinomas. Oncol. Rep. 18: 885–891. [PubMed] [Google Scholar]
- 4.Saito K., Chen M., Bard F., Chen S., Zhou H., Woodley D., Polischuk R., Schekman R., Malhotra V. 2009. TANGO1 facilitates cargo loading at endoplasmic reticulum exit sites. Cell. 136: 891–902. [DOI] [PubMed] [Google Scholar]
- 5.Samani N. J., Erdmann J., Hall A. S., Hengstenberg C., Mangino M., Mayer B., Dixon R. J., Meitinger T., Braund P., Wichmann H. E., et al. 2007. Genomewide association analysis of coronary artery disease. N. Engl. J. Med. 357: 443–453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kathiresan S., Voight B. F., Purcell S., Musunuru K., Ardissino D., Mannucci P. M., Anand S., Engert J. C., Samani N. J., Schunkert H., et al. 2009. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat. Genet. 41: 334–341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hellerbrand C., Bataille F., Schlegel J., Hartmann A., Muhlbauer M., Scholmerich J., Buttner R., Hofstadter F., Bosserhoff A. K. 2005. In situ expression patterns of melanoma inhibitory activity 2 in healthy and diseased livers. Liver Int. 25: 357–366. [DOI] [PubMed] [Google Scholar]
- 8.Hellerbrand C., Amann T., Schlegel J., Wild P., Bataille F., Spruss T., Hartmann A., Bosserhoff A. K. 2008. The novel gene MIA2 acts as a tumour suppressor in hepatocellular carcinoma. Gut. 57: 243–251. [DOI] [PubMed] [Google Scholar]
- 9.Bard F., Casano L., Mallabiabarrena A., Wallace E., Saito K., Kitayama H., Guizzunti G., Hu Y., Wendler F., Dasgupta R., et al. 2006. Functional genomics reveals genes involved in protein secretion and Golgi organization. Nature. 439: 604–607. [DOI] [PubMed] [Google Scholar]
- 10.Heckel D., Brass N., Fischer U., Blin N., Steudel I., Tureci O., Fackler O., Zang K. D., Meese E. 1997. cDNA cloning and chromosomal mapping of a predicted coiled-coil proline-rich protein immunogenic in meningioma patients. Hum. Mol. Genet. 6: 2031–2041. [DOI] [PubMed] [Google Scholar]
- 11.Lloyd D. J., Hall F. W., Tarantino L. M., Gekakis N. 2005. Diabetes insipidus in mice with a mutation in aquaporin-2. PLoS Genet. 1: e20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wen B. G., Pletcher M. T., Warashina M., Choe S. H., Ziaee N., Wiltshire T., Sauer K., Cooke M. P. 2004. Inositol (1,4,5) trisphosphate 3 kinase B controls positive selection of T cells and modulates Erk activity. Proc. Natl. Acad. Sci. USA. 101: 5604–5609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lloyd D. J., Wheeler M. C., Gekakis N. 2010. A point mutation in Sec61alpha1 leads to diabetes and hepatosteatosis in mice. Diabetes. 59: 460–470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Vojtek A. B., Hollenberg S. M., Cooper J. A. 1993. Mammalian Ras interacts directly with the serine/threonine kinase Raf. Cell. 74: 205–214. [DOI] [PubMed] [Google Scholar]
- 15.Gekakis N., Saez L., Delahaye-Brown A. M., Myers M. P., Sehgal A., Young M. W., Weitz C. J. 1995. Isolation of timeless by PER protein interaction: defective interaction between timeless protein and long-period mutant PERL. Science. 270: 811–815. [DOI] [PubMed] [Google Scholar]
- 16.Lee J. M., Lee Y. K., Mamrosh J. L., Busby S. A., Griffin P. R., Pathak M. C., Ortlund E. A., Moore D. D. 2011. A nuclear-receptor-dependent phosphatidylcholine pathway with antidiabetic effects. Nature. 474: 506–510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Stoll R., Renner C., Zweckstetter M., Bruggert M., Ambrosius D., Palme S., Engh R. A., Golob M., Breibach I., Buettner R., et al. 2001. The extracellular human melanoma inhibitory activity (MIA) protein adopts an SH3 domain-like fold. EMBO J. 20: 340–349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lougheed J. C., Holton J. M., Alber T., Bazan J. F., Handel T. M. 2001. Structure of melanoma inhibitory activity protein, a member of a recently identified family of secreted proteins. Proc. Natl. Acad. Sci. USA. 98: 5515–5520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Comtesse N., Niedermayer I., Glass B., Heckel D., Maldener E., Nastainczyk W., Feiden W., Meese E. 2002. MGEA6 is tumor-specific overexpressed and frequently recognized by patient-serum antibodies. Oncogene. 21: 239–247. [DOI] [PubMed] [Google Scholar]
- 20.Usener D., Schadendorf D., Koch J., Dubel S., Eichmuller S. 2003. cTAGE: a cutaneous T cell lymphoma associated antigen family with tumor-specific splicing. J. Invest. Dermatol. 121: 198–206. [DOI] [PubMed] [Google Scholar]
- 21.Su A. I., Wiltshire T., Batalov S., Lapp H., Ching K. A., Block D., Zhang J., Soden R., Hayakawa M., Kreiman G., et al. 2004. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA. 101: 6062–6067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Eichmuller S., Usener D., Dummer R., Stein A., Thiel D., Schadendorf D. 2001. Serological detection of cutaneous T-cell lymphoma-associated antigens. Proc. Natl. Acad. Sci. USA. 98: 629–634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wendler F., Gillingham A. K., Sinka R., Rosa-Ferreira C., Gordon D. E., Franch-Marro X., Peden A. A., Vincent J. P., Munro S. 2010. A genome-wide RNA interference screen identifies two novel components of the metazoan secretory pathway. EMBO J. 29: 304–314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Stone S. J., Levin M. C., Farese R. V., Jr 2006. Membrane topology and identification of key functional amino acid residues of murine acyl-CoA:diacylglycerol acyltransferase-2. J. Biol. Chem. 281: 40273–40282. [DOI] [PubMed] [Google Scholar]
- 25.Hannah M. J., Weiss U., Huttner W. B. 1998. Differential extraction of proteins from paraformaldehyde-fixed cells: lessons from synaptophysin and other membrane proteins. Methods. 16: 170–181. [DOI] [PubMed] [Google Scholar]
- 26.Olofsson S. O., Wiklund O., Boren J. 2007. Apolipoproteins A-I and B: biosynthesis, role in the development of atherosclerosis and targets for intervention against cardiovascular disease. Vasc. Health Risk Manag. 3: 491–502. [PMC free article] [PubMed] [Google Scholar]
- 27.Brown M. S., Goldstein J. L. 2009. Cholesterol feedback: from Schoenheimer's bottle to Scap's MELADL. J. Lipid Res. 50(Suppl.): 15–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Samani N. J., Deloukas P., Erdmann J., Hengstenberg C., Kuulasmaa K., McGinnis R., Schunkert H., Soranzo N., Thompson J., Tiret L., et al. 2009. Large scale association analysis of novel genetic loci for coronary artery disease. Arterioscler. Thromb. Vasc. Biol. 29: 774–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Saito K., Yamashiro K., Ichikawa Y., Erlmann P., Kontani K., Malhotra V., Katada T. 2011. cTAGE5 mediates collagen secretion through interaction with TANGO1 at endoplasmic reticulum exit sites. Mol. Biol. Cell. 22: 2301–2308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ridsdale A., Denis M., Gougeon P. Y., Ngsee J. K., Presley J. F., Zha X. 2006. Cholesterol is required for efficient endoplasmic reticulum-to-Golgi transport of secretory membrane proteins. Mol. Biol. Cell. 17: 1593–1605. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.