Abstract
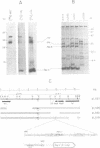
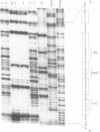
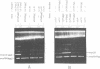
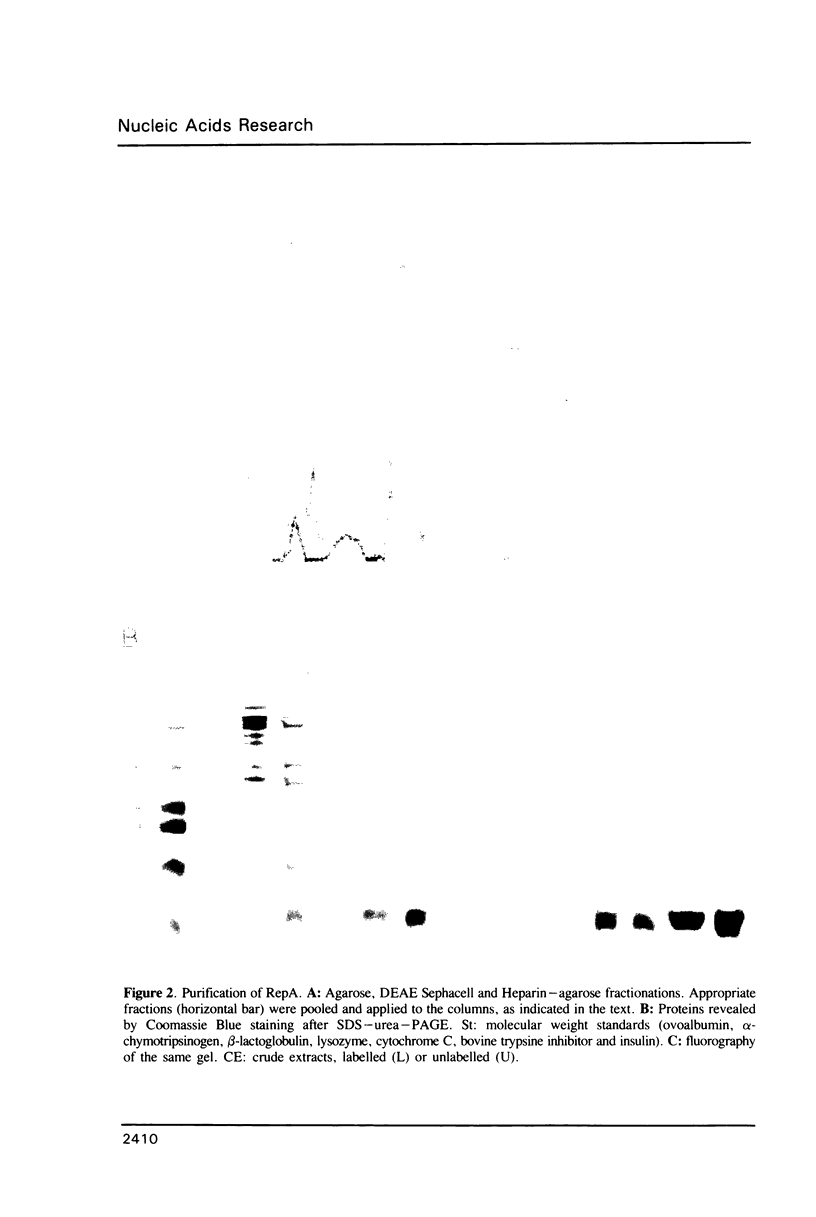
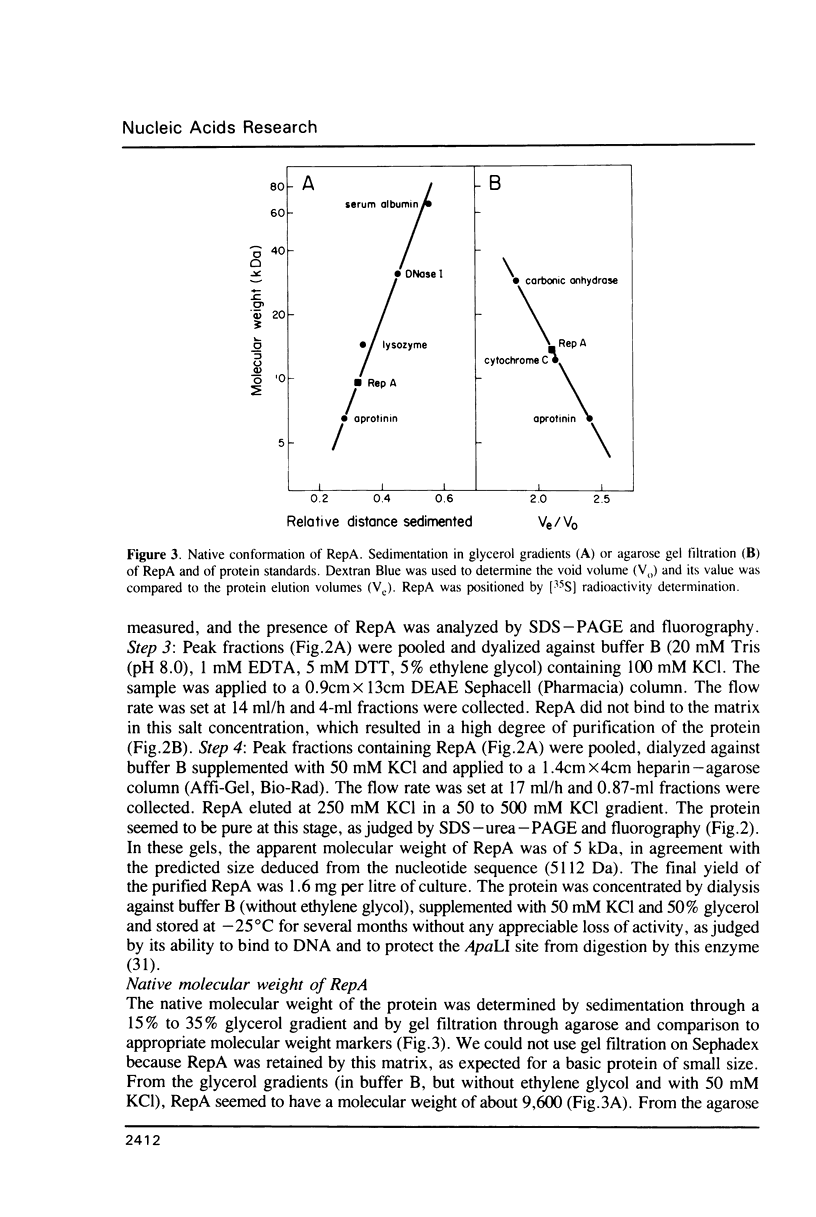
The promiscuous streptococcal plasmid pLS1 encodes for the 5.1 kDa RepA protein, involved in the regulation of the plasmid copy number. Synthesis of RepA was observed both in Bacillus subtilis minicells and in an Escherichia coli expression system. From this system, the protein has been purified and it appears to be a dimer of identical subunits. The amino acid sequence of RepA has been determined. RepA shows the alpha helix-turn-alpha helix motif typical of many DNA-binding proteins and it shares homology with a number of repressors, specially with the TrfB repressor encoded by the broad-host-range plasmid RK2. DNase I footprinting revealed that the RepA target is located in the region of the promoter for the repA and repB genes. Trans-complementation analysis showed that in vivo, RepA behaves as a repressor by regulating the plasmid copy number. We propose that the regulatory role of RepA is by limitation of the synthesis of the initiator protein RepB.
Full text
PDF











Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Chou P. Y., Fasman G. D. Prediction of the secondary structure of proteins from their amino acid sequence. Adv Enzymol Relat Areas Mol Biol. 1978;47:45–148. doi: 10.1002/9780470122921.ch2. [DOI] [PubMed] [Google Scholar]
- Dodd I. B., Egan J. B. Systematic method for the detection of potential lambda Cro-like DNA-binding regions in proteins. J Mol Biol. 1987 Apr 5;194(3):557–564. doi: 10.1016/0022-2836(87)90681-4. [DOI] [PubMed] [Google Scholar]
- Espinosa M., López P., Lacks S. A. Transfer and expression of recombinant plasmids carrying pneumococcal mal genes in Bacillus subtilis. Gene. 1984 Jun;28(3):301–310. doi: 10.1016/0378-1119(84)90147-1. [DOI] [PubMed] [Google Scholar]
- Gribskov M., Burgess R. R., Devereux J. PEPPLOT, a protein secondary structure analysis program for the UWGCG sequence analysis software package. Nucleic Acids Res. 1986 Jan 10;14(1):327–334. doi: 10.1093/nar/14.1.327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hewick R. M., Hunkapiller M. W., Hood L. E., Dreyer W. J. A gas-liquid solid phase peptide and protein sequenator. J Biol Chem. 1981 Aug 10;256(15):7990–7997. [PubMed] [Google Scholar]
- Jordan S. R., Pabo C. O. Structure of the lambda complex at 2.5 A resolution: details of the repressor-operator interactions. Science. 1988 Nov 11;242(4880):893–899. doi: 10.1126/science.3187530. [DOI] [PubMed] [Google Scholar]
- Kok J., van der Vossen J. M., Venema G. Construction of plasmid cloning vectors for lactic streptococci which also replicate in Bacillus subtilis and Escherichia coli. Appl Environ Microbiol. 1984 Oct;48(4):726–731. doi: 10.1128/aem.48.4.726-731.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Lacks S. A., Lopez P., Greenberg B., Espinosa M. Identification and analysis of genes for tetracycline resistance and replication functions in the broad-host-range plasmid pLS1. J Mol Biol. 1986 Dec 20;192(4):753–765. doi: 10.1016/0022-2836(86)90026-4. [DOI] [PubMed] [Google Scholar]
- Lehming N., Sartorius J., Oehler S., von Wilcken-Bergmann B., Müller-Hill B. Recognition helices of lac and lambda repressor are oriented in opposite directions and recognize similar DNA sequences. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7947–7951. doi: 10.1073/pnas.85.21.7947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopez P., Espinosa M., Lacks S. A. Physical structure and genetic expression of the sulfonamide-resistance plasmid pLS80 and its derivatives in Streptococcus pneumoniae and Bacillus subtilis. Mol Gen Genet. 1984;195(3):403–410. doi: 10.1007/BF00341440. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Meyer B. J., Kleid D. G., Ptashne M. Lambda repressor turns off transcription of its own gene. Proc Natl Acad Sci U S A. 1975 Dec;72(12):4785–4789. doi: 10.1073/pnas.72.12.4785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molin S., Stougaard P., Light J., Nordström M., Nordström K. Isolation and characterization of new copy mutants of plasmid R1, and identification of a polypeptide involved in copy number control. Mol Gen Genet. 1981;181(1):123–130. doi: 10.1007/BF00339015. [DOI] [PubMed] [Google Scholar]
- Nordström K., Molin S., Aagaard-Hansen H. Partitioning of plasmid R1 in Escherichia coli. II. Incompatibility properties of the partitioning system. Plasmid. 1980 Nov;4(3):332–339. doi: 10.1016/0147-619x(80)90071-2. [DOI] [PubMed] [Google Scholar]
- Nordström K., Molin S., Light J. Control of replication of bacterial plasmids: genetics, molecular biology, and physiology of the plasmid R1 system. Plasmid. 1984 Sep;12(2):71–90. doi: 10.1016/0147-619x(84)90054-4. [DOI] [PubMed] [Google Scholar]
- Novick R. P. Plasmid incompatibility. Microbiol Rev. 1987 Dec;51(4):381–395. doi: 10.1128/mr.51.4.381-395.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohlendorf D. H., Anderson W. F., Matthews B. W. Many gene-regulatory proteins appear to have a similar alpha-helical fold that binds DNA and evolved from a common precursor. J Mol Evol. 1983;19(2):109–114. doi: 10.1007/BF02300748. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Piggot P. J., Hoch J. A. Revised genetic linkage map of Bacillus subtilis. Microbiol Rev. 1985 Jun;49(2):158–179. doi: 10.1128/mr.49.2.158-179.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poteete A. R., Hehir K., Sauer R. T. Bacteriophage P22 Cro protein: sequence, purification, and properties. Biochemistry. 1986 Jan 14;25(1):251–256. doi: 10.1021/bi00349a035. [DOI] [PubMed] [Google Scholar]
- Projan S. J., Carleton S., Novick R. P. Determination of plasmid copy number by fluorescence densitometry. Plasmid. 1983 Mar;9(2):182–190. doi: 10.1016/0147-619x(83)90019-7. [DOI] [PubMed] [Google Scholar]
- Projan S. J., Novick R. Comparative analysis of five related Staphylococcal plasmids. Plasmid. 1988 May;19(3):203–221. doi: 10.1016/0147-619x(88)90039-x. [DOI] [PubMed] [Google Scholar]
- Puyet A., del Solar G. H., Espinosa M. Identification of the origin and direction of replication of the broad-host-range plasmid pLS1. Nucleic Acids Res. 1988 Jan 11;16(1):115–133. doi: 10.1093/nar/16.1.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pérez-Martín J., del Solar G. H., de la Campa A. G., Espinosa M. Three regions in the DNA of plasmid pLS1 show sequence-directed static bending. Nucleic Acids Res. 1988 Oct 11;16(19):9113–9126. doi: 10.1093/nar/16.19.9113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riise E., Molin S. Purification and characterization of the CopB replication control protein, and precise mapping of its target site in the R1 plasmid. Plasmid. 1986 May;15(3):163–171. doi: 10.1016/0147-619x(86)90034-x. [DOI] [PubMed] [Google Scholar]
- Rosenberg A. H., Lade B. N., Chui D. S., Lin S. W., Dunn J. J., Studier F. W. Vectors for selective expression of cloned DNAs by T7 RNA polymerase. Gene. 1987;56(1):125–135. doi: 10.1016/0378-1119(87)90165-x. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Moffatt B. A. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986 May 5;189(1):113–130. doi: 10.1016/0022-2836(86)90385-2. [DOI] [PubMed] [Google Scholar]
- Theophilus B. D., Thomas C. M. Nucleotide sequence of the transcriptional repressor gene korB which plays a key role in regulation of the copy number of broad host range plasmid RK2. Nucleic Acids Res. 1987 Sep 25;15(18):7443–7450. doi: 10.1093/nar/15.18.7443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas C. M. Recent studies on the control of plasmid replication. Biochim Biophys Acta. 1988 Mar 31;949(3):253–263. doi: 10.1016/0167-4781(88)90150-9. [DOI] [PubMed] [Google Scholar]
- Van Kaer L., Van Montagu M., Dhaese P. Transcriptional control in the EcoRI-F immunity region of Bacillus subtilis phage phi 105. Identification and unusual structure of the operator. J Mol Biol. 1987 Sep 5;197(1):55–67. doi: 10.1016/0022-2836(87)90609-7. [DOI] [PubMed] [Google Scholar]
- de la Campa A. G., Springhorn S. S., Kale P., Lacks S. A. Proteins encoded by the DpnI restriction gene cassette. Hyperproduction and characterization of the DpnI endonuclease. J Biol Chem. 1988 Oct 15;263(29):14696–14702. [PubMed] [Google Scholar]
- del Solar G. H., Puyet A., Espinosa M. Initiation signals for the conversion of single stranded to double stranded DNA forms in the streptococcal plasmid pLS1. Nucleic Acids Res. 1987 Jul 24;15(14):5561–5580. doi: 10.1093/nar/15.14.5561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- te Riele H., Michel B., Ehrlich S. D. Are single-stranded circles intermediates in plasmid DNA replication? EMBO J. 1986 Mar;5(3):631–637. doi: 10.1002/j.1460-2075.1986.tb04257.x. [DOI] [PMC free article] [PubMed] [Google Scholar]