Abstract
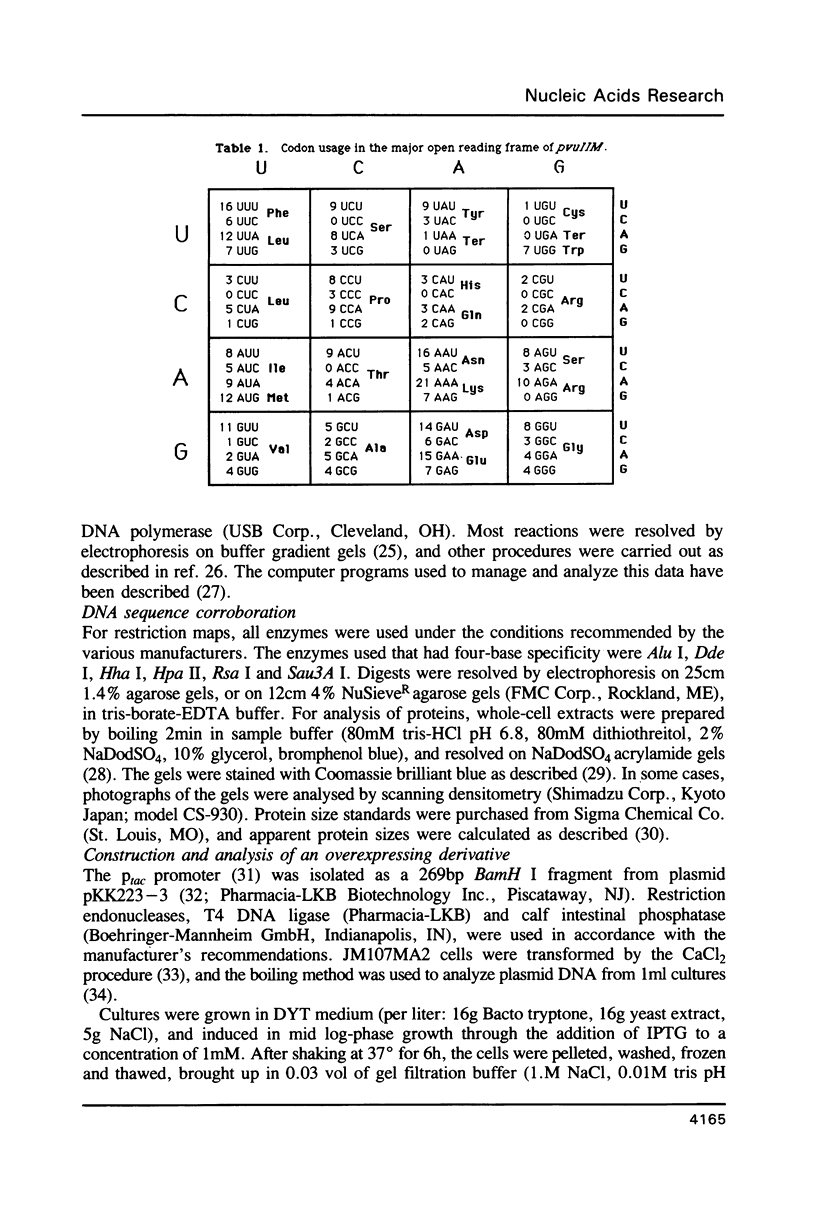
The base sequence of the pvuIIM gene has been determined. This gene codes for a DNA-(cytosine N4)-methyltransferase, M.Pvu II. The base sequence contains a single large open reading frame that predicts a 38.3kDa polypeptide, consistent with experimental data. The pvuIIM gene contains some sequences common to DNA methyltransferases in general, but includes none of the sequences specifically conserved among DNA-(cytosine 5)-methyltransferases. The pvuIIM sequence also reveals an internal homology at the amino acid level, each half of which spans over 100 amino acids and is itself homologous to the sequences of some DNA-(adenine N6)-methyltransferases. A derivative of the pvuIIM plasmid was constructed to allow high-level production of M.Pvu II. Specifically, the composite Ptac promoter was inserted 5' to pvuIIM, intervening DNA was deleted, and the resulting construct was used to transform an mcrB laclq strain of Escherichia coli. When this transformant was induced with isopropyl-B-D-galactopyranoside (IPTG), growth rapidly ceased and M.Pvu II accumulated to the point of comprising over 10% of the total soluble protein.
Full text
PDF










Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amann E., Brosius J. "ATG vectors' for regulated high-level expression of cloned genes in Escherichia coli. Gene. 1985;40(2-3):183–190. doi: 10.1016/0378-1119(85)90041-1. [DOI] [PubMed] [Google Scholar]
- Bestor T., Laudano A., Mattaliano R., Ingram V. Cloning and sequencing of a cDNA encoding DNA methyltransferase of mouse cells. The carboxyl-terminal domain of the mammalian enzymes is related to bacterial restriction methyltransferases. J Mol Biol. 1988 Oct 20;203(4):971–983. doi: 10.1016/0022-2836(88)90122-2. [DOI] [PubMed] [Google Scholar]
- Biggin M. D., Gibson T. J., Hong G. F. Buffer gradient gels and 35S label as an aid to rapid DNA sequence determination. Proc Natl Acad Sci U S A. 1983 Jul;80(13):3963–3965. doi: 10.1073/pnas.80.13.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumenthal R. M., Gregory S. A., Cooperider J. S. Cloning of a restriction-modification system from Proteus vulgaris and its use in analyzing a methylase-sensitive phenotype in Escherichia coli. J Bacteriol. 1985 Nov;164(2):501–509. doi: 10.1128/jb.164.2.501-509.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumenthal R. M., Reeh S., Pedersen S. Regulation of transcription factor rho and the alpha subunit of RNA polymerase in Escherichia coli B/r. Proc Natl Acad Sci U S A. 1976 Jul;73(7):2285–2288. doi: 10.1073/pnas.73.7.2285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumenthal R. M., Rice P. J., Roberts R. J. Computer programs for nucleic acid sequence manipulation. Nucleic Acids Res. 1982 Jan 11;10(1):91–101. doi: 10.1093/nar/10.1.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butkus V., Klimasauskas S., Petrauskiene L., Maneliene Z., Lebionka A., Janulaitis A. Interaction of AluI, Cfr6I and PvuII restriction-modification enzymes with substrates containing either N4-methylcytosine or 5-methylcytosine. Biochim Biophys Acta. 1987 Aug 25;909(3):201–207. doi: 10.1016/0167-4781(87)90078-9. [DOI] [PubMed] [Google Scholar]
- DOSKOCIL J., SORMO'VA Z. THE OCCURRENCE OF 5-METHYLCYTOSINE IN BACTERIAL DEOXYRIBONUCLEIC ACIDS. Biochim Biophys Acta. 1965 Mar 15;95:513–515. [PubMed] [Google Scholar]
- DUNN D. B., SMITH J. D. The occurrence of 6-methylaminopurine in deoxyribonucleic acids. Biochem J. 1958 Apr;68(4):627–636. doi: 10.1042/bj0680627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrlich M., Gama-Sosa M. A., Carreira L. H., Ljungdahl L. G., Kuo K. C., Gehrke C. W. DNA methylation in thermophilic bacteria: N4-methylcytosine, 5-methylcytosine, and N6-methyladenine. Nucleic Acids Res. 1985 Feb 25;13(4):1399–1412. doi: 10.1093/nar/13.4.1399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrlich M., Norris K. F., Wang R. Y., Kuo K. C., Gehrke C. W. DNA cytosine methylation and heat-induced deamination. Biosci Rep. 1986 Apr;6(4):387–393. doi: 10.1007/BF01116426. [DOI] [PubMed] [Google Scholar]
- Ehrlich M., Wilson G. G., Kuo K. C., Gehrke C. W. N4-methylcytosine as a minor base in bacterial DNA. J Bacteriol. 1987 Mar;169(3):939–943. doi: 10.1128/jb.169.3.939-943.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gamas P., Chandler M. G., Prentki P., Galas D. J. Escherichia coli integration host factor binds specifically to the ends of the insertion sequence IS1 and to its major insertion hot-spot in pBR322. J Mol Biol. 1987 May 20;195(2):261–272. doi: 10.1016/0022-2836(87)90648-6. [DOI] [PubMed] [Google Scholar]
- Gingeras T. R., Greenough L., Schildkraut I., Roberts R. J. Two new restriction endonucleases from Proteus vulgaris. Nucleic Acids Res. 1981 Sep 25;9(18):4525–4536. doi: 10.1093/nar/9.18.4525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greene P. J., Gupta M., Boyer H. W., Brown W. E., Rosenberg J. M. Sequence analysis of the DNA encoding the Eco RI endonuclease and methylase. J Biol Chem. 1981 Mar 10;256(5):2143–2153. [PubMed] [Google Scholar]
- Hattman S., Wilkinson J., Swinton D., Schlagman S., Macdonald P. M., Mosig G. Common evolutionary origin of the phage T4 dam and host Escherichia coli dam DNA-adenine methyltransferase genes. J Bacteriol. 1985 Nov;164(2):932–937. doi: 10.1128/jb.164.2.932-937.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattori M., Sakaki Y. Dideoxy sequencing method using denatured plasmid templates. Anal Biochem. 1986 Feb 1;152(2):232–238. doi: 10.1016/0003-2697(86)90403-3. [DOI] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Janulaitis A., Klimasauskas S., Petrusyte M., Butkus V. Cytosine modification in DNA by BcnI methylase yields N4-methylcytosine. FEBS Lett. 1983 Sep 5;161(1):131–134. doi: 10.1016/0014-5793(83)80745-5. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lauster R., Kriebardis A., Guschlbauer W. The GATATC-modification enzyme EcoRV is closely related to the GATC-recognizing methyltransferases DpnII and dam from E. coli and phage T4. FEBS Lett. 1987 Aug 10;220(1):167–176. doi: 10.1016/0014-5793(87)80897-9. [DOI] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Miller H. I., Nash H. A. Direct role of the himA gene product in phage lambda integration. Nature. 1981 Apr 9;290(5806):523–526. doi: 10.1038/290523a0. [DOI] [PubMed] [Google Scholar]
- Nardone G., George J., Chirikjian J. G. Sequence-specific BamHI methylase. Purification and characterization. J Biol Chem. 1984 Aug 25;259(16):10357–10362. [PubMed] [Google Scholar]
- Newman A. K., Rubin R. A., Kim S. H., Modrich P. DNA sequences of structural genes for Eco RI DNA restriction and modification enzymes. J Biol Chem. 1981 Mar 10;256(5):2131–2139. [PubMed] [Google Scholar]
- Pogolotti A. L., Jr, Ono A., Subramaniam R., Santi D. V. On the mechanism of DNA-adenine methylase. J Biol Chem. 1988 Jun 5;263(16):7461–7464. [PubMed] [Google Scholar]
- Pósfai J., Bhagwat A. S., Roberts R. J. Sequence motifs specific for cytosine methyltransferases. Gene. 1988 Dec 25;74(1):261–265. doi: 10.1016/0378-1119(88)90299-5. [DOI] [PubMed] [Google Scholar]
- Rubin R. A., Modrich P. EcoRI methylase. Physical and catalytic properties of the homogeneous enzyme. J Biol Chem. 1977 Oct 25;252(20):7265–7272. [PubMed] [Google Scholar]
- Sanchez-Pescador R., Urdea M. S. Use of unpurified synthetic deoxynucleotide primers for rapid dideoxynucleotide chain termination sequencing. DNA. 1984 Aug;3(4):339–343. doi: 10.1089/dna.1.1984.3.339. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seeman N. C., Rosenberg J. M., Rich A. Sequence-specific recognition of double helical nucleic acids by proteins. Proc Natl Acad Sci U S A. 1976 Mar;73(3):804–808. doi: 10.1073/pnas.73.3.804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shepherd J. C. From primeval message to present-day gene. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1099–1108. doi: 10.1101/sqb.1983.047.01.124. [DOI] [PubMed] [Google Scholar]
- Smith D. I., Blattner F. R., Davies J. The isolation and partial characterization of a new restriction endonuclease from Providencia stuartii. Nucleic Acids Res. 1976 Feb;3(2):343–353. doi: 10.1093/nar/3.2.343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Som S., Bhagwat A. S., Friedman S. Nucleotide sequence and expression of the gene encoding the EcoRII modification enzyme. Nucleic Acids Res. 1987 Jan 12;15(1):313–332. doi: 10.1093/nar/15.1.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sznyter L. A., Slatko B., Moran L., O'Donnell K. H., Brooks J. E. Nucleotide sequence of the DdeI restriction-modification system and characterization of the methylase protein. Nucleic Acids Res. 1987 Oct 26;15(20):8249–8266. doi: 10.1093/nar/15.20.8249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szybalski W., Blumenthal R. M., Brooks J. E., Hattman S., Raleigh E. A. Nomenclature for bacterial genes coding for class-II restriction endonucleases and modification methyltransferases. Gene. 1988 Dec 25;74(1):279–280. doi: 10.1016/0378-1119(88)90303-4. [DOI] [PubMed] [Google Scholar]
- Theriault G., Roy P. H., Howard K. A., Benner J. S., Brooks J. E., Waters A. F., Gingeras T. R. Nucleotide sequence of the PaeR7 restriction/modification system and partial characterization of its protein products. Nucleic Acids Res. 1985 Dec 9;13(23):8441–8461. doi: 10.1093/nar/13.23.8441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tjian R. T antigen binding and the control of SV40 gene expression. Cell. 1981 Oct;26(1 Pt 1):1–2. doi: 10.1016/0092-8674(81)90026-x. [DOI] [PubMed] [Google Scholar]
- Walder R. Y., Walder J. A., Donelson J. E. The organization and complete nucleotide sequence of the PstI restriction-modification system. J Biol Chem. 1984 Jun 25;259(12):8015–8026. [PubMed] [Google Scholar]
- Weber K., Osborn M. The reliability of molecular weight determinations by dodecyl sulfate-polyacrylamide gel electrophoresis. J Biol Chem. 1969 Aug 25;244(16):4406–4412. [PubMed] [Google Scholar]
- Wilke K., Rauhut E., Noyer-Weidner M., Lauster R., Pawlek B., Behrens B., Trautner T. A. Sequential order of target-recognizing domains in multispecific DNA-methyltransferases. EMBO J. 1988 Aug;7(8):2601–2609. doi: 10.1002/j.1460-2075.1988.tb03110.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodbury C. P., Jr, von Hippel P. H. Relaxed sequence specificities of Eco RI endonuclease and methylase: mechanisms, possible practical applications, and uses in defining protein-nucleic acid recognition mechanisms. Gene Amplif Anal. 1981;1:181–207. [PubMed] [Google Scholar]
- Wu J. C., Santi D. V. Kinetic and catalytic mechanism of HhaI methyltransferase. J Biol Chem. 1987 Apr 5;262(10):4778–4786. [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- de Boer H. A., Comstock L. J., Vasser M. The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc Natl Acad Sci U S A. 1983 Jan;80(1):21–25. doi: 10.1073/pnas.80.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]



