Abstract
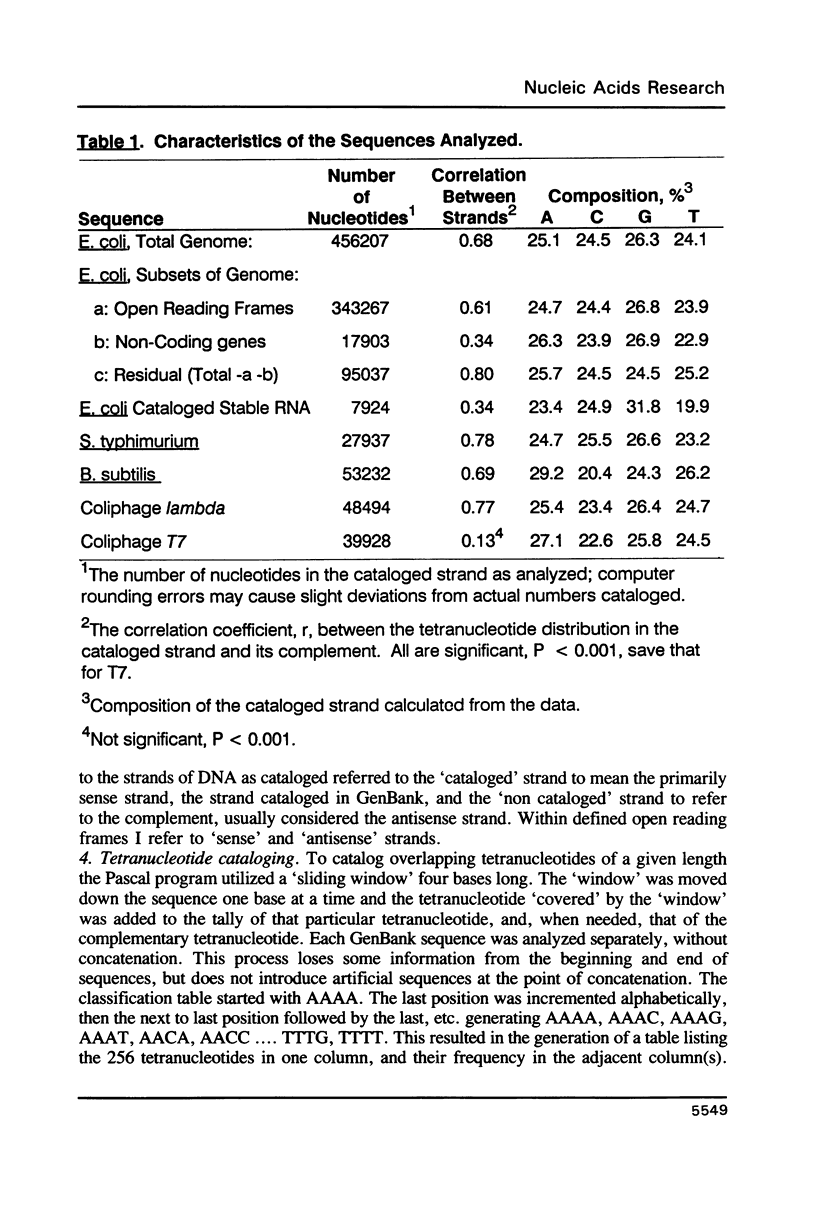
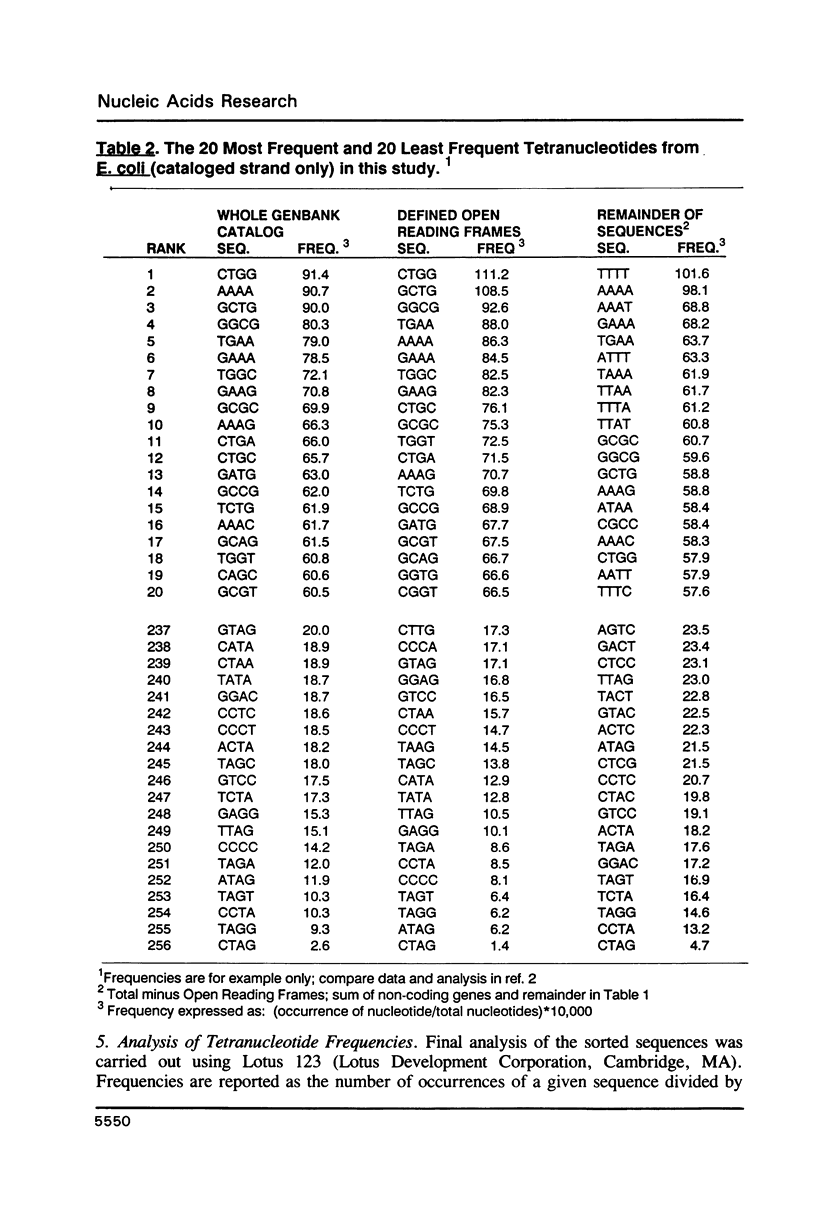
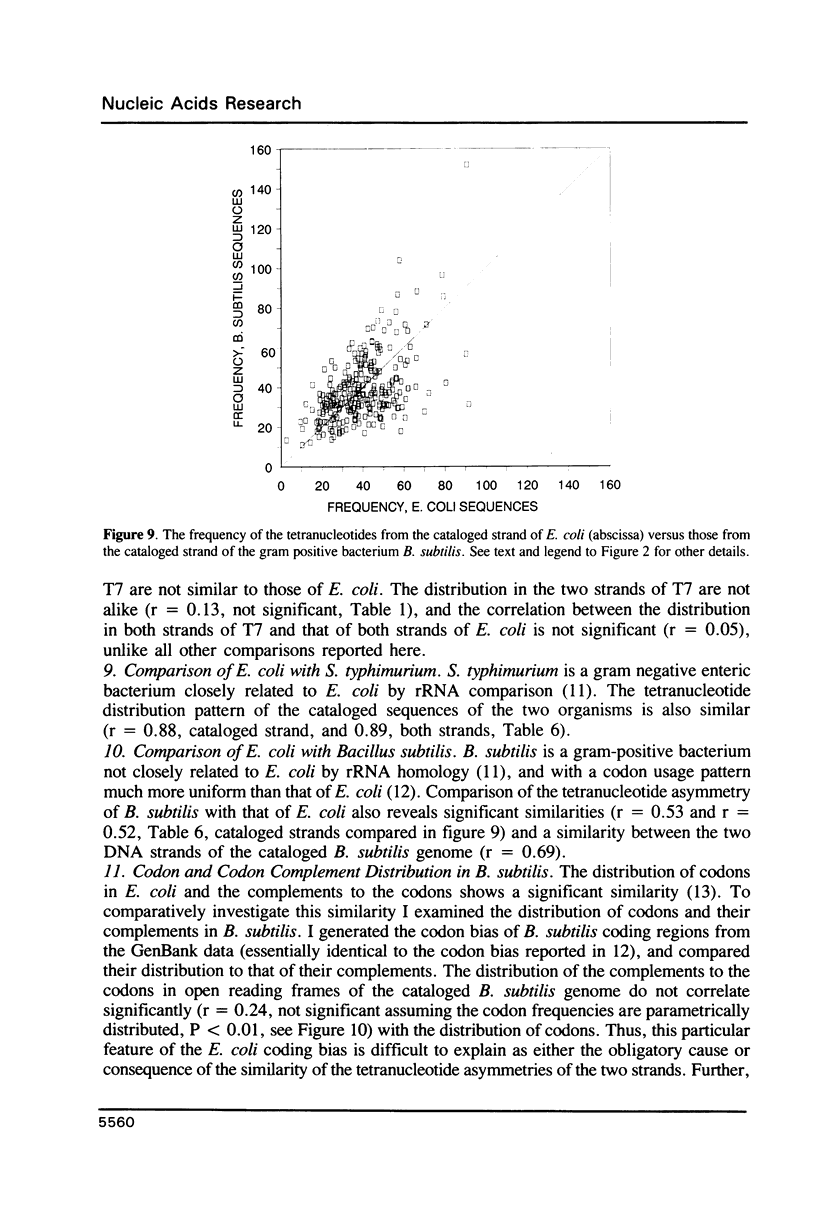
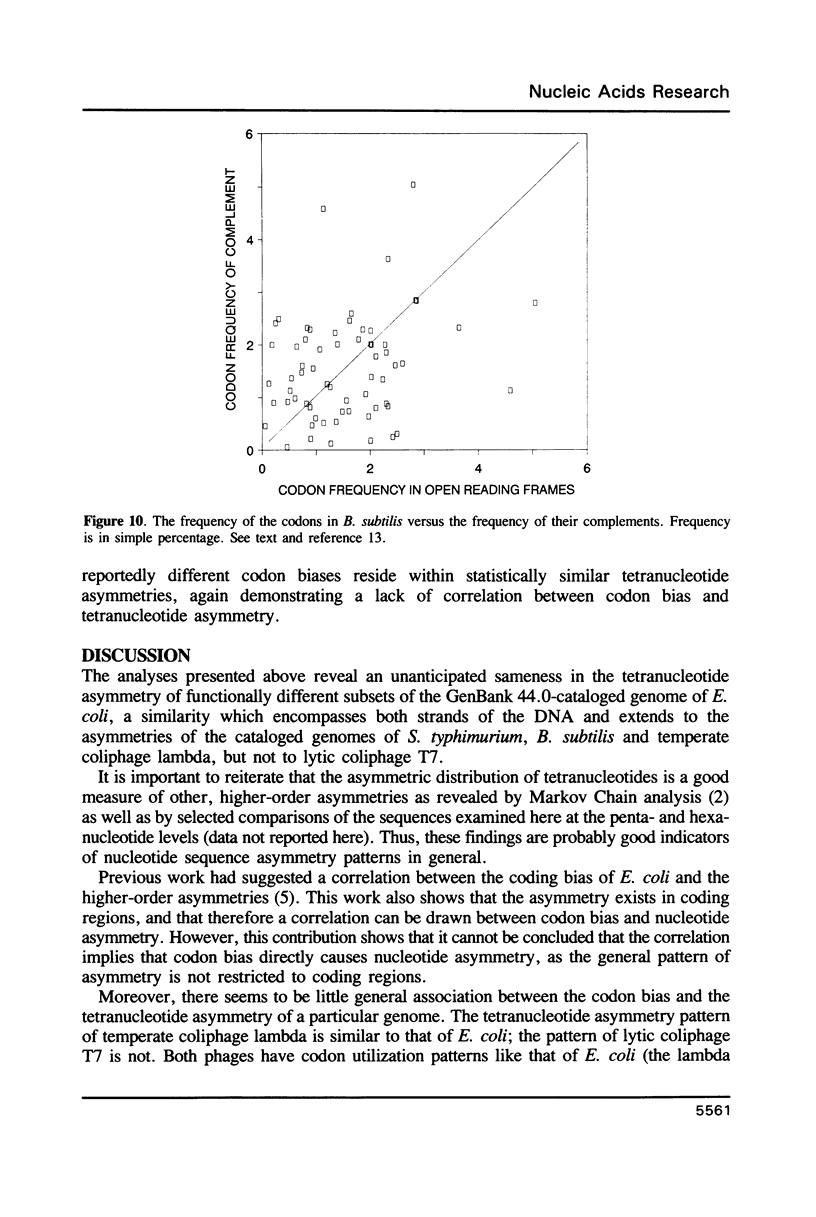
I have examined potential determinants of the asymmetric distribution of nucleotide sequences in the genome of Escherichia coli as cataloged in GenBank release 44. I have used the frequency of occurrence of all possible tetranucleotides in a given sequence catalog or derivative as a comparative measure of asymmetry. The GenBank-cataloged strand and its complement show statistically similar (not complementary) distributions. The distribution is statistically similar in comparisons between the protein coding subset and the total genome, the coding subset and selected non-coding genes, the coding subset and the remainder of the DNA, and the coding subset and stable RNA sequences. I have compared the distribution in the genome of E. coli with the distributions found in the cataloged genomes of Salmonella typhimurium, Bacillus subtilis, and of coliphages lambda and T7. The distribution summed in both strands of the cataloged DNA differs statistically only in comparisons with lytic bacteriophage T7 because only the two strands of T7 show statistically dissimilar distributions. Despite similarities in tetranucleotide distribution, the pattern of codon complementarity in B. subtilis is different than that documented for E. coli. Thus, sequence asymmetry does not seem related to specific DNA function or to documented similarities or differences in codon bias. The sequence asymmetry of the E. coli genome may thus reflect a hitherto unsuspected pattern impressed on both strands of DNA which is or can be packaged into bacterial genomes.
Full text
PDF

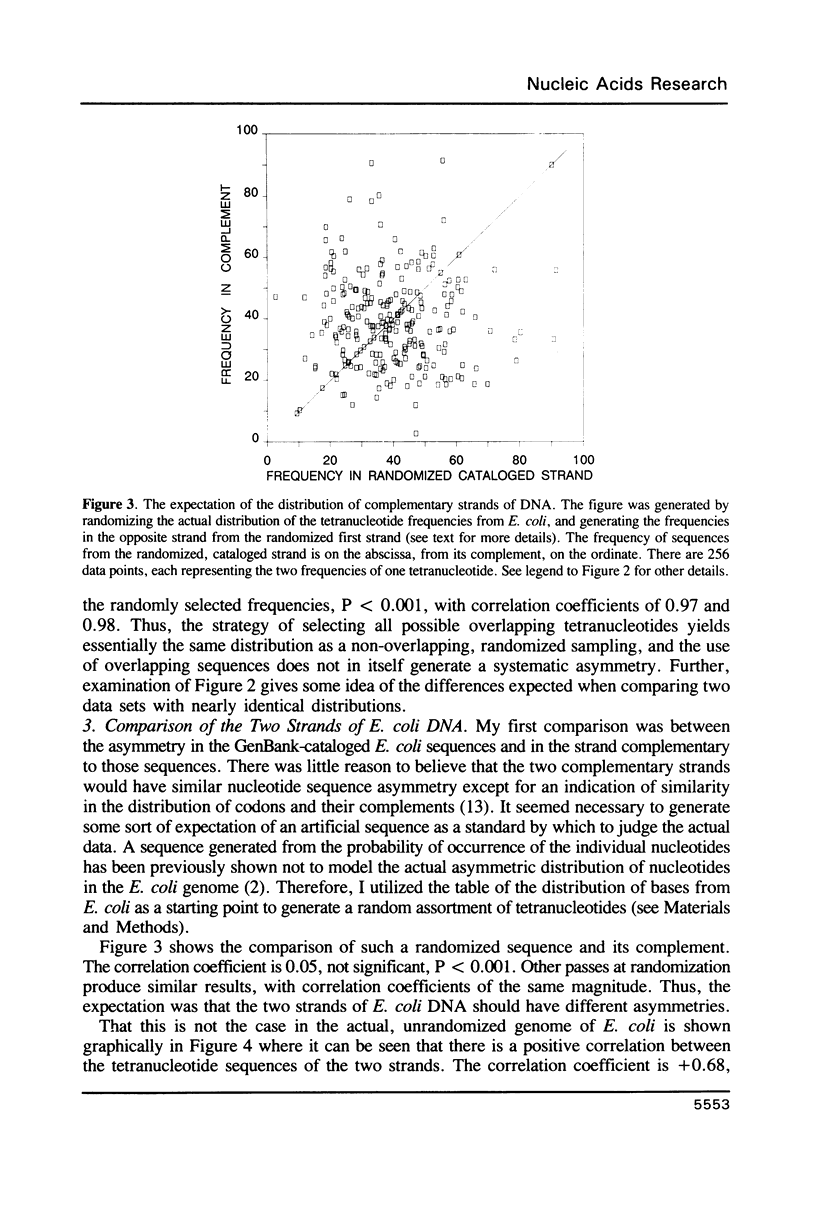
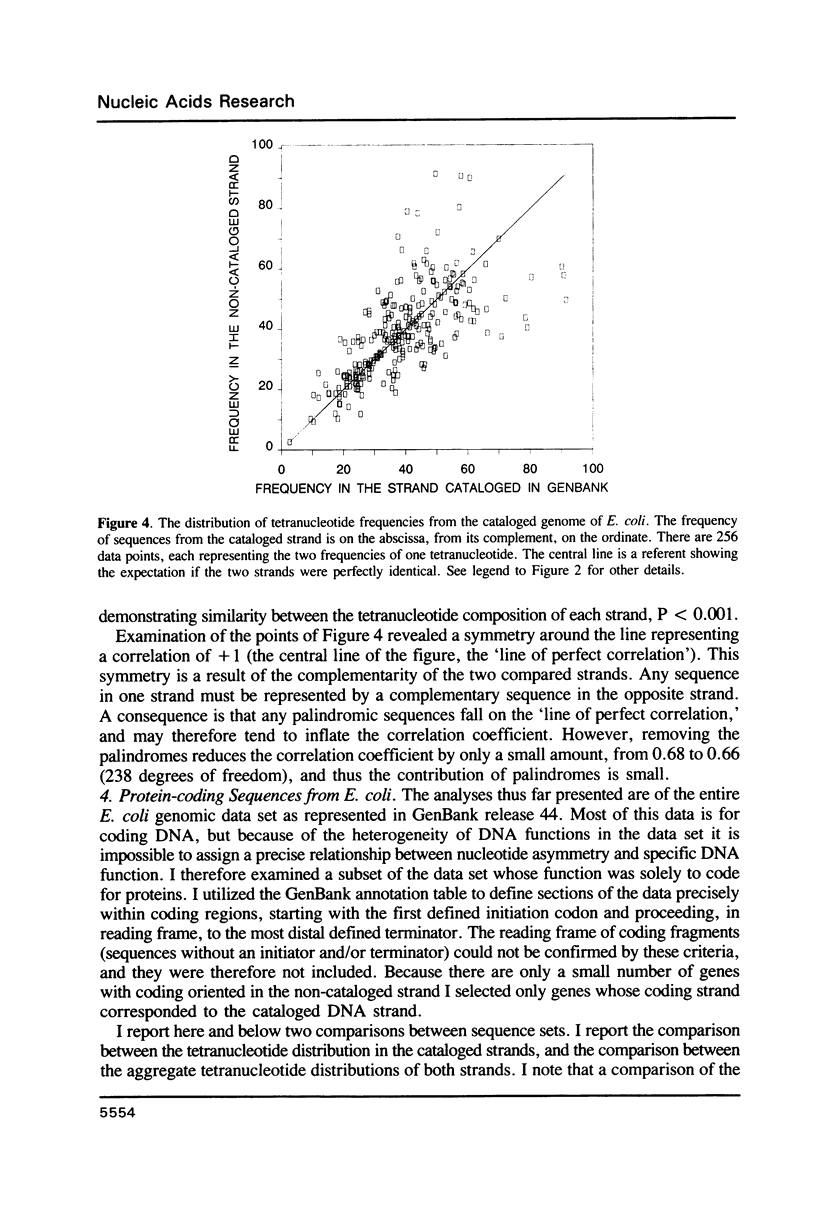
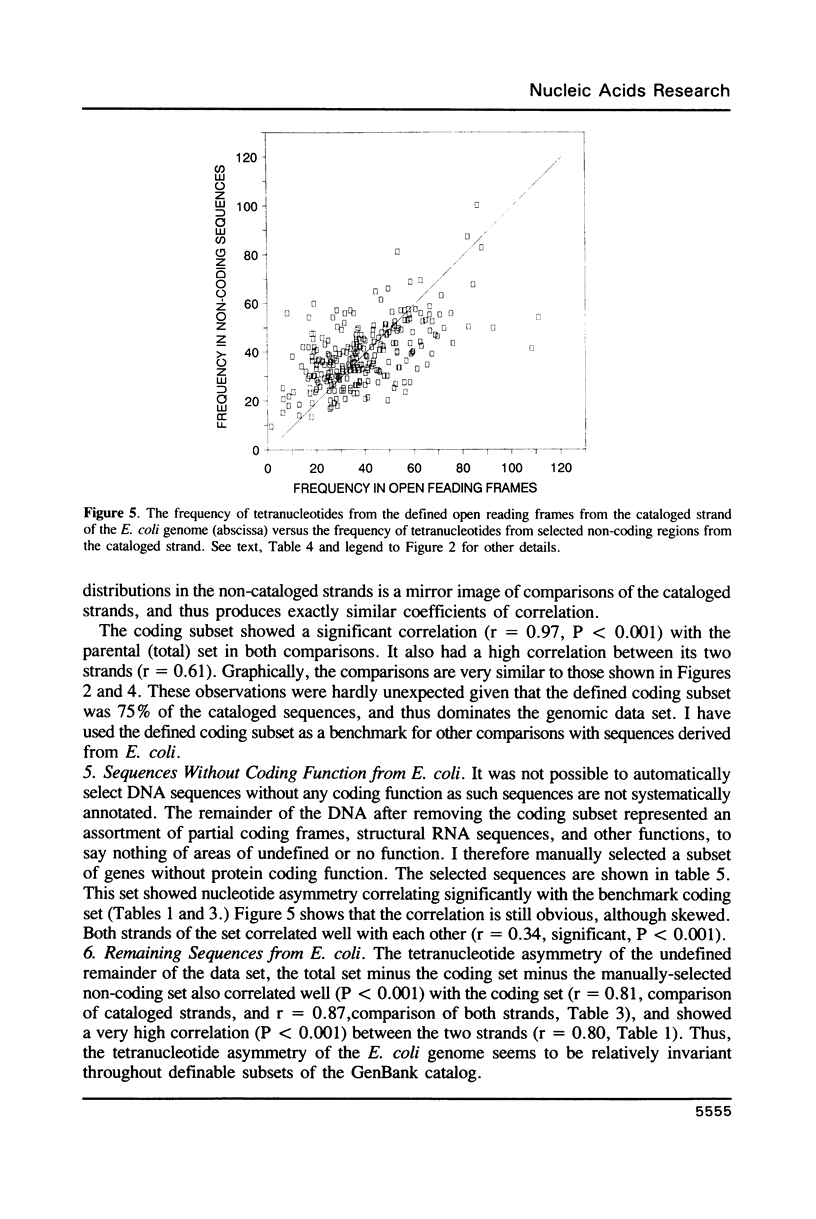



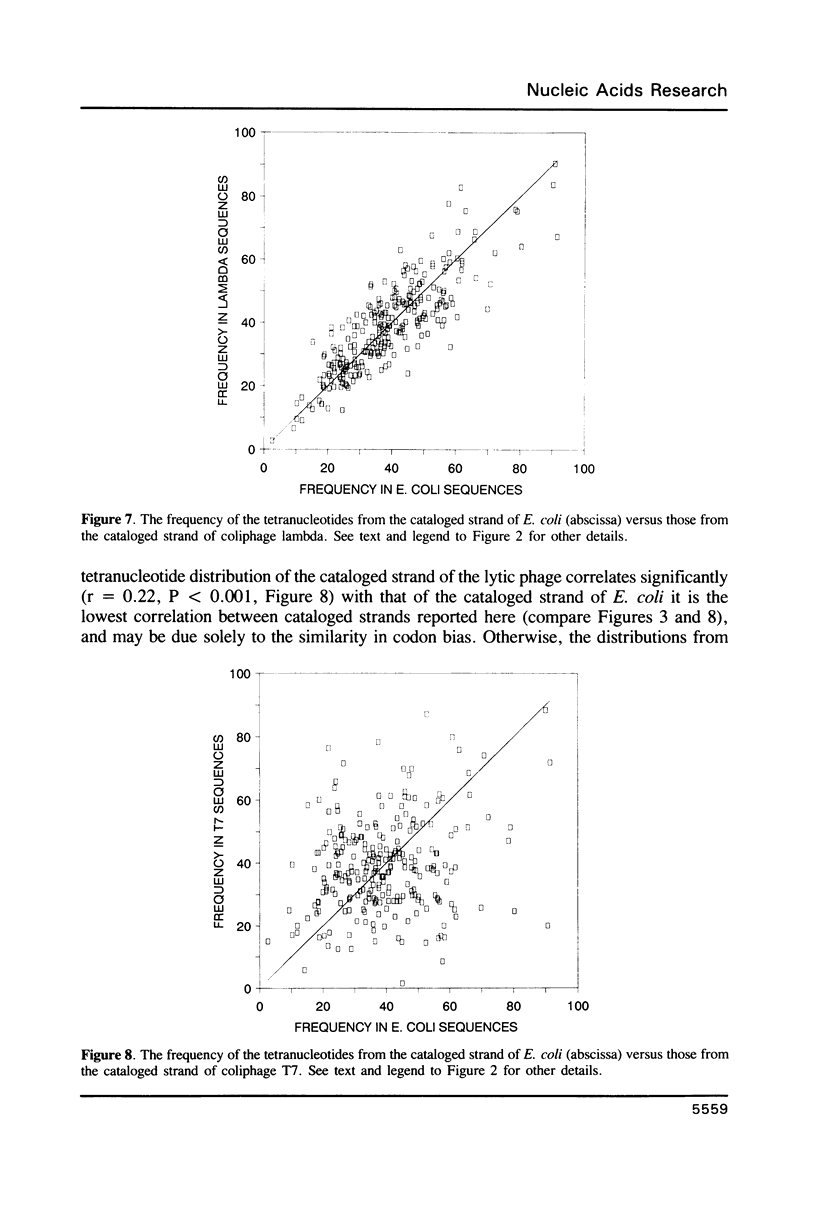


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alff-Steinberger C. Evidence for a coding pattern on the non-coding strand of the E. coli genome. Nucleic Acids Res. 1984 Mar 12;12(5):2235–2241. doi: 10.1093/nar/12.5.2235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernardi G., Bernardi G. Compositional constraints and genome evolution. J Mol Evol. 1986;24(1-2):1–11. doi: 10.1007/BF02099946. [DOI] [PubMed] [Google Scholar]
- Daniels D. L., Sanger F., Coulson A. R. Features of bacteriophage lambda: analysis of the complete nucleotide sequence. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1009–1024. doi: 10.1101/sqb.1983.047.01.115. [DOI] [PubMed] [Google Scholar]
- McClelland M., Jones R., Patel Y., Nelson M. Restriction endonucleases for pulsed field mapping of bacterial genomes. Nucleic Acids Res. 1987 Aug 11;15(15):5985–6005. doi: 10.1093/nar/15.15.5985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogasawara N. Markedly unbiased codon usage in Bacillus subtilis. Gene. 1985;40(1):145–150. doi: 10.1016/0378-1119(85)90035-6. [DOI] [PubMed] [Google Scholar]
- Phillips G. J., Arnold J., Ivarie R. Mono- through hexanucleotide composition of the Escherichia coli genome: a Markov chain analysis. Nucleic Acids Res. 1987 Mar 25;15(6):2611–2626. doi: 10.1093/nar/15.6.2611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips G. J., Arnold J., Ivarie R. The effect of codon usage on the oligonucleotide composition of the E. coli genome and identification of over- and underrepresented sequences by Markov chain analysis. Nucleic Acids Res. 1987 Mar 25;15(6):2627–2638. doi: 10.1093/nar/15.6.2627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Rogers M. S., McConnell D. J. Selection pressures on codon usage in the complete genome of bacteriophage T7. J Mol Evol. 1984;21(2):150–160. doi: 10.1007/BF02100089. [DOI] [PubMed] [Google Scholar]
- Staden R. A computer program to search for tRNA genes. Nucleic Acids Res. 1980 Feb 25;8(4):817–825. [PMC free article] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Gutell R., Gupta R., Noller H. F. Detailed analysis of the higher-order structure of 16S-like ribosomal ribonucleic acids. Microbiol Rev. 1983 Dec;47(4):621–669. doi: 10.1128/mr.47.4.621-669.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yarus M., Folley L. S. Sense codons are found in specific contexts. J Mol Biol. 1985 Apr 20;182(4):529–540. doi: 10.1016/0022-2836(85)90239-6. [DOI] [PubMed] [Google Scholar]


