Abstract
The rat growth hormone (rGH) gene contains two classes of repetitive DNA arranged as clusters within intron B and the 3' flanking region. The major family is equivalent to the CHO type 2 DNA. The second ("truncated repeat", TR) is a truncated version of the first and occurs in certain neural-specific transcripts and genes ("identifier" elements, ID). Here we report, using the HeLa cell-free transcription assay, that RNA polymerase III (Pol III) efficiently initiates at internal promoters within a tandem array of rGH gene repetitive DNA monomers and results in a novel organization of overlapping Class III transcription units. Transcription competition studies revealed that the rat type 2 structures share Pol III transcription factors with a tRNA gene, a human Alu repeat, and a mutant VA1 gene. Also, the rGH type 2 but not the TR DNA efficiently promotes Pol III initiation, yet other TR members, which differ only in flanking DNA, are transcribed. Thus, the rGH gene is strikingly enriched with 10 repetitive DNA monomers; multimeric type 2 elements are actively transcribed; rGH-TR sequences are expressed only as part of larger transcripts promoted by type 2 DNA; and, type 2 DNA uses tRNA gene transcription factors. These studies show that flanking sequences, promoter organization and factor competition may all affect rat repetitive DNA expression.
Full text
PDF



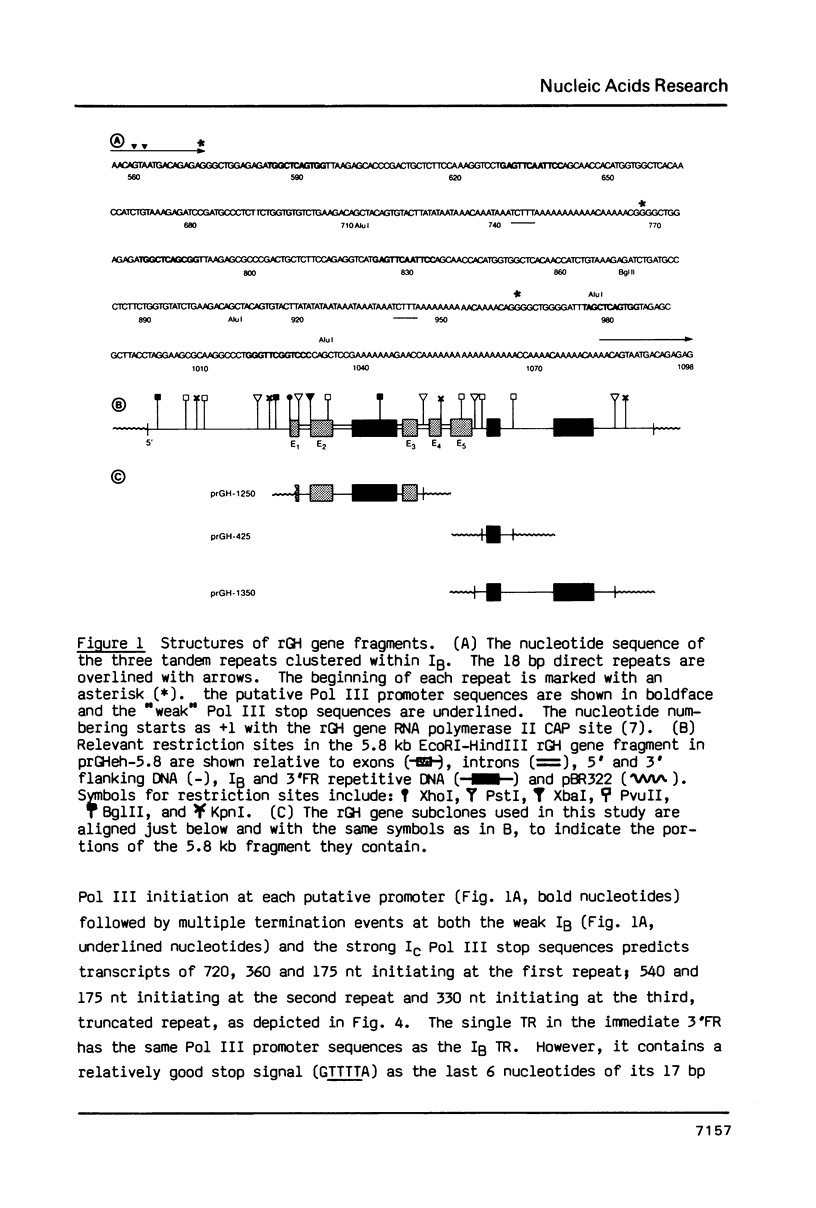
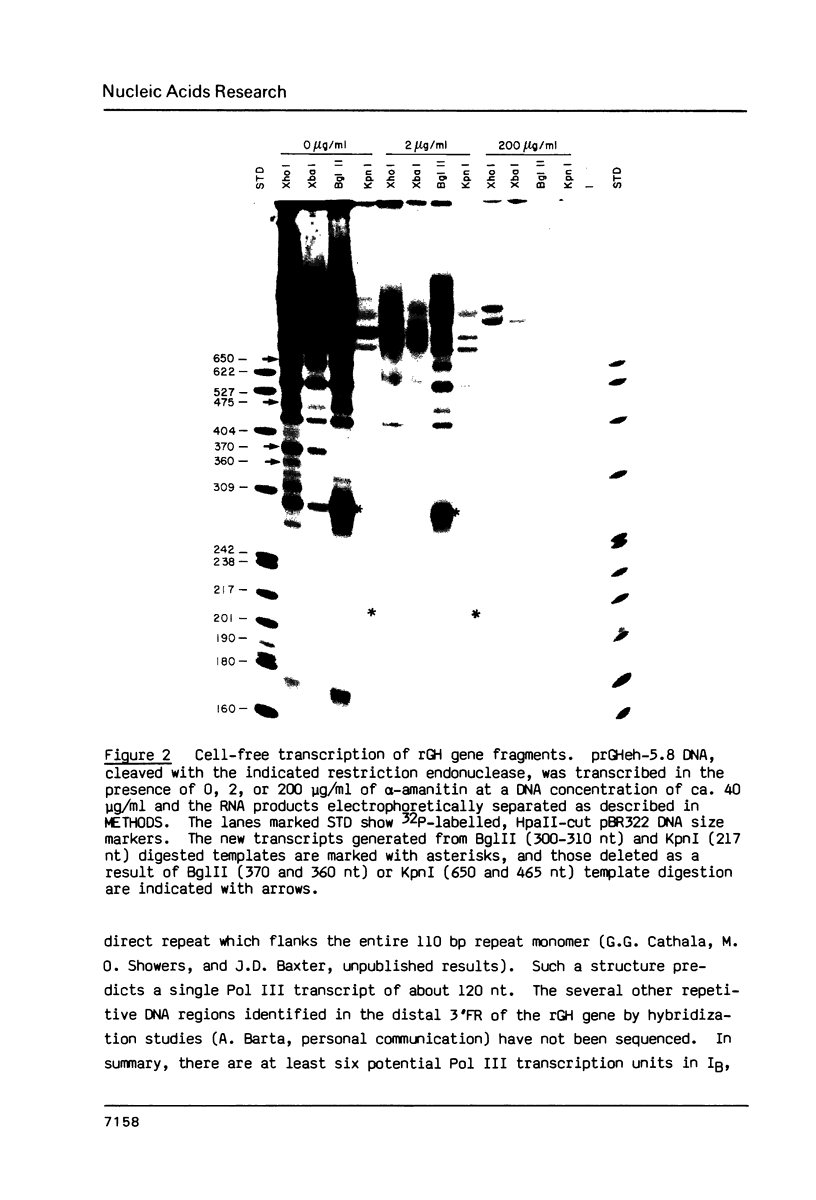

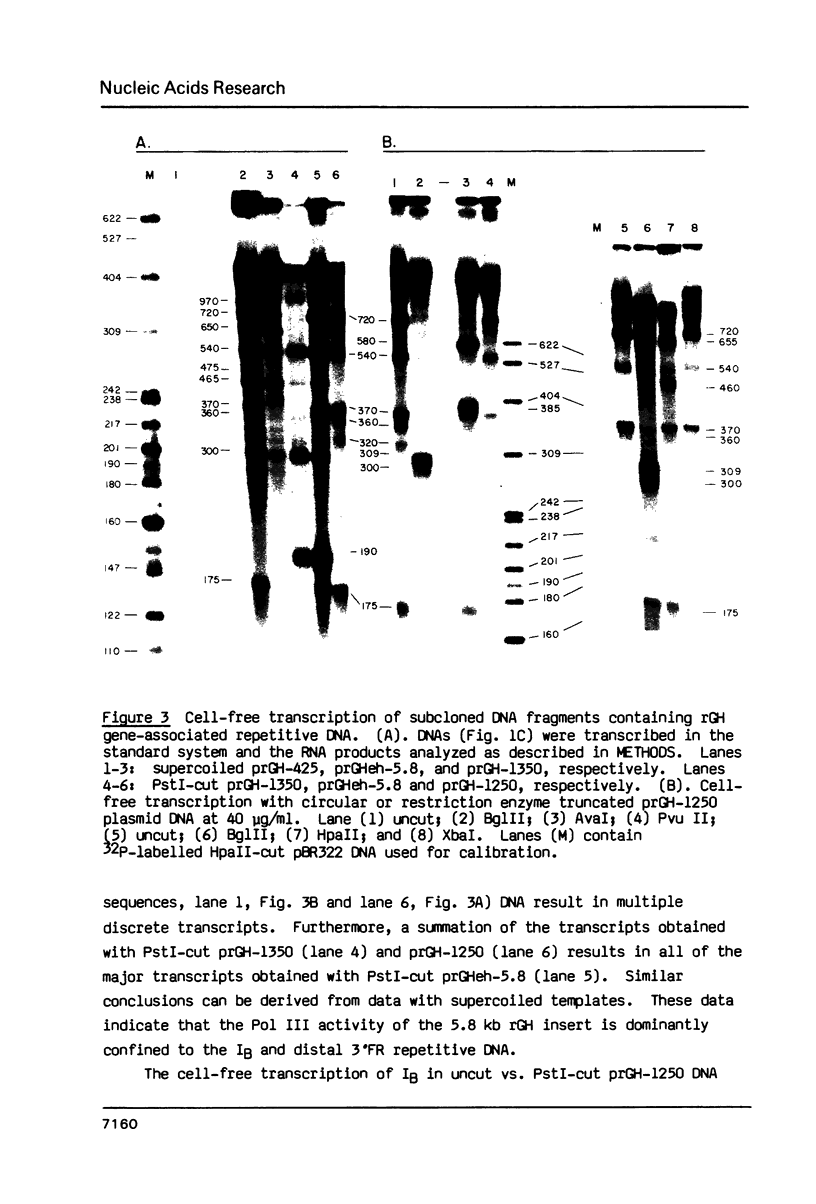

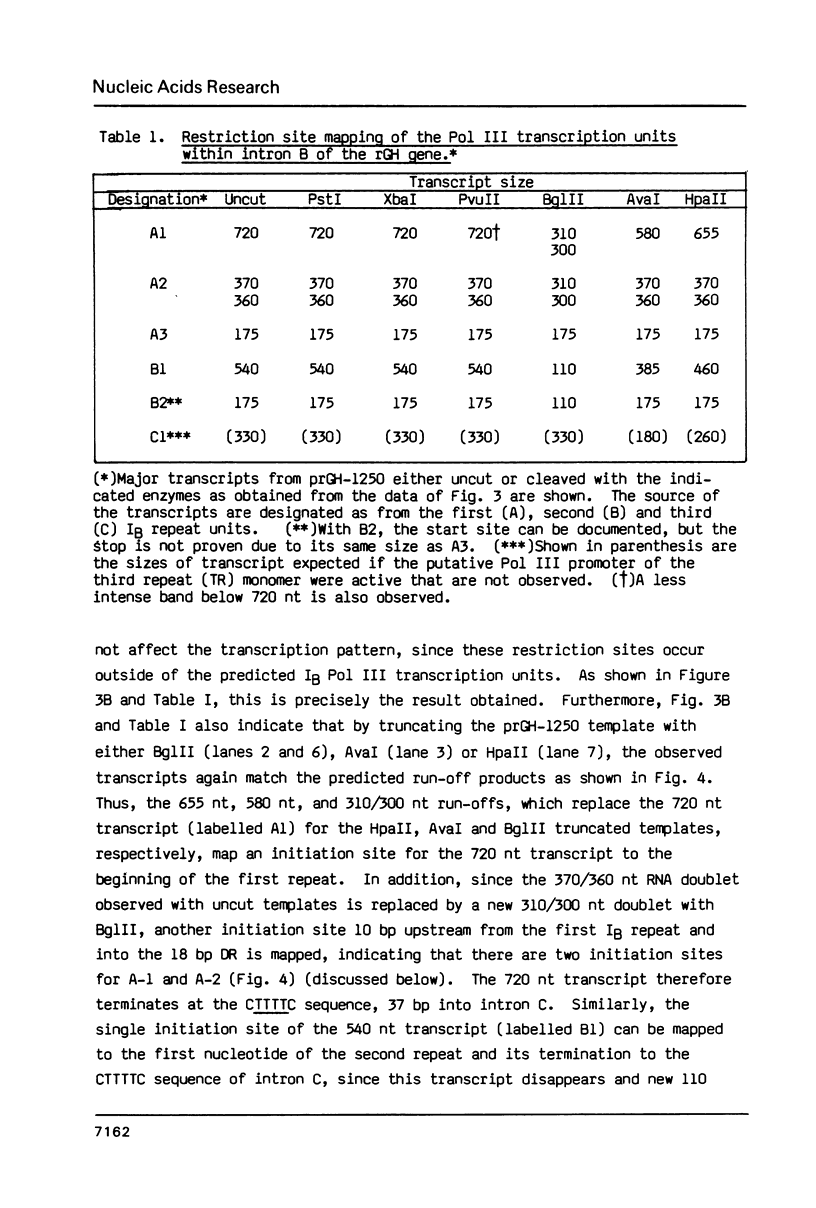




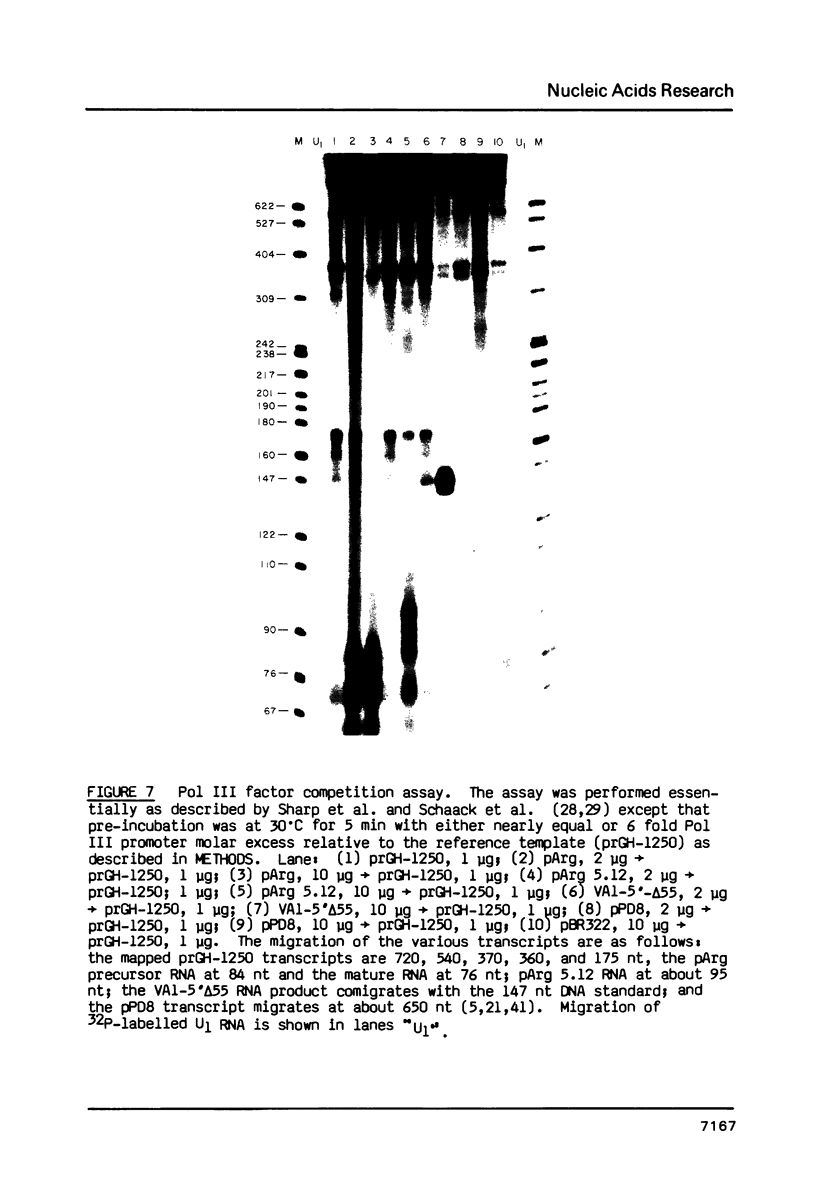






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barta A., Richards R. I., Baxter J. D., Shine J. Primary structure and evolution of rat growth hormone gene. Proc Natl Acad Sci U S A. 1981 Aug;78(8):4867–4871. doi: 10.1073/pnas.78.8.4867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogenhagen D. F., Brown D. D. Nucleotide sequences in Xenopus 5S DNA required for transcription termination. Cell. 1981 Apr;24(1):261–270. doi: 10.1016/0092-8674(81)90522-5. [DOI] [PubMed] [Google Scholar]
- Ciliberto G., Traboni C., Cortese R. Transcription of multimeric tRNA genes. Nucleic Acids Res. 1984 Jan 25;12(2):1277–1285. doi: 10.1093/nar/12.2.1277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B., Helinski D. R. Supercoiled circular DNA-protein complex in Escherichia coli: purification and induced conversion to an opern circular DNA form. Proc Natl Acad Sci U S A. 1969 Apr;62(4):1159–1166. doi: 10.1073/pnas.62.4.1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooke N. E., Baxter J. D. Structural analysis of the prolactin gene suggests a separate origin for its 5' end. Nature. 1982 Jun 17;297(5867):603–606. doi: 10.1038/297603a0. [DOI] [PubMed] [Google Scholar]
- DeFranco D., Sharp S., Söll D. Identification of regulatory sequences contained in the 5'-flanking region of Drosophila lysine tRNA2 genes. J Biol Chem. 1981 Dec 10;256(23):12424–12429. [PubMed] [Google Scholar]
- Dhar R., Ellis R. W., Shih T. Y., Oroszlan S., Shapiro B., Maizel J., Lowy D., Scolnick E. Nucleotide sequence of the p21 transforming protein of Harvey murine sarcoma virus. Science. 1982 Sep 3;217(4563):934–936. doi: 10.1126/science.6287572. [DOI] [PubMed] [Google Scholar]
- Dingermann T., Burke D. J., Sharp S., Schaack J., Söll D. The 5- flanking sequences of Drosophila tRNAArg genes control their in vitro transcription in a Drosophila cell extract. J Biol Chem. 1982 Dec 25;257(24):14738–14744. [PubMed] [Google Scholar]
- Duncan C. H., Jagadeeswaran P., Wang R. R., Weissman S. M. Structural analysis of templates and RNA polymerase III transcripts of Alu family sequences interspersed among the human beta-like globin genes. Gene. 1981 Mar;13(2):185–196. doi: 10.1016/0378-1119(81)90007-x. [DOI] [PubMed] [Google Scholar]
- England T. E., Bruce A. G., Uhlenbeck O. C. Specific labeling of 3' termini of RNA with T4 RNA ligase. Methods Enzymol. 1980;65(1):65–74. doi: 10.1016/s0076-6879(80)65011-3. [DOI] [PubMed] [Google Scholar]
- Folk W. R., Hofstetter H. A detailed mutational analysis of the eucaryotic tRNAmet1 gene promoter. Cell. 1983 Jun;33(2):585–593. doi: 10.1016/0092-8674(83)90439-7. [DOI] [PubMed] [Google Scholar]
- Fowlkes D. M., Shenk T. Transcriptional control regions of the adenovirus VAI RNA gene. Cell. 1980 Nov;22(2 Pt 2):405–413. doi: 10.1016/0092-8674(80)90351-7. [DOI] [PubMed] [Google Scholar]
- Fox J. L. Gene splicers contemplate the rat brain. Science. 1983 Nov 18;222(4625):828–829. doi: 10.1126/science.6138857. [DOI] [PubMed] [Google Scholar]
- Fuhrman S. A., Deininger P. L., LaPorte P., Friedmann T., Geiduschek E. P. Analysis of transcription of the human Alu family ubiquitous repeating element by eukaryotic RNA polymerase III. Nucleic Acids Res. 1981 Dec 11;9(23):6439–6456. doi: 10.1093/nar/9.23.6439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruissem W., Prescott D. M., Greenberg B. M., Hallick R. B. Transcription of E. coli and Euglena chloroplast tRNA gene clusters and processing of polycistronic transcripts in a HeLa cell-free system. Cell. 1982 Aug;30(1):81–92. doi: 10.1016/0092-8674(82)90014-9. [DOI] [PubMed] [Google Scholar]
- Guilfoyle R., Weinmann R. Control region for adenovirus VA RNA transcription. Proc Natl Acad Sci U S A. 1981 Jun;78(6):3378–3382. doi: 10.1073/pnas.78.6.3378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall B. D., Clarkson S. G., Tocchini-Valentini G. Transcription initiation of eucaryotic transfer RNA genes. Cell. 1982 May;29(1):3–5. doi: 10.1016/0092-8674(82)90083-6. [DOI] [PubMed] [Google Scholar]
- Haynes S. R., Jelinek W. R. Low molecular weight RNAs transcribed in vitro by RNA polymerase III from Alu-type dispersed repeats in Chinese hamster DNA are also found in vivo. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6130–6134. doi: 10.1073/pnas.78.10.6130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ish-Horowicz D., Burke J. F. Rapid and efficient cosmid cloning. Nucleic Acids Res. 1981 Jul 10;9(13):2989–2998. doi: 10.1093/nar/9.13.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jat P., Arrand J. R. In vitro transcription of two Epstein-Barr virus specified small RNA molecules. Nucleic Acids Res. 1982 Jun 11;10(11):3407–3425. doi: 10.1093/nar/10.11.3407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jelinek W. R., Schmid C. W. Repetitive sequences in eukaryotic DNA and their expression. Annu Rev Biochem. 1982;51:813–844. doi: 10.1146/annurev.bi.51.070182.004121. [DOI] [PubMed] [Google Scholar]
- Klemenz R., Stillman D. J., Geiduschek E. P. Specific interactions of Saccharomyces cerevisiae proteins with a promoter region of eukaryotic tRNA genes. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6191–6195. doi: 10.1073/pnas.79.20.6191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koski R. A., Clarkson S. G., Kurjan J., Hall B. D., Smith M. Mutations of the yeast SUP4 tRNATyr locus: transcription of the mutant genes in vitro. Cell. 1980 Nov;22(2 Pt 2):415–425. doi: 10.1016/0092-8674(80)90352-9. [DOI] [PubMed] [Google Scholar]
- Lassar A. B., Martin P. L., Roeder R. G. Transcription of class III genes: formation of preinitiation complexes. Science. 1983 Nov 18;222(4625):740–748. doi: 10.1126/science.6356356. [DOI] [PubMed] [Google Scholar]
- Lemischka I., Sharp P. A. The sequences of an expressed rat alpha-tubulin gene and a pseudogene with an inserted repetitive element. Nature. 1982 Nov 25;300(5890):330–335. doi: 10.1038/300330a0. [DOI] [PubMed] [Google Scholar]
- Manley J. L., Fire A., Cano A., Sharp P. A., Gefter M. L. DNA-dependent transcription of adenovirus genes in a soluble whole-cell extract. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3855–3859. doi: 10.1073/pnas.77.7.3855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao J., Schmidt O., Söll D. Dimeric transfer RNA precursors in S. pombe. Cell. 1980 Sep;21(2):509–516. doi: 10.1016/0092-8674(80)90488-2. [DOI] [PubMed] [Google Scholar]
- Maxwell I. H., Maxwell F., Hahn W. E. Removal of RNase activity from DNase by affinity chromatography on agarose coupled aminophenylphosphoryl-uridine-2' (3')-phosphate. Nucleic Acids Res. 1977 Jan;4(1):241–246. doi: 10.1093/nar/4.1.241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milner R. J., Bloom F. E., Lai C., Lerner R. A., Sutcliffe J. G. Brain-specific genes have identifier sequences in their introns. Proc Natl Acad Sci U S A. 1984 Feb;81(3):713–717. doi: 10.1073/pnas.81.3.713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murphy M. H., Baralle F. E. Directed semisynthetic point mutational analysis of an RNA polymerase III promoter. Nucleic Acids Res. 1983 Nov 25;11(22):7695–7700. doi: 10.1093/nar/11.22.7695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newman A. J., Ogden R. C., Abelson J. tRNA gene transcription in yeast: effects of specified base substitutions in the intragenic promoter. Cell. 1983 Nov;35(1):117–125. doi: 10.1016/0092-8674(83)90214-3. [DOI] [PubMed] [Google Scholar]
- Page G. S., Smith S., Goodman H. M. DNA sequence of the rat growth hormone gene: location of the 5' terminus of the growth hormone mRNA and identification of an internal transposon-like element. Nucleic Acids Res. 1981 May 11;9(9):2087–2104. doi: 10.1093/nar/9.9.2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raymond G. J., Johnson J. D. The role of non-coding DNA sequences in transcription and processing of a yeast tRNA. Nucleic Acids Res. 1983 Sep 10;11(17):5969–5988. doi: 10.1093/nar/11.17.5969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaack J., Sharp S., Dingermann T., Söll D. Transcription of eukaryotic tRNA genes in vitro. II. Formation of stable complexes. J Biol Chem. 1983 Feb 25;258(4):2447–2453. [PubMed] [Google Scholar]
- Schmidt O., Mao J., Ogden R., Beckmann J., Sakano H., Abelson J., Söll D. Dimeric tRNA precursors in yeast. Nature. 1980 Oct 23;287(5784):750–752. doi: 10.1038/287750a0. [DOI] [PubMed] [Google Scholar]
- Schuler L. A., Weber J. L., Gorski J. Polymorphism near the rat prolactin gene caused by insertion of an Alu-like element. Nature. 1983 Sep 8;305(5930):159–160. doi: 10.1038/305159a0. [DOI] [PubMed] [Google Scholar]
- Sharp P. A. Conversion of RNA to DNA in mammals: Alu-like elements and pseudogenes. Nature. 1983 Feb 10;301(5900):471–472. doi: 10.1038/301471a0. [DOI] [PubMed] [Google Scholar]
- Sharp S., DeFranco D., Dingermann T., Farrell P., Söll D. Internal control regions for transcription of eukaryotic tRNA genes. Proc Natl Acad Sci U S A. 1981 Nov;78(11):6657–6661. doi: 10.1073/pnas.78.11.6657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp S., Dingermann T., Schaack J., DeFranco D., Söll D. Transcription of eukaryotic tRNA genes in vitro. I. Analysis of control regions using a competition assay. J Biol Chem. 1983 Feb 25;258(4):2440–2446. [PubMed] [Google Scholar]
- Sutcliffe J. G., Milner R. J., Bloom F. E., Lerner R. A. Common 82-nucleotide sequence unique to brain RNA. Proc Natl Acad Sci U S A. 1982 Aug;79(16):4942–4946. doi: 10.1073/pnas.79.16.4942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sutcliffe J. G., Milner R. J., Gottesfeld J. M., Lerner R. A. Identifier sequences are transcribed specifically in brain. Nature. 1984 Mar 15;308(5956):237–241. doi: 10.1038/308237a0. [DOI] [PubMed] [Google Scholar]
- Traboni C., Ciliberto G., Cortese R. Mutations in Box B of the promoter of a eucaryotic tRNAPro gene affect rate of transcription, processing, and stability of the transcripts. Cell. 1984 Jan;36(1):179–187. doi: 10.1016/0092-8674(84)90087-4. [DOI] [PubMed] [Google Scholar]
- Watanabe-Nagasu N., Itoh Y., Tani T., Okano K., Koga N., Okada N., Ohshima Y. Structural analysis of gene loci for rat U1 small nuclear RNA. Nucleic Acids Res. 1983 Mar 25;11(6):1791–1801. doi: 10.1093/nar/11.6.1791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zassenhaus H. P., Farrelly F., Hudspeth M. E., Grossman L. I., Butow R. A. Transcriptional analysis of the Saccharomyces cerevisiae mitochondrial var1 gene: anomalous hybridization of RNA from AT-rich regions. Mol Cell Biol. 1983 Sep;3(9):1615–1624. doi: 10.1128/mcb.3.9.1615. [DOI] [PMC free article] [PubMed] [Google Scholar]