Abstract
A spontaneous point mutation in pilQ (pilQ1) resulted in phenotypic suppression of a hemoglobin (Hb) receptor mutant (hpuAB mutant), allowing gonococci to grow on Hb as the sole source of iron. PilQ, formerly designated OMP-MC, is a member of the secretin family of proteins located in the outer membrane and is required for pilus biogenesis. The pilQ1 mutant also showed decreased piliation and transformation efficiency. Insertional inactivation of pilQ1 resulted in the loss of the Hb utilization phenotype and decreased entry of free heme. Despite the ability of the pilQ1 mutant to use Hb for iron acquisition and porphyrin, there was no demonstrable binding of Hb to the cell surface. The pilQ1 mutant was more sensitive to the toxic effect of free heme in growth medium and hypersensitive to the detergent Triton X-100 and multiple antibiotics. Double mutation in pilQ1 and tonB had no effect on these phenotypes, but a double pilQ1 pilT mutant showed a reduction in Hb-dependent growth and decreased sensitivity to heme and various antimicrobial agents. Insertional inactivation of wild-type pilQ also resulted in reduced entry of heme, Triton X-100, and some antibiotics. These results show that PilQ forms a channel that allows entry of heme and certain antimicrobial compounds and that a gain-of function point mutation in pilQ results in TonB-independent, PilT-dependent increase of entry.
Mammalian hosts use iron-binding proteins and iron-sequestering compounds to maintain free iron at a level that is too low for the growth of invading pathogens. Many pathogenic bacteria have evolved mechanisms for scavenging essential iron in vivo from transferrin, lactoferrin, hemoglobin (Hb), and other sources. The ability to scavenge iron is important to bacterial growth in vivo and therefore virulence (49). Many bacteria acquire iron by secretion of iron siderophores, but pathogenic neisseriae do not produce siderophores, relying instead on specific receptors for binding host iron compounds, including transferrin, lactoferrin, and Hb (12, 35). Either the transferrin or the lactoferrin receptor is essential for gonococci to cause urethral infection in male volunteers (1, 11). Moreover, the presence of both the transferrin and lactoferrin receptors provides a selective advantage during experimental male urethral infection (1, 11). The phase-variable gonococcal HpuAB Hb receptor apparently is not involved in infection of the male urethra but is selected in vivo in women during the first half of the menstrual cycle (2) and therefore also is important to infection.
Mechanisms of iron uptake from Hb by the pathogenic neisseriae are well studied (5, 6, 7, 21, 22, 23, 24, 41, 42), but many details remain unclear. Meningococci express two Hb receptors: HmbR for binding Hb or HpuAB for binding either Hb alone or Hb bound to its serum binding protein haptoglobin (22). Gonococci express only the HpuAB receptor, although they do possess an hmbR pseudogene (42). HpuA is a lipoprotein, and HpuB is an integral outer membrane protein. Both HpuA and -B are essential for use of Hb as an iron source (5). Construction of hemA mutants that cannot synthesize heme proved that gonococci which express HpuAB can utilize Hb-bound heme as a source of porphyrins, and thus heme enters the cytoplasm intact (48). The function of the HpuAB receptor is dependent on activation by TonB (3). HpuA facilitates binding of Hb to the receptor and protects free heme released on the outer surface of the bacterium from binding by human serum albumin (HSA) (6). Heme released from Hb bound to the HpuAB receptor apparently traverses a pore within HpuB in a TonB-dependent manner (6) and then crosses the periplasmic space and cytoplasmic membrane by unknown mechanisms.
Gonococci and meningococci also can utilize free heme for growth in the absence of an Hb receptor (16, 44). Gonococcal growth with free heme does not require TonB (3, 7, 44, 48) and, although the existence of a heme receptor has been suggested (19, 20), one has never been clearly identified in either meningococci or gonococci. At present, pathways for entry and transport of free heme in the absence of an Hb receptor are not well understood.
Recently, we utilized an hpuA deletion mutant of gonococcal strain FA1090 to isolate two classes of Hb-utilizing (Hb+) mutants. One class required TonB and had point mutations in hpuB that restored entry of heme from Hb in the absence of HpuA, whereas the other required neither TonB nor HpuAB (6). The HpuAB- and TonB-independent Hb+ mutants were designated hgbX, in recognition of the lack of understanding of the molecular basis for the Hb-utilizing phenotype. The hgbX mutants exhibited increased sensitivity to certain antimicrobial agents and grew normally on both Hb and low concentrations of free heme.
We report here that two of the mutants formerly designated hgbX are due to a specific point mutation in the pilQ gene (pilQ1) that allows gonococci to utilize Hb as a source of essential heme iron and porphyrins in the absence of an Hb receptor. Formerly designated the major subunit of outer membrane protein-macromolecular complex (OMP-MC) (18, 47), PilQ is a member of the secretin family of proteins (34), which functions in type IV pilus organelle biogenesis in Neisseria gonorrhoeae (13, 52). The type IV pilus is required for twitching motility and DNA transformation competence (30, 50), and the pilQ1 point mutation allows for both pilus biogenesis and DNA transformation but at slightly reduced levels. We demonstrate that the pilQ1 point mutation that results in use of Hb as a heme and iron source in the absence of an Hb receptor also results in elevated sensitivities to various antimicrobial compounds. The phenotype is dependent on the expression of the mutant form of PilQ and in part on expression of PilT. Thus, this mutant PilQ allows for increased entry of exogenous compounds into the bacterial cell in addition to its role in pilus fiber extension.
MATERIALS AND METHODS
Bacterial strains and growth conditions.
The parent strain used in this study was FA1090. Plasmids and gonococcal strains constructed and/or utilized in this study are listed in Table 1. The growth conditions of gonococcal and Escherichia coli strains were as described previously (43). Strains unable to utilize Hb as the sole source of iron (Hb−) were grown on Bacto GC medium base (Difco/Becton Dickinson, Sparks, Md.) plates (GCB plates) containing Kellogg's Supplement I plus 12 μM ferric nitrate. Strains that were able to grow on Hb (Hb+) were grown on GCB plates or modified GCB plates (Hb-Des plates) containing human Hb at a concentration of 2 μM and desferoxamine mesylate (Desferal) at 100 μM instead of ferric nitrate.
TABLE 1.
Plasmids and gonococcal strains constructed and/or used in this study
| Plasmid or strain | Genotype and phenotype | Source or reference |
|---|---|---|
| Plasmids | ||
| pHP45Ω | Source for Ω (Spr Smr) cassette | 32 |
| pNC40 | Source for cat (Cmr) cassette | 46 |
| pUNCH173 | tonB::Ω | 3 |
| pUNCH1306 | hemA::Ω | 48 |
| pUNCH288 | Partial FA7168 pilQ1 in pCR2.1-TOPO | This study |
| pUNCH290 | pilQ1::Ω of p288 | This study |
| pUNCH291 | pilQ1::cat of p288 | This study |
| Strains | ||
| FA1090 | Parent strain; wild type | 15 |
| FA1090 H+ | HpuAB phase on variant of FA1090 | 7 |
| FA7167 | ΔhpuA hpuB::cat; Cmr Hb− | 6 |
| FA7168 | Cms Hb+ revertant of FA7167; pilQ1 | 6 |
| FA7169 | ΔhpuA hpuB+ derivative of FA7167; Cms Hb− | 6 |
| FA7186 | hmbR::Ω transformant of FA7169; Spr Hb− | 6 |
| FA7186H1 | hgbX2; Spr Hb+ revertant of FA7186 | 6 |
| FA7186H12 | hgbX3; Spr Hb+ revertant of FA7186 | This study |
| FA7186H15 | hgbX4; Spr Hb+ revertant of FA7186 | This study |
| FA7186H19 | Spr Hb+ revertant of FA7186; pilQ1 | This study |
| FA7186H20 | hgbX6; Spr Hb+ revertant of FA7186 | This study |
| FA7294 | FA7186H19 pilQ1 transformant of FA7167; Cms Sps Hb+ | This study |
| FA7311 | pilQ::Ω transformant of FA1090 H+; Spr Hb+ | This study |
| FA7312 | pilQ1::Ω transformant of FA7294; Spr Hb− | This study |
| FA7317 | FA7168 pilQ1 transformant of FA7167; Cms Sps Hb+ | This study |
| FA7318 | FA7186H19 pilQ1 transformant of FA7167; Cms Sps Hb+ | This study |
| FA7319 | pilQ::Ω transformant of FA7167; Spr Hb− | This study |
| FA7320 | pilQ1::Ω transformant of FA7317; Spr Hb− | This study |
| FA7321 | tonB::Ω transformant of FA7317; Spr Hb+ | This study |
| FA7322 | hemA::Ω transformant of FA7317; Spr Hb+ | This study |
| RM11.2recA6 | Defined P+ variant of FA1090 with IPTG-regulated recA | 25 |
| FA7332 (RM7167) | ΔhpuA hpuB::cat transformant of RM11.2recA6; Hb− | This study |
| FA7333 (RM7168) | FA7168 pilQ1 transformant of FA7332; Hb+ | This study |
| FA7334 | pilT::erm transformant of RM11.2recA6 | 26 |
| FA7335 | pilQ1 pilT::erm transformant of RM11.2recA6 | This study |
| FA7336 | ΔpilE of RM11.2recA6 | This study |
| FA7337 | ΔpilE transformant of FA7332 | This study |
| FA7338 | ΔpilE transformant of FA7333 | This study |
| FA7339 | pilQ1 transformant of RM11.2recA6 | This study |
| FA7340 | pilT::erm transformant of FA7333 | This study |
| FA7341 | pilT::erm transformant of FA1090 | This study |
| FA7342 | pilT::erm transformant t of FA7168 | This study |
| FA7343 | pilT::erm transformant of FA7186H19 | This study |
| FA7344 | pilT::erm transformant of FA7317 | This study |
| FA7345 | pilT::erm transformant of FA7318 | This study |
| FA7348 | pilT::tet transformant of FA1090 | This study |
| FA7349 | pilT::tet transformant of FA7167 | This study |
| FA7350 | pilT::tet transformant of FA7186H19 | This study |
Spr, SPT resistance; Smr, streptomycin (100 μg/ml) resistance; Cmr, CHL resistance.
Antibiotics in growth media were used at the indicated concentrations unless otherwise noted: for E. coli strains, ampicillin (AMP) at 100 μg/ml, erythromycin (ERY) at 100 μg/ml, chloramphenicol (CHL) at 30 μg/ml, kanamycin (KAN) at 30 μg/ml; for gonococcal strains: erythromycin at 1 μg/ml (ERY1), chloramphenicol (CHL) at 1 μg/ml (CHL1), spectinomycin (SPT) at 100 μg/ml (SPT100), and tetracycline (TET) at 2 μg/ml (TET2). A stock solution of heme was prepared by dissolving 10 mg of bovine hemin chloride in 1 ml of 0.1 N NaOH. A stock solution of Hb was prepared by dissolving 100 mg of human Hb in 10 ml of 10 mM HEPES (pH 7.4). The solution was rocked overnight at 4°C and then filtered through a 0.45-μm-pore size Acrodisc (Pall Gelman Laboratory, Ann Arbor, Mich.).
Reagents, oligonucleotides, and DNA sequencing.
All chemicals were purchased from Sigma (St. Louis, Mo.) unless otherwise indicated. Oligonucleotides were synthesized at the Lineberger Comprehensive Cancer Center DNA Synthesis Facility of the University of North Carolina-Chapel Hill. Sequencing was carried out at the Automated DNA Sequencing Facility of the University of North Carolina-Chapel Hill with an Applied Biosystems model 373 DNA sequencer by use of the Taq dye terminator cycle sequencing kit (Applied Biosystems/Perkin-Elmer, Foster City, Calif.) or in the Seifert laboratory on a Beckman Coulter CEQ L system. All oligonucleotides used as primers for PCR or DNA sequencing of pilQ are listed in Table 2.
TABLE 2.
Oligonucleotides used in genomic PCR and DNA sequencing of pilQ
| Oligonucleotide | Sequence |
|---|---|
| hgbX06 | 5′-ACCGACGACAGCATCATCCT-3′ |
| hgbX08 | 5′-CACCGGCAAAACAACAGGCT-3′ |
| hgbX10 | 5′-TGCGCCAGCAAGGGAACATC-3′ |
| hgbX04 | 5′-GGCTGGGGCGTGAACTC-3′ |
| hgbX02 | 5′-CTGAATACGCAGGCTATGGT-3′ |
| hgbX03 | 3′-CGGGAATGACGGCTCAAAAG-5′ |
| hgbX01 | 3′-AGCCGTATCCGACAGATTCC-5′ |
Construction of FA7168, FA7186 and mutagenesis of tonB and hemA.
Construction of hpuA deletion (ΔhpuA) mutants which express HpuB under the control of the hpuA promoter and isolation of FA7168, a ΔhpuA hpuB::cat hgbX mutant that grew on Hb, were described previously (6). Isolation of the Hb+ FA7168 from the Hb− FA7167 (also ΔhpuA hpuB::cat) prompted us to seek Hb+ revertants among derivatives of FA7169 (ΔhpuA hpuB+) that were unable to grow on Hb. A hmbR::Ω derivative of FA7169 that cannot revert to Hb+ by point mutations in the pseudogene, hmbR, was named FA7186. Hb+ mutants of FA7186 were identified by plating cell suspensions on Hb-Des plates and picking colonies that grew well when passed onto fresh Hb-Des plates.
TonB dependence of Hb+, hgbX mutants was tested by insertional inactivation of tonB with the Ω cassette. This procedure utilized the tonB::Ω plasmid pUNCH173 (3) to transform various ΔhpuA mutants with selection for resistance to SPT. The ability of various mutants to utilize Hb as a heme source was tested by the construction of hemA mutants utilizing the hemA::Ω plasmid pUNCH1306 for transformation as described by Turner et al. (48).
Cloning the pilQ1 gene.
FA7168 chromosomal DNA fragments of various sizes were obtained after Sau3AI partial digestion and agarose gel electrophoresis fractionation (10). Fragments of 0.5 to 4 kb eluted from gel slices were divided into five fractions and transformed into FA7167. The fraction with DNA fragments of 1.6 to 3 kb was enriched in the gene (see below) that transformed FA7167 to Hb+. A plasmid DNA library of this fraction was prepared by inserting DNA fragments into BamHI-digested, shrimp alkaline phosphatase (Boehringer Mannheim)-treated, pBluescript II KS(+) (Stratagene, La Jolla, Calif.) and cloning them in E. coli DH5αMCR (Gibco-BRL, Gaithersburg, Md.). Pools of plasmid DNA were generated and used to individually transform Hb− FA7167 to Hb+. Active pools were subdivided and screened again until eventually a single 5.2 kb clone was identified that conferred the ability to utilize Hb. The 2.2-kb gonococcal insert in this clone contained 862 bp of the 3′ end of pilQ, and comparison to the gonococcal genomic database (GenBank accession NC_002946) showed that it contained a point mutation 412 bp from the 3′ end of the open reading frame of pilQ. This allele was designated pilQ1.
Oligonucleotides were designed (Table 2) to amplify and sequence pilQ from the genomes of hgbX mutants derived from FA7186. The DNA sequences revealed that FA7186H19 had precisely the same pilQ point mutation as that of FA7168. Of six mutants with the hgbX phenotype, FA7168 carried a cat cassette in hpuB; the rest were derivatives of FA7186 and carried the Ω cassette in hmbR. They were all sensitive to CHL1 despite expression of the cat resistance cassette in the hpuB of FA7168.
In order to use the Ω cassette to inactivate pilQ and to provide further proof that the point mutation in pilQ was responsible for the Hb+ phenotype, FA7186H19 pilQ1 was moved by transformation into FA7167 (ΔhpuA hpuB::cat, Hb−), with selection for Hb+. A parallel transformation was done with the chromosomal DNA of FA7168. All Hb+ transformants from these crosses were SPT100 and CHL1 sensitive (SPTs CHLs). Genomic PCR and DNA sequencing confirmed the presence of pilQ1 point mutation in the Hb+ transformants. The transformant with chromosomal DNA from FA7168 was named FA7317. Two transformants obtained from two transformations by FA7186H19 DNA were named FA7294 and FA7318.
Mutagenesis of pilQ.
A shortened version of pilQ was made by genomic-PCR with the upstream PCR primer hgbX08, which started 434 bp from the ATG site, and the downstream primer hgbX03, which ends 52 bp after the TGA site. The 1,872-bp PCR product from FA7168 was inserted into the vector PCR2.1-TOPO (Invitrogen) and cloned in E. coli DH5αMCR to generate pUNCH 288. A 30-bp segment of pUNCH 288 between two HincCII sites was removed and replaced with a Ω cassette to become pUNCH 290. Replacing the same segment with a cat cassette generated pUNCH 291. Both pUNCH 290 and pUNCH291 were used to create pilQ insertional knockout mutants. Western blots were performed to confirm loss of PilQ by using a rabbit antiserum against OMP-MC kindly supplied by Charles E. Wilde III (Indiana University School of Medicine).
Construction of gonococcal pilQ mutants in RM11.2recA6 background.
The FA1090 derivative RM11.2recA6 (25, 31) was induced with isopropyl-β-d-thiogalactopyranoside (IPTG) and transformed with FA7167 chromosomal DNA, which contains the ΔhpuA hpuB::cat mutations. CHL1-resistant (CHLr) transformants were screened by Southern blot hybridization and PCR analysis, and the pilE sequence was determined. The ΔhpuA hpuB::cat mutant of RM11.2recA6 was named FA7332. FA7332 was transformed with FA7168 chromosomal DNA that contains pilQ1. Transformants with the pilQ1 point mutation (FA7333) were identified by growth on Hb-Des plates and were authenticated by genomic PCR and DNA sequencing of pilQ and pilE.
RM11.2recA6 ΔpilE and RM11.2recA6 ΔhpuA hpuB::cat ΔpilE mutants were generated by induction of recA with IPTG and selection of nonpiliated areas of growth on colonies. To delete the pilE of FA7333, chromosomal DNA from RM11.2recA6 ΔpilE was used to transform in the deletion. The pilE deletions were verified with Southern blot and PCR analysis. DNA sequencing determined that the 924-bp deletion spans between upstream sequence and the cys2 sequence in pilE, which deletes the pilE promoter and ribosomal binding site.
Construction of pilT mutants.
In order to make pilT mutants of strains in the RM11.2recA6 background, FA7332 and FA7333 were transformed with RM11.2recA6 pilT::erm (26) chromosomal DNA. ERY1-resistant transformants of FA7332 and ERY0.05-resistant transformants of FA7333 were screened by Southern blot analysis. The insertional inactivation of pilT was identified by screening for the characteristic colony morphology differences of pilT (dud1) mutants (4). These pilT mutants carried the parental pilE sequence. The pilT::erm mutants of FA1090, FA7168, FA7186H19, FA7317, and FA7318 were constructed by moving chromosomal DNA of RM11.2recA6 pilT::erm into the recipients. Transformants of FA1090 were selected on ERY1, and transformants of the pilQ1 mutants were selected on ERY0.5 plates. In order to evaluate the effect of PilT inactivation on ERY sensitivity, FA1090, FA7167, and FA7186H19 were also transformed with chromosomal DNA of an MS11 mutant carrying an inducible pilT promoter. (The mutation was moved into MS11 by Laura Potter of Oregon Health Sciences University from a strain originally constructed in the laboratory of Michael Koomey [50], University of Oslo, Oslo, Norway.) Transformants were selected on TET2. Preservation of the hpuA deletion and hpuB insertion in these pilT mutants were confirmed by genomic PCR and DNA sequencing.
Immunoelectron microscopy.
For analysis of gonococcal piliation states, Formvar-coated grids were used to lift cells directly from 18-h-old colonies grown on GCB plates. Grids were fixed for 15 min by floating on a drop of 0.2% glutaraldehyde-4% paraformaldehyde (Fisher Scientific). After fixation, the grids were rinsed three times with Dulbecco phosphate-buffered saline (PBS; Gibco-BRL) with 1% bovine serum albumin (BSA; Sigma) and then incubated in 0.1% gelatin (Aurion, Inc.) in PBS for 30 min. The grids were rinsed once with PBS-BSA and then incubated with a 1:10 dilution of rabbit α-pilin peptide antiserum (26) for 1 h. The antiserum was raised against a synthetic peptide found in the hypervariable loop region of the pilin variant expressed on RM11.2recA6 (KRDAGAKTGADDVKADGN), synthesized on keyhole limpet hemocyanin resin and used to immunize New Zealand White female rabbits (Research Genetics). Grids were rinsed three times with PBS-BSA and incubated with 0.1% gelatin in PBS for 30 min. The grids were rinsed once with PBS-BSA and incubated with goat α-rabbit immunoglobulin G antibody conjugated to 12-nm gold (1:20 dilution; Jackson Immunolabs) for 1 h. Grids were rinsed five times in water for 3 min each step. Excess water was carefully wicked away, and grids were negatively stained with 1% uranyl acetate for 1 min. All incubations were performed at 25°C. Grids were viewed by using a JEOL JEM-1220 transmission electron microscope at 60 kV.
Transformation competence assays for strains in RM11.2recA6 background.
Gonococci were grown on GCB plates for 18 h, collected with dacron swabs, and suspended to a density of 108 CFU/ml in liquid GCB (GCBL). Then, 20 μl of cells was added to 200 μl of GCBL that contained 5 mM MgSO4 (35), Kellogg supplements, 1 mM IPTG, and 50 ng of pSY6 (37). After 15 min at 37°C, DNase I (Promega) was added, and the incubation was continued for 5 min at 37°C to remove exogenous DNA. Then, transformation mixes were diluted into 2 ml of GCBL plus 5 mM MgSO4, Kellogg supplements, and 1 mM IPTG, followed by incubation at 37°C in 5% CO2 for 4 h without agitation. Transformants were selected on GCB plates containing 2 μg of nalidixic acid (NAL)/ml.
Hb-binding assays.
These procedures were as described previously (6). Briefly, the binding was measured by 125I-labeled anti-human Hb antibodies bound to buffer-washed whole cells previously incubated in Chelex-treated defined medium (CDM) supplemented with Hb.
Growth and microbial sensitivity assays.
The ability of a gonococcal strain to use Hb or heme for growth was measured in liquid culture with iron-chelated CDM or on GCB-Des agar plates into which defined amounts of Hb or heme were placed into wells cut into the agar, as described previously (6). Disk assays were used to test antimicrobial sensitivities on GCB agar plates. Single, phenotypically piliated colonies of strains to be tested were passed from selection plates to GCB plates for the following day's experiment. Overnight plate-grown gonococci were suspended in GCB to an optical density at 600 nm (OD600) of 0.1. Then, 50 μl of the cell suspension was mixed with 5 ml of liquid GCB agar, and the mixture was poured over a plate containing 15 ml of solidified GC agar. Four filter paper disks (6 mm in diameter; Schleicher & Schuell, Dassel, Germany) were then placed on top of the solidified GCB agar-gonococci mix. The agents tested on disks included AMP at 1 to 5 μg per disk, CHL at 0.25 μg per disk, ERY at 0.1 to 0.5 μg per disk, rifampin (RIF) at 0.05 to 0.5 μg per disk and 2.5 to 10 μl of Triton X-100 (TX-100) at 1:100 dilution.
Plating efficiency and determination of the MIC.
The plating efficiency was determined by plating dilutions of cell suspensions at an OD600 of 0.18 onto GCB plates or Hb-Des plates. MICs were determined by plating diluted cell suspensions on GCB plates containing doubling dilutions of antimicrobial agents; the MIC was the lowest concentration that prevented growth.
RESULTS
Isolation of novel Hb+ mutants.
We previously reported spontaneously arising Hb-utilizing (Hb+) colonies from hpuA deletion mutants of FA1090, FA7167 (Hb−, ΔhpuA hpuB::cat) and its derivative FA7186 (Hb−, ΔhpuA hpuB+ hmbR::Ω) (6). Some of them carried a point mutation in hpuB, whereas others had mutations originally designated ΔhpuA hgbX that mapped outside the hpuAB locus (6). All of the hgbX mutants grew well on Hb-Des plates and in liquid culture with Hb as the sole source of iron (data not shown). The hgbX mutant strain isolated from FA7167, FA7168, retained the cat cassette in hpuB but became sensitive to CHL1. Nevertheless, it showed a much smaller zone of growth inhibition, compared to wild-type FA1090, when tested with disk loaded with CHL (Table 3). Screening of its sensitivities to several other antimicrobial agents revealed that FA7168 was also more sensitive to heme, AMP, ERY, RIF, and TX-100 than FA7167 and FA1090 (Table 3).
TABLE 3.
The pilQ1 point mutation in FA7168 increased sensitivity to certain antibiotics and the detergent TX-100
| Strain | Diam (mm)a with:
|
|||||||
|---|---|---|---|---|---|---|---|---|
| AMP (5 μg) | CHL (1.5 μg) | ERY (0.5 μg) | RIF (0.5 μg) | TX-100 (5 μl of a 1:100 dilution) | KAN (1.5 μg) | NAL (1.5 μg) | SPT (5 μg) | |
| FA1090 wild type, Hb+ | 22 | 26 | 14 | 22 | 8 | 10 | 16 | 16 |
| FA7167 (ΔhpuA hpuB::cat), Hb− | 22 | 8 | 12 | 24 | 8 | 12 | 16 | 18 |
| FA7168 (ΔhpuA hpuB::cat pilQ1), Hb+ | 28 | 16 | 26 | 36 | 22 | 10 | 14 | 16 |
Values are diameters of the zone of growth inhibition around each 6-mm disk loaded with the indicated amount of antimicrobial agent measured after 24 h. Data were from one representative experiment.
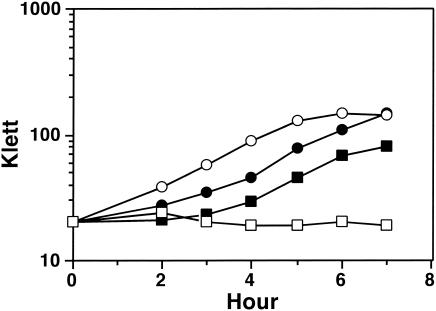
In liquid culture with Hb (1 μM) as the sole source of iron, the growth of FA7168 was inhibited by the addition of HSA which binds free heme (16) (Fig. 1). This result is in contrast to the behavior of FA1090, which is able to compete for heme complexed to HSA by the expression of the HpuAB receptor (6).
FIG. 1.
Representative growth curves of the ΔhpuA hpuB::cat pilQ1 point mutant, FA7168, and HpuAB phase on FA1090 in liquid cultures, as measured by the OD600 in the Klett colorimeter. Gonococci were grown in liquid cultures of CDM-Des supplemented with Hb (1 μM) or Hb and HSA (16 μM). Symbols: •, FA1090 in Hb; ○, FA1090 in Hb-HSA; ▪, FA7168 in Hb; □, FA7168 in Hb-HSA.
The Hb+ phenotype in FA7168 is due to a point mutation in pilQ.
Transformation of the Hb− recipient FA7167 with pooled plasmids containing chromosomal DNA from one hgbX mutant, FA7168, resulted in isolation of a plasmid that contained a point mutation in pilQ. Subsequent studies showed that this point mutation was responsible for the growth on Hb in the absence of an Hb receptor and for increased sensitivity to various antimicrobial agents. A T-to-G mutation occurred at base 1761 of FA7168 pilQ coding sequence, resulting in an F563L mutation in the predicted mature PilQ protein. Sequence from pilQ genomic PCR products from the other five hgbX mutants—H1, H12, H15, H19, and H20—derived from FA7186 (Hb− ΔhpuA hpuB+ hmbR::Ω) revealed precisely the same point mutation in one, i.e., FA7186H19. The other four hgbX mutants had the wild-type pilQ sequence. Like FA7168, FA7186H19 showed elevated sensitivities to various antimicrobial agents (data not shown). Therefore, the same F563L mutation was independently isolated in two strain backgrounds and designated pilQ1.
Moving FA7168 and FA7186H19 back into FA7167 background generated the Hb+ transformants FA7317, FA7294, and FA7318 (all ΔhpuA hpuB::cat pilQ1), supporting the notion that their growth on Hb was dependent on preservation of the pilQ1 point mutation. The necessity of PilQ for Hb utilization in the ΔhpuA pilQ1 mutants was also confirmed by the insertional inactivation of pilQ. The pilQ::Ω transformants of FA7294 and FA7317 were both completely unable to utilize Hb for growth. As a control, a pilQ::Ω mutant of FA1090 Hb+ still grew well on Hb-Des plates, as expected, because of the presence of the HpuAB receptor in FA1090 (data not shown).
The Hb+ phenotype is independent of TonB and PilQ1 is not an Hb receptor.
The HpuAB and HmbR Hb receptors, as well as the transferrin and the lactoferrin receptors, are dependent on TonB for function. To test for possible TonB dependence of the Hb supported growth of pilQ1 mutant, FA7317 tonB were insertionally inactivated with the Ω cassette. Although the previously obtained ΔhpuA hpuB point mutants still required TonB to utilize Hb as the sole source of iron (6), the pilQ1 tonB double mutants grew on iron-chelated medium containing Hb but not transferrin (data not shown). Thus, the Hb+ phenotype conferred by pilQ1 was independent of TonB.
The possibility that the pilQ1 mutation might restore Hb receptor activity to strains carrying mutations in hpuAB was examined by radioimmunoassay. FA7168 exhibited no iron-dependent binding of Hb and very low levels of Hb binding compared to HpuAB phase on FA1090 that expressed a wild-type form of the HpuAB Hb receptor (Table 4). The Hb binding was also low in parent FA7167 (ΔhpuA hpuB::cat pilQ+) (Table 4). The minor differences between FA7167 and FA7168 were not significant (Student t test).
TABLE 4.
The pilQ point mutation does not increase Hb binding to whole cells
| Strain | Phenotype | Growth condition | % Total cpm loadeda ± SD |
|---|---|---|---|
| FA1090 (HpuAB phase on) | Hb+ | Iron replete | 0.24 ± 0.03 |
| Iron limited | 1.51 ± 0.36 | ||
| FA7167 (ΔhpuA hpuB::cat) | Hb− | Iron replete | 0.07 ± 0.02 |
| Iron limited | 0.05 ± 0.03 | ||
| FA7168 (ΔhpuA hpuB::cat pilQ1) | Hb+ | Iron replete | 0.08 ± 0.04 |
| Iron limited | 0.08 ± 0.05 |
125I-labeled anti-human Hb antibody bound to gonococcal bound Hb is expressed as a percentage of the total counts per minute added to each well. Data are averages of two experiments, each done in quadruplicate.
Different classes of ΔhpuA hgbX mutants.
Compared to their immediate parents, FA7167 and FA7186, the hgbX mutants were more sensitive to the same set of antimicrobial compounds and protoporphyrin IX (PPIX) and FePPIX. However, among six mutants identified, the two that were subsequently determined to be pilQ mutants, FA7168 and FA7186H19, were relatively less sensitive to TX-100, PPIX, and FePPIX but more sensitive to the antibiotics RIF and ERY (data not shown). The MIC of PPIX was 250 μg/ml for FA1090 compared to ∼4 μg/ml for FA7168 and FA7186H19 and <2 μg/ml for the other hgbX mutants. These phenotypic differences suggested that there might be different genotypic classes of hgbX mutants with a similar Hb+ phenotype.
According to Drake et al. (14), PilP, a pilus biogenesis lipoprotein, affects the expression of PilQ as a macromolecular multimer. To check for the possibility that a mutation in pilP resulted in the Hb+ phenotype of other hgbX mutants, genomic PCR products of the pilP gene of all hgbX mutants were sequenced. Their sequences were the same as wild-type FA1090. The increased sensitivity to TX-100 and RIF of the hgbX mutants prompted an attempt to determine whether the mtr CDE-encoded efflux pump might be involved (17). Introduction of mtrC::Km, mtrD::Km, and mtrE::Km mutations (38) by transformation into Hb− strain FA7167 (ΔhpuA hpuB::cat) resulted in increased sensitivity to TX-100 and RIF as expected but none of the insertional mutants acquired the ability to grow on Hb-Des plates (data not shown). Thus, it seemed unlikely that the mtr genes were involved in the mutations that resulted in ability to grow on Hb in the absence of a Hb receptor. The possible involvement of another efflux pump, NorM (33), that recognizes cationic antimicrobial agents was also studied. Sequencing the gene norM also revealed no difference between the hgbX mutants and the wild type. At present, the identities of the other hgbX mutations are unknown.
Increased entry of free heme due to pilQ1.
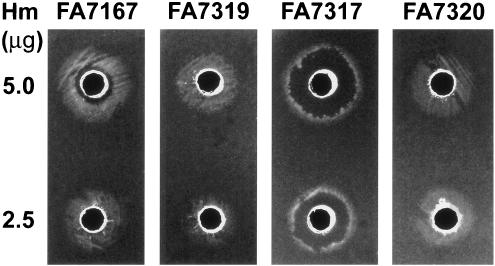
When 5 μg of heme was placed in a well cut into a GCB-Des agar plate previously spread with wild-type strain FA1090 (data not shown) or ΔhpuA hpuB::cat strain FA7167, these strains exhibited a ring of dense growth surrounding an inner zone of growth inhibition (Fig. 2). In contrast, the pilQ1 mutants exhibited a thin ring of growth with a larger inner zone of growth inhibition (Fig. 2), indicating increased sensitivity to heme toxicity in pilQ1 mutants. This result is consistent with these mutants' increased sensitivity to antibiotics and the porphyrins PPIX and FePPIX (40) (Table 3 and data not shown). With reduced amount of heme (2.5 μg) in the central wells, zones of growth inhibition decreased in both the pilQ+ and the pilQ1 strains (Fig. 2). The dose dependence of zone of growth inhibition confirmed that excess free heme was the cause of growth inhibition and that the entry of free heme was augmented in pilQ1 mutants.
FIG. 2.
Free heme supported growth of hpuAB mutants and their pilQ inactivation mutants. GCB-Des plates were spread with cell suspensions of respective strains: FA7167, ΔhpuA hpuB::cat pilQ+ parent; FA7319, pilQ::Ω transformant of FA7167; FA7317, pilQ1 transformant of FA7167; FA7320, pilQ1::Ω transformant of FA7317. Wells were cut into the agar and loaded with 50 μl of heme at 0.1 or 0.05 μg/μl. Images were obtained after 48 h of incubation. At 5.0 μg of heme per well, FA7167 grew in a wide growth ring, whereas the pilQ1 mutant, FA7317, showed a thin growth ring with a wide zone of growth inhibition. Decreasing the amount of heme decreased the growth inhibition zones. Insertional inactivation of pilQ abolished heme toxicity.
The increased sensitivity to porphyrins of strains with the pilQ1 point mutation strongly suggested that entry of intact porphyrins was increased in the mutant background. To further address the question of whether the pilQ1 mutation resulted in entry of heme or merely iron released from heme into the cell, hemA mutants were constructed in the relevant strains. Mutation in hemA of FA1090 results in inability to grow without exogenous heme source, since the hemA mutants are unable to synthesize their own heme (48). The hemA::Ω transformant, FA7322, of FA7317 (Hb+, pilQ1) grew well on Hb (data not shown), indicating that heme, or at least an intact porphyrin ring, entered the pilQ1 mutant cells.
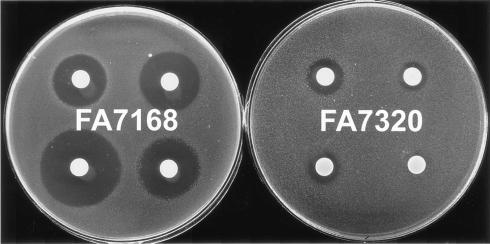
Additional evidence for the role of PilQ in the entry of free heme and antimicrobial compounds was provided by analysis of the effects of pilQ inactivation on both the pilQ+ and the pilQ1 strains. Inactivation of PilQ completely prevented the zone of heme toxicity in both FA7319 (FA7167 pilQ::Ω) and FA7320 (FA7317 pilQ1::Ω) (Fig. 2). Reversal of hypersensitivities to AMP, ERY, RIF, and TX-100 was also obvious in FA7320 compared to the parent pilQ1 mutant (Fig. 3). Slight decrease of sensitivities to ERY, RIF, and TX-100 was also observed in the pilQ::Ω mutant of FA1090, FA7311 (Table 5). Moreover, introduction of pilQ1 by transformation of RM11.2 recA6 (25) (FA7339) resulted in hypersensitivity to the same agents (data not shown), showing that the effects of pilQ1 were not dependent on a mutation in the HpuAB receptor. We concluded that PilQ forms an entry channel for free heme and certain other antimicrobial compounds in wild-type gonococci, which is accentuated by the pilQ1 point mutation. Since there still was growth around wells containing free heme in FA1090 and FA7167 derivatives lacking PilQ, there also must be additional routes for entry of heme that do not depend on PilQ.
FIG. 3.
Effects of pilQ inactivation on pilQ1 mutant's sensitivity to antimicrobial agents. Filter paper disks placed on the surface of FA7168 (ΔhpuA hpuB::cat pilQ1)- and FA7320 (ΔhpuA hpuB::cat pilQ1::Ω)-coated plates were loaded with AMP (1 μg per disk, top left), ERY (0.25 μg, top right), RIF (0.05 μg, bottom left), and TX-100 (5 μl of a 1:100 dilution, bottom right). Images were obtained after 24 h of incubation.
TABLE 5.
Insertional inactivation of pilQ and pilT in FA1090 decreased sensitivities to ERY, RIF, and TX-100
| Straina | Mean zone of growth inhibition (mm)b ± SD with:
|
|||
|---|---|---|---|---|
| AMP (2 μg) | ERY (0.5 μg) | RIF (0.1 μg) | TX-100 (10 μl of a 1:100 dilution) | |
| FA1090 P+, Hb+ | 17.6 ± 1.1 | 19.8 ± 3.5 | 18.0 ± 1.0 | 11.0 ± 1.0 |
| FA1090 P−, Hb+ | 18.2 ± 1.5 | 20.2 ± 3.3 | 18.4 ± 1.5 | 11.0 ± 1.6 |
| FA7311 (FA1090 pilQ::Ω), Hb+ | 18.0 ± 1.1 | 18.8 ± 4.3* | 16.0 ± 2.1* | 9.3 ± 1.0** |
| FA7348 (FA1090 pilT::tet), Hb+ | 18.3 ± 1.5 | 18.7 ± 3.0* | 16.2 ± 1.0* | 9.5 ± 0.8* |
For FA1090, phenotypically piliated (P+) and nonpiliated (P−) colonies were selectively passed in preparation for the following day's experiment.
Each value is an average of five zones of growth inhibition obtained with the indicated amount of antimicrobial agent measured from five experiments after 48 h of incubation. A paired t test showed significant differences (*, P < 0.05; **, P < 0.01) as indicated compared to FA1090 P+.
The pilQ1 mutation decreases pilus expression and transformation efficiency.
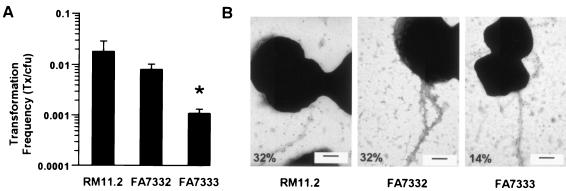
The product of pilQ is essential for the biogenesis of type IV pili in Neisseria gonorrhoeae (13). To ascertain the role of the PilQ point mutation in pilus assembly and function, the ΔhpuA hpuB::cat pilQ1 mutation was transformed into RM11.2recA6, a defined pilin variant of strain FA1090 with a regulatable recA that allows for IPTG control of DNA transformation and pilin antigenic variation (34). All RM11.2recA6 derivatives were shown to carry identical pilE sequences, ensuring that differences in piliation were not due to changes in pilus expression due to pilin variation (data not shown). To directly examine piliation of the RM11.2recA6 ΔhpuA hpuB::cat and RM11.2recA6 ΔhpuA hpuB::cat pilQ1 transformants, FA7332 and FA7333, immunogold electron microscopy with an anti-pilin peptide antibody was performed. Compared to the wild-type RM11.2recA6, the number of cells expressing pili was unchanged in FA7332 but was reduced by ∼2-fold in the pilQ1 mutant FA7333 (Fig. 4), which nevertheless retained a piliated colony morphology. The pilQ1 mutant also showed sevenfold-lower transformation efficiency than FA7332 (Fig. 4), although there is still substantial transformation competence at 10−3 transformants/CFU.
FIG. 4.
Bar graph illustrating transformation efficiencies (A) and electron micrographs comparing the piliation (B) of RM11.2recA6 (wild type), FA7332 (ΔhpuA hpuB::cat), and FA7333 (ΔhpuA hpuB::cat pilQ1). The piliation of these three strains is given as the percentage of cells expressing pili, as visualized by immunogold electron microscopy with anti-pilin sera, and the percent piliation is indicated on each electron micrograph. The bar in each electron micrograph is 500 nm. Two grids for each strain were examined. The numbers of cells examined were 216, 346, and 558, respectively. All three strains exhibited predominantly bundled pili. FA7333 showed decreased piliation as well as decreased transformation efficiency. The asterisk in panel A indicates a significant difference by using the Student t test (P < 0.05) compared to RM11.2 recA6 and FA7332.
To determine the effects of pilus expression on the entry of antimicrobial compounds through PilQ macromolecular complex, ΔpilE mutants were made from FA7332 and FA7333 and compared to RM11.2recA6 ΔpilE (FA7336). The transformation efficiencies for each of the three ΔpilE mutants were similar and reduced ca. 5 to 6 logs compared to the isogenic Pil+ strains (data not shown). Loss of pilus expression in pilQ1 mutant had little effect on Hb-supported growth, but the ΔpilE pilQ1 double mutant, FA7338, showed increased sensitivity to heme toxicity (Fig. 5) and slightly increased sensitivity to AMP and TX-100 (Table 6). Thus, pilus expression might compete with certain molecules for the PilQ macromolecular complex.
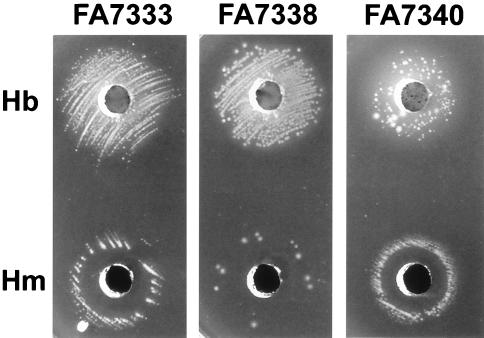
FIG. 5.
Effects of pilE deletion and pilT inactivation on Hb (50 μl at 10 μg/μl) and heme (50 μl at 0.1 μg/μl) supported growth of a pilQ1 mutant, FA7333 (ΔhpuA hpuB::cat pilQ1, Hb+). FA7338 is the ΔpilE transformant and FA7340 is the pilT::erm transformant of FA7333. Images were obtained after 48 h of incubation. The pilQ1 pilT double mutant gave only isolated large colonies around the Hb well. The zone of growth inhibition around the heme well was wider in the pilQ1 ΔpilE double mutant but was reduced in the pilQ1 pilT double mutant compared to that of FA7333.
TABLE 6.
Deletion of pilE and inactivation of pilT changed sensitivities to TX-100 and certain antibiotics in pilQ1 mutants
| Strain | Mean zone of growth inhibition (mm)a ± SD with:
|
|||
|---|---|---|---|---|
| AMP (1 μg) | ERY (0.1 μg) | RIF (0.05 μg) | TX-100 (5 μl of a 1:100 dilution) | |
| FA7332 (RM11.2recA6 hpuAB−), Hb− | 16.3 ± 2.9** | 7.7 ± 3.3** | 14.3 ± 2.3** | 7.0 ± 2.4** |
| FA7333 (pilQ1 mutant of FA7332), Hb+ | 22.2 ± 1.9 | 19.5 ± 3.3 | 30.0 ± 2.3 | 16.2 ± 2.9 |
| FA7338 (FA7333 ΔpilE) | 23.7 ± 1.8* | 19.1 ± 2.8 | 30.0 ± 1.8 | 20.6 ± 1.2* |
| FA7340 (FA7333 pilT::erm) | 21.3 ± 1.8* | NA | 23.4 ± 2.8** | 7.4 ± 2.3** |
Each value is an average five zones of growth inhibition obtained with the indicated amount of antimicrobial agent measured from five experiments after 48 h of incubation. NA, not applicable. A paired t test showed a significant difference (*, P < 0.05; **, P < 0.01) as indicated compared to FA7333.
PilT is essential for the observed phenotypic effects of the pilQ1 mutation.
The pilus accessory gene pilT plays a crucial role in twitching motility and pilus retraction (27, 30), acting as a conditional antagonist of stable pilus fiber formation (50, 51, 52). In order to examine the influence of PilT in a pilQ1 background, we introduced two kinds of pilT loss-of-function mutations into pilQ wild-type strains FA1090, FA7167, and pilQ1 mutants FA7168, FA7186H19, FA7317, and FA7318, some with pilT::erm and the others with pilT::tet. The pilQ1 pilT::erm double mutants showed low plating efficiencies on Hb-Des plates (ca. 5 × 10−3 Hb-Des CFU/GCB CFU). After 48 h of incubation on Hb-Des plates, they exhibited a mixture of a few regular-sized and many tiny colonies. Passage of the regular sized colonies to fresh Hb-Des plates yielded uniformly regular-sized colonies. Passage of tiny colonies on the same media continued to yield the same proportions of tiny and normal-sized colonies. The rates of colony variation suggested possible phase variation of another surface component, but no differences were found in PilC expression (determined by Western blots with anti-PilC polyclonal sera; data not shown).
Similar low plating efficiencies and apparent phase variation from tiny colonies to large normally growing colonies were noted in earlier studies of double mutants that expressed neither PilQ nor PilT, which was shown to be the result of toxicity from assembled intracellular pilus fibers that could neither be extruded through PilQ, nor broken down due to loss of PilT function (52). In that case, the large-colony variants were due to pilin mutations that resulted in the inability to assemble toxic fibers. To further address whether a similar phenomenon in the pilQ1 pilT::erm double mutants might account for low plating efficiencies and frequent reversion to large-colony variants on Hb-Des plates, the pilQ1 pilT::erm mutations were introduced into FA7332 (RM11.2recA6 ΔhpuA hpuB::cat), which does not undergo pilin variations due to the recA6 mutation in the absence of induction of expression of recA function by IPTG (36). The results showed that there were similar low plating efficiencies on Hb-Des plates and the appearance of large and small colony variants on these plates (data not shown). This strongly suggested that recombination pathways were not necessary to the reversions to normal growth and therefore that pilin variations were not responsible for the restored growth.
Loss of PilT expression decreased the sensitivity of pilQ1 mutants to heme toxicity. The pilQ1 pilT::erm double mutant, FA7340 (in RM11.2recA6 ΔhpuA hpuB::cat background), exhibited a heavier ring of growth and a smaller inner zone of growth inhibition around the central well containing heme, compared to the parent strain with intact PilT function (Fig. 5). FA7340 was less sensitive to AMP, RIF, and TX-100 than its parent, FA7333 (Table 6), whereas the pilT::tet mutant of FA7186H19 (ΔhpuA hpuB::cat pilQ1), FA7350, demonstrated decreased sensitivity to ERY (data not shown). Loss of PilT also reduced the entry of some antimicrobial compounds in pilQ wild-type strains. Although the effect of pilT inactivation in FA1090 was small, differences were significant for sensitivities to ERY, RIF, and TX-100 (Table 5). Thus, PilT was required for the susceptibility to antibiotics, detergents, and heme/porphyrins that can utilize both wild-type and PilQ1 macromolecular complex for entry into the bacterial cell.
DISCUSSION
We initiated this study to elucidate the mechanisms responsible for receptor-independent uptake of iron and heme from Hb, based on the previous isolation of gonococcal mutants that were able to grow on Hb despite mutational loss of HpuA, the lipoprotein coreceptor that normally is essential for the function of the HpuAB receptor (6). The hgbX mutants fell into two general classes based on differences in their susceptibilities to a variety of antimicrobial agents. Although all were more susceptible to generally hydrophobic antibiotics and TX-100 than the wild type, some were particularly susceptible. One of the two classes of mutants turned out to have a point mutation near the 3′ end of pilQ (pilQ1), whereas the other was normal in all tested genes. Efforts to clone the gene(s) responsible for the remaining uncharacterized hgbX mutants other than pilQ by the same strategy used to isolate the pilQ1 mutation have been unsuccessful. The results described in this communication showed that the Hb+ phenotype of the pilQ1 mutants was neither dependent on HpuAB or TonB nor on Hb binding to whole cells. Rather, growth on Hb was due to the release of free heme from Hb and entry through a channel or channels that did not depend on the Hb receptor. We presume that heme was released from Hb on the cell surface or in the extracellular medium because HSA prevented the entry of heme. So far, there has not been any report of Hb entry in any bacterial species. The relatively large sizes of albumin and Hb (molecular weights of ca. 69,000 and 64,500, respectively) and the funnel-shaped cavity in the PilQ macromolecule (9) make it unlikely that either could enter through the PilQ channel.
Demonstration that the pilQ1 mutation resulted in increased entry of heme, TX-100, and a variety of antibiotics was surprising, since PilQ is not known to facilitate the entry of exogenous compounds. It is likely that only a few mutations in pilQ result in this phenotype, since the same F563L mutation was isolated independently twice. Very recently, an additional and different pilQ mutation has been isolated that results in increased resistance to antibiotics, further suggesting that wild-type PilQ allows the entry of a variety of compounds (R. Nicholas et al., unpublished data). Our demonstration that the phenotype was very closely linked to pilQ1 and was lost by insertional inactivation of pilQ1 suggests that the effects were mediated by changes within PilQ. We cannot absolutely exclude the possibility that the PilQ F563L mutation provides these phenotypes by affecting membrane integrity or another secondary effect. The simplest hypothesis is that PilQ can directly allow entry of these molecules into the bacterial cell and that the F563L mutation increases the number of molecules that can efficiently utilize this route.
The conclusion that PilQ forms not only a pore for exit of the assembled pilus fibril (13, 52) but also a channel for entry of heme and various antimicrobial agents is strengthened by the additional observation that loss of pilT function decreased the effects of the pilQ1 mutation. PilT is responsible for twitching motility mediated by retraction of the pilus fibril (27, 30). It also is necessary for degradation of the pilus fibril when PilQ is not available for export of the intracellular assembled pilus fibril; in the absence of both PilQ and PilT, cell viability is lost unless other secondary mutations in pilE prevent formation of what would otherwise be toxic intracellular pilus fibrils (52). PilT is also required for the transport of DNA into the cell for genetic transformation (48). Our observation that inactivation of pilT in strains carrying the pilQ1 missense mutation resulted in decreased sensitivity to heme, TX-100, and antibiotics argues that PilT normally increases the entry properties of the PilQ macromolecular complex. Perhaps the PilQ-dependent entry of small molecules shares mechanistic properties with pilus retraction, where the assembled pilus is imported back into the cell. It is not clear whether heme and antimicrobial agents bind to pili or to PilQ, but modestly increased sensitivity to heme and certain antimicrobial agents in a ΔpilE construct (Fig. 5 and Table 6) suggests that either extruded pili compete with exogenous compounds for PilQ or pili partially impeded passage of these compounds through the PilQ channels.
Many bacteria possess receptors for binding and utilizing heme-carrying compounds (43). The entrance of heme in gonococci is known to be TonB independent (3, 7, 44, 48), and no clear evidence for a heme receptor in the pathogenic Neisseria spp. has been produced. Our results demonstrated that some heme enters through a PilT- and PilQ-dependent pathway, but there almost certainly must be an additional mechanism for heme entry as well, since strains lacking both an Hb receptor and PilQ were able to grow on free heme (Fig. 2). It was curious that there was no toxicity when Hb was the source of heme, as opposed to adding free heme to the media. This may reflect relatively low concentrations of free heme released by Hb, compared to the amounts added to the wells in the assays used. Heme is known to be toxic for bacteria when in excess (40). Indeed, use of iron regulated receptors for obtaining heme from Hb under physiological circumstances may help to control the amount of heme that enters the cell, thereby avoiding toxicity.
Recent structural studies have utilized electron microscopy to establish proposed quaternary structures for members of the secretin family. According to Collins et al. (8, 9), the assembled meningococcal PilQ product takes the form of a doughnut composed of 12 identical subunits, surrounding a central cavity. When viewed from the side, the assembly is proposed to have a rounded, conical profile. Viewed from the top, the open end of the central channel apparently is just large enough to accommodate the pilus fibril, and the tapered end presumably blocks the channel (9). How the PilQ macromolecular assembly might be gated open so as to allow exit of pilus fibrils has been the subject of speculation involving the propulsive force of the pilus, which is sufficient to cause membrane protrusions in cells lacking PilQ and PilT (52). Such a mechanism might account for the specificity of the channel for pili. The only other function associated with the pilus-PilQ channel has been uptake of transforming DNA, which depends on pili, PilC, PilT, and other members of the complex (51). How DNA enters through this channel remains unclear; however, our data suggest that pilQ1 changes the specificity of the channel to allow increased entry of other molecules into the cell.
Studies on certain other members of the secretin family (34) have demonstrated gated channel properties. The pIV secretin for the filamentous phage f1 forms an ion conductive pore in planar lipid bilayers, and mutant forms of the protein allow import of molecules in whole cells to which E. coli normally is resistant (28). Entry through the pIV channel was decreased by production of phage that could not be released from the cell surface (29). The Klebsiella oxytoca secretin PulD also forms ion conductive channels in planar lipid bilayers (31). Each of these secretins has a quaternary shape similar to that proposed for PilQ (8). Future studies on PilQ might be expected to show voltage-dependent ion conductive channels in planar lipid bilayers, and mutants of PilQ such as the one we describe should alter the properties of the channel. These mutants hopefully also will help us to understand how PilQ interacts with other members of the complex (PilT, PilP, and PilC) in the secretion of pili and the mechanisms and physiological roles for entry of exogenous compounds, including heme and antibiotics through PilQ.
Acknowledgments
We thank Charles E. Wilde III (Indiana University School of Medicine) for providing the anti-N. gonorrhoeae OMP-MC antibody and Magdalene So and Laura Potter (Oregon Health Sciences University) for providing the MS11 inducible pilT mutant. We thank current and previous members of the Sparling laboratory: James Anderson, Gour Biswas, Christopher Elkins, Dalton Mclean, and Annice Rountree and Corinne Rouquette-Loughlin of the Shafer laboratory for help with this study. We also thank Jadranka Bozja and Igor Stojiljkovic (Emory University School of Medicine) for testing the gonococcal strains for their susceptibility to PPIX and FePPIX. We acknowledge the assistance of the Gonococcal Genome Sequencing Project supported by USPHS/NIH grant AI38399 and B. A. Roe, L. Song, S. P. Lin, X. Yuan, S. Clifton, T. Ducey, L. Lewis, and D. W. Dyer at the University of Oklahoma.
This study was supported by R37 grant 5-30837 and STD CRC grant 5-31660 to P.F.S., grants R01 AI33493 and R01 AI44239 to H.S.S., and grants AI21150 and AI45883 (R. C. Compans, Emory University) to W.M.S. W.M.S. is the recipient of a Senior Research Career Scientist Award from the VA Medical Research Service. D.M.T. was partially supported by NSRA F32 AI10568.
Footnotes
This paper is dedicated to the memory of the late Igor Stojiljkovic.
REFERENCES
- 1.Anderson, J. E., M. M. Hobbs, G. D. Biswas, and P. F. Sparling. 2003. Opposing selective forces for expression of the gonococcal lactoferrin receptor. Mol. Microbiol. 48:1325-1337. [DOI] [PubMed] [Google Scholar]
- 2.Anderson, J. E., P. A. Leone, W. C. Miller, C.-J. Chen, M. M. Hobbs, and P. F. Sparling. 2001. Selection of the gonococcal hemoglobin receptor during menses. J. Infect. Dis. 184:1621-1623. [DOI] [PubMed] [Google Scholar]
- 3.Biswas, G. D., J. E. Anderson, and P. F. Sparling. 1997. Cloning and functional characterization of Neisseria gonorrhoeae tonB, exbB, and exbD genes. Mol. Microbiol. 24:169-179. [DOI] [PubMed] [Google Scholar]
- 4.Biswas, G. D., S. A Lacks, and P. F. Sparling. 1989. Transformation-deficient mutants of piliated Neisseria gonorrhoeae. J. Bacteriol. 171:657-664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chen, C.-J., C. Elkins, and P. F. Sparling. 1998. Phase variation of hemoglobin utilization in Neisseria gonorrhoeae. Infect. Immun. 66:987-993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chen, C.-J., D. Mclean, C. E. Thomas, J. E. Anderson, and P. F. Sparling. 2002. Point mutations in HpuB enable gonococcal HpuA deletion mutants to grow on hemoglobin. J. Bacteriol. 184:420-426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chen, C.-J., P. F. Sparling, L. A. Lewis, D. W. Dyer, and C. Elkins. 1996. Identification and purification of a hemoglobin-binding outer membrane protein from Neisseria gonorrhoeae. Infect. Immun. 64:5008-5014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Collins, R. F., L. Davidsen, J. P. Derrick, R. C. Ford, and T. Tonjum. 2001. Analysis of the PilQ secretin from Neisseria meningitidis by transmission electron microscopy reveals a dodecameric quaternary structure. J. Bacteriol. 183:3825-3832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Collins, R. F., R. C. Ford, A. Kimitto, R. O. Olsen, T. Tonjum, and J. P. Derrick. 2003. Three-dimensional structure of the Neisseria meningitidis secretin PilQ determined from negative-stain transmission electron microscopy. J. Bacteriol. 185:2611-2617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cornelissen, C. N., G. D. Biswas, J. Tsai, D. K. Paruchuri, S. A. Thompson, and P. F. Sparling. 1992. Gonococcal transferrin binding protein 1 is required for transferrin utilization and is homologous to Ton-B-dependent outer membrane receptors. J. Bacteriol. 174:5788-5797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cornelissen, C. N., M. Kelly, M. M. Hobbs, J. E. Anderson, J. G. Cannon, M. S. Cohen, and P. F. Sparling. 1998. The transferrin receptor expressed by gonococcal strain FA1090 is required for the experimental infection of human male volunteers. Mol. Microbiol. 27:611-616. [DOI] [PubMed] [Google Scholar]
- 12.Cornelissen, C. N., and P. F. Sparling. 1994. Iron piracy: acquisition of transferrin-bound iron by bacterial pathogens. Mol. Microbiol. 14:843-850. [DOI] [PubMed] [Google Scholar]
- 13.Drake, S. L., and M. Koomey. 1995. The product of the pilQ gene is essential for the biogenesis of type IV pili in Neisseria gonorrhoeae. Mol. Microbiol. 18:975-986. [DOI] [PubMed] [Google Scholar]
- 14.Drake, S. L., S. A., Sandstedt, and M. Koomey. 1997. PilP, a pilus biogenesis lipoprotein in Neisseria gonorrhoeae, affects expression of PilQ as a high-molecular-mass multimer. Mol. Microbiol. 23:657-668. [DOI] [PubMed] [Google Scholar]
- 15.Dumpsey, J. F., W. Litaker, A. Madure, T. L. Snodgrass, and J. G. Cannon. 1991. Physical map of the chromosome of Neisseria gonorrhoeae FA1090 with locations of genetic markers, including opa and pil genes. J. Bacteriol. 173:5476-5486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dyer, D. W., E. P. West, and P. F. Sparling. 1987. Effects of serum carrier proteins on the growth of pathogenic neisseriae with heme-bound iron. Infect. Immun. 55:2171-2175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hagaman, K. E., W. Pan, B. G. Spratt, J. T. Balthazar, R. C. Judd, and W. M. Shafer. 1995. Resistance of Neisseria gonorrhoeae to antimicrobial hydrophobic agent is modulated by the mtrCDE efflux system. Microbiology 141:611-622. [DOI] [PubMed] [Google Scholar]
- 18.Hansen, M. V., and C. E. Wilde III. 1984. Conservation of peptide structure of outer membrane protein-macromolecular complex from Neisseria gonorrhoeae. Infect. Immun. 43:839-845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lee, B. C. 1992. Isolation of haemin-binding proteins of Neisseria gonorrhoeae. J. Med. Microbiol. 36:121-127. [DOI] [PubMed] [Google Scholar]
- 20.Lee, B. C., and S. Levesque. 1997. A monoclonal antibody directed against the 97-kilodalton gonococcal hemin-binding protein inhibits hemin utilization by Neisseria gonorrhoeae. Infect. Immun. 65:2970-2974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lewis, L. A., and D. W. Dyer. 1995. Identification of an iron-regulated outer membrane protein of Neisseria meningitidis involved in the utilization of haemoglobin complexed to haptoglobin. J. Bacteriol. 177:1299-1306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lewis, L. A., M. Gipson, K. Hartman, T. Ownbey, J. Vaughn, and D. W. Dyer. 1999. Phase variation of HpuAB and HmbR, two distinct hemoglobin receptors of Neisseria meningitidis DNM2. Mol. Microbiol. 32:977-989. [DOI] [PubMed] [Google Scholar]
- 23.Lewis, L. A., E. Gray, Y.-P. Wang, B. A. Roe, and D. W. Dyer. 1997. Molecular characterization of hpuAB, the haemoglobin-haptoglobin-utilization operon of Neisseria meningitidis. Mol. Microbiol. 23:737-749. [DOI] [PubMed] [Google Scholar]
- 24.Lewis, L. A., M.-H. Sung, M. Gipson, K. Hartman, and D. W. Dyer. 1998. Transportation of intact porphrin by HpuAB, the hemoglobin-haptoglobin utilization system of Neisseria meningitidis. J. Bacteriol. 180:6043-6047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Long, C. D., R. N. Madraswala, and H. S. Seifert. 1998. Comparisons between phase variations of Neisseria gonorrhoeae FA1090 and pilus, pilin and S-pilin expression. Infect. Immun. 66:1918-1927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Long, C. D., D. M. Tobiason, M. Lazio, K. A. Klin, and H. S. Seifert. 2003. Low level pilin expression allows for substantial DNA transformation competence in Neisseria gonorrhoeae. Infect. Immun. 71:6279-6291. [DOI] [PMC free article] [PubMed]
- 27.Maier, B., L. Potter, M. So, H. S. Seifert, and M. P. Sheetz. 2002. Single pilus motor forces exceed 100 pN. Proc. Natl. Acad. Sci. USA 99:16012-16017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Marciano, D. K., M. Russel, and S. M. Simon. 1999. An aqueous channel for filamentous phage export. Science 284:1516-1519. [DOI] [PubMed] [Google Scholar]
- 29.Marciano, D. K., M. Russel, and S. M. Simon. 2001. Assembling filamentous phage occlude pIV channels. Proc. Natl. Acad. Sci. USA 98:9359-9364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Merz, J. A., M. So, and M. P. Sheetz. 2000. Pilus retraction powers bacterial twitching motility. Nature 407:98-102. [DOI] [PubMed] [Google Scholar]
- 31.Nouwen, N., N. Ranson, H. Saibil, B. Wolpensinger, A. Engel, A. Ghazi, and A. P. Pugsley. 1999. Secretin PulD: association with pilot PulS, structure, and ion-conducting channel formation. Proc. Natl. Acad. Sci. USA 96:8173-8177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Prentki, P., and H. M. Kirsch. 1984. In vitro insertional mutagenesis with a selectable DNA fragment. Gene 29:303-313. [DOI] [PubMed] [Google Scholar]
- 33.Rouguette-Loughlin, C., S. A. Dunham, M. Kuhn, J. T. Balthazar, and W. M. Shafer. 2003. The NorM efflux pump of Neisseria gonorrhoeae and Neisseria meningitidis recognizes antimicrobial cationic compounds. J. Bacteriol. 185:1101-1106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Russel, M. 1998. Macromolecular assembly and secretion across the bacterial cell envelope: type II protein secretion systems. J. Mol. Biol. 279:485-499. [DOI] [PubMed] [Google Scholar]
- 35.Schryvers, A. B., and I. Stojiljkovic. 1999. Iron acquisition systems in the pathogenic Neisseria. Mol. Microbiol. 32:1113-1117. [DOI] [PubMed] [Google Scholar]
- 36.Seifert, H. S. 1997. Insertionally inactivated and inducible recA6 alleles for use in Neisseria. Gene 188:215-220. [DOI] [PubMed] [Google Scholar]
- 37.Seifert, H. S., R. S. Ajioka, D. Paruchuri, F. Heffron, and M. So. 1990. Shuttle mutagenesis of Neisseria gonorrhoeae: pilin null mutation lower DNA transformation competence. J. Bacteriol. 172:40-46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Shafer, W. M., W. L. Veal, E.-H. Lee, L. Zarantonelli, J. T. Balthazar, and C. Rouguette-Loughlin. 2001. Genetic organization and regulation of antimicrobial efflux systems possessed by Neisseria gonorrhoeae and Neisseria meningitidis. J. Mol. Microbiol. Biotechnol. 3:219-224. [PubMed] [Google Scholar]
- 39.Stein, D. C. 1991. Transformation of Neisseria gonorrhoeae: physical requirements of the transforming DNA. Can. J. Microbiol. 37:345-349. [DOI] [PubMed] [Google Scholar]
- 40.Stojiljkovic, I., B. D. Evanvold, and V. Kumar. 2001. Antimicrobial properties of porphyrins. Expert Opin. Investig. Drugs 10:309-320. [DOI] [PubMed] [Google Scholar]
- 41.Stojiljkovic, I., V. Hwa, L. D. S. Martin, P. O'Gaora, X. Nassif, F. Heffron, and M. So. 1995. The Neisseria meningitidis hemoglobin receptor: its role in iron utilization and virulence. Mol. Microbiol. 15:531-541. [DOI] [PubMed] [Google Scholar]
- 42.Stojiljkovic, I., J. Larson, V. Hwa, S. Anic, and M. So. 1996. HmbR outer membrane receptors of pathogenic Neisseria spp.: iron-regulated, hemoglobin-binding protein with a high level of primary structure conservation. J. Bacteriol. 178:4670-4678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Stojiljkovic, I., and D. Perkins-Balding. 2002. Processing of heme and heme-containing proteins by bacteria. DNA Cell Biol. 21:281-295. [DOI] [PubMed] [Google Scholar]
- 44.Stojiljkovic, I., and N. Srinivasan. 1997. Neisseria meningitidis tonB, exbB, and exbD genes: Ton-dependent utilization of protein-bound iron in neisseriae. J. Bacteriol. 179:805-812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Swanson, J. 1973. Studies on gonococcus infection. IV. Pili: their role in attachment of gonococci to tissue culture cells. J. Exp. Med. 137:571-589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Thomas, C. E., N. H. Carbonetti, and P. F. Saprling. 1996. Pseudo-transposition of a Tn5 derivative in Neisseria gonorrhoeae. FEMS Microbiol. Lett. 145:371-376. [DOI] [PubMed] [Google Scholar]
- 47.Tsai, W.-M., S. H. Larsen, and C. E. Wilde III. 1989. Cloning and DNA sequence of the omc gene encoding the outer membrane protein-macromolecular complex from Neisseria gonorrhoeae. Infect. Immun. 57:2653-2659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Turner, P. C., C. E. Thomas, C. Elkins, S. Clary, and P. F. Sparling. 1998. Neisseria gonorrhoeae heme biosynthesis mutants utilize heme and hemoglobin as a heme source but fail to grow within epithelial cells. Infect. Immun. 66:5215-5223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Weinberg, E. D. 1978. Iron and infection. Microbiol. Rev. 42:45-66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wolfgang, M., P. Lauer, H. P. Park, L. Brossay, J. Herbert, and M. Koomey. 1998. PilT mutations lead to simultaneous defects in competence for natural transformation and twitching motility in piliated Neisseria gonorrhoeae. Mol. Microbiol. 29:321-330. [DOI] [PubMed] [Google Scholar]
- 51.Wolfgang, M., H. P. Park, S. F. Hayes, J. P. M. van Putten, S. F. Hayes, and M. Koomey. 1998. Suppression of an absolute defect in type IV pilus biogenesis by loss-of-function mutations in pilT, a twitching motility gene in Neisseria gonorrhoeae. Proc. Natl. Acad. Sci. USA 95:14973-14978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wolfgang, M. J. P. M. van Putten, S. F. Hayes, D. Dorward, and M. Koomey. 2000. Component and dynamics of fiber formation define a ubiquitous biogenesis pathway of bacterial pili. EMBO J. 19:6408-6418. [DOI] [PMC free article] [PubMed] [Google Scholar]