Summary
The assembly of synapses and neuronal circuits relies on an array of molecular recognition events and their modification by neuronal activity. Neurexins are a highly polymorphic family of synaptic receptors diversified by extensive alternative splicing. Neurexin variants exhibit distinct isoform-specific biochemical interactions and synapse assembly functions but the mechanisms governing splice isoform choice are not understood. We demonstrate that Nrxn1 alternative splicing is temporally and spatially controlled in the mouse brain. Neuronal activity triggers a shift in Nrxn1 splice isoform choice via calcium/calmodulin-dependent kinase IV signaling. Activity-dependent alternative splicing of Nrxn1 requires the KH-domain RNA binding protein SAM68 which associates with RNA response elements in the Nrxn1 pre-mRNA. Our findings uncover SAM68 as a key regulator of dynamic control of Nrxn1 molecular diversity and activity-dependent alternative splicing in the central nervous system.
Introduction
Neuronal activity-dependent signaling pathways play a central role in circuit assembly and plasticity processes in the brain (Wong and Ghosh, 2002; Davis, 2006). Neurons modify their molecular repertoire in response to cellular depolarization through transcriptional programs and fine-tuning of protein turnover (Cohen and Greenberg, 2008; Bingol and Sheng, 2011). More recently, activity-dependent alternative splicing has emerged as an additional mechanism for the dynamic modification of neuronal function where production of specific splice variants modifies cellular trafficking, signaling properties or function of synaptic proteins (An and Grabowski, 2007; Li et al., 2007b).
Signaling pathways for activity-dependent splicing regulation are only beginning to be uncovered. Calcium/calmodulin-dependent kinase IV (CaMKIV) activation modifies alternative splicing in cultured neuronal cells. Some pre-mRNAs contain CaMKIV-responsive-elements (CaRRE1 and 2) and UAGG motifs which impose depolarization-dependent regulation (Xie and Black, 2001; An and Grabowski, 2007; Lee et al., 2007). However, these RNA elements are not found in all pre-mRNAs subject to depolarization-dependent alternative splicing and additional response elements remain to be uncovered.
One class of synaptic cell surface receptors that is extensively regulated at the level of alternative splicing are the Neurexins (Ushkaryov et al., 1992). Neurexins contribute to the assembly of functional presynaptic terminals (Dean et al., 2003; Missler et al., 2003; Li et al., 2007a) and mutations in the human neurexins are associated with neurodevelopmental disorders (Sudhof, 2008; Rujescu et al., 2009). Three neurexin genes (Nrxn1,2,3) each encode two primary neurexin transcripts (alpha and beta) that are subject to alternative splicing at five positions yielding more than 3,000 protein-coding mRNAs (Rowen et al., 2002; Tabuchi and Sudhof, 2002; Baudouin and Scheiffele, 2010). This molecular diversity as well as the capability of neurexins to organize synaptic structures led to the hypothesis that alternative splicing may give rise to cell type and/or synapse-specific functions (Chih et al., 2006; Craig and Kang, 2007; Huang and Scheiffele, 2008).
Most attention of functional studies has focused on the Nrxn alternatively spliced segment 4 (AS4), which is highly conserved in all Nrxn pre-mRNAs. Incorporation of exon 20 at AS4 generates NRX 4(+) protein variants containing a 30 amino acid insertion, whereas skipping results in the 4(−) variant. Importantly, alternative splicing of Nrxns may underlie an adhesive code: The NRX 4(+) and 4(−) variants exhibit differential interactions with several ligands that are critical mediators of synaptogenesis, including neuroligins, leucine-rich repeat proteins (LRRTMs), and the Cbln1-GluD2 complex (Ichtchenko et al., 1995; Boucard et al., 2005; Chih et al., 2006; de Wit et al., 2009; Koehnke et al., 2010; Siddiqui et al., 2010; Uemura et al., 2010; Matsuda and Yuzaki, 2011). For example, the NRX1 4(+) isoform exhibits only weak adhesion and binding with the neuroligin-1B postsynaptic receptor (NL1B), whereas the NRX1 4(−) isoform engages in strong adhesive interactions (Dean et al., 2003; Chih et al., 2006; Reissner et al., 2008). Therefore, alternative splicing at AS4 controls a fundamental switch in neurexin function at synapses.
The critical functional consequences of Nrxn alternative splicing at AS4 raise the question of how alternative splice isoform choice is regulated. Genome-wide identification of mRNA targets for Nova or nPTB proteins has not yielded insights into Nrxn1 AS4 regulation {Boutz, 2007 #1988; Ule, 2005 #1375}. Other studies reported modifications of splicing patterns after long-term manipulation of neuronal activity and in response to fear conditioning (Rozic-Kotliroff and Zisapel, 2007; Kang et al., 2008; Rozic et al., 2011). However, the RNA response elements and RNA-binding proteins conferring alternative splice isoform choice in Nrxns have not been identified. In fact, the RNA-binding proteins controlling activity-dependent alternative splicing of any neuronal pre-mRNA are not understood. Here, we describe a signaling pathway and RNA response elements for Nrxn1 and demonstrate that the KH-domain RNA-binding protein SAM68 (Src-associated in mitosis 68 kDa protein) is a key regulator of activity-dependent alternative splicing in the CNS.
Results
Neurexin alternative splicing is regulated by neuronal activity
Incorporation of the cassette exon at AS4 (exon 20) in Nrxn1,2,3 varies across brain regions (Figure 1A, Figure S1). Moreover, for some regions exon 20 inclusion differs between the three Nrxns indicating gene-specific regulation. In the developing cerebellar cortex Nrxn 4(−) mRNAs declines successively from postnatal day 0 to postnatal day 21 (Figure 1B). This reveals spatial and developmental regulation of Nrxn alternative splicing in the mouse brain.
Figure 1. Depolarization-dependent alternative splicing of Neurexins in cerebellar neurons.
(A) Schematic diagram outlining organization of Neurexin alpha and neurexin beta protein variants. The positions of alternatively spliced segments (AS1,2,3,4,5) are indicated by arrows. LamininG-domains are shown as ovals, EGF-domains as rectangles, black line is the trans-membrane domain. The cartoon on the right illustrates the 90bp cassette exon (20) at AS4. Position of primers used for analysis indicated by arrows.
(B) Developmental regulation of Nrxns at AS4 in mouse cerebellum (n=3).
(C) Dissociated cultures of mouse cerebellar neurons maintained in vitro for 14 days were depolarized by addition of 25 mM KCl for 6 hours. Semi-quantitative PCR analysis with primers for AS4, c-Fos or Gapdh.
(D) Quantitative real-time PCR for total Nrxn1, Nrxn1 4(+) and Nrxn1 4(−). Nrxn mRNA / Gapdh values in untreated control cells were set to 1.0 and compared to cells stimulated (25 mM KCl) for 6 hours (n=4).
(E) Analysis of semi-quantitative PCR data from panel C. The fraction of Nrxn 4(−) compared to total Nrxn is plotted (n=4).
(F) The glutamate receptor agonist kainic acid (50 µM) was applied for 6 hours in presence or absence of the AMPA/kainate receptor antagonist CNQX (50 µM) (n=4–5 independent cultures). Electrical field stimulation was applied in three trains (3 min of 200Hz) spaced by 7 mins and RNA was isolated 5.5 hrs after stimulation (n=6–8 independent cultures per condition).
Mean and SEM. See also Figure S1 and S2.
The changes in Nrxn splicing regulation during the first two postnatal weeks parallel the developmental shift in resting membrane potential of cerebellar granule cells (−25 mV to −55 mV)(Rossi et al., 1998). To test whether alternative splicing is altered by membrane depolarization we stimulated cultured granule cells by addition of 25 mM potassium chloride which shifts the membrane potential to approximately −36 mV (Mellor et al., 1998) (Figure 1C–E). After 6 hrs the abundance of Nrxn1 4(−) mRNA was increased nearly tenfold as measured by quantitative PCR (n= 4, p<0.01, Figure 1D). This was accompanied by a decrease in the Nrxn1 4(+) and total Nrxn1 transcript indicating that in addition to the increase of Nrxn1 4(−) also the turnover of the pre-existing Nrxn1 mRNA might be altered by depolarization (Figure 1D). Using a semi-quantitative PCR assay with flanking primers we found that transcripts from all three neurexin genes (Nrxn1,2,3) each responded similarly to depolarization (n= 4, p<0.001, Figure 1C,E). The shift in Nrxn1 alternative splicing was detectable after 30 minutes, and was reversible, as Nrxn1 4(−) levels returned to baseline 48 hrs after stimulation (Figure S2A,B).
To explore whether alternative splicing was controlled by neuronal activity we stimulated glutamate receptors by application of kainic acid (50 µM for 6hrs) or applied electrical field stimulation (3 × 3min at 200Hz spaced by 7 min intervals). Both paradigms produced a significant increase in Nrxn1 and Nrxn2 4(−) mRNA (for electrical stimulation n≥7, p<0.05, Figure 1F). Nrxn3 4(−) was not significantly up-regulated in response to electrical stimulation. The activity-dependent alternative splicing shift in Nrxn1 AS4 was further confirmed by quantitative PCR (Rq Nrxn1 4(−)/Gapdh increased 2.4 ± 0.4 fold compared to non-stimulated control, n=6, p=0.004). Importantly, depolarization did not significantly alter exon incorporation at the neighboring alternatively spliced segments 3 and 5 in Nrxn1 (Figure S2C). This highlights site-selective activity-dependent splicing regulation for Nrxn1 AS4 in cerebellar neurons.
We assessed the effects of pharmacological inhibitors to obtain insight into the signaling mechanisms underlying the regulation of Nrxns at AS4. Blockade of voltage-gated calcium channels by 20 µM Cd2+ or the L-type voltage-gated calcium channel blocker nifedipine abolished the depolarization-dependent splicing shifts (n=4, p<0.01, Figure 2A). The depolarization-induced increase in the Nrxn1 4(−) isoform was abolished by application of the CaMK inhibitors KN-62 and KN-93 (n=5, p<0.01, Figure 2B). By contrast, inhibitors of the MAP-Kinases MEK1 and MEK2 (U0126) and calcineurin (FK506) had no significant effect on depolarization-dependent Nrxn splicing regulation (Figure S2D). When cells were treated with actinomycin D during stimulation no increase in the Nrxn 4(−) splice variant was observed, demonstrating a requirement for ongoing transcription (Figure S2D). KN-62 and KN-93 inhibit CaMKs but also affect other kinases and neuronal receptors (Wayman et al., 2008). Therefore, we further examined the involvement of the CaMK pathway by inhibiting CaMKKs with STO-609 (Tokumitsu et al., 2002) or by RNA interference knock-down of CaMKIV, the dominant nuclear CaMK isoform in granule neurons (Ohmstede et al., 1989). Both treatments reduced depolarization-induced alternative splicing (n≥7, STO609: p<0.01, siCaMKIV: p=0.02, Figure 2B). These experiments uncover a depolarization-induced, calcium- and CaMKIV-dependent mechanism for alternative splicing of Nrxn1.
Figure 2. Depolarization-dependent alternative splicing of neurexin requires CaMKIV and controls trans-synaptic signaling.
(A) Dissociated cultures of mouse cerebellar neurons (DIV14 days) were depolarized (25 mM KCl) for 6 hours and CdCl2 (20 µM) or nifedipine (10 µM) were added as indicated. Nrxn 4(−) levels were quantified by semi-quantitative PCR (n=4).
(B) CaMK signaling in depolarization-dependent alternative splicing of Nrxn1 was inhibited by co-application of pharmacological inhibitors or RNAi knockdown of CaMKIV (n=4–7). c-Fos activation was probed by RT-PCR to confirm immediate-early gene induction.
(C) Schematic diagram illustrating binding preferences of neurexin splice isoforms. Beta-NRX1 4(−) interacts similarly with the NL1A and NL1B splice variants, the 4(+) insertion in the beta-NRX1 4(+) variant (red insert) prevents strong adhesive interactions with NL1B but still allows for interactions with NL1A. The Cbln1/GluD2 complex specifically interacts with the 4(+) insertion in beta-NRX1 4(+).
(D) HEK293 cells co-expressing GFP and NL1B (left) or GluD2 (right) were added to cerebellar neurons at DIV9 and analyzed 2 days later. For depolarizing conditions, 25mM KCl was added to the culture medium at DIV 6. Presynaptic differentiation assessed by vGluT1 staining.
(E) HEK293 cells co-expressing GFP and NL1B (left) or NL1A (right) were added at DIV7 under depolarizing or control conditions. Pharmacological inhibitors (5 µM STO609 or 10 µM FK605) were applied at DIV 6.
(F) Morphometric analysis of vGluT1 staining at neuron-HEK293 contact sites analyzing the fraction of the vGluT1-positive area of the GFP-positive neuroligin-expressing cell (n>30 cells per condition, 3 independent experiments).
Mean and SEM. Scale bars are 5µm. See also Figure S2.
Depolarization-dependent alternative splicing alters response to synaptogenic receptors
Alternative splicing of Nrxns at AS4 regulates adhesive interactions with synaptic receptors (Figure 2C). NRX1 4(−) but not NRX1 4(+) forms strong trans-synaptic adhesion complexes with the neuroligin-1B receptor (NL1B) (Dean et al., 2003; Boucard et al., 2005; Chih et al., 2006). Conversely, NRX1 4(+) but not NRX1 4(−) binds the synaptic Cbln1-GluD2 complex (Uemura et al., 2010; Matsuda and Yuzaki, 2011). To test whether the depolarization-induced alternative splicing switch in Nrxns translates into altered trans-synaptic signaling we employed a co-culture system where primary neurons are combined with non-neuronal cells expressing a single postsynaptic receptor isoform (Scheiffele et al., 2000). NL1B-expressing HEK293 cells induced only low levels of presynaptic differentiation as measured by the accumulation of the synaptic vesicle marker vGluT1 (Figure 2D). By contrast, GluD2-expressing cells triggered robust recruitment of this marker. These findings are consistent with weak functional coupling between NL1B and NRX1 4(+), the primary Nrxn1 isoform expressed in unstimulated granule cells, and strong coupling of NRX1 4(+) to the Cbln1-GluD2 complex. When granule cells were maintained under depolarizing conditions, the response to NL1B-expressing cells was strongly increased and responses to GluD2 were reduced (Figure 2D, F and Figure S2E). Interestingly, this reduction in GluD2 responses is likely due to both, a change in Nrxn1 alternative splicing but also reduced expression of Cbln1, a critical component of the tripartite NRX-Cbln1-GluD2 complex (Figure S2E and see (Iijima et al., 2009)).
We further examined the mechanisms of altered NL1B responses in this assay. Inhibition of the CaMK pathway by addition of STO609 suppressed not only Nrxn1 4(−) production but also depolarization-induced responsiveness to NL1B (n=35 cells per condition, p<0.001, Figure 2E,F). By contrast, inhibition of calcineurin, another calcium-dependent signaling molecule activated in depolarized granule cells had no effect on NL1B-induced presynaptic differentiation (FK506, Figure 2E,F). To confirm that the altered presynaptic assembly in depolarized cells was indeed due to neurexin-receptor binding specificities we compared the induction of presynaptic structures with the NL1A isoform which exhibits similar adhesion and binding affinities for both, NRX1 4(+) and NRX1 4(−) (Boucard et al., 2005; Chih et al., 2006; Koehnke et al., 2010). Presynaptic differentiation induced by NL1A did not differ significantly between non-depolarized and depolarized granule cells (n=37 cells per condition, p=0.29, Figure 2E,F), highlighting that the depolarization-induced increase in presynaptic assembly in response to NL1 is dependent on the identity of the postsynaptic neuroligin receptor isoform.
SAM68 regulates Nrxn1 AS4 alternative splicing in vitro
To identify RNA-binding proteins that regulate Nrxn1 alternative splicing we established an assay in heterologous cells. A splice reporter spanning Nrxn1 AS4 (Nrxn1–4, exon 19 – 21, Figure 3A) primarily produced the Nrxn1 4(+) splice isoform when transfected into HEK293 cells, therefore, providing an assay to identify proteins that promote exon 20 skipping. A survey of the intronic sequences flanking exon 20 revealed AU-rich sequence elements reminiscent of binding sites for the RNA-binding protein SAM68 (Lin et al., 1997; Galarneau and Richard, 2009) Therefore, we examined whether SAM68 could regulate alternative splicing of the Nrxn1–4 reporter. In addition, we tested several other RNA proteins implicated in neuronal alternative splicing (Nova-1, Nova-2, nPTB, hnRNPA1, hnRNPH) (Matter et al., 2002; Ule et al., 2005; Chawla et al., 2009; Fisette et al., 2010). Of these proteins examined, only SAM68 effectively shifted the reporter splicing pattern to produce Nrxn1 4(−) (n=3, p<0.001, Figure 3B). SAM68 is a member of the STAR (“signal transduction and activation of RNA”) family of RNA binding proteins containing a single KH-domain (Volk et al., 2008). SLM1 and SLM2 are two SAM68-like mammalian proteins, which share 70–80% amino acid sequence identity with SAM68 in their RNA-binding domains (Di Fruscio et al., 1999). Both, SLM1 and SLM2 exhibited similar activities as SAM68 in shifting alternative splicing to the 4(−) isoform (Figure 3B). SAM68 function in this assay was dose-dependent as increasing amounts of SAM68 resulted in increased exclusion of exon 20 (Figure 3C).
Figure 3. SAM68-dependent alternative splicing of Neurexins in vitro.
(A) Schematic diagram of Nrxn1–4 splice reporter construct containing AS4 with constitutive exons (orange), alternative exon 20 (red), and introns shown as lines. Two AU-rich sequence stretches are highlighted with arrows. Intron 19 (>40 kbp) was truncated leaving >550 bp adjacent to the splice donor and the acceptor sites intact.
(B) Splice reporter expression vector was co-transfected into HEK293 cells with negative control DNA or epitope-tagged expression constructs for GFP-SAM68, T7-Nova1, T7-Nova2, Xpress-nPTB, YFP-hnRNPA1, YFP-hnRNPH1, GFP-SLM1, GFPSLM2. Alternative splice isoform choice was measured by semi-quantitative RT-PCR with primers flanking the alternative splice site (n=3). Expression of all transfected RNA-binding proteins was confirmed by Western blotting with anti-GFP, -T7 or -Xpress antibodies. Molecular weights in kDa.
(C) Co-transfection of increasing amounts of a SAM68 expression vector with the Nrxn1–4 splice reporter. Alternative splicing was assayed by semi-quantitative RT-PCR and HA-tagged SAM68 detected by immunoblot.
(D) GFP-tagged point mutants in SAM68 KH-domain and nuclear localization signal were co-expressed with the Nrxn1-4 splice reporter and alternative splicing was analyzed by semi-quantitative RT-PCR. Protein expression levels were compared using western blotting.
(E) Co-transfection of plasmids encoding constitutively active Fyn-kinase and Nrxn1–4 reporter.
Mean and SEM.
SAM68 has been characterized as a nuclear RNA-binding protein but also non-nuclear functions have been described (Grange et al., 2004; Henao-Mejia et al., 2009; Huot et al., 2009). SAM68 point mutants (I184N and G178E) that abolish RNA-binding (Lin et al., 1997; Chawla et al., 2009) or nuclear targeting (Y440F) had strongly reduced function in the splice reporter assay (Figure 3D). Similarly, SAM68 function was inhibited when SAM68 was co-transfected with constitutively active Fyn kinase (Figure 3E) which inactivates the nuclear-localization signal of SAM68 (Paronetto et al., 2007). Thus, SAM68-mediated splicing regulation of Nrxn1 AS4 relies on RNA-binding and SAM68 transport to the nucleus.
Nrxn1 contains multiple intronic SAM68-response elements
We sought to identify the RNA elements which control SAM68-dependent alternative splicing of Nrxn1. Deletion analysis in intron 19 and intron 20 identified sequences in both introns that significantly reduce SAM68-induced skipping of exon 20 when deleted from the reporter construct (Figure 4A and S3). A maximum reduction in SAM68-dependent splicing shifts was obtained with deletions 19-c and 20-c, respectively (Figure 4A). When deletion 19-c and deletions in intron 20 were combined (constructs 19/20-a,b,c,d), SAM68-induced exon skipping was completely abolished for constructs 19/20-c and 19/20-d (n=4, p<0.001, Figure 4A). This suggests that multiple intronic SAM68-response elements located in introns 19 and 20 cooperate in regulating Nrxn1 AS4. Notably, the apparent SAM68-response element between deletion sites 20-b and 20-c contains multiple AU-rich sequence elements conforming to SAM68 RNA binding sites in other pre-mRNAs (Lin et al., 1997; Galarneau and Richard, 2009). Therefore, we tested binding of SAM68 to synthetic RNA-oligonucleotides (probes A and B) each spanning 30 bases of the presumptive binding region (Figure 4B). In pull-down assays, probe A yielded efficient binding of endogenous SAM68 from brain extracts whereas only background binding was observed for probe B (Figure 4B). SLM1 and SLM2 exhibited similar interaction with probe A as SAM68. Importantly, under the same conditions, SF2 and NeuN/Fox3, two other RNA binding proteins (Kim et al., 2009) were not recovered in the precipitates. Mutation of two nucleotides in a presumptive SAM68 binding site of probe A (probe A*) (Figure 4B) significantly reduced the recovery of SAM68 in the pull-down experiment (Figure 4B). Therefore, in heterologous cells, SAM68 promotes skipping of exon 20 through association with intronic SAM68 response elements in the Nrxn1 pre-mRNA.
Figure 4. Mapping of SAM68-response elements in Nrxn1.
(A) Intronic sequences preceding (19-a, 19-b, 19-c) or following exon 20 (20-a to 20-d) were deleted from Nrxn1–4. Intron 19-c deletion was combined with deletions 20-a to 20-d in intron 20. Panels show semi-quantitative PCR results from HEK293 cells without and with co-expression of SAM68 and Nrxn1–4, respectively. Quantitation was performed for products in presence of SAM68 (n=4).
(B) Biotinylated RNA-oligonucleotides probes covering intronic sequence between 20-b and 20-c deletions used in pull-down experiments with mouse adult brain extracts. Probe A* contains two nucleotide changes (red). Bound proteins were analyzed by western blotting with anti-SAM68, anti-SLM1, anti-SLM2, anti-Fox3/NeuN, and anti-SF2/ASF antibodies. SAM68 binding was quantified by densitometric scanning of Western blot signals (n=4).
(C) Splice reporters were delivered into cortical pyramidal cell precursors of E14.5 embryos by in utero electroporation and reporter processing was examined at P6. Reporter expression levels were examined with primers specific for constitutive reporter sequence elements and RNA input levels were monitored by semi-quantitative PCR for endogenous Gapdh (n=3–6).
Mean and SEM. See also Figure S3.
Next we developed an in vivo RNA Response Element assay (iRRE) to test whether the SAM68 response elements are essential for Nrxn1 AS4 splicing regulation in the intact nervous system. We delivered the Nrxn1–4 reporters specifically into mouse cortical pyramidal cells in vivo by in utero electroporation followed by analysis at postnatal day six. Histochemical and semi-quantitative PCR analysis of cortical sections confirmed similar levels of mutant and wild-type splice reporter expression in cortical layer 2/3 pyramidal cells (Figure 4C). Alternative splicing of the Nrxn1–4 reporter was comparable to the products detected for endogenous Nrxn1 in total cortical RNA samples (compare Figure S1). Notably, deletion of the SAM68-response elements abolished Nrxn1 4(−) production in the iRRE assay (n=3 to 6 animals per construct, p<0.01 and 0.001 for 19-c and 20-c, respectively; Figure 4C). This demonstrates that the intronic response elements are essential for Nrxn1 4(−) production in the context of the endogenous splicing machinery in cortical pyramidal cells in vivo.
Cell type-specific expression of STAR proteins in the mouse cerebellum
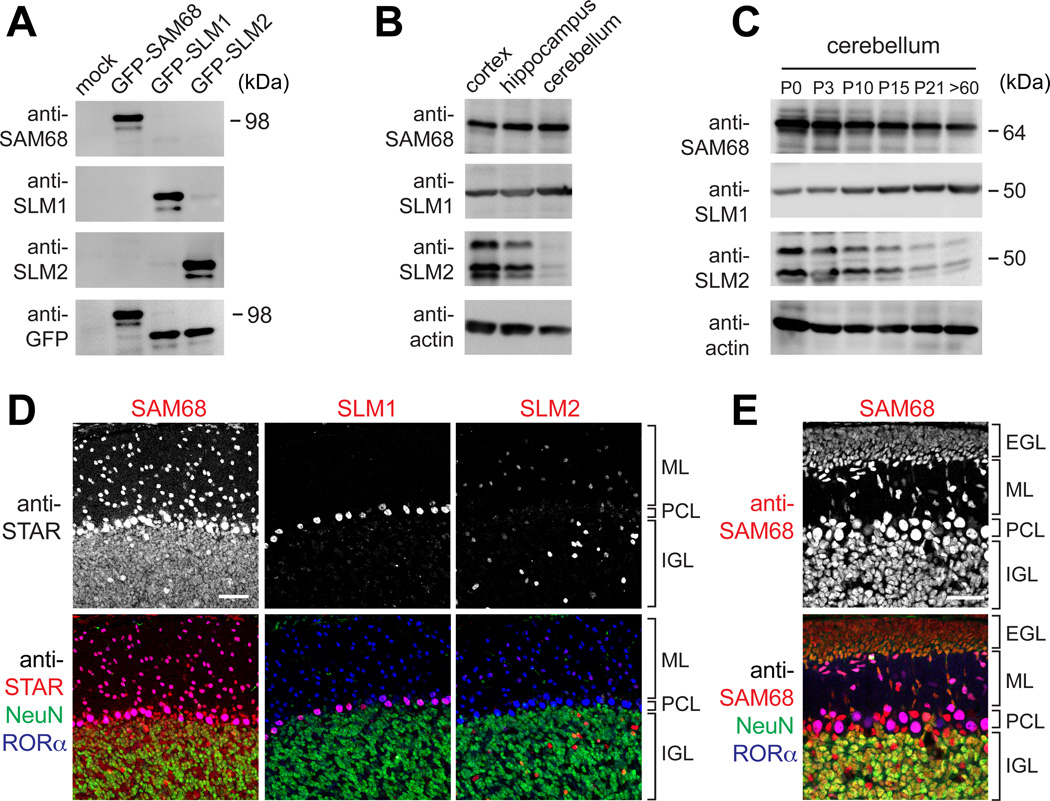
To begin to examine whether endogenous STAR proteins are critical regulators of Nrxn1 alternative splicing in vivo we examined SAM68, SLM1, and SLM2 expression patterns. Affinity-purified antibodies specifically recognized the respective STAR family proteins in Western blots and identified significant expression across various brain areas (Figure 5A,B). In the developing cerebellum SAM68 immune-reactivity was high at birth and slightly declined over the first three weeks. SLM1-immune-reactivity increased over the first three postnatal weeks whereas SLM2 protein levels decreased (Figure 5C). Notably, SAM68, SLM1 and SLM2 differed significantly in their cellular expression patterns in the cerebellar cortex (Figure 5D). SAM68 was detected in most neuronal cell populations with high immune-reactivity in cerebellar granule cells of the internal granular layer (IGL), RORα-positive Purkinje cells, as well as IGL and molecular layer interneurons. SLM1 immunoreactivity was predominantly observed in Purkinje cells, whereas SLM2 antibodies stained selectively IGL and (less significantly) molecular layer interneurons (Figure 5D). Considering that of all STAR family proteins only SAM68 was significantly expressed in cerebellar granule cells (the major source of cerebellar mRNA) we further examined its distribution during development. At postnatal day 7, SAM68 was highly expressed in granule cell precursor cells in the EGL (external granular layer) and mature granule cells of the IGL (Figure 5E). Therefore, SAM68 is well positioned to regulate Nrxn1 alternative splicing in granule cells during development and in the adult cerebellum.
Figure 5. Cell-type specific STAR protein expression in the cerebellum.
(A) Specificity of anti-STAR protein antibodies confirmed by probing lysates from HEK293 cells overexpressing GFP-tagged SAM68, SLM1, or SLM2.
(B) Immunoblot of adult cortical, hippocampal, and cerebellar tissue extracts probed with STAR protein antibodies.
(C) Developmental regulation of STAR protein expression in the cerebellum from postnatal day 0 (P0) to adult (>P60).
(D,E) Para-sagittal sections through adult cerebellum triple stained with anti-NeuN, anti-RORα, and anti-SAM68, anti-SLM1, or anti-SLM2 antibodies. In E, P7 cerebellum was analyzed using anti-SAM68 antibodies. ML, molecular layer; PCL, Purkinje cell layer; IGL, internal granular layer; EGL, external granular layer.
Scale bars are 50 µm. See also Figure S4.
SAM68 regulates Nrxn1 alternative splicing in vivo
To explore how loss of SAM68 affects neuronal alternative splicing regulation we examined SAM68 knockout (KO) mice (Richard et al., 2005). In the mutant mice, SAM68 protein was undetectable whereas expression of SLM1 and SLM2 was not significantly altered (Figure 6A and Figure S4A). Notably, SAM68 KO mice exhibit defects in basal motor performance (Lukong and Richard, 2008). To further assess a potential impairment of cerebellar function in SAM68 knock-out mice we tested their motor learning on an accelerating rotarod. SAM68 KO mice performed significantly worse than wild-type mice (p<0.01). Behavioral defects persisted over a 7-day training period demonstrating that KO mice have significant motor coordination deficits (Figure 6B).
Figure 6. Cerebellar alterations in SAM68 KO mice.
(A) Low magnification view of para-sagittal section of brain stem and cerebellum reveals anti-SAM68 immune-reactivity which is lost in SAM68 KO (−/−) tissue.
(B) Motor performance of male control mice and SAM68 KO littermates were tested on an accelerating rotarod. The latency to fall is plotted for days 1,3, and 7 of training (n=5–8 mice per genotype).
(C) Immunoreactivity for synaptic vesicle proteins VGluT1 (magenta) and VAMP2 (green) in the internal granular layer (IGL) of adult wild-type and SAM68 KO cerebellar cortex. The right panel shows single rosettes at high magnification.
Scale bars are 20µm in (left) and 5µm in (right).
(D) Representative line scans from wild-type and SAM68 KO cerebellum displaying immunoreactivity for vGluT1 in mossy fiber rosettes in C. Cumulative probability of VGluT1 and VAMP2 average intensities in cerebellar glomeruli compared between wild-type (blue line) and SAM68 KO mice (orange) (n> 438 rosettes from 3 animals per genotype, Komolgorov-Smirnov test).
(E) Quantitative PCR performed on cortical, cerebellar, brainstem cDNAs from control (+/+, black) and SAM68 KO mice (−/−, white; n=3 animals per genotype). Abundance of total Nrxn1 mRNA, Nrxn1 4(+) and Nrxn1 4(−) transcripts is expressed compared to Gapdh (values for control tissue were set to 1.0).
Mean and SEM. See also Figure S5.
SAM68 KO cerebella do not show gross anatomical alterations as judged by histology (Figure 6A) and quantitative western blotting for a panel of synaptic markers (Figure S4B). However, synaptic glomeruli formed by mossy fiber axons showed an abnormal accumulation of the presynaptic vesicle marker vGluT1 (Figure 6C,D). Quantitative analysis of vGluT1 staining intensity revealed a shift in the distribution of mossy fiber rosettes towards higher staining intensities (n>438 rosettes per genotype, p<0.05, Komolgorov-Smirnov test, Figure 6D). By contrast, staining intensity for the presynaptic marker VAMP2 was unaltered (p>0.1, Komolgorov-Smirnov test, Figure 6D). Consistent with a significantly shifted distribution also the mean staining intensity for vGluT1 within mossy fiber rosettes was significantly increased in SAM68 KO brains (31±7% increase in the KO, n=5 animals per genotype, p<0.001). While the exact molecular and anatomical underpinnings of this synaptic phenotype remain to be defined, these findings indicate that SAM68 KO mice exhibit alterations in cerebellar mossy fiber synapses in vivo.
Alternative splicing of Nrxn1 at AS4 in SAM68 KO mice was severely perturbed. The abundance of the Nrxn1 4(−) form in the cerebellum was reduced fourfold at P7, a developmental stage where wild-type cerebellar cells express higher levels of Nrxn1 4(−) and threefold in the adult (n=3, p<0.001, Figure 6E). Importantly, the overall level of Nrxn1 mRNA was not altered indicating a change in alternative splicing rather than mRNA stability. Analysis of brainstem and cortical mRNAs confirmed a similar reduction in Nrxn1 4(−) and a corresponding increase in Nrxn1 4(+) (n=3, p<0.001, Figure 6E). There is accumulating evidence for coupling between transcription and splicing regulation resulting in promoter-specific splicing patterns (Batsche et al., 2006). However, the reduction in the Nrxn1 4(−) splice variant was detected for both, the Nrxn1 alpha and beta transcripts (Figure S4C). The splice isoform choice in the neighboring AS3 or alternative splicing of Nlgn1 and Nlgn2, which encode NRX1 binding partners, was unaltered in the SAM68 KO tissues analyzed (Figure S4C). Therefore, SAM68 is an essential regulator of Nrxn1 alternative splicing in vivo.
SAM68 controls activity-dependent splicing of Nrxn1
To explore whether SAM68 regulates activity-dependent Nrxn1 alternative splicing, we first assessed whether SAM68 is modified by neuronal depolarization. We did not observe any significant increase in SAM68 transcript, protein level, or change in sub-cellular localization in response to neuronal depolarization (Figure S5). Therefore, we explored post-translational modifications of SAM68. We analyzed SAM68 immunoprecipitated from control and depolarized cerebellar granule cells by tandem mass-spectrometry (Figure 7A). A phosphopeptide modified at serine 20 was eightfold enriched in the depolarized sample, whereas total SAM68 levels were unchanged (n=5, p<0.01 Figure 7B). Notably, serine 20 conforms to the consensus for CaMK-mediated phosphorylation (R-X-X-S/T) (Songyang et al., 1996) consistent with the regulation of SAM68 downstream of the CaMK pathway in response to neuronal activity.
Figure 7. SAM68 is essential for regulation of activity-dependent splicing of Nrxn1.
(A) Relative abundances of phosphorylated and non-phosphorylated SAM68 were assessed by liquid-chromatography-tandem mass spectrometry following SAM68-IP from control and depolarized cerebellar granule cells in culture.
(B) The CaMK consensus motif in the amino acid sequence surrounding ser 20 is highlighted in bold. Ratios of the normalized peptide abundances (depolarized / control) are shown (total SAM68 (depolarized/control) levels (phospho-serine 20: 8.39 ± 0.90, p < 0.01).
(C) Heterozygous control (+/−) or homozygous SAM68 KO neurons (−/−) were incubated in depolarizing medium for 6 hours and exon 20 inclusion was monitored by semi-quantitative PCR with flanking primers for Nrxn1, Nrxn2, or Nrxn3 (n=4). Loss of Sam68 expression was confirmed with oligonucleotides amplifying the Sam68 cDNA.
(D) Depolarization-dependent alternative splicing of the calcium-response element (CaRRe) containing pre-mRNAs Grin1 exon 5 and Kcnma1 STREX exon in control and SAM68 KO neurons (n=4).
(E) Neuronal activity-dependent alternative splicing in vivo examined by stereotaxic injection of 50 µM kainic acid (or saline as control) into the mouse cerebellum. Animals were sacrificed 5 hours after injection. Neuronal stimulation was confirmed by the increase in the expression of c-Fos mRNA in the injected ipsilateral versus non-injected contralateral side by semi-quantitative (middle panel) and quantitative PCR (right panel, Rq(c-Fos/Gapdh) for contralateral side set to 1.0, log scale, n=3 animals).
(F) Inclusion of exon 20 in Nrxn1 mRNA probed using quantitative PCR on cDNA derived from the injected cerebella (n=5 animals per genotype). Rq(Nrxn/Gapdh) values for the ipsilateral (injected) hemisphere (white columns) are expressed relative to the Rq(Nrxn/Gapdh) values obtained for the uninjected contralateral side from the same animals (black columns).
Mean and SEM. See also Figure S6.
To test whether SAM68 is required for depolarization-dependent alternative splicing regulation in cerebellar granule cells, cultures from control (SAM68 +/−) and knock-out mice (SAM68 −/−) were stimulated followed by transcript analysis. Control neurons (SAM68+/−) exhibited a significant up-regulation of the Nrxn1 4(−) splice variant but in SAM68 −/− neurons the depolarization-dependent alternative splicing shift at Nrxn1 and Nrxn3 AS4 was impaired (n=4 per genotype p<0.01, Figure 7C). Loss of depolarization-dependent alternative splicing was further confirmed by quantitative PCR for Nrxn1 which revealed a 7.4fold upregulation in SAM68 +/− cells but no significant difference in SAM68−/− cultures (RQ Nrxn1 4(−)/Gapdh: 7.4±3.1 in +/− vs 1.05±0.06 in −/−, n=4 per genotype, p<0.02). c-Fos upregulation was similar in control and homozygous knock-out neurons. Interestingly, depolarization-dependent alternative splicing of Nrxn2 AS4 remained intact in SAM68 KO neurons (Figure 7C). Moreover, Nrxn2 4(−) splice variant abundance was only very modestly altered in SAM68 KO brains (Figure S6). These findings point to unexpected gene-specific mechanisms for alternative splicing regulation of Nrxn1 and Nrxn2 at AS4 in vivo.
We further explored the importance of SAM68 for activity-dependent alternative splicing programs by monitoring the CaRRE-containing pre-mRNAs Grin1 exon 5 and the STREX exon of Kcnma1. For both pre-mRNAs, activity-dependent splicing was unaltered in SAM68 KO cerebellar neurons (Figure 7D). Therefore, SAM68 is essential for a specific sub-program of activity-dependent alternative splicing that includes regulation of Nrxn1 but not Grin1 or Kcnma1 CaRRE-containing mRNAs.
Finally, we examined whether SAM68 is required for activity-dependent alternative splicing of Nrxn1 AS4 in vivo. We stimulated cerebellar neurons by injection of the glutamate receptor agonist kainic acid directly into one cerebellar hemisphere of adult wild-type and SAM68 KO mice. Five hours after injection, animals were sacrificed and tissue was isolated from the injection site and the corresponding area of the uninjected hemisphere (Figure 7E). Transcript levels confirmed c-Fos induction in the injected hemispheres (Figure 7E). We assessed stimulation-induced alternative splicing of Nrxn1 at AS4 by comparison of the injected versus non-injected hemisphere using quantitative PCR (Figure 7F). Nrxn1 4(−) transcripts were significantly increased in the stimulated hemisphere whereas 4(+) containing transcripts were decreased (n=5 animals, p=0.002 for Nrxn1 4(−) and p=0.003 for Nrxn1 4(+)). Interestingly, total Nrxn1 mRNA transcripts were reduced indicating that Nrxn1 4(+) mRNA destabilization may occur simultaneously with the activity-dependent increase of Nrxn1 4(−) production. In injected SAM68 KO cerebellum, the Nrxn1 4(−) upregulation was strongly attenuated (Figure 7F). The same KO samples showed normal c-Fos induction confirming successful KA injection and immediate early gene induction (Figure 7E). In summary, these findings demonstrate that SAM68 is required for activity-dependent alternative splicing of Nrxn1 AS4 in vivo.
Discussion
Alternative splicing of neuronal pre-mRNAs has emerged as an important mechanism for the functional diversification and dynamic regulation of neuronal proteins (Wojtowicz et al., 2004; Ule et al., 2005; Zhang et al., 2008). For neurexins, alternative splicing of AS4 represents a key mechanism for the regulation of NRX-ligand interactions. Our analysis identifies the KH-domain RNA-binding protein SAM68 as a critical regulator of this splice isoform choice that is essential for the activity-dependent regulation of Nrxn1 in cerebellar neurons.
Alternative splicing-dependent regulation of neurexin function
Alternative splicing of Nrxn1 AS4 controls multiple splice-isoform specific ligand interactions, including the binding to NL1B, Cbln1, LRRTM1 and 2. Our results demonstrate that the depolarization-dependent Nrxn isoform switch upregulates transsynaptic signaling via NL1B and at the same time, reduces the response to GluD2 which interacts through CBLN1 with NRX 4(+) variants (Uemura et al., 2010). Notably, Cbln1 expression in granule cells is itself down-regulated by neuronal activity (Iijima et al., 2009) and the reduction in Cbln1 levels also contribute to the loss of GluD2 responses. Therefore, depolarization-dependent alternative splicing of Nrxn1 and downregulation of Cbln1 synergize to mediate a trans-synaptic ligand switch from interactions with GluD2 towards NL1B.
LRRTM proteins, a third group of NRX ligands, lack high affinity interactions with the NRX 4(+) variants and bind exclusively to the NRX 4(−) isoforms (Siddiqui et al., 2010). In SAM68 KO mice the Nrxn1 4(−) isoform is significantly reduced and our morphological analysis of SAM68 KO mice reveals abnormal distribution of the presynaptic marker vGluT1 in mossy fiber glomeruli, similar to a phenotype reported for Lrrtm1 knock-out mice (Linhoff et al., 2009). Further studies will be required to elucidate whether the synaptic phenotype in SAM68 KO mice is due to a change in Nrxn1 alternative splicing. However, our work provides a first indication for synaptic abnormalities caused by the loss of SAM68 in vivo.
Regulators of Nrxn AS4 alternative splicing
Cell- and tissue-specific alternative splicing are thought to be dependent on differentially expressed splicing factors or ubiquitous factors at different concentrations or activity (Chen and Manley, 2009). SAM68 is detected in most neuronal cells of the mouse cerebellum and throughout the nervous system (Figure 5 and data not shown). Therefore, presence of SAM68 in a cell population is not predictive of the outcome of alternative splicing regulation of Nrxn1 but SAM68 activity depends on neuronal depolarization (or additional stimuli). Resnick and colleagues implicated neuronal polypyrimidine tract binding protein (nPTB) in the regulation of Nrxn2 alternative splicing in cell lines (Resnick et al., 2008). Notably, our results indicate that the requirements for activity-dependent alternative splicing at AS4 differ between the Nrxn1 and Nrxn2 genes. Nrxn2 alternative splicing at AS4 is not altered in most SAM68 KO neurons (Figure S6) and we did not detect an activity of nPTB towards Nrxn1 AS4 alternative splicing in the reporter assays. Therefore, nPTB represents a candidate factor for selective regulation of Nrxn2 AS4 in vivo although additional work will be required to explore this.
The severity of Nrxn1 alternative splicing perturbation differed between brain areas with most significant effects in caudal brain areas and less dramatic alterations in hippocampus and cortex. Notably, SLM1 and SLM2 exhibited potent activities towards the Nrxn1 splice reporters in vitro and might compensate for the loss of SAM68 in those neuronal populations. Alternatively, additional RNA-binding proteins might co-operate with SAM68 in the regulation of Nrxn1 alternative splicing in these cells.
Neural activity-dependent regulation of neurexins by SAM68
SAM68 and other STAR-family proteins have been established as RNA regulators downstream of growth factor signaling in non-neuronal cells (Matter et al., 2002; Stoss et al., 2004; Valacca et al., 2010). Our work uncovered a novel mode of SAM68 regulation by neuronal activity and CaMKIV and identified serine 20 as a depolarization-dependent phosphorylation site. Intriguingly, SAM68 shares sequence similarity with SF1, a component of the constitutive splicing machinery. In SF1 serine 20 phosphorylation controls interaction with U2AF65 and, thereby, regulates spliceosome assembly (Wang et al., 1999; Selenko et al., 2003). Whether SAM68 phosphorylation at serine 20 functions in an analogous manner remains to be examined but, notably, SAM68 has been suggested to interact with U2AF65 in nonneuronal cells (Tisserant and Konig, 2008).
CaMKIV is a major regulator of depolarization-dependent gene expression and alternative splicing but the RNA-binding proteins acting downstream of CaMKIV have remained unknown (Xie and Black, 2001; Xie et al., 2005). Loss of depolarization-dependent Nrxn1 splicing regulation in SAM68 KO neurons establishes SAM68 as a key mediator of this pathway. Notably, Nrxn1 exon 20 does not contain recognizable CaRRE sequences (unpublished observation) and the alternative splicing of several CaRRE containing pre-mRNAs was not impaired in SAM68 KO neurons. This suggests that CaMKIV-induced regulation of alternative splicing is relayed via multiple different RNA-binding proteins (or combinations thereof) that bind different recognition elements in their respective target pre-mRNAs. Genome-wide mapping of targets for the neuronal splicing factors NOVA1 and -2 has revealed coordinate regulation of a specific set of mRNAs encoding synaptic proteins (Ule et al., 2005). Given the significant number of mRNAs that undergo depolarization-dependent alternative splicing it will be important to map the entire SAM68-dependent program of activity-dependent alternative splicing in the future.
Experimental Procedures
Alternative splicing analysis
Oligonucleotide primers used for semi-quantitative PCR are listed in Table S1. Input cDNA amounts were titrated to ensure linear relationships between templates and amplification products. DNA fragment intensities were quantified by image analyzer (FAS-III, Toyobo) and ImageGauge software (Fujifilm). Quantitative PCR was performed on a StepOnePlus qPCR system (Applied Biosystems). Gene expression assays (Nrxn1:Mm00660298m1, Gapdh:Mm99999915g1, Khdrbs1:Mm00516130_m1 Applied Biosystems), or custom designed assays (Table S2) were used with TaqMan Master Mix (Applied Biosystems) and comparative CT method. The mRNA levels were normalized to that of Gapdh mRNA.
For in vivo stimulation by kainate, adult mice (2–3 month old) were anesthetized by intraperitoneal injection of 100 mg/ml ketamine and 10 mg/ml xylazine. A small hole in the occipital bone was made with a dental drill. A 33 gauge microsyringe was inserted into one hemisphere of the cerebellar cortex, and 2–3µl of 50 µM kainic acid in saline containing bromphenol blue dye (0.5 mg/ml) was infused (0.4 µl/min). Five hours after injection, tissue was dissected and subjected to quantitative PCR analysis.
In utero RNA Response Element (iRRE) assay
Splice reporter and EGFP-expression vector (2:1 ratio) were delivered into E14.5 precursor cells of cortical pyramidal cells by in utero electroporation with five pulses (50V, 50 msec, each spaced by 950 msec). Dams were brought to term and EGFP-positive cortical tissue was dissected from P6 pups and processed for splicing analysis with primers unique to the reporter sequences.
Neuronal cell culture
Cerebellar cultures and mixed culture assays were performed essentially as described (Scheiffele et al., 2000). Briefly, cerebellar neurons (1 × 105/cm2) were maintained for 9 days in vitro and NL- or GluD2-expressing HEK293 cells (2 × 104/cm2) were added to the culture for an additional two days. Cells were fixed and processed for immunohistochemistry and image analysis. For depolarization, the potassium concentration in the medium was elevated by addition of 25 mM KCl from DIV6.
Statistical analysis
Pairwise comparisons were performed using Student's t-test. For multiple comparisons, analysis of variance (ANOVA) followed by Bonferroni test was used. Data are represented as the mean ± SEM. Statistical comparison of vGluT1 staining intensity distributions in wild-type and SAM68 KO cerebellum was done using the Komolgorov-Smirnov test. Significance is indicated as follows: *** p < 0.001, ** p < 0.01, * p < 0.05.
See Supplementary procedures for further details.
Supplementary Material
Acknowledgements
We are grateful to Dr. Jane Dodd for generously sharing lab space and resources, to Dr. Gollan who contributed important initial findings in the early phase of this project, to Drs. Yves Barde, Olivier Pertz, Oguz Kanca, Stéphane Baudouin, and Thi-Minh-Phuc Nguyen for comments on the manuscript, to Drs. Boukhtouche, Cocas, Sommer, and Ms. Stiefvater for help with in utero electroporations, to Laetitia Burklé and Caroline Bornmann for excellent technical support, to Prof. Michisuke Yuzaki for advice and for generously sharing a GluD2 expression vector, to Dr. Kyo-ichi Emi for advice on cerebellar injections, and to Gillian Vogel and members of the Scheiffele Lab for valuable discussions. We thank Drs. Chabot, Darnell, Ghosh, König, Smith, Sette, Soderling, and the Developmental Studies Hybridoma Bank, University of Iowa, for making important reagents available. K.W. is the recipient of a Jane Coffin Childs Fund postdoctoral fellowship award. T.I. was supported by the Uehara Memorial Foundation, KANAE Foundation for the Promotion of Medical Science and Astellas Foundation for Research on Metabolic Disorders. S.R. is funded by the Canadian Institutes of Health Research, and the project was funded by grants from NINDS (R21 NS53936) and an award from the Swiss National Science Foundation (to P.S.).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- An P, Grabowski PJ. Exon silencing by UAGG motifs in response to neuronal excitation. PLoS Biol. 2007;5:e36. doi: 10.1371/journal.pbio.0050036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batsche E, Yaniv M, Muchardt C. The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nat Struct Mol Biol. 2006;13:22–29. doi: 10.1038/nsmb1030. [DOI] [PubMed] [Google Scholar]
- Baudouin S, Scheiffele P. SnapShot: Neuroligin-neurexin complexes. Cell. 2010;141 doi: 10.1016/j.cell.2010.05.024. 908, 908 e901. [DOI] [PubMed] [Google Scholar]
- Bingol B, Sheng M. Deconstruction for reconstruction: the role of proteolysis in neural plasticity and disease. Neuron. 2011;69:22–32. doi: 10.1016/j.neuron.2010.11.006. [DOI] [PubMed] [Google Scholar]
- Boucard AA, Chubykin AA, Comoletti D, Taylor P, Sudhof TC. A splice code for trans-synaptic cell adhesion mediated by binding of neuroligin 1 to alpha- and beta-neurexins. Neuron. 2005;48:229–236. doi: 10.1016/j.neuron.2005.08.026. [DOI] [PubMed] [Google Scholar]
- Chawla G, Lin CH, Han A, Shiue L, Ares M, Jr, Black DL. Sam68 regulates a set of alternatively spliced exons during neurogenesis. Mol Cell Biol. 2009;29:201–213. doi: 10.1128/MCB.01349-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M, Manley JL. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat Rev Mol Cell Biol. 2009;10:741–754. doi: 10.1038/nrm2777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chih B, Gollan L, Scheiffele P. Alternative splicing controls selective trans-synaptic interactions of the neuroligin-neurexin complex. Neuron. 2006;51:171–178. doi: 10.1016/j.neuron.2006.06.005. [DOI] [PubMed] [Google Scholar]
- Cohen S, Greenberg ME. Communication between the synapse and the nucleus in neuronal development, plasticity, and disease. Annu Rev Cell Dev Biol. 2008;24:183–209. doi: 10.1146/annurev.cellbio.24.110707.175235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craig AM, Kang Y. Neurexin-neuroligin signaling in synapse development. Curr Opin Neurobiol. 2007;17:43–52. doi: 10.1016/j.conb.2007.01.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis GW. Homeostatic control of neural activity: from phenomenology to molecular design. Annu Rev Neurosci. 2006;29:307–323. doi: 10.1146/annurev.neuro.28.061604.135751. [DOI] [PubMed] [Google Scholar]
- de Wit J, Sylwestrak E, O'Sullivan ML, Otto S, Tiglio K, Savas JN, Yates JR, 3rd, Comoletti D, Taylor P, Ghosh A. LRRTM2 interacts with Neurexin1 and regulates excitatory synapse formation. Neuron. 2009;64:799–806. doi: 10.1016/j.neuron.2009.12.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean C, Scholl FG, Choih J, DeMaria S, Berger J, Isacoff E, Scheiffele P. Neurexin mediates the assembly of presynaptic terminals. Nat Neurosci. 2003;6:708–716. doi: 10.1038/nn1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Fruscio M, Chen T, Richard S. Characterization of Sam68-like mammalian proteins SLM-1 and SLM-2: SLM-1 is a Src substrate during mitosis. Proc Natl Acad Sci U S A. 1999;96:2710–2715. doi: 10.1073/pnas.96.6.2710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisette JF, Toutant J, Dugre-Brisson S, Desgroseillers L, Chabot B. hnRNP A1 and hnRNP H can collaborate to modulate 5' splice site selection. RNA. 2010;16:228–238. doi: 10.1261/rna.1890310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galarneau A, Richard S. The STAR RNA binding proteins GLD-1, QKI, SAM68 and SLM-2 bind bipartite RNA motifs. BMC Mol Biol. 2009;10:47. doi: 10.1186/1471-2199-10-47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grange J, Boyer V, Fabian-Fine R, Fredj NB, Sadoul R, Goldberg Y. Somatodendritic localization and mRNA association of the splicing regulatory protein Sam68 in the hippocampus and cortex. J Neurosci Res. 2004;75:654–666. doi: 10.1002/jnr.20003. [DOI] [PubMed] [Google Scholar]
- Henao-Mejia J, Liu Y, Park IW, Zhang J, Sanford J, He JJ. Suppression of HIV-1 Nef translation by Sam68 mutant-induced stress granules and nef mRNA sequestration. Mol Cell. 2009;33:87–96. doi: 10.1016/j.molcel.2008.11.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang ZJ, Scheiffele P. GABA and neuroligin signaling: linking synaptic activity and adhesion in inhibitory synapse development. Curr Opin Neurobiol. 2008;18:77–83. doi: 10.1016/j.conb.2008.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huot ME, Brown CM, Lamarche-Vane N, Richard S. An adaptor role for cytoplasmic Sam68 in modulating Src activity during cell polarization. Mol Cell Biol. 2009;29:1933–1943. doi: 10.1128/MCB.01707-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ichtchenko K, Hata Y, Nguyen T, Ullrich B, Missler M, Moomaw C, Sudhof TC. Neuroligin 1: a splice site-specific ligand for beta-neurexins. Cell. 1995;81:435–443. doi: 10.1016/0092-8674(95)90396-8. [DOI] [PubMed] [Google Scholar]
- Iijima T, Emi K, Yuzaki M. Activity-dependent repression of Cbln1 expression: mechanism for developmental and homeostatic regulation of synapses in the cerebellum. J Neurosci. 2009;29:5425–5434. doi: 10.1523/JNEUROSCI.4473-08.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang Y, Zhang X, Dobie F, Wu H, Craig AM. Induction of GABAergic postsynaptic differentiation by alpha-neurexins. J Biol Chem. 2008;283:2323–2334. doi: 10.1074/jbc.M703957200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim KK, Adelstein RS, Kawamoto S. Identification of neuronal nuclei (NeuN) as Fox-3, a new member of the Fox-1 gene family of splicing factors. J Biol Chem. 2009;284:31052–31061. doi: 10.1074/jbc.M109.052969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koehnke J, Katsamba PS, Ahlsen G, Bahna F, Vendome J, Honig B, Shapiro L, Jin X. Splice form dependence of beta-neurexin/neuroligin binding interactions. Neuron. 2010;67:61–74. doi: 10.1016/j.neuron.2010.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JA, Xing Y, Nguyen D, Xie J, Lee CJ, Black DL. Depolarization and CaM kinase IV modulate NMDA receptor splicing through two essential RNA elements. PLoS Biol. 2007;5:e40. doi: 10.1371/journal.pbio.0050040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J, Ashley J, Budnik V, Bhat MA. Crucial role of Drosophila neurexin in proper active zone apposition to postsynaptic densities, synaptic growth, and synaptic transmission. Neuron. 2007a;55:741–755. doi: 10.1016/j.neuron.2007.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Q, Lee JA, Black DL. Neuronal regulation of alternative pre-mRNA splicing. Nat Rev Neurosci. 2007b;8:819–831. doi: 10.1038/nrn2237. [DOI] [PubMed] [Google Scholar]
- Lin Q, Taylor SJ, Shalloway D. Specificity and determinants of Sam68 RNA binding. Implications for the biological function of K homology domains. J Biol Chem. 1997;272:27274–27280. doi: 10.1074/jbc.272.43.27274. [DOI] [PubMed] [Google Scholar]
- Linhoff MW, Lauren J, Cassidy RM, Dobie FA, Takahashi H, Nygaard HB, Airaksinen MS, Strittmatter SM, Craig AM. An unbiased expression screen for synaptogenic proteins identifies the LRRTM protein family as synaptic organizers. Neuron. 2009;61:734–749. doi: 10.1016/j.neuron.2009.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lukong KE, Richard S. Motor coordination defects in mice deficient for the Sam68 RNA-binding protein. Behav Brain Res. 2008;189:357–363. doi: 10.1016/j.bbr.2008.01.010. [DOI] [PubMed] [Google Scholar]
- Matsuda K, Miura E, Miyazaki T, Kakegawa W, Emi K, Narumi S, Fukazawa Y, Ito-Ishida A, Kondo T, Shigemoto R, et al. Cbln1 is a ligand for an orphan glutamate receptor delta2, a bidirectional synapse organizer. Science (New York, NY. 2010;328:363–368. doi: 10.1126/science.1185152. [DOI] [PubMed] [Google Scholar]
- Matsuda K, Yuzaki M. Cbln family proteins promote synapse formation by regulating distinct neurexin signaling pathways in various brain regions. Eur J Neurosci. 2011;33:1447–1461. doi: 10.1111/j.1460-9568.2011.07638.x. [DOI] [PubMed] [Google Scholar]
- Matter N, Herrlich P, Konig H. Signal-dependent regulation of splicing via phosphorylation of Sam68. Nature. 2002;420:691–695. doi: 10.1038/nature01153. [DOI] [PubMed] [Google Scholar]
- Mellor JR, Merlo D, Jones A, Wisden W, Randall AD. Mouse cerebellar granule cell differentiation: electrical activity regulates the GABAA receptor alpha 6 subunit gene. J Neurosci. 1998;18:2822–2833. doi: 10.1523/JNEUROSCI.18-08-02822.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Missler M, Zhang W, Rohlmann A, Kattenstroth G, Hammer RE, Gottmann K, Sudhof TC. Alpha-neurexins couple Ca2+ channels to synaptic vesicle exocytosis. Nature. 2003;423:939–948. doi: 10.1038/nature01755. [DOI] [PubMed] [Google Scholar]
- Ohmstede CA, Jensen KF, Sahyoun NE. Ca2+/calmodulin-dependent protein kinase enriched in cerebellar granule cells. Identification of a novel neuronal calmodulin-dependent protein kinase. J Biol Chem. 1989;264:5866–5875. [PubMed] [Google Scholar]
- Paronetto MP, Achsel T, Massiello A, Chalfant CE, Sette C. The RNA-binding protein Sam68 modulates the alternative splicing of Bcl-x. The Journal of cell biology. 2007;176:929–939. doi: 10.1083/jcb.200701005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reissner C, Klose M, Fairless R, Missler M. Mutational analysis of the neurexin/neuroligin complex reveals essential and regulatory components. Proc Natl Acad Sci U S A. 2008;105:15124–15129. doi: 10.1073/pnas.0801639105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick M, Segall A, G GR, Lupowitz Z, Zisapel N. Alternative splicing of neurexins: a role for neuronal polypyrimidine tract binding protein. Neuroscience letters. 2008;439:235–240. doi: 10.1016/j.neulet.2008.05.034. [DOI] [PubMed] [Google Scholar]
- Richard S, Torabi N, Franco GV, Tremblay GA, Chen T, Vogel G, Morel M, Cleroux P, Forget-Richard A, Komarova S, et al. Ablation of the Sam68 RNA binding protein protects mice from age-related bone loss. PLoS Genet. 2005;1:e74. doi: 10.1371/journal.pgen.0010074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rossi P, De Filippi G, Armano S, Taglietti V, D'Angelo E. The weaver mutation causes a loss of inward rectifier current regulation in premigratory granule cells of the mouse cerebellum. J Neurosci. 1998;18:3537–3547. doi: 10.1523/JNEUROSCI.18-10-03537.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowen L, Young J, Birditt B, Kaur A, Madan A, Philipps DL, Qin S, Minx P, Wilson RK, Hood L, et al. Analysis of the human neurexin genes: alternative splicing and the generation of protein diversity. Genomics. 2002;79:587–597. doi: 10.1006/geno.2002.6734. [DOI] [PubMed] [Google Scholar]
- Rozic G, Lupowitz Z, Piontkewitz Y, Zisapel N. Dynamic changes in neurexins' alternative splicing: role of Rho-associated protein kinases and relevance to memory formation. PLoS One. 2011;6:e18579. doi: 10.1371/journal.pone.0018579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rozic-Kotliroff G, Zisapel N. Ca2+ -dependent splicing of neurexin IIalpha. Biochem Biophys Res Commun. 2007;352:226–230. doi: 10.1016/j.bbrc.2006.11.008. [DOI] [PubMed] [Google Scholar]
- Rujescu D, Ingason A, Cichon S, Pietilainen OP, Barnes MR, Toulopoulou T, Picchioni M, Vassos E, Ettinger U, Bramon E, et al. Disruption of the neurexin 1 gene is associated with schizophrenia. Hum Mol Genet. 2009;18:988–996. doi: 10.1093/hmg/ddn351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scheiffele P, Fan J, Choih J, Fetter R, Serafini T. Neuroligin expressed in nonneuronal cells triggers presynaptic development in contacting axons. Cell. 2000;101:657–669. doi: 10.1016/s0092-8674(00)80877-6. [DOI] [PubMed] [Google Scholar]
- Selenko P, Gregorovic G, Sprangers R, Stier G, Rhani Z, Kramer A, Sattler M. Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP. Mol Cell. 2003;11:965–976. doi: 10.1016/s1097-2765(03)00115-1. [DOI] [PubMed] [Google Scholar]
- Siddiqui TJ, Pancaroglu R, Kang Y, Rooyakkers A, Craig AM. LRRTMs and neuroligins bind neurexins with a differential code to cooperate in glutamate synapse development. J Neurosci. 2010;30:7495–7506. doi: 10.1523/JNEUROSCI.0470-10.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Songyang Z, Lu KP, Kwon YT, Tsai LH, Filhol O, Cochet C, Brickey DA, Soderling TR, Bartleson C, Graves DJ, et al. A structural basis for substrate specificities of protein Ser/Thr kinases: primary sequence preference of casein kinases I and II, NIMA, phosphorylase kinase, calmodulin-dependent kinase II, CDK5, and Erk1. Mol Cell Biol. 1996;16:6486–6493. doi: 10.1128/mcb.16.11.6486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stoss O, Novoyatleva T, Gencheva M, Olbrich M, Benderska N, Stamm S. p59(fyn)-mediated phosphorylation regulates the activity of the tissue-specific splicing factor rSLM-1. Mol Cell Neurosci. 2004;27:8–21. doi: 10.1016/j.mcn.2004.04.011. [DOI] [PubMed] [Google Scholar]
- Sudhof TC. Neuroligins and neurexins link synaptic function to cognitive disease. Nature. 2008;455:903–911. doi: 10.1038/nature07456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabuchi K, Sudhof TC. Structure and evolution of neurexin genes: insight into the mechanism of alternative splicing. Genomics. 2002;79:849–859. doi: 10.1006/geno.2002.6780. [DOI] [PubMed] [Google Scholar]
- Tisserant A, Konig H. Signal-regulated Pre-mRNA occupancy by the general splicing factor U2AF. PLoS One. 2008;3:e1418. doi: 10.1371/journal.pone.0001418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tokumitsu H, Inuzuka H, Ishikawa Y, Ikeda M, Saji I, Kobayashi R. STO-609, a specific inhibitor of the Ca(2+)/calmodulin-dependent protein kinase kinase. J Biol Chem. 2002;277:15813–15818. doi: 10.1074/jbc.M201075200. [DOI] [PubMed] [Google Scholar]
- Uemura T, Lee SJ, Yasumura M, Takeuchi T, Yoshida T, Ra M, Taguchi R, Sakimura K, Mishina M. Trans-synaptic interaction of GluRdelta2 and Neurexin through Cbln1 mediates synapse formation in the cerebellum. Cell. 2010;141:1068–1079. doi: 10.1016/j.cell.2010.04.035. [DOI] [PubMed] [Google Scholar]
- Ule J, Ule A, Spencer J, Williams A, Hu JS, Cline M, Wang H, Clark T, Fraser C, Ruggiu M, et al. Nova regulates brain-specific splicing to shape the synapse. Nat Genet. 2005;37:844–852. doi: 10.1038/ng1610. [DOI] [PubMed] [Google Scholar]
- Ushkaryov YA, Petrenko AG, Geppert M, Sudhof TC. Neurexins: synaptic cell surface proteins related to the alpha- latrotoxin receptor and laminin. Science (New York, NY. 1992;257:50–56. doi: 10.1126/science.1621094. [DOI] [PubMed] [Google Scholar]
- Valacca C, Bonomi S, Buratti E, Pedrotti S, Baralle FE, Sette C, Ghigna C, Biamonti G. Sam68 regulates EMT through alternative splicing-activated nonsense-mediated mRNA decay of the SF2/ASF proto-oncogene. The Journal of cell biology. 2010;191:87–99. doi: 10.1083/jcb.201001073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Volk T, Israeli D, Nir R, Toledano-Katchalski H. Tissue development and RNA control: "HOW" is it coordinated? Trends Genet. 2008;24:94–101. doi: 10.1016/j.tig.2007.11.009. [DOI] [PubMed] [Google Scholar]
- Wang X, Bruderer S, Rafi Z, Xue J, Milburn PJ, Kramer A, Robinson PJ. Phosphorylation of splicing factor SF1 on Ser20 by cGMP-dependent protein kinase regulates spliceosome assembly. EMBO J. 1999;18:4549–4559. doi: 10.1093/emboj/18.16.4549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wayman GA, Lee YS, Tokumitsu H, Silva AJ, Soderling TR. Calmodulin-kinases: modulators of neuronal development and plasticity. Neuron. 2008;59:914–931. doi: 10.1016/j.neuron.2008.08.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wojtowicz WM, Flanagan JJ, Millard SS, Zipursky SL, Clemens JC. Alternative splicing of Drosophila Dscam generates axon guidance receptors that exhibit isoform-specific homophilic binding. Cell. 2004;118:619–633. doi: 10.1016/j.cell.2004.08.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong RO, Ghosh A. Activity-dependent regulation of dendritic growth and patterning. Nat Rev Neurosci. 2002;3:803–812. doi: 10.1038/nrn941. [DOI] [PubMed] [Google Scholar]
- Xie J, Black DL. A CaMK IV responsive RNA element mediates depolarization-induced alternative splicing of ion channels. Nature. 2001;410:936–939. doi: 10.1038/35073593. [DOI] [PubMed] [Google Scholar]
- Xie J, Jan C, Stoilov P, Park J, Black DL. A consensus CaMK IV-responsive RNA sequence mediates regulation of alternative exons in neurons. RNA. 2005;11:1825–1834. doi: 10.1261/rna.2171205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang C, Zhang Z, Castle J, Sun S, Johnson J, Krainer AR, Zhang MQ. Defining the regulatory network of the tissue-specific splicing factors Fox-1 and Fox-2. Genes Dev. 2008;22:2550–2563. doi: 10.1101/gad.1703108. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.