Abstract
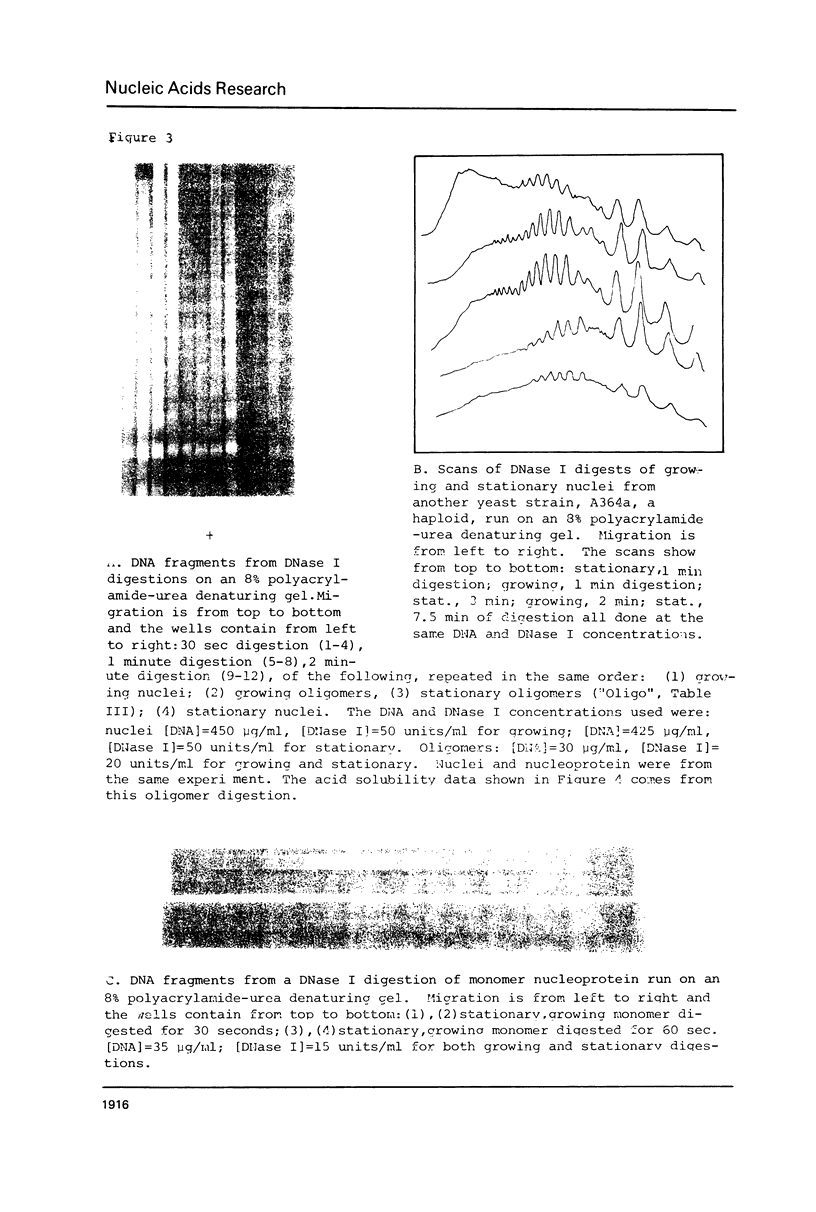
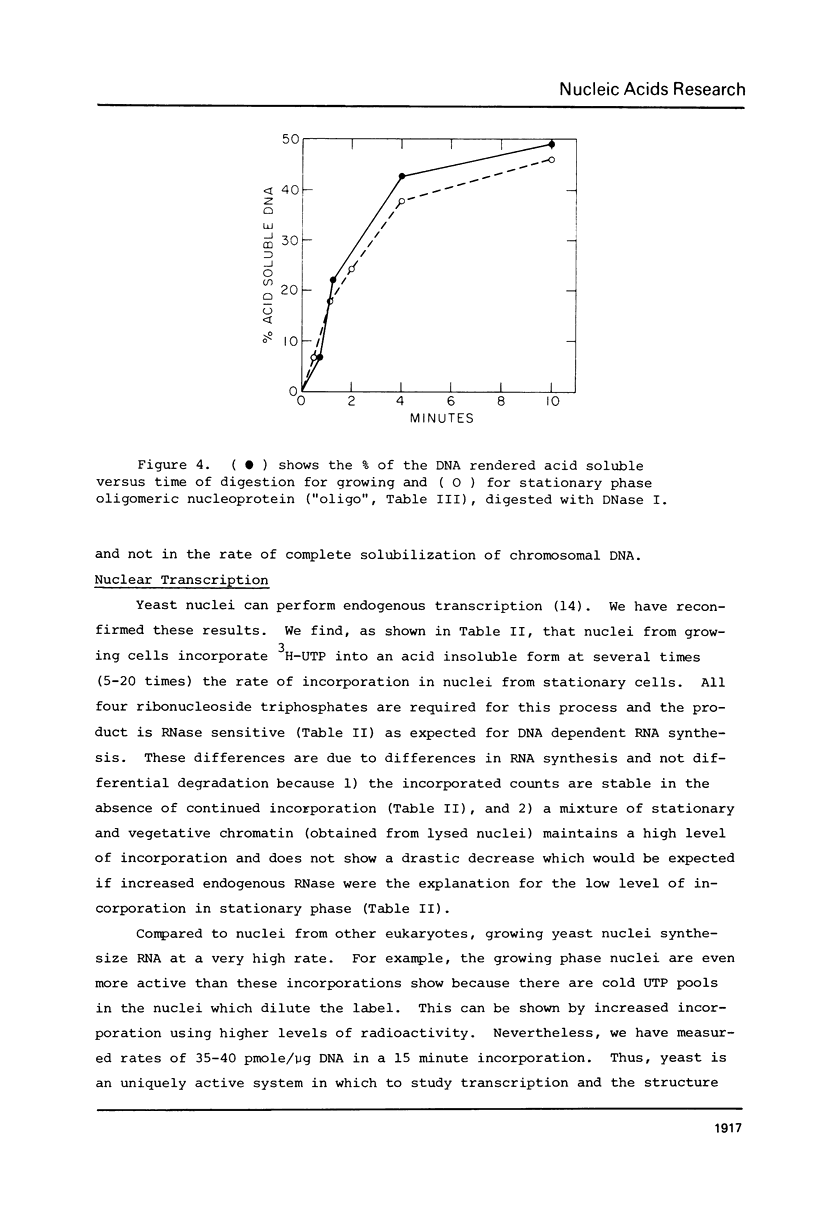
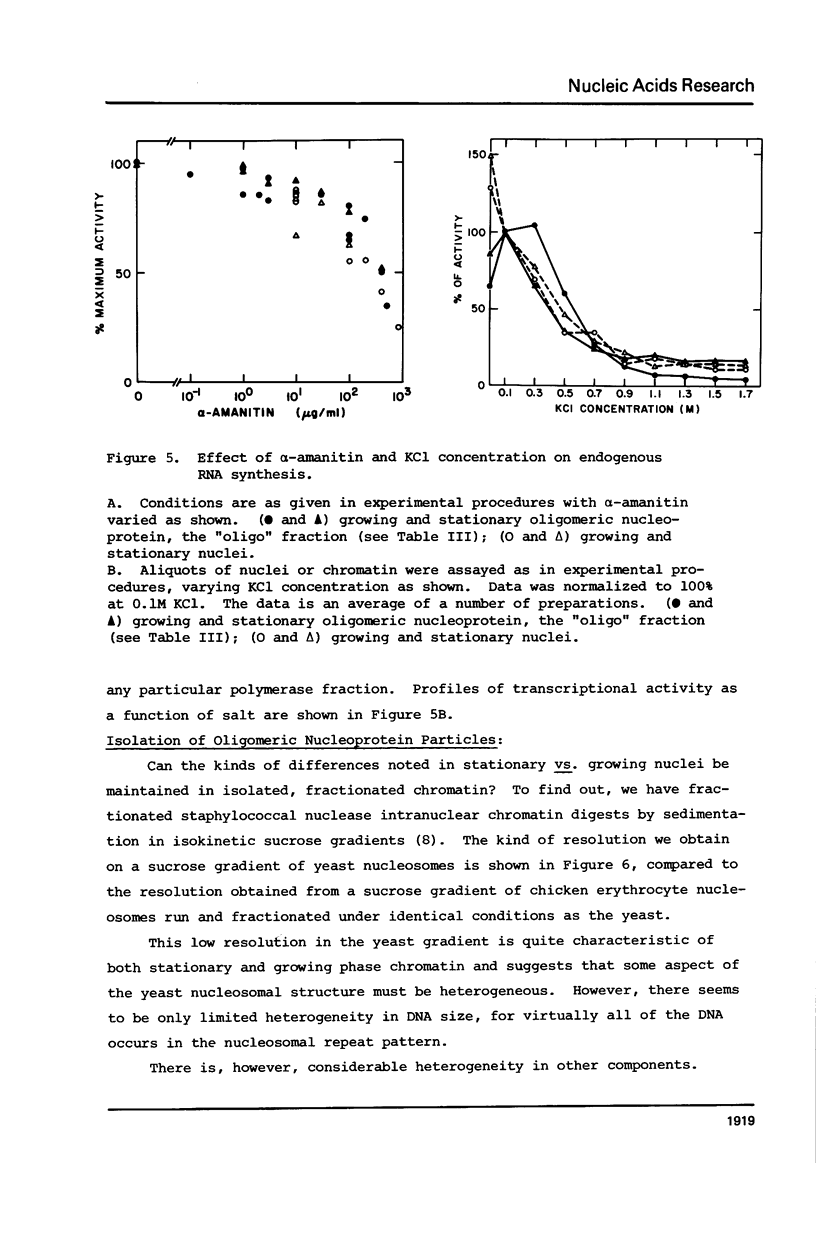
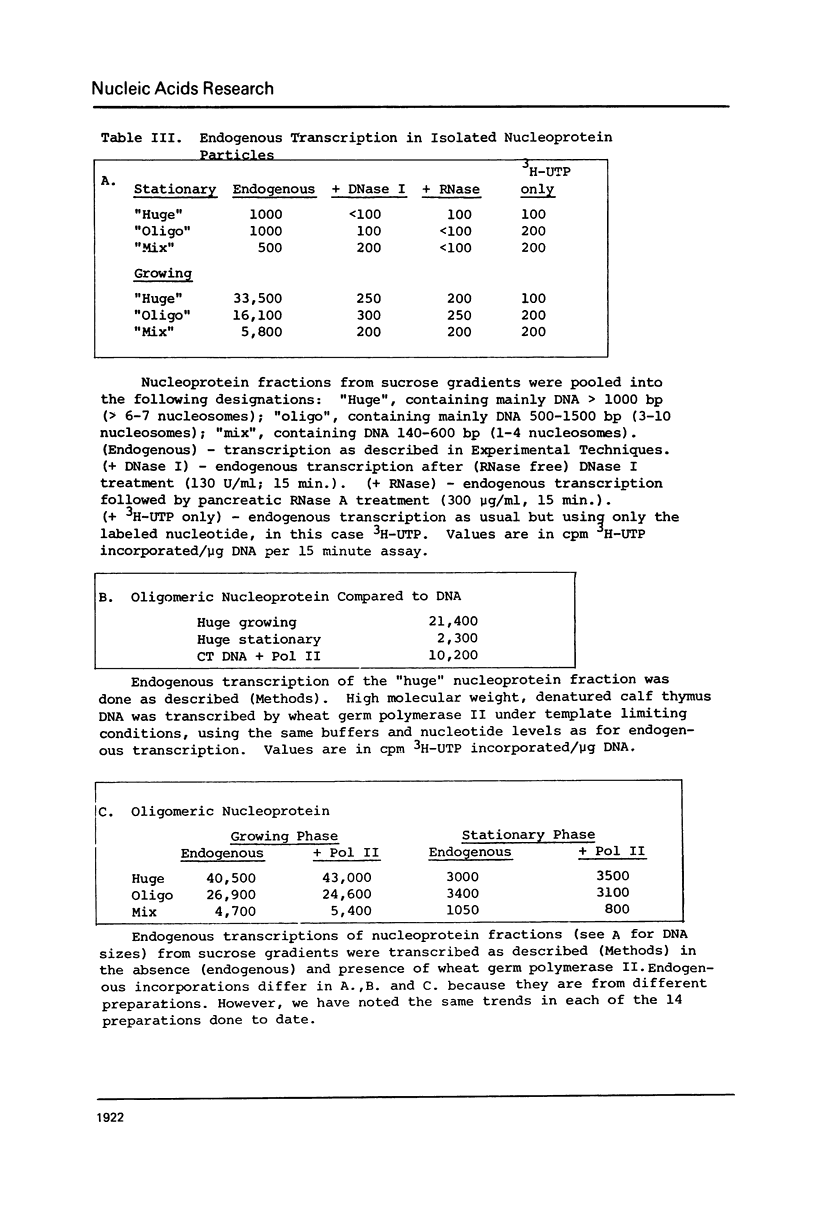
We have compared the structure of intra-nuclear and isolated chromatin from logarithmically growing yeast cells to chromatin from cells which had entered the stationary phase and ceased growing. Both chromatins show a similar nucleosomal repeat pattern, 160 bp repeat size, with staphylococcal nuclease and similar variability in repeat sizes within the genome. DNase I produces the same ladder (less than 120 b) and a quite similar extended ladder (120-300 b) which shows that both chromatins have phased nucleosomes. However, the rate of DNase I digestion of growing phase is greater than in stationary. Functionally speaking, growing phase nuclei are 5-20 times as active in the rate of endogenous transcription (all three polymerases are involved). The transcriptional and DNase I susceptibility differences noted in nuclei are maintained in sucrose gradient isolated oligonucleosomes and mononucleosomes from the two states.
Full text
PDF












Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adman R., Schultz L. D., Hall B. D. Transcription in yeast: separation and properties of multiple FNA polymerases. Proc Natl Acad Sci U S A. 1972 Jul;69(7):1702–1706. doi: 10.1073/pnas.69.7.1702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Axel R. Cleavage of DNA in nuclei and chromatin with staphylococcal nuclease. Biochemistry. 1975 Jul;14(13):2921–2925. doi: 10.1021/bi00684a020. [DOI] [PubMed] [Google Scholar]
- Bellard M., Gannon F., Chambon P. Nucleosome structure III: the structure and transcriptional activity of the chromatin containing the ovalbumin and globin genes in chick oviduct nuclei. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):779–791. doi: 10.1101/sqb.1978.042.01.078. [DOI] [PubMed] [Google Scholar]
- Camerini-Otero R. D., Sollner-Webb B., Simon R. H., Williamson P., Zasloff M., Felsenfeld G. Nucleosome structure, DNA folding, and gene activity. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 1):57–75. doi: 10.1101/sqb.1978.042.01.008. [DOI] [PubMed] [Google Scholar]
- Felsenfeld G. Chromatin. Nature. 1978 Jan 12;271(5641):115–122. doi: 10.1038/271115a0. [DOI] [PubMed] [Google Scholar]
- Frank K. R., Mills D. Ribosome activity and degradation in meiotic cells of Saccharomyces cerevisiae. Mol Gen Genet. 1978 Mar 20;160(1):59–65. doi: 10.1007/BF00275119. [DOI] [PubMed] [Google Scholar]
- Garel A., Axel R. Selective digestion of transcriptionally active ovalbumin genes from oviduct nuclei. Proc Natl Acad Sci U S A. 1976 Nov;73(11):3966–3970. doi: 10.1073/pnas.73.11.3966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottesfeld J. M., Butler P. J. Structure of transcriptionally-active chromatin subunits. Nucleic Acids Res. 1977 Sep;4(9):3155–3173. doi: 10.1093/nar/4.9.3155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hereford L. M., Rosbash M. Number and distribution of polyadenylated RNA sequences in yeast. Cell. 1977 Mar;10(3):453–462. doi: 10.1016/0092-8674(77)90032-0. [DOI] [PubMed] [Google Scholar]
- Kornberg R. D. Structure of chromatin. Annu Rev Biochem. 1977;46:931–954. doi: 10.1146/annurev.bi.46.070177.004435. [DOI] [PubMed] [Google Scholar]
- Lacy E., Axel R. Analysis of DNA of isolated chromatin subunits. Proc Natl Acad Sci U S A. 1975 Oct;72(10):3978–3982. doi: 10.1073/pnas.72.10.3978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lohr D., Corden J., Tatchell K., Kovacic R. T., Van Holde K. E. Comparative subunit structure of HeLa, yeast, and chicken erythrocyte chromatin. Proc Natl Acad Sci U S A. 1977 Jan;74(1):79–83. doi: 10.1073/pnas.74.1.79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohr D., Kovacic R. T., Van Holde K. E. Quantitative analysis of the digestion of yeast chromatin by staphylococcal nuclease. Biochemistry. 1977 Feb 8;16(3):463–471. doi: 10.1021/bi00622a020. [DOI] [PubMed] [Google Scholar]
- Lohr D., Tatchell K., Van Holde K. E. On the occurrence of nucleosome phasing in chromatin. Cell. 1977 Nov;12(3):829–836. doi: 10.1016/0092-8674(77)90281-1. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Jeffrey A., van deSande H. Chain length determination of small double- and single-stranded DNA molecules by polyacrylamide gel electrophoresis. Biochemistry. 1975 Aug 26;14(17):3787–3794. doi: 10.1021/bi00688a010. [DOI] [PubMed] [Google Scholar]
- McMaster-Kaye R., Kaye J. S. Staining of histones on polyacrylamide gels with amido black and fast green. Anal Biochem. 1974 Sep;61(1):120–132. doi: 10.1016/0003-2697(74)90339-x. [DOI] [PubMed] [Google Scholar]
- Nelson D. A., Beltz W. R., Rill R. L. Chromatin subunits from baker's yeast: isolation and partial characterization. Proc Natl Acad Sci U S A. 1977 Apr;74(4):1343–1347. doi: 10.1073/pnas.74.4.1343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prunell A., Kornberg R. D. Relation of nucleosomes to DNA sequences. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 1):103–108. doi: 10.1101/sqb.1978.042.01.011. [DOI] [PubMed] [Google Scholar]
- Riley D., Weintraub H. Nucleosomal DNA is digested to repeats of 10 bases by exonuclease III. Cell. 1978 Feb;13(2):281–293. doi: 10.1016/0092-8674(78)90197-6. [DOI] [PubMed] [Google Scholar]
- Roeder R. G. Multiple forms of deoxyribonucleic acid-dependent ribonucleic acid polymerase in Xenopus laevis. Isolation and partial characterization. J Biol Chem. 1974 Jan 10;249(1):241–248. [PubMed] [Google Scholar]
- Scheer U. Changes of nucleosome frequency in nucleolar and non-nucleolar chromatin as a function of transcription: an electron microscopic study. Cell. 1978 Mar;13(3):535–549. doi: 10.1016/0092-8674(78)90327-6. [DOI] [PubMed] [Google Scholar]
- Schultz L. D., Hall B. D. Transcription in yeast: alpha-amanitin sensitivity and other properties which distinguish between RNA polymerases I and III. Proc Natl Acad Sci U S A. 1976 Apr;73(4):1029–1033. doi: 10.1073/pnas.73.4.1029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz L. D. Transcriptional role of yeast deoxyribonucleic acid dependent ribonucleic acid polymerase III. Biochemistry. 1978 Feb 21;17(4):750–758. doi: 10.1021/bi00597a031. [DOI] [PubMed] [Google Scholar]
- Sollner-Webb B., Felsenfeld G. A comparison of the digestion of nuclei and chromatin by staphylococcal nuclease. Biochemistry. 1975 Jul;14(13):2915–2920. doi: 10.1021/bi00684a019. [DOI] [PubMed] [Google Scholar]
- Tata J. R., Baker B. Enzymatic fractionation of nuclei: polynucleosomes and RNA polymerase II as endogenous transcriptional complexes. J Mol Biol. 1978 Jan 25;118(3):249–272. doi: 10.1016/0022-2836(78)90227-9. [DOI] [PubMed] [Google Scholar]
- Tatchell K., Van Holde K. E. Compact oligomers and nucleosome phasing. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3583–3587. doi: 10.1073/pnas.75.8.3583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valenzuela P., Hager G. L., Weinberg F., Rutter W. J. Molecular structure of yeast RNA polymerase III: demonstration of the tripartite transcriptive system in lower eukaryotes. Proc Natl Acad Sci U S A. 1976 Apr;73(4):1024–1028. doi: 10.1073/pnas.73.4.1024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weintraub H., Groudine M. Chromosomal subunits in active genes have an altered conformation. Science. 1976 Sep 3;193(4256):848–856. doi: 10.1126/science.948749. [DOI] [PubMed] [Google Scholar]