Abstract
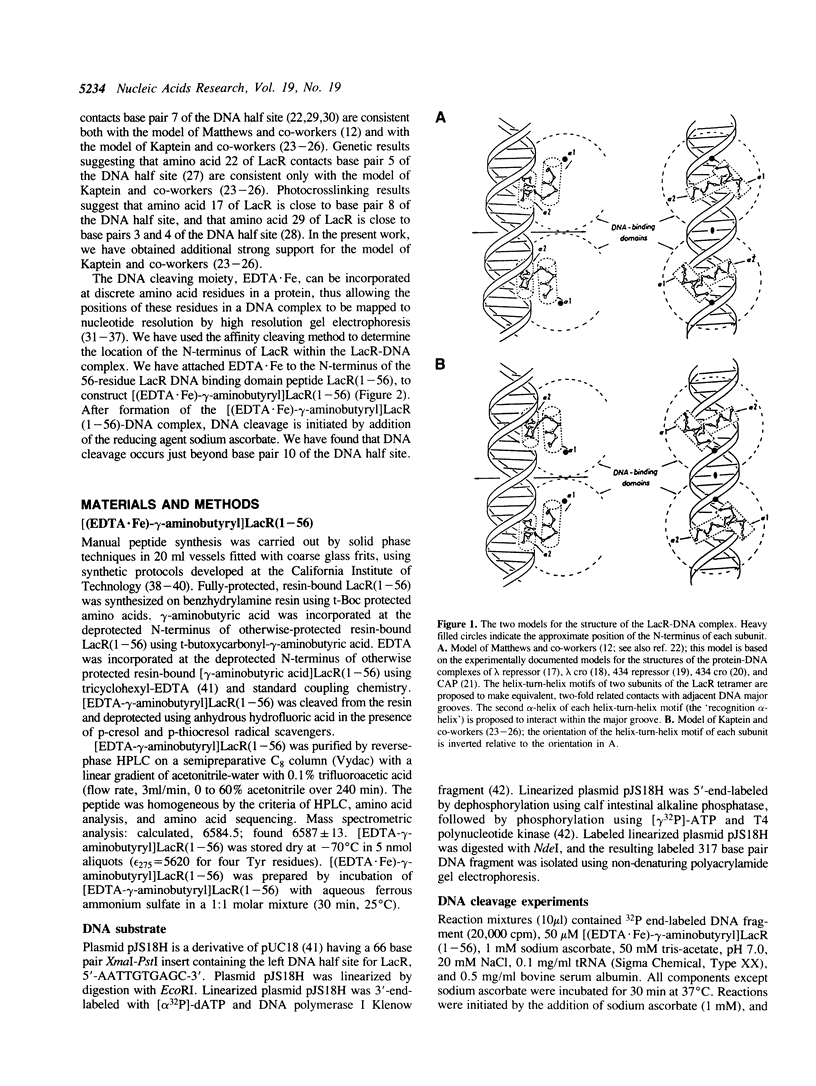
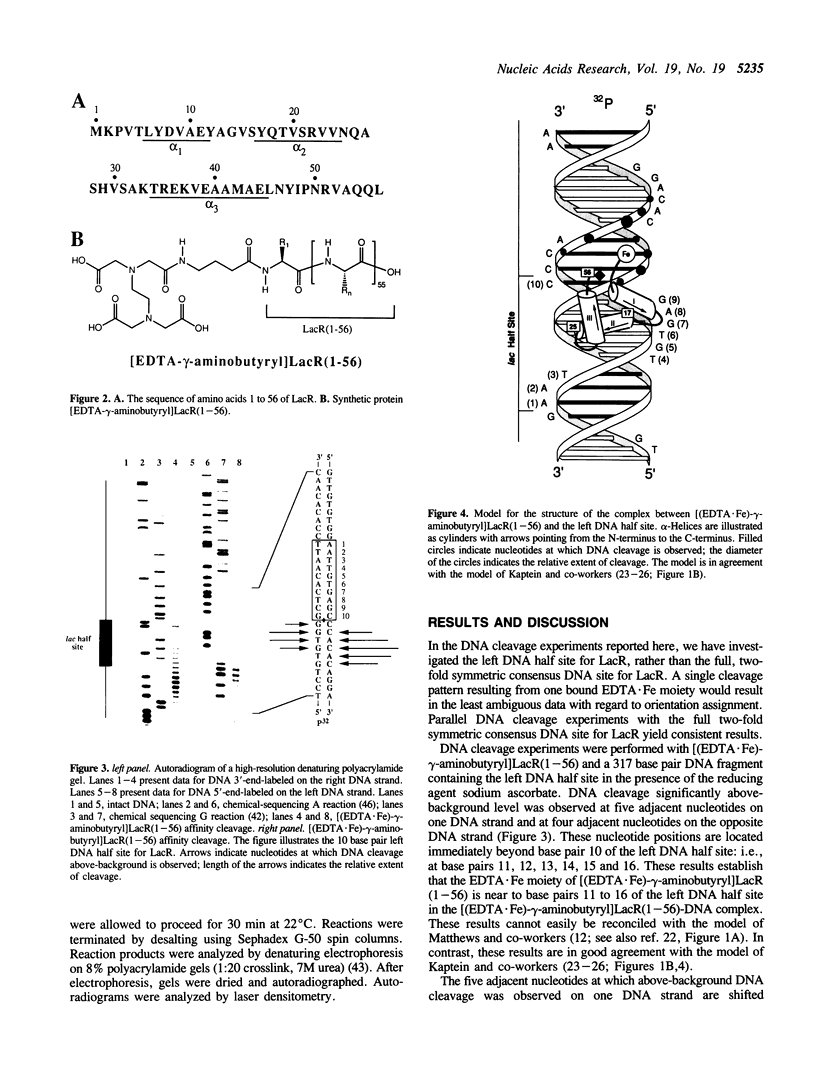
Lac repressor (LacR) is a helix-turn-helix motif sequence-specific DNA binding protein. Based on proton NMR spectroscopic investigations, Kaptein and co-workers have proposed that the helix-turn-helix motif of LacR binds to DNA in an orientation opposite to that of the helix-turn-helix motifs of lambda repressor, lambda cro, 434 repressor, 434 cro, and CAP [Boelens, R., Scheek, R., van Boom, J. and Kaptein, R., J. Mol. Biol. 193, 1987, 213-216]. In the present work, we have determined the orientation of the helix-turn-helix motif of LacR in the LacR-DNA complex by the affinity cleaving method. The DNA cleaving moiety EDTA.Fe was attached to the N-terminus of a 56-residue synthetic protein corresponding to the DNA binding domain of LacR. We have formed the complex between the modified protein and the left DNA half site for LacR. The locations of the resulting DNA cleavage positions relative to the left DNA half site provide strong support for the proposal of Kaptein and co-workers.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aggarwal A. K., Rodgers D. W., Drottar M., Ptashne M., Harrison S. C. Recognition of a DNA operator by the repressor of phage 434: a view at high resolution. Science. 1988 Nov 11;242(4880):899–907. doi: 10.1126/science.3187531. [DOI] [PubMed] [Google Scholar]
- Allen T. D., Wick K. L., Matthews K. S. Identification of amino acids in lac repressor protein cross-linked to operator DNA specifically substituted with bromodeoxyuridine. J Biol Chem. 1991 Apr 5;266(10):6113–6119. [PubMed] [Google Scholar]
- Arndt K. T., Boschelli F., Lu P., Miller J. H. lac Repressor: a proton magnetic resonance look at the deoxyribonucleic acid binding fragment. Biochemistry. 1981 Oct 13;20(21):6109–6118. doi: 10.1021/bi00524a030. [DOI] [PubMed] [Google Scholar]
- Boelens R., Scheek R. M., van Boom J. H., Kaptein R. Complex of lac repressor headpiece with a 14 base-pair lac operator fragment studied by two-dimensional nuclear magnetic resonance. J Mol Biol. 1987 Jan 5;193(1):213–216. doi: 10.1016/0022-2836(87)90638-3. [DOI] [PubMed] [Google Scholar]
- Brennan R. G., Roderick S. L., Takeda Y., Matthews B. W. Protein-DNA conformational changes in the crystal structure of a lambda Cro-operator complex. Proc Natl Acad Sci U S A. 1990 Oct;87(20):8165–8169. doi: 10.1073/pnas.87.20.8165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark-Lewis I., Aebersold R., Ziltener H., Schrader J. W., Hood L. E., Kent S. B. Automated chemical synthesis of a protein growth factor for hemopoietic cells, interleukin-3. Science. 1986 Jan 10;231(4734):134–139. doi: 10.1126/science.3079915. [DOI] [PubMed] [Google Scholar]
- Dervan P. B. Design of sequence-specific DNA-binding molecules. Science. 1986 Apr 25;232(4749):464–471. doi: 10.1126/science.2421408. [DOI] [PubMed] [Google Scholar]
- Ebright R. H. Evidence for a contact between glutamine-18 of lac repressor and base pair 7 of lac operator. Proc Natl Acad Sci U S A. 1986 Jan;83(2):303–307. doi: 10.1073/pnas.83.2.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ebright R. H. Use of "loss-of-contact" substitutions to identify residues involved in an amino acid-base pair contact: effect of substitution of Gln18 of lac repressor by Gly, Ser, and Leu. J Biomol Struct Dyn. 1985 Oct;3(2):281–297. doi: 10.1080/07391102.1985.10508417. [DOI] [PubMed] [Google Scholar]
- Geisler N., Weber K. Escherichia coli lactose repressor: isolation of two different homogeneous headpieces and the existence of a hinge region between residues 50 and 60 in the repressor molecule. FEBS Lett. 1978 Mar 15;87(2):215–218. doi: 10.1016/0014-5793(78)80335-4. [DOI] [PubMed] [Google Scholar]
- Graham K. S., Dervan P. B. Structural motif of the DNA binding domain of gamma delta-resolvase characterized by affinity cleaving. J Biol Chem. 1990 Sep 25;265(27):16534–16540. [PubMed] [Google Scholar]
- Iverson B. L., Dervan P. B. Adenine specific DNA chemical sequencing reaction. Nucleic Acids Res. 1987 Oct 12;15(19):7823–7830. doi: 10.1093/nar/15.19.7823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- JACOB F., MONOD J. Genetic regulatory mechanisms in the synthesis of proteins. J Mol Biol. 1961 Jun;3:318–356. doi: 10.1016/s0022-2836(61)80072-7. [DOI] [PubMed] [Google Scholar]
- Jordan S. R., Pabo C. O. Structure of the lambda complex at 2.5 A resolution: details of the repressor-operator interactions. Science. 1988 Nov 11;242(4880):893–899. doi: 10.1126/science.3187530. [DOI] [PubMed] [Google Scholar]
- Kaptein R., Lamerichs R. M., Boelens R., Rullmann J. A. Two-dimensional NMR study of a protein-DNA complex. lac repressor headpiece-operator interaction. Biochem Pharmacol. 1990 Jul 1;40(1):89–96. doi: 10.1016/0006-2952(90)90183-l. [DOI] [PubMed] [Google Scholar]
- Kaptein R., Zuiderweg E. R., Scheek R. M., Boelens R., van Gunsteren W. F. A protein structure from nuclear magnetic resonance data. lac repressor headpiece. J Mol Biol. 1985 Mar 5;182(1):179–182. doi: 10.1016/0022-2836(85)90036-1. [DOI] [PubMed] [Google Scholar]
- Kent S. B. Chemical synthesis of peptides and proteins. Annu Rev Biochem. 1988;57:957–989. doi: 10.1146/annurev.bi.57.070188.004521. [DOI] [PubMed] [Google Scholar]
- Lamerichs R. M., Boelens R., van der Marel G. A., van Boom J. H., Kaptein R., Buck F., Fera B., Rüterjans H. H NMR study of a complex between the lac repressor headpiece and a 22 base pair symmetric lac operator. Biochemistry. 1989 Apr 4;28(7):2985–2991. doi: 10.1021/bi00433a037. [DOI] [PubMed] [Google Scholar]
- Lehming N., Sartorius J., Oehler S., von Wilcken-Bergmann B., Müller-Hill B. Recognition helices of lac and lambda repressor are oriented in opposite directions and recognize similar DNA sequences. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7947–7951. doi: 10.1073/pnas.85.21.7947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mack D. P., Sluka J. P., Shin J. A., Griffin J. H., Simon M. I., Dervan P. B. Orientation of the putative recognition helix in the DNA-binding domain of Hin recombinase complexed with the hix site. Biochemistry. 1990 Jul 17;29(28):6561–6567. doi: 10.1021/bi00480a003. [DOI] [PubMed] [Google Scholar]
- Matthews B. W., Ohlendorf D. H., Anderson W. F., Takeda Y. Structure of the DNA-binding region of lac repressor inferred from its homology with cro repressor. Proc Natl Acad Sci U S A. 1982 Mar;79(5):1428–1432. doi: 10.1073/pnas.79.5.1428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Mondragón A., Harrison S. C. The phage 434 Cro/OR1 complex at 2.5 A resolution. J Mol Biol. 1991 May 20;219(2):321–334. doi: 10.1016/0022-2836(91)90568-q. [DOI] [PubMed] [Google Scholar]
- Nick H., Arndt K., Boschelli F., Jarema M. A., Lillis M., Sadler J., Caruthers M., Lu P. lac repressor-lac operator interaction: NMR observations. Proc Natl Acad Sci U S A. 1982 Jan;79(2):218–222. doi: 10.1073/pnas.79.2.218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oakley M. G., Dervan P. B. Structural motif of the GCN4 DNA binding domain characterized by affinity cleaving. Science. 1990 May 18;248(4957):847–850. doi: 10.1126/science.2111578. [DOI] [PubMed] [Google Scholar]
- Ogata R. T., Gilbert W. An amino-terminal fragment of lac repressor binds specifically to lac operator. Proc Natl Acad Sci U S A. 1978 Dec;75(12):5851–5854. doi: 10.1073/pnas.75.12.5851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogata R. T., Gilbert W. DNA-binding site of lac repressor probed by dimethylsulfate methylation of lac operator. J Mol Biol. 1979 Aug 25;132(4):709–728. doi: 10.1016/0022-2836(79)90384-x. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Sadler J. R., Sasmor H., Betz J. L. A perfectly symmetric lac operator binds the lac repressor very tightly. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6785–6789. doi: 10.1073/pnas.80.22.6785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasmor H. M., Betz J. L. Symmetric lac operator derivatives: effects of half-operator sequence and spacing on repressor affinity. Gene. 1990 Apr 30;89(1):1–6. doi: 10.1016/0378-1119(90)90198-z. [DOI] [PubMed] [Google Scholar]
- Simons A., Tils D., von Wilcken-Bergmann B., Müller-Hill B. Possible ideal lac operator: Escherichia coli lac operator-like sequences from eukaryotic genomes lack the central G X C pair. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1624–1628. doi: 10.1073/pnas.81.6.1624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sluka J. P., Horvath S. J., Bruist M. F., Simon M. I., Dervan P. B. Synthesis of a sequence-specific DNA-cleaving peptide. Science. 1987 Nov 20;238(4830):1129–1132. doi: 10.1126/science.3120311. [DOI] [PubMed] [Google Scholar]
- Steitz T. A. Structural studies of protein-nucleic acid interaction: the sources of sequence-specific binding. Q Rev Biophys. 1990 Aug;23(3):205–280. doi: 10.1017/s0033583500005552. [DOI] [PubMed] [Google Scholar]
- Takeda Y., Ohlendorf D. H., Anderson W. F., Matthews B. W. DNA-binding proteins. Science. 1983 Sep 9;221(4615):1020–1026. doi: 10.1126/science.6308768. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Zuiderweg E. R., Billeter M., Boelens R., Scheek R. M., Wüthrich K., Kaptein R. Spatial arrangement of the three alpha helices in the solution conformation of E. coli lac repressor DNA-binding domain. FEBS Lett. 1984 Sep 3;174(2):243–247. doi: 10.1016/0014-5793(84)81166-7. [DOI] [PubMed] [Google Scholar]
- Zuiderweg E. R., Kaptein R., Wüthrich K. Secondary structure of the lac repressor DNA-binding domain by two-dimensional 1H nuclear magnetic resonance in solution. Proc Natl Acad Sci U S A. 1983 Oct;80(19):5837–5841. doi: 10.1073/pnas.80.19.5837. [DOI] [PMC free article] [PubMed] [Google Scholar]



