Abstract
We evaluated a new line probe assay (LiPA) kit to identify Mycobacterium species and to detect mutations related to drug resistance in Mycobacterium tuberculosis. A total of 554 clinical isolates of Mycobacterium tuberculosis (n = 316), Mycobacterium avium (n = 71), Mycobacterium intracellulare (n = 51), Mycobacterium kansasii (n = 54), and other Mycobacterium species (n = 62) were tested with the LiPA kit in six hospitals. The LiPA kit was also used to directly test 163 sputum specimens. The results of LiPA identification of Mycobacterium species in clinical isolates were almost identical to those of conventional methods. Compared with standard drug susceptibility testing results for the clinical isolates, LiPA showed a sensitivity and specificity of 98.9% and 97.3%, respectively, for detecting rifampin (RIF)-resistant clinical isolates; 90.6% and 100%, respectively, for isoniazid (INH) resistance; 89.7% and 96.0%, respectively, for pyrazinamide (PZA) resistance; and 93.0% and 100%, respectively, for levofloxacin (LVX) resistance. The LiPA kit could detect target species directly in sputum specimens, with a sensitivity of 85.6%. Its sensitivity and specificity for detecting RIF-, PZA-, and LVX-resistant isolates in the sputum specimens were both 100%, and those for detecting INH-resistant isolates were 75.0% and 92.9%, respectively. The kit was able to identify mycobacterial bacilli at the species level, as well as drug-resistant phenotypes, with a high sensitivity and specificity.
INTRODUCTION
The emergence of multidrug-resistant (MDR) Mycobacterium tuberculosis, resistant to at least rifampin (RIF) and isoniazid (INH), markedly hinders the control of tuberculosis (8). Nontuberculous mycobacteria (NTM) are also associated with pulmonary diseases (2, 16). Drug resistance in M. tuberculosis is due to mutations, including rpoB mutations, associated with RIF resistance; mutations in katG, the promoter region of the fabG1-inhA operon, fabG1, furA, and inhA, associated with INH resistance; pncA mutations, associated with pyrazinamide (PZA) resistance; and gyrA mutations, associated with resistance to fluoroquinolones (FQ) (47). Hybridization-based line probe assays (LiPAs) detect mutations associated with resistance to RIF (12, 21, 33, 38), INH (3), PZA (42), and FQ (15).
A new LiPA kit was recently developed to identify clinically important Mycobacterium species and to detect drug resistance mutations in M. tuberculosis. Evaluation of this kit in six independent hospitals in Japan showed that this assay is promising for the rapid detection of drug-resistant tuberculosis and for identification of major NTM.
MATERIALS AND METHODS
Clinical isolates.
A total of 554 clinical isolates of M. tuberculosis and NTM were obtained between January 2005 and December 2009 from 554 patients with pulmonary tuberculosis or NTM-related disease in the following six hospitals in Japan: Japan Anti-Tuberculosis Association Fukujuji Hospital (hospital A), National Hospital Organization (NHO) Tokyo Hospital (hospital B), NHO Kinki-Chuo Chest Medical Center (hospital C), NHO Ibaraki Higashi Hospital (hospital D), Tokyo Metropolitan Tama Medical Center (hospital E), and Osaka Prefectural Medical Center for Respiratory and Allergic Diseases (hospital F). Each participating hospital provided 79 to 109 isolates, all of which were subjected to species identification and drug susceptibility testing (DST). The M. tuberculosis isolates included 160 that were susceptible to all drugs tested and 156 that were resistant to at least one of the drugs tested (see Table S1 in the supplemental material). Of the drug-resistant isolates, 88 were resistant to RIF, 138 were resistant to INH, 58 were resistant to PZA, and 57 were resistant to levofloxacin (LVX) (data not shown). Other isolates included Mycobacterium avium (n = 71), Mycobacterium intracellulare (n = 51), and Mycobacterium kansasii (n = 54), as well as other NTM (n = 62) (see Table S1).
Clinical specimens.
A total of 163 sputum specimens were obtained from patients suspected to have or previously diagnosed with tuberculosis or NTM disease (one specimen each) in the hospitals during the period from June 2009 to April 2010. These specimens were transported to the National Reference Laboratory of Tuberculosis (RIT) and stored at −80°C until tested. Each specimen was smeared and stained according to the Ziehl-Neelsen method, followed by treatment with an N-acetyl-l-cysteine–NaOH solution as described previously (41). Each pretreated specimen was resuspended in 1.5 ml of phosphate buffer (pH 6.8). Aliquots of 0.2 ml of each suspension were transferred to 1.5-ml tubes and subjected to LiPA. Further aliquots of 0.2 ml of each suspension were transferred to fresh 1.5-ml tubes and subjected to the PCR-based Cobas Amplicor MTB test (Roche Diagnostics, Basel, Switzerland) (9, 26, 31) and the Cobas Amplicor M. avium and M. intracellulare tests (Roche Diagnostics). Aliquots of 0.1 ml and 0.5 ml of each specimen were inoculated into egg-based modified Ogawa medium (27) containing 2% (wt/vol) KH2PO4 and into MGIT broth (Bactec MGIT 960; BD Biosciences, Sparks, MD), respectively, for mycobacterial examination. Aliquots of 1 ml of the suspension for LiPA were centrifuged for 15 min at 13,000 × g, and the supernatant was removed with a pipette. Tris-EDTA (TE) buffer (100 μl) was added to the pellet, and the solution was again centrifuged for 15 min at 13,000 × g. The pellet was suspended in 50 μl of TE buffer and incubated at 95°C for 30 min. Aliquots of the supernatant (5 μl) were used for each LiPA. The total time for all procedures was about 3 h.
Species identification.
M. tuberculosis was identified at hospitals A and B by use of TRCRapid M.TB kits (Tosoh Bioscience, Tokyo, Japan), based on the transcription-reverse transcription concerted reaction (13, 44), and at hospitals C to F and RIT by use of the Cobas Amplicor MTB test. M. avium and M. intracellulare were identified at hospitals C to F and RIT by use of the Cobas Amplicor M. avium and M. intracellulare tests, respectively. The M. avium complex (MAC) was identified at hospitals A and B by use of TRCRapid MAC kits (Tosoh Bioscience); isolates identified as MAC species were heat-killed and transported to RIT for species identification. The other NTM were identified at hospitals A, B, and D to F and at RIT by use of the DNA-DNA hybridization technique (DDH Mycobacteria Kyokuto; Kyokuto Pharmaceutical Industrial Co., Tokyo, Japan) (28) and at hospital C by using AccuProbe (Gen-Probe, San Diego, CA) (17, 18). NTM isolates that were not identified by commercial kits were subjected to 16S rRNA gene sequencing at RIT.
DST and pyrazinamidase activity assay.
DSTs for RIF, INH, PZA, and LVX were performed at each participating hospital. At hospitals A and B, the MGIT AST (BD Biosciences) test was performed to detect RIF, INH, and PZA resistance, and an egg-based Ogawa medium (24) (1% KH2PO4) method (Welpack S test; Nihon BCG Inc., Tokyo, Japan) was used to detect RIF, INH, and LVX resistance. At hospital C, MGIT AST, Welpack S, and a broth microdilution method (broth MIC MTB-I; Kyokuto Pharmaceutical Industrial Co.) were performed to detect RIF, INH, and LVX resistance. At hospital D, the egg-based Ogawa medium (1% KH2PO4) method (Bit Spectre-SR; Kyokuto Pharmaceutical Industrial Co., Tokyo, Japan) was performed to detect RIF, INH, and LVX resistance, and a broth method was used to detect PZA resistance (PZA broth; Kyokuto Pharmaceutical Industrial Co.). At hospital E, the MGIT AST test was used to test for RIF, INH, and PZA resistance (LVX resistance was not tested at this hospital). At hospital F, the broth MIC MTB-I test was used to test for RIF, INH, and LVX resistance, and PZA broth (Kyokuto) was used to detect PZA resistance. At RIT, the standard proportion method using Ogawa medium (1% KH2PO4) was used to test for RIF, INH, and LVX resistance, and the MGIT AST test was used to test for PZA resistance. Isolates showing discordant results between phenotypic and genotypic DSTs for PZA were transferred to RIT and their pyrazinamidase activities tested (45), except for six isolates that had not been stored at the hospital. The INH resistance levels were as follows: isolates were considered resistant to INH at 0.2 μg/ml when they were resistant to INH at 0.2 μg/ml and susceptible to INH at 1.0 μg/ml; isolates were considered resistant to INH at 1.0 μg/ml when they were resistant to INH at 1.0 μg/ml. All kits for identification of mycobacteria and DSTs used in this study were recommended by the Japanese Society for Tuberculosis and approved as diagnosis reagents by the Ministry of Health, Labor and Welfare, Japan.
LiPA.
LiPA was performed as described previously (3, 42), using 121 oligonucleotide probes (see Table S2 in the supplemental material) immobilized onto four strips, called the NTM/MDR-TB, INH, PZA, and FQ strips (Nipro Co., Osaka, Japan). All clinical isolates and all sputum specimens were tested by LiPAs using all four strips, regardless of the results of any particular strip. The NTM/MDR-TB strip was designed to identify four Mycobacterium species—M. tuberculosis, M. avium, M. intracellulare, and M. kansasii—and to detect mutations associated with RIF and INH resistance in M. tuberculosis. The INH, PZA, and FQ strips were designed to detect mutations associated with INH, PZA, and FQ resistance of M. tuberculosis, respectively. The corresponding regions and mutations for each probe are shown in Table S2. A probe designed to detect the wild-type sequence of M. tuberculosis was designated as S probe, whereas a probe designed to detect a mutant sequence frequently found in drug-resistant M. tuberculosis was designated as R probe. On the INH strip, 46 S probes covered various regions of the following M. tuberculosis genes: PfabG1-inhA (inhA-1), inhA (inhA-2), fabG1 (fabG1-1 and -2), furA (furA-1 and -2), and katG (katG-1 to -40) (3). The katG probes covered 90 mutations related to INH resistance. On the PZA strip, 47 S probes covered regions of M. tuberculosis pncA (pncA-1 to -47), with 2 probes (pncA-16 and -17) containing a silent mutation in pncA (42). Probes inhA-S6 and -S7 and katG-S8 to -S11 on the NTM/MDR-TB strip were the same as inhA-1 and -2 and katG-20, -22, -23, and -24 on the INH strip, respectively.
Using biotinylated primers, the following products were obtained by nested PCR: rpoB (290 bp), PfabG1-inhA (477 bp), and katG (248 bp) for the NTM/MDR-TB strip; PfabG1-inhA (477 bp), fabG1 (209 bp), furA (256 bp), and katG (612 bp, 698 bp, and 907 bp) for the INH strip; pncA (641 bp) for the PZA strip; and gyrA (379 bp) for the FQ strip. The immobilized probes on each strip were hybridized with the PCR products at 62°C for 30 min and then incubated with streptavidin labeled with alkaline phosphatase at room temperature for 30 min. Color was developed by incubation with 5-bromo-4-chloro-3-indolylphosphate p-toluidine and nitroblue tetrazolium.
The presence or absence of bands, i.e., hybridization signals, on all strips was judged independently by three different observers. The results of LiPA were interpreted as follows. For identification of Mycobacterium species, when a signal was observed on the NTM/MDR-TB strip with any of the four probes (rpoB-AVI, rpoB-INT, rpoB-KAN, and rpoB-TB), the sample was thought to contain the corresponding Mycobacterium species. Conversely, when no signals were observed, the sample contained none of these four species. For detection of drug-resistant M. tuberculosis, when no signal was observed with any of the S probes, M. tuberculosis in the sample was considered resistant to the corresponding drug. In addition, when a signal(s) was observed with any of the R probes, the samples contained drug-resistant M. tuberculosis with the corresponding mutation(s). It took about 7 h to complete all procedures of the LiPA method.
DNA sequencing.
The PCR products were sequenced. The sequenced samples were as follows: 1 isolate and 1 clinical specimen that showed discrepancies in species identification between conventional methods and LiPA and 40 isolates and 4 clinical specimens that showed discrepancies in drug susceptibility between phenotypic DST and LiPA. DNA sequences were compared with the sequence of M. tuberculosis H37Rv by using Genetyx-Mac, version 14.0.2 (Genetyx Corporation, Tokyo, Japan). We also sequenced the 16S rRNA genes of NTM isolates when they could not be identified by conventional identification kits. The sequences of the 16S rRNA genes were analyzed with software for DNA sequence-based diagnosis, published by the Ribosomal Differentiation of Microorganisms Project (RIDOM) (19), or with the Basic Local Alignment Search Tool (BLAST) to identify the species.
Ethical considerations.
The study protocol was carefully reviewed and approved by the ethics committee of each participating hospital (hospital A approval date, 29 January 2009; hospital B approval date, 30 April 2009 [approval number 21-02-Da]; hospital C approval date, 14 November 2008 [approval number 20-18]; hospital D approval date, 18 September 2008; hospital E approval date, 28 November 2008; hospital F approval date, 28 March 2009 [approval number 5-84]). All clinical sputum specimens were collected after obtaining written informed consent from the participants.
RESULTS
Identification of clinical isolates.
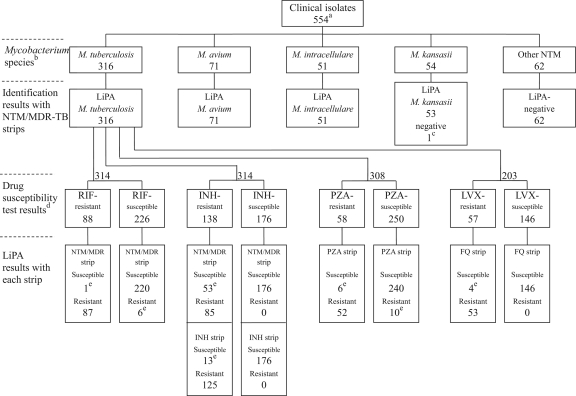
Among 554 isolates, LiPA results for species identified as M. tuberculosis, M. avium, and M. intracellulare showed 100% agreement with those of conventional genetic methods (Fig. 1; see Table S3 in the supplemental material). Of 54 M. kansasii isolates, 53 were identified as M. kansasii by the LiPA kit. The one discrepant isolate of M. kansasii carried an rpoB sequence identical to that of subtype III of the seven subtypes of M. kansasii, defined by sequence polymorphisms in hsp65 (1, 43). LiPA results were negative for 62 isolates of other NTM species.
Fig 1.
Distribution of LiPA results for 554 clinical isolates. a, number of clinical isolates. b, Mycobacterium species identified by conventional methods. c, M. kansasii subtype III. d, drug susceptibility testing and assays for pyrazinamidase activity were performed at each hospital (see the supplemental material). Of 316 M. tuberculosis isolates, 314 were subjected to RIF and INH susceptibility testing, 308 to PZA susceptibility testing, and 203 to LVX susceptibility testing. e, some isolates showed different results between DST and LiPA. DNA sequences of each target gene were determined for these discrepant isolates (see the footnote in Table S4 in the supplemental material).
Correlation between conventional DST and LiPA results.
LiPA results were compared with those of DST (Table 1; see Table S4 in the supplemental material).
Table 1.
Diagnostic performance of LiPA in comparison with drug susceptibility testing
| Antituberculosis drug (strip used in LiPAc) | Clinical isolates |
Clinical samples (sputa) |
||
|---|---|---|---|---|
| Sensitivitya | Specificityb | Sensitivitya | Specificityb | |
| RIF (NTM/MDR-TB strip) | 98.9 (87/88) | 97.3 (220/226) | 100 (3/3) | 100 (52/52) |
| INH (INH strip) | 90.6 (125/138) | 100 (176/176) | 75.0 (3/4) | 92.9 (39/42) |
| INH (NTM/MDR-TB strip) | 61.6 (85/138) | 100 (176/176) | 50.0 (3/6) | 97.8 (45/46) |
| PZA (PZA strip) | 89.7 (52/58) | 96.0 (240/250) | 100 (4/4) | 100 (52/52) |
| LVX (FQ strip) | 93.0 (53/57) | 100 (146/146) | 100 (7/7) | 100 (48/48) |
Data are percentages (no. of drug-resistant samples by LiPA/no. of drug-resistant samples by DST).
Data are percentages (no. of drug-susceptible samples by LiPA/no. of drug-susceptible samples by DST).
LiPA was performed using four strips, namely, NTM/MDR-TB, INH, PZA, and FQ strips (see the supplemental material). The NTM/MDR-TB strip was designed to identify four Mycobacterium species and to detect mutations associated with RIF resistance and INH resistance (C-15T and T-8C mutations in PfabG1-inhA and S315T and S315N mutations in katG). The INH, PZA, and FQ strips were designed to detect mutations associated with INH, PZA, and FQ resistance of M. tuberculosis, respectively. The corresponding regions and mutations for each probe are shown in Table S1 in the supplemental material. The INH strip covered 46 regions of the following M. tuberculosis genes: PfabG1-inhA, inhA, fabG1, furA, and katG (3). The PZA strip covered pncA (40), and the FQ strip covered gyrA (4).
(i) RIF resistance and rpoB mutations.
LiPA identified 98.9% (87/88 isolates) of RIF-resistant isolates and 97.3% (220/226 isolates) of RIF-susceptible isolates when M. tuberculosis isolates were tested using NTM/MDR-TB strips designed to detect rpoB mutations (Table 1). Of all the isolates tested, seven showed discrepancies between DST and LiPA for RIF susceptibility testing. One isolate, identified as RIF resistant by DST but RIF susceptible by LiPA, had an I572F substitution. The remaining six were identified as RIF susceptible by DST but RIF resistant by LiPA. Of these, three had an H526S substitution, while the other three had an L511P mutation, a D516Y mutation, and a silent mutation at codon 516 (GAC → GAT). Of these six isolates, the three with the H526S mutation and the one with the L511P mutation were reported by hospital C as RIF susceptible by the MGIT AST and Welpack S tests but as RIF “intermediate” (MICs of 0.25 mg/liter and 0.5 mg/liter, respectively) by the broth MIC MTB-I test.
(ii) INH resistance and mutations of PfabG1-inhA, fabG1, furA, and katG.
The INH strip was designed to detect mutations associated with INH resistance in M. tuberculosis, including mutations in PfabG1-inhA (C-15T and T-8C), fabG1 (G609A [L203L]), furA (C41T [A14V]), and katG (see Table S2 in the supplemental material). The strips identified 90.6% (125/138 isolates) of INH-resistant isolates and 100% (176/176 isolates) of INH-susceptible isolates (Table 1). Thirteen isolates were found to be INH resistant by DST but INH susceptible by LiPA (see Table S4). Of these, 10 had no mutations in the amplified regions for the INH strip, while the other 3 had S17N, G206S, and E340Q substitutions in katG.
The NTM/MDR-TB strip was designed to detect mutations of PfabG1-inhA (C-15T and T-8C) and katG (S315T and S315N) (see Table S2 in the supplemental material) which are frequently detected in INH-resistant clinical isolates (41, 47). NTM/MDR-TB strips identified 61.6% (85/138 isolates) of INH-resistant isolates and 100% (176/176 isolates) of INH-susceptible isolates (Table 1). Fifty-three isolates were identified as INH resistant by DST but INH susceptible by LiPA using NTM/MDR-TB strips. Of these, 13 isolates and the remaining 40 isolates were identified as INH susceptible and INH resistant, respectively, by LiPA using INH strips (see Table S4). Of these 40 INH-resistant isolates, 21 were resistant to INH at 1.0 μg/ml, and the remaining 19 isolates were resistant to INH at 0.2 μg/ml. Of the 21 isolates resistant to INH at 1.0 μg/ml, 5 showed no hybridization with the fabG1-1 probe (G609A [L203L]); 12 showed no hybridization with any of the katG probes, including katG-1 (1 isolate with a Δ152A mutation [frameshift]), katG-5 (1 isolate with an A338C [Y113S] mutation), katG-6 (2 isolates with a Δ367G mutation [frameshift]), katG-8 (1 isolate with a G412T [N138Y] mutation), katG-9 (1 isolate with an A425G [D142G] mutation), katG-10 (1 isolate with an A454C [K152Q] mutation), katG-11 (1 isolate with a G487A [D163N] mutation), katG-15 (1 isolate with a T571G [W191G] mutation), katG-29 (1 isolate with an A1382C [Q461P] mutation), katG-37 (1 isolate with a G1795T [G599stop] mutation), and katG-39 (1 isolate with a T2093C [F698S] mutation); 2 showed no hybridization with two katG probes, either katG-21 and katG-25 (A922C [T308P] and G1037C [S346T] mutations) or katG-39 and katG-40 (Δ1991-2173 [frameshift] mutation); 1 showed no hybridization with katG-26 to -40 (the DNA sequence was not determined); and 1 showed no hybridization with the fabG1-1 (G609A [L203L] mutation) and katG-6 (G378T [M126I] mutation) probes. Of the 19 isolates resistant to INH at 0.2 μg/ml, 12 showed no hybridization with the fabG1-1 probe (G609A [L203L] mutation), 6 showed no hybridization with the katG-28 probe (G1255C [D419H] mutation), and 1 showed no hybridization with the katG-28 and -34 probes (sequence not determined).
(iii) PZA resistance and pncA mutations.
The PZA strip was designed to detect pncA mutations associated with PZA resistance in M. tuberculosis (42). The LiPA test identified 89.7% (52/58 isolates) of PZA-resistant and 96.0% (240/250 isolates) of PZA-susceptible isolates (Table 1). Sixteen isolates showed discrepancies between DST and LiPA results (see Table S4 in the supplemental material). Six isolates found to be PZA resistant by DST but PZA susceptible by LiPA had no mutations in pncA. Ten other isolates were PZA susceptible by DST but PZA resistant by LiPA. Of these 10 isolates, 4 had G162S, 2 had G17D, 2 had T168I, 2 had G132D, and 2 had V147I substitutions.
(iv) FQ resistance and gyrA mutations.
The FQ strip was designed to detect gyrA mutations associated with FQ resistance in M. tuberculosis (see Table S2 in the supplemental material). FQ strips identified 93.0% (53/57 isolates) of LVX-resistant and 100% (146/146 isolates) of LVX-susceptible isolates (Table 1). Four isolates found to be LVX resistant by DST but LVX susceptible by LiPA had no mutations in gyrA. These isolates also had no mutations in gyrB.
Direct identification of Mycobacterium species and detection of drug-resistant M. tuberculosis in sputum specimens.
A total of 163 sputum specimens were collected from patients who had been diagnosed with or were suspected to have pulmonary tuberculosis or NTM diseases.
(i) Detection and identification of Mycobacterium species in sputum specimens.
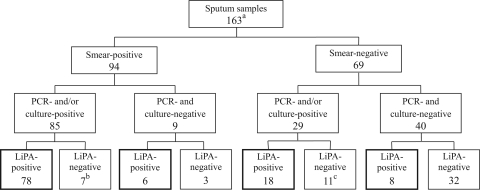
Direct application of the LiPA kit to sputum samples for species identification showed high degrees of consistency and efficiency that were comparable with those of conventional methods. The sensitivity of LiPA with NTM/MDR-TB strips was 90.2% (74/82 specimens) for M. tuberculosis, 84.6% (11/13 specimens) for M. avium, 54.5% (6/11 specimens) for M. intracellulare, and 80.0% (4/5 specimens) for M. kansasii (see Table S5 in the supplemental material). One specimen, which was misidentified by LiPA, was found to be Mycobacterium rhodesiae by DNA sequence analysis. The overall sensitivity of LiPA with NTM/MDR-TB strips for detection of target species was 85.6% (95/111 specimens) (see Table S5). Eighteen samples were LiPA negative despite being PCR and/or culture positive (7 smear-positive and 11 smear-negative samples) (Fig. 2), whereas 14 samples were LiPA positive despite being PCR and culture negative (6 smear-positive and 8 smear-negative samples) (Fig. 2).
Fig 2.
Distribution of LiPA results obtained with NTM/MDR-TB strips for the detection of target species in 163 sputum samples. a, number of clinical samples. b, one of these isolates was Mycobacterium fortuitum. c, one of these isolates was Mycobacterium abscessus.
(ii) Correlation of conventional DST and LiPA results for sputum specimens.
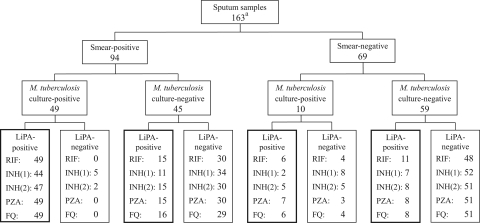
Among 163 samples, 49 smear-positive and 10 smear-negative samples were culture positive for M. tuberculosis (Fig. 3). For the 49 smear-positive samples, LiPA results for any drug susceptibility ranged from 89.8% (44/49 specimens) to 100% (49/49 specimens); for the 10 smear-negative samples, LiPA results ranged from 20% (2/10 specimens) to 70% (7/10 specimens). LiPA results were even obtained for some culture-negative samples, although these results could not be compared with those of DST. LiPA results were obtained for 11 to 16 of 45 smear-positive and M. tuberculosis culture-negative samples and for 7 to 11 of 59 smear-negative and M. tuberculosis culture-negative samples (Fig. 3). Direct application of the LiPA kit to sputum samples showed high sensitivities and specificities for detection of resistance to RIF, PZA, and LVX, whereas LiPA for detection of INH resistance showed a relatively low sensitivity (Table 1; see Table S6 in the supplemental material). However, its sensitivity and specificity were improved by using the INH strip.
Fig 3.
Distribution of LiPA results obtained with four strips for detection of a mutation(s) associated with drug resistance in 163 sputum samples. a, number of clinical samples. RIF, RIF susceptibility with NTM/MDR-TB strips; INH(1), INH susceptibility with INH strips; INH(2), INH susceptibility with NTM/MDR-TB strips; PZA, PZA susceptibility with PZA strips; FQ, LVX susceptibility with FQ strips.
Eight specimens showed discordance between DST and LiPA results (see Table S6 in the supplemental material). Four showed discordance between DST results for INH susceptibility and LiPA results obtained using INH strips. Of these, isolates from two specimens that were INH resistant by LiPA had a fabG1 (G609A [L203L]) mutation, and one had a PfabG1-inhA (C-15T) mutation. Four specimens showed discordance between DST results for INH susceptibility and LiPA results obtained using NTM/MDR-TB strips (see Table S6). Of these, two specimens indicated as INH susceptible with NTM/MDR-TB strips were identified as INH resistant with INH strips, and isolates from these two specimens had katG mutations, i.e., G1795T (G599stop) and T2093C (F698S) mutations. One specimen was also identified as INH susceptible with the INH strip, and DNA sequencing revealed that an isolate from the specimen had two katG mutations (T571C [W191R] and G1079A [G360D]). One specimen showing INH resistance with the NTM/MDR-TB strip had a PfabG1-inhA (C-15T) mutation.
As shown in Fig. 3, for the culture-negative specimens, LiPA results were obtained for 26 specimens (15 smear-positive and 11 smear-negative specimens) for RIF susceptibility, 18 specimens (11 smear-positive and 7 smear-negative specimens) for INH susceptibility with INH strips, 23 specimens (15 smear-positive and 8 smear-negative specimens) for INH susceptibility with NTM/MDR-TB strips, 23 specimens (15 smear-positive and 8 smear-negative specimens) for PZA susceptibility, and 24 specimens (16 smear-positive and 8 smear-negative specimens) for LVX susceptibility. Of these, no specimens were found to be RIF resistant, three were INH resistant, one was PZA resistant, and none were LVX resistant (data not shown).
DISCUSSION
The newly developed LiPA kit successfully identified important Mycobacterium species, including M. kansasii, except for subtype III of M. kansasii. The rpoB-KAN probe on the NTM/MDR-TB strip is compatible with the rpoB genes of subtypes I, II, IV, and V but not III and VI, perhaps explaining why this LiPA kit was unable to identify a subtype III M. kansasii isolate (Fig. 1; see Table S3 in the supplemental material). Among the isolates of M. kansasii obtained in four European countries, the majority belonged to subtypes I (68%) and II (31%), with only 1% belonging to subtype III (1). Similar distributions of subtypes were reported in Switzerland (subtype I, 67%; subtype II, 21%; subtype III, 8%; and other subtypes, 4%) (43) and in Catalonia, Spain (subtype I, 98%; subtype VI, 2%) (39). These epidemiological results indicate that subtype III of M. kansasii causes significantly fewer human infections than subtypes I and II. Although the LiPA kit showed a significantly efficient performance, improvements are required for detection of subtype III.
Seven isolates showing discrepancies between DST and LiPA for detection of RIF resistance had an I572F, D516D (a silent mutation), L511P, D516Y, or H526S mutation in rpoB. With the exception of the D516D mutation, these mutations are associated with RIF resistance. The I572F mutation has been reported to be associated with RIF resistance (46), although this mutation was not covered by the probes on the NTM/MDR-TB strip. The L511P and D516Y mutations have been reported to be associated with RIF resistance (25). RIF-resistant isolates have been reported to possess at least 11 mutations in codon 526, resulting in amino acid mutation of H to C, D, E, G, L, N, P, R, Q, T, or Y but not to S (47). An H526S mutation would be associated with RIF resistance.
LiPA using NTM/MDR-TB strips to detect mutations associated with INH resistance showed a low sensitivity (61.6%) among the isolates, although the strips were able to detect the most frequent mutations found in INH-resistant isolates, including S315T and S315N mutations in katG and C-15T and T-8C mutations in the promoter region of inhA (34, 35, 47). The Genotype MTBDRplus kit (Hain Lifescience, Nehren, Germany), a commercially available LiPA kit that uses a strip to detect these mutations, showed various degrees of sensitivity to INH-resistant M. tuberculosis isolates, including MDR isolates, in several countries, i.e., 92% in Germany (20), 82% in Taiwan (23), 73% in Spain (29), 67% in Italy (30), and 66% in Japan (10). The frequency of INH-resistant clinical isolates with S315T and S315N katG mutations and C-15T and T-8C mutations in the promoter region of inhA depends on the geographical origin of isolates. INH-resistant isolates with these mutations make up relatively small populations in Japan and Italy.
LiPA using INH strips, which covered more mutations, showed greater sensitivity than that with NTM/MDR-TB strips. Thirteen isolates were found to be INH resistant by DST but INH susceptible by LiPA using INH strips (see Table S4 in the supplemental material). No mutations were detected in 10 isolates, indicating that mutations in other genes may be associated with INH resistance. One isolate had a mutation of katG (S17N) which has been reported to confer INH resistance (11) but which is located away from the target sites. Of the remaining two isolates, one each had katG G206S and katG E340Q mutations, neither of which has been reported previously, to our knowledge. Both may be associated with INH resistance.
LiPA using PZA strips to detect mutations associated with PZA resistance showed high sensitivity and specificity. However, discrepancies between LiPA and DST were observed for 16 isolates. Six isolates were identified as PZA resistant by DST but PZA susceptible by LiPA, with none of these having a mutation in pncA, although one was positive in the pyrazinamidase test. Pyrazinamidase-positive but PZA-resistant strains are very rare and usually show a low level of resistance (36). The PZA resistance of the pyrazinamidase-positive strain may have been due to a mechanism other than pncA mutation (37). Ten isolates were identified as PZA susceptible by DST but PZA resistant by LiPA. Of these, two had pncA mutations causing T168I substitution, and one had a V147I mutation. These three isolates were positive in the pyrazinamidase test, suggesting that these mutations are not related to PZA resistance. Four isolates had a pncA G162S mutation, two had a G17D mutation, and one had a G132D mutation. These isolates were PZA susceptible by DST, but they were not tested for pyrazinamidase activity. To our knowledge, the G162S mutation has not been reported previously. The G17D and G132D mutations have been reported to confer PZA resistance (22). These discrepancies may have been due to the limited efficiency of DST methods (6).
LiPA with FQ strips for detection of mutations associated with FQ resistance showed high sensitivity and specificity, with only four isolates showing discordant results. None of these four had a mutation in gyrA, indicating that the FQ strips could detect all known mutations associated with FQ resistance. These four isolates had no mutation in gyrB, which was recently reported to confer FQ resistance in clinical isolates without gyrA mutations (7, 14, 32). Alternatively, the results of DST may show false resistance.
Of 163 sputum samples, 14 were LiPA positive but PCR and culture negative (Fig. 2). The results of LiPA for these 14 samples are likely correct assignments, as all came from patients previously diagnosed by culture methods or PCR as having tuberculosis or NTM diseases, showing 100% agreement. However, many of these results could have come from the shedding of nonviable bacilli from previously treated patients. Therefore, nucleic acid amplification methods, including LiPA, need to be interpreted carefully for previously treated tuberculosis patients. LiPA was always performed with a negative control and repeated when the results were in discordance with those of conventional methods. However, the discrepancies may be explained by cross-contamination during LiPA procedures.
The LiPA kit might be useful for rapid diagnosis of MDR tuberculosis, especially in Asian countries, where the genetic characteristics of INH resistance are unique (36). It is also important to detect resistance to PZA and LVX in MDR tuberculosis, as the majority of MDR M. tuberculosis isolates have been reported to be resistant to either PZA or LVX in Japan (4, 5).
The LiPA kit reported here is the first genetic diagnosis kit that can simultaneously identify the major clinical isolates of Mycobacterium species and detect mutations associated with resistance to INH, RIF, PZA, and FQ. The present study provides a unique perspective for assessing the overall reliability, specificity, and sensitivity of this kit in comparison with conventional tests. The LiPA kit may also be useful in laboratories in developing countries where mycobacterial culture cannot be performed. However, a follow-up culture-based DST is recommended where resources permit.
Supplementary Material
ACKNOWLEDGMENTS
This study was supported by grants from the Ministry of Health, Labor, and Welfare of Japan (H21-SHINKO-IPPAN-016) and the Nipro Corporation (3R07020).
Footnotes
Published ahead of print 28 December 2011
Supplemental material for this article may be found at http://jcm.asm.org/.
REFERENCES
- 1. Alcaide F, et al. 1997. Heterogeneity and clonality among isolates of Mycobacterium kansasii: implications for epidemiological and pathogenicity studies. J. Clin. Microbiol. 35:1959–1964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Alvarez-Uria G. 2010. Lung disease caused by nontuberculous mycobacteria. Curr. Opin. Pulm. Med. 16:251–256 [DOI] [PubMed] [Google Scholar]
- 3. Ando H, et al. 2010. Identification of katG mutations associated with high-level isoniazid resistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 54:1793–1799 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Ando H, et al. 2011. Evaluation of a line probe assay for the rapid detection of gyrA mutations associated with fluoroquinolone resistance in multidrug-resistant Mycobacterium tuberculosis. J. Med. Microbiol. 60:184–188 [DOI] [PubMed] [Google Scholar]
- 5. Ando H, et al. 2010. Pyrazinamide resistance in multidrug-resistant Mycobacterium tuberculosis isolates in Japan. Clin. Microbiol. Infect. 16:1164–1168 [DOI] [PubMed] [Google Scholar]
- 6. Aono A, Hirano K, Hamasaki S, Abe C. 2002. Evaluation of BACTEC MGIT 960 PZA medium for susceptibility testing of Mycobacterium tuberculosis to pyrazinamide (PZA): compared with the results of pyrazinamidase assay and Kyokuto PZA test. Diagn. Microbiol. Infect. Dis. 44:347–352 [DOI] [PubMed] [Google Scholar]
- 7. Aubry A, et al. 2006. Novel gyrase mutations in quinolone-resistant and -hypersusceptible clinical isolates of Mycobacterium tuberculosis: functional analysis of mutant enzymes. Antimicrob. Agents Chemother. 50:104–112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Aziz MA, et al. 2006. Epidemiology of antituberculosis drug resistance (the Global Project on Anti-Tuberculosis Drug Resistance Surveillance): an updated analysis. Lancet 368:2142–2154 [DOI] [PubMed] [Google Scholar]
- 9. Bodmer T, Gurtner A, Scholkmann M, Matter L. 1997. Evaluation of the COBAS AMPLICOR MTB system. J. Clin. Microbiol. 35:1604–1605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Chikamatsu K, et al. 2011. Evaluation of GenoTypeR MTBDRplus for the detection of multi-drug resistant Mycobacterium tuberculosis strains. Kekkaku 86:697–702 [PubMed] [Google Scholar]
- 11. Cockerill FR, III, et al. 1995. Rapid identification of a point mutation of the Mycobacterium tuberculosis catalase-peroxidase (katG) gene associated with isoniazid resistance. J. Infect. Dis. 171:240–245 [DOI] [PubMed] [Google Scholar]
- 12. Cooksey RC, Morlock GP, Glickman S, Crawford JT. 1997. Evaluation of a line probe assay kit for characterization of rpoB mutations in rifampin-resistant Mycobacterium tuberculosis isolates from New York City. J. Clin. Microbiol. 35:1281–1283 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Drouillon V, et al. 2009. Multicenter evaluation of a transcription-reverse transcription concerted assay for rapid detection of Mycobacterium tuberculosis complex in clinical specimens. J. Clin. Microbiol. 47:3461–3465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Duong DA, et al. 2009. Beijing genotype of Mycobacterium tuberculosis is significantly associated with high-level fluoroquinolone resistance in Vietnam. Antimicrob. Agents Chemother. 53:4835–4839 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Giannoni F, et al. 2005. Evaluation of a new line probe assay for rapid identification of gyrA mutations in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 49:2928–2933 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Glassroth J. 2008. Pulmonary disease due to nontuberculous mycobacteria. Chest 133:243–251 [DOI] [PubMed] [Google Scholar]
- 17. Gonzalez R, Hanna BA. 1987. Evaluation of Gen-Probe DNA hybridization systems for the identification of Mycobacterium tuberculosis and Mycobacterium avium-intracellulare. Diagn. Microbiol. Infect. Dis. 8:69–77 [DOI] [PubMed] [Google Scholar]
- 18. Goto M, Oka S, Okuzumi K, Kimura S, Shimada K. 1991. Evaluation of acridinium-ester-labeled DNA probes for identification of Mycobacterium tuberculosis and Mycobacterium avium-Mycobacterium intracellulare complex in culture. J. Clin. Microbiol. 29:2473–2476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Harmsen D, et al. 2003. RIDOM: comprehensive and public sequence database for identification of Mycobacterium species. BMC Infect. Dis. 3:26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Hillemann D, Rusch-Gerdes S, Richter E. 2007. Evaluation of the GenoType MTBDRplus assay for rifampin and isoniazid susceptibility testing of Mycobacterium tuberculosis strains and clinical specimens. J. Clin. Microbiol. 45:2635–2640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Hirano K, Abe C, Takahashi M. 1999. Mutations in the rpoB gene of rifampin-resistant Mycobacterium tuberculosis strains isolated mostly in Asian countries and their rapid detection by line probe assay. J. Clin. Microbiol. 37:2663–2666 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Hirano K, Takahashi M, Kazumi Y, Fukasawa Y, Abe C. 1997. Mutation in pncA is a major mechanism of pyrazinamide resistance in Mycobacterium tuberculosis. Tuber. Lung Dis. 78:117–122 [DOI] [PubMed] [Google Scholar]
- 23. Huang WL, Chen HY, Kuo YM, Jou R. 2009. Performance assessment of the GenoType MTBDRplus test and DNA sequencing in detection of multidrug-resistant Mycobacterium tuberculosis. J. Clin. Microbiol. 47:2520–2524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Kantor IN, et al. 1998. Laboratory services in tuberculosis control. III. Culture. World Health Organization, Geneva, Switzerland [Google Scholar]
- 25. Kapur V, et al. 1994. Characterization by automated DNA sequencing of mutations in the gene (rpoB) encoding the RNA polymerase beta subunit in rifampin-resistant Mycobacterium tuberculosis strains from New York City and Texas. J. Clin. Microbiol. 32:1095–1098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Katila ML, Katila P, Erkinjuntti-Pekkanen R. 2000. Accelerated detection and identification of mycobacteria with MGIT 960 and COBAS AMPLICOR systems. J. Clin. Microbiol. 38:960–964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Kudoh S, Kudoh T. 1974. A simple technique for culturing tubercle bacilli. Bull. World Health Organ. 51:71–82 [PMC free article] [PubMed] [Google Scholar]
- 28. Kusunoki S, et al. 1991. Application of colorimetric microdilution plate hybridization for rapid genetic identification of 22 Mycobacterium species. J. Clin. Microbiol. 29:1596–1603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Lacoma A, et al. 2008. GenoType MTBDRplus assay for molecular detection of rifampin and isoniazid resistance in Mycobacterium tuberculosis strains and clinical samples. J. Clin. Microbiol. 46:3660–3667 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Miotto P, et al. 2006. Use of genotype MTBDR assay for molecular detection of rifampin and isoniazid resistance in Mycobacterium tuberculosis clinical strains isolated in Italy. J. Clin. Microbiol. 44:2485–2491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Mitarai S, Kurashima A, Tamura A, Nagai H, Shishido H. 2001. Clinical evaluation of Amplicor Mycobacterium detection system for the diagnosis of pulmonary mycobacterial infection using sputum. Tuberculosis (Edinburgh) 81:319–325 [DOI] [PubMed] [Google Scholar]
- 32. Mokrousov I, et al. 2008. Molecular characterization of ofloxacin-resistant Mycobacterium tuberculosis strains from Russia. Antimicrob. Agents Chemother. 52:2937–2939 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Morgan M, Kalantri S, Flores L, Pai M. 2005. A commercial line probe assay for the rapid detection of rifampicin resistance in Mycobacterium tuberculosis: a systematic review and meta-analysis. BMC Infect. Dis. 5:62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Morris S, et al. 1995. Molecular mechanisms of multiple drug resistance in clinical isolates of Mycobacterium tuberculosis. J. Infect. Dis. 171:954–960 [DOI] [PubMed] [Google Scholar]
- 35. Musser JM. 1995. Antimicrobial agent resistance in mycobacteria: molecular genetic insights. Clin. Microbiol. Rev. 8:496–514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Park H, et al. 2006. Comparison of a conventional antimicrobial susceptibility assay to an oligonucleotide chip system for detection of drug resistance in Mycobacterium tuberculosis isolates. J. Clin. Microbiol. 44:1619–1624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Raynaud C, et al. 1999. Mechanisms of pyrazinamide resistance in mycobacteria: importance of lack of uptake in addition to lack of pyrazinamidase activity. Microbiology 145:1359–1367 [DOI] [PubMed] [Google Scholar]
- 38. Rossau R, et al. 1997. Evaluation of the INNO-LiPA Rif.TB assay, a reverse hybridization assay for the simultaneous detection of Mycobacterium tuberculosis complex and its resistance to rifampin. Antimicrob. Agents Chemother. 41:2093–2098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Santin M. 2004. Incidence and molecular typing of Mycobacterium kansasii in a defined geographical area in Catalonia, Spain. Epidemiol. Infect. 132:425–432 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Scorpio A, et al. 1997. Characterization of pncA mutations in pyrazinamide-resistant Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 41:540–543 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Sekiguchi J, et al. 2007. Detection of multidrug resistance in Mycobacterium tuberculosis. J. Clin. Microbiol. 45:179–192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Sekiguchi J, et al. 2007. Development and evaluation of a line probe assay for rapid identification of pncA mutations in pyrazinamide-resistant Mycobacterium tuberculosis strains. J. Clin. Microbiol. 45:2802–2807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Taillard C, et al. 2003. Clinical implications of Mycobacterium kansasii species heterogeneity: Swiss National Survey. J. Clin. Microbiol. 41:1240–1244 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Takakura S, et al. 2005. Rapid detection of Mycobacterium tuberculosis in respiratory samples by transcription-reverse transcription concerted reaction with an automated system. J. Clin. Microbiol. 43:5435–5439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Wayne LG. 1974. Simple pyrazinamidase and urease tests for routine identification of mycobacteria. Am. Rev. Respir. Dis. 109:147–151 [DOI] [PubMed] [Google Scholar]
- 46. Yuen LK, Leslie D, Coloe PJ. 1999. Bacteriological and molecular analysis of rifampin-resistant Mycobacterium tuberculosis strains isolated in Australia. J. Clin. Microbiol. 37:3844–3850 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Zang Y, Telenti A. 2000. Genetics of drug resistance in Mycobacterium tuberculosis, p.235–254 In Hatfull G. F., Jabobs W. R. (ed), Molecular genetics of mycobacteria. American Society for Microbiology, Washington, DC [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.